Abstract
Low-GC Actinobacteria are among the most abundant and widespread microbes in freshwaters and have largely resisted all cultivation efforts. Consequently, their phages have remained totally unknown. In this work, we have used deep metagenomic sequencing to assemble eight complete genomes of the first tailed phages that infect freshwater Actinobacteria. Their genomes encode the actinobacterial-specific transcription factor whiB, frequently found in mycobacteriophages and also in phages infecting marine pelagic Actinobacteria. Its presence suggests a common and widespread strategy of modulation of host transcriptional machinery upon infection via this transcriptional switch. We present evidence that some whiB-carrying phages infect the acI lineage of Actinobacteria. At least one of them encodes the ADP-ribosylating component of the widespread bacterial AB toxins family (for example, clostridial toxin). We posit that the presence of this toxin reflects a ‘trojan horse’ strategy, providing protection at the population level to the abundant host microbes against eukaryotic predators.
Similar content being viewed by others
Introduction
Studies on specific phages infecting dominant freshwater microbes remain scarce due to the obstacle of obtaining them in axenic culture. Only freshwater cyanophage genomes have been described, six from culture (Yoshida et al., 2008; Chenard et al., 2015) and a few from metagenomic assembly (Skvortsov et al., 2016). However, phages infecting all other major abundant phyla that are typical inhabitants of freshwaters (for example, Actinobacteria) remain totally unexplored. Metagenomics provides a culture-free approach to recover phage genomes and has been successful in several environments ranging from hypersaline (Garcia-Heredia et al., 2012) and marine waters (Mizuno et al., 2013a, 2013b) to the human gut (Dutilh et al., 2014).
Recovery of phage genomes
We were interested in a first glimpse of phages infecting freshwater Actinobacteria, the most abundant microbes in this habitat. We have assembled sequence data from a metagenome and also sequenced a fosmid library (ca. 5500 fosmids) from the freshwater reservoir Amadorio (Alicante, Spain; Supplementary Methods). We focussed only on assembled contigs >10 Kb (both from the metagenomic assembly and from the fosmids) that provided evidence of a circular genome (that is, exact overlapping ends of at least 50 bp). Only contigs that appeared to be Caudovirales phages were retained (as these can be reliably identified by characteristic genes (for example, capsid, terminase, portal protein and tail proteins). Finally, we selected phage contigs that specifically carried a whiB transcription factor found exclusively in genomes of Actinobacteria (Ventura et al., 2007), marking them as putative actinobacterial phages. This gene has been found in several actinobacterial phages, for example, mycobacteriophages (Morris et al., 2008) and streptomyces phages (Van Dessel et al., 2005). It has also been found in a genomic fragment of an uncultured phage predicted to infect the marine actinobacterium Ca. Actinomarina minuta (Ghai et al., 2013; Mizuno et al., 2013b). WhiB is involved in a wide range of cellular processes (Gomez and Bishai, 2000; Alam et al., 2007). In the mycobacteriophage TM4, whiB has been shown to function as a dominant negative regulator by binding specifically to the host whiB promoter resulting in an aberrant septum, similar to a host whiB gene knockout (Rybniker et al., 2010). It also leads to an alteration in cell wall lipids resulting in superinfection exclusion. Thus, we obtained eight complete phage genomes predicted to infect Actinobacteria (Supplementary Table S1).
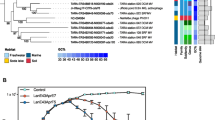
The longest genome was 61.3 Kb and the smallest only 14.4 Kb (the smallest siphoviral genome described yet) and all others were 39–42 Kb. The GC content for all varied from 41 to 49%, as expected from the low-GC content of abundant freshwater Actinobacteria (Ghai et al., 2012). In addition, principal component analyses of oligonucleotide profiles and codon usages of the actinophage genomes displayed good correspondence with known freshwater actinobacterial genomes (Supplementary Figures S1 and S2). Large fractions of their genes (upto 40%) gave best BLAST hits to existing actinobacterial genomes (likely to phages inserted in them) or to actinobacterial phages. Specific comparisons with the latter suggested that these new phages are likely Siphoviruses (Supplementary Table S2). Whole-proteome comparisons revealed a total of four distinct groups (G1–G4), two groups of three phages each with >20% conserved proteins (G2 and G3) and two singletons (Figure 1; Supplementary Figures S3 and S4). Nucleotide identity was only detectable in few short sequence matches within a group, and was non-existent between groups. Phylogenetic analysis of the terminase large subunit (TerL; Casjens et al., 2005) confirmed the same clusters (G1–G4; Supplementary Figure S5). G2 and G3 terminases appear to be related to either siphoviruses or myoviruses, whereas G4 appeared related to lambda-like siphoviruses with 3′-cos ends supporting a siphoviral morphology. Overall, these terminase sequences represent at least three novel evolutionarily distinct lineages of phage terminases.
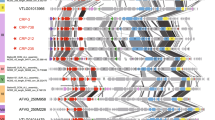
Phage groups and complete genomes of G1 and G4 actinophages. (a) All-vs-all comparison matrix of all proteins in all eight phage genomes. A color scale for the percentage of conserved protein hits is shown below. The four groups of phages, G1, G2, G3 and G4 are shown alongside the matrix. (b) Complete genomes of the shortest (G4) and (c) the longest actinophages (G1) are shown. All proteins for which function could be assigned are labeled and colored in blue. Hypothetical proteins are shown in gray. A length scale is shown at middle right. Genes discussed in the text are underlined.
The three phages of G2 gave potentially several significant hits using both nucleotide and translated nucleotide searches to one contig from a single-cell-amplified genome identified by its 16 S rRNA sequence as belonging to the acI-A7 lineage (acIA-AAA024-D14; Supplementary Figures S6 and S7). This contig represents an incomplete phage genome that is distantly related but still syntenic to all G2 actinophages. Most protein sequence identities are in the range of 30–40% (similar to those among the G2 actinophages). These results indicate that G2-like phages actually infect freshwater Actinobacteria belonging to the acI-A lineage.
Abundance and global distribution
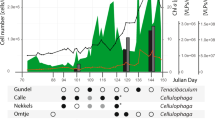
The abundance of these phages is comparable and sometimes higher than actinobacterial genomes assembled from the same reservoir (for example, acMicro1, acAMD2 and acAcidi). All phages recruit maximum reads from the Amadorio reservoir (Figure 2; Supplementary Figure S8) and at very low levels from other sites, suggesting a somewhat endemic character. In contrast, although some actinobacterial genomes recruit mostly from their site of origin (Figure 2), others, for example, acIA-AAA023-D18 obtained from Lake Damariscotta can be found at high levels in multiple locations, for example, Yellowstone lake and Amadorio (Supplementary Figure S9). However, comparisons at the protein level suggest that phages similar to the ones described here can be found globally (Supplementary Figure S10). Moreover, related phages appear to inhabit spatiotemporal regimes similar to their predicted hosts (for example, higher abundances in epilimnion in Yellowstone Lake and in spring compared with summer in Lake Mendota) (Supplementary Figure S11; Allgaier and Grossart, 2006; Salcher et al., 2010). The restricted distribution of some of these phages is in stark contrast to marine phages that may be retrieved from geographically distant locations (Breitbart et al., 2004; Mizuno et al., 2013b). However, the oceans are inter-connected by a defined circulation, whereas freshwater habitats are disconnected and considerably heterogenous in the local environmental conditions.
Fragment recruitment of freshwater Actinobacteria and actinophages in metagenomes and metaviromes. Abundance is expressed as RPKG (reads recruited per Kb of genome per Gb of metagenome). Only hits with >95% identity and >50 bp length were considered. Microbial genomes are grouped according to how they were obtained, single-cell genomics, metagenomic assembly or from pure culture. Genomes are also grouped by phylogeny, acI-A, acI-B, acAMD, acIV and Micrococcineae. The actinophage genomes are shown to the right, and the groups (G1–G4) are indicated within boxes. LF, large 5 μm fraction metagenome; SF, small 0.1 μm fraction metagenome.
A phage-encoded ADP-ribosyltransferase toxin
The G1 actinophage genome (uvFW-CGR-AMD-COM-C203) encodes a protein that shows all the hallmark characteristics and active site motifs of ADP-ribosyltransferases found in AB toxins, also called binary toxins (Figure 1; Supplementary Figures S12 and S13). In these toxins, a receptor-binding B-component facilitates entry of the A-component (the toxin) into the cytoplasm of eukaryotic cells, where it exerts its toxic effect by ADP-ribosylating its target protein. AB toxins are frequently encoded by free-living phages (Casas et al., 2006), or by prophages or plasmids in pathogenic bacteria, for example, Shiga toxin (Obrien et al., 1984), cholera toxin (Waldor and Mekalanos, 1996) or anthrax toxin (Helgason et al., 2000). Phagocytic eukaryotic predators have been suggested as their natural targets (Lainhart et al., 2009). The G1 actinophage genome encodes only the toxic A-component of VIP2-like toxins, a special class of AB toxins, similar to Bacillus cereus VIP2 and Clostridium perfringens iota toxins, that inhibit polymerization of the major eukaryotic cytoskeletal protein actin, leading to cell death (Barth et al., 2004). It is likely that this phage toxin acts in similar manner, but without the requirement for a receptor-binding B-subunit. Remarkably, it has been shown that Escherichia coli cells with phages encoding only the toxic component of the Shiga toxin (Stx), can still cause significant mortality when phagocytosed by Acanthamoeba cells (ca. 25%), nearly all Acanthamoeba internalizing Stx-encoding bacteria being killed (Arnold and Koudelka, 2014). Given the high frequency of phage infections, it is not unlikely that abundant phages (like the G1 actinophage) may infect significant numbers of actinobacterial cells that may comprise more than half of the total microbial community (Glockner et al., 2000). Given that a specific receptor is not needed, this could provide a degree of protection to the host population under the paradigm that a proportion of infected cells will be rendered toxic to a broad range of eukaryotic predators. This altruistic strategy executed in tandem by the host and its phage against eukaryotic predators transforms the infected sub-population into ‘trojan horses’. The phage becomes ‘public goods’ by helping eliminate the predator. In spite of the infected cell being killed by the phage, the host-population density would increase. Thus, the use of such phage-encoded toxins is expected to contribute towards increased fitness both for the phage and its host from a group (kin) selection perspective (Smith, 1976).
References
Alam MS, Garg SK, Agrawal P . (2007). Molecular function of WhiB4/Rv3681c of Mycobacterium tuberculosis H37Rv: a [4Fe-4S] cluster co-ordinating protein disulphide reductase. Mol Microbiol 63: 1414–1431.
Allgaier M, Grossart HP . (2006). Diversity and seasonal dynamics of Actinobacteria populations in four lakes in northeastern Germany. App Environ Microbiol 72: 3489–3497.
Arnold JW, Koudelka GB . (2014). The Trojan Horse of the microbiological arms race: phage-encoded toxins as a defence against eukaryotic predators. Environ Microbiol 16: 454–466.
Barth H, Aktories K, Popoff MR, Stiles BG . (2004). Binary bacterial toxins: biochemistry, biology, and applications of common Clostridium and Bacillus proteins. Microbiol Mol Biol Rev 68: 373–402.
Breitbart M, Miyake JH, Rohwer F . (2004). Global distribution of nearly identical phage‐encoded DNA sequences. FEMS Microbiol Lett 236: 249–256.
Casas V, Miyake J, Balsley H, Roark J, Telles S, Leeds S et al. (2006). Widespread occurrence of phage-encoded exotoxin genes in terrestrial and aquatic environments in Southern California. FEMS Microbiol Lett 261: 141–149.
Casjens SR, Gilcrease EB, Winn-Stapley DA, Schicklmaier P, Schmieger H, Pedulla ML et al. (2005). The generalized transducing Salmonella bacteriophage ES18: complete genome sequence and DNA packaging strategy. J Bacteriol 187: 1091–1104.
Chenard C, Chan AM, Vincent WF, Suttle CA . (2015). Polar freshwater cyanophage S-EIV1 represents a new widespread evolutionary lineage of phages. ISME J 9: 2046–2058.
Dutilh BE, Noriko CW, McNair K, Sanchez SE, Silva GGZ, Boling L et al. (2014). A highly abundant bacteriophage discovered in the unknown sequences of human faecal metagenomes. Nat Commun 5: 4498.
Garcia-Heredia I, Martin-Cuadrado A-B, Mojica FJ, Santos F, Mira A, Antón J et al. (2012). Reconstructing viral genomes from the environment using fosmid clones: the case of haloviruses. PloS ONE 7: e33802.
Ghai R, McMahon KD, Rodriguez-Valera F . (2012). Breaking a paradigm: cosmopolitan and abundant freshwater actinobacteria are low GC. Environ Microbiol Rep 4: 29–35.
Ghai R, Mizuno CM, Picazo A, Camacho A, Rodriguez-Valera F . (2013). Metagenomics uncovers a new group of low GC and ultra-small marine Actinobacteria. Sci Rep 3: 2471.
Glockner FO, Zaichikov E, Belkova N, Denissova L, Pernthaler J, Pernthaler A et al. (2000). Comparative 16 S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria. Appl Environ Microbiol 66: 5053–5065.
Gomez JE, Bishai WR . (2000). whmD is an essential mycobacterial gene required for proper septation and cell division. Proc Natl Acad Sci USA 97: 8554–8559.
Helgason E, Okstad OA, Caugant DA, Johansen HA, Fouet A, Mock M et al. (2000). Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis - One species on the basis of genetic evidence. Appl Environ Microbiol 66: 2627–2630.
Lainhart W, Stolfa G, Koudelka GB . (2009). Shiga Toxin as a Bacterial Defense against a Eukaryotic Predator, Tetrahymena thermophila. J Bacteriol 191: 5116–5122.
Mizuno CM, Rodriguez-Valera F, Garcia-Heredia I, Martin-Cuadrado AB, Ghai R . (2013a). Reconstruction of novel cyanobacterial siphovirus genomes from Mediterranean metagenomic fosmids. Appl Environ Microbiol 79: 688–695.
Mizuno CM, Rodriguez-Valera F, Kimes NE, Ghai R . (2013b). Expanding the marine virosphere using metagenomics. PLoS Genet 9: e1003987.
Morris P, Marinelli LJ, Jacobs-Sera D, Hendrix RW, Hatfull GF . (2008). Genomic characterization of mycobacteriophage Giles: evidence for phage acquisition of host DNA by illegitimate recombination. J Bacteriol 190: 2172–2182.
Obrien AD, Newland JW, Miller SF, Holmes RK, Smith HW, Formal SB . (1984). Shiga-Like Toxin-Converting Phages from Escherichia-Coli Strains That Cause Hemorrhagic Colitis or Infantile Diarrhea. Science 226: 694–696.
Rybniker J, Nowag A, van Gumpel E, Nissen N, Robinson N, Plum G et al. (2010). Insights into the function of the WhiB-like protein of mycobacteriophage TM4-a transcriptional inhibitor of WhiB2. Mol Microbiol 77: 642–657.
Salcher MM, Pernthaler J, Posch T . (2010). Spatiotemporal distribution and activity patterns of bacteria from three phylogenetic groups in an oligomesotrophic lake. Limnol Oceanogr 55: 846–856.
Skvortsov T, de Leeuwe C, Quinn JP, McGrath JW, Allen CC, McElarney Y et al. (2016). Metagenomic characterisation of the viral community of Lough Neagh, the largest freshwater lake in Ireland. PloS One 11: e0150361.
Smith JM . (1976). Commentary: group selection. Q Rev Biol 51: 277–283.
Van Dessel W, Van Mellaert L, Liesegang H, Raasch C, De Keersmaeker S, Geukens N et al. (2005). Complete genomic nucleotide sequence and analysis of the temperate bacteriophage VWB. Virology 331: 325–337.
Ventura M, Canchaya C, Tauch A, Chandra G, Fitzgerald GF, Chater KF et al. (2007). Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol Mol Biol Rev 71: 495–548.
Waldor MK, Mekalanos JJ . (1996). Lysogenic conversion by a filamentous phage encoding cholera toxin. Science 272: 1910–1914.
Yoshida T, Nagasaki K, Takashima Y, Shirai Y, Tomaru Y, Takao Y et al. (2008). Ma-LMM01 infecting toxic Microcystis aeruginosa illuminates diverse cyanophage genome strategies. J Bacteriol 190: 1762–1772.
Acknowledgements
This work was supported by projects PROMETEO II/2014/012 project AQUAMET from the Generalitat Valenciana, MEDIMAX BFPU2013-48007-P from the Spanish Ministerio de Economía y Competitividad and MaCuMBA Project 311975 of the European Commission FP7 to FR-V. RG was partially supported by the Grant Agency of the Czech Science Foundation under the research grant 13-00243 S. We thank Michaela M. Salcher for discussions and helpful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Ghai, R., Mehrshad, M., Mizuno, C. et al. Metagenomic recovery of phage genomes of uncultured freshwater actinobacteria. ISME J 11, 304–308 (2017). https://doi.org/10.1038/ismej.2016.110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.110
This article is cited by
-
COBRA improves the completeness and contiguity of viral genomes assembled from metagenomes
Nature Microbiology (2024)
-
High-resolution metagenomic reconstruction of the freshwater spring bloom
Microbiome (2023)
-
Isolation of phages infecting the abundant freshwater Actinobacteriota order ‘Ca. Nanopelagicales’
The ISME Journal (2023)
-
Comparative genomic analysis of five freshwater cyanophages and reference-guided metagenomic data mining
Microbiome (2022)
-
Diversity and distribution of viruses inhabiting the deepest ocean on Earth
The ISME Journal (2021)