Abstract
Rapid warming in the highly productive western Antarctic Peninsula (WAP) region of the Southern Ocean has affected multiple trophic levels, yet viral influences on microbial processes and ecosystem function remain understudied in the Southern Ocean. Here we use cultivation-independent quantitative ecological and metagenomic assays, combined with new comparative bioinformatic techniques, to investigate double-stranded DNA viruses during the WAP spring–summer transition. This study demonstrates that (i) temperate viruses dominate this region, switching from lysogeny to lytic replication as bacterial production increases, and (ii) Southern Ocean viral assemblages are genetically distinct from lower-latitude assemblages, primarily driven by this temperate viral dominance. This new information suggests fundamentally different virus–host interactions in polar environments, where intense seasonal changes in bacterial production select for temperate viruses because of increased fitness imparted by the ability to switch replication strategies in response to resource availability. Further, temperate viral dominance may provide mechanisms (for example, bacterial mortality resulting from prophage induction) that help explain observed temporal delays between, and lower ratios of, bacterial and primary production in polar versus lower-latitude marine ecosystems. Together these results suggest that temperate virus–host interactions are critical to predicting changes in microbial dynamics brought on by warming in polar marine systems.
Similar content being viewed by others
Introduction
Polar marine ecosystems are highly susceptible to the effects of warming and climate change because of the significant influence of sea ice in ecosystem dynamics (reviewed by Doney et al., 2012). A focal point of this polar research has been the marine region of the western Antarctic Peninsula (WAP) as it is one of the most rapidly warming regions on the planet (reviewed by Doney et al., 2012; Ducklow et al., 2012a) and is part of the highly productive Southern Ocean, which contributes an estimated 20% of global oceanic CO2 uptake (Takahashi et al., 2002). Long-term research in the WAP demonstrates that rapid warming has already reduced sea ice extent and duration, affecting multiple trophic levels ranging from ecosystem foundational microbes to high-level consumers such as krill and penguins (Ducklow et al., 2012a, b; Saba et al., 2014). On land, microorganisms are having significant roles in polar ecosystem changes brought on by thawing permafrost, with some cases showing that the abundance of key microorganisms best predicts carbon cycling and climate feedbacks for global-scale models (McCalley et al., 2014; Mondav et al., 2014). Similarly, marine microbes are critical to the functioning of the WAP food web, and with warming are expected to have increasingly significant ecosystem roles (Sailley et al., 2013).
Viruses are known to have substantial roles in marine ecosystems through their alteration of microbial community dynamics and processes. Viral infections cause ca 10–40% of bacterial mortality in marine systems and alter biogeochemical cycling through the release of cellular contents during lysis (reviewed by Fuhrman, 1999; Suttle, 2007; Brum et al., 2014). Further, viruses can possess metabolic genes including those involved in photosynthesis, sulfur metabolism and carbon metabolism, thus directly affecting ecosystem productivity and biogeochemical cycling through expression of these genes during infection of microorganisms (reviewed by Breitbart, 2012; Brum and Sullivan, 2015). Quantitative examination of these viral roles in nature has been challenging, but recent methodological advances have drastically expanded our ability to investigate viral ecology and their effects on ecosystem processes (reviewed by Brum and Sullivan, 2015). Critically, these advances include an optimized sample-to-sequence pipeline to generate quantitative double-stranded DNA (dsDNA) viral metagenomes (viromes; reviewed by Solonenko and Sullivan, 2013), which has facilitated leaps in our knowledge of viral genomic diversity, niche differentiation and ecological drivers of community variability (reviewed by Brum and Sullivan, 2015).
Although the above advances facilitate understanding of viruses in marine ecosystems, knowledge of viral roles in polar regions remains quite limited. In the Southern Ocean, the majority of studies to date have focused on spatial and temporal variability of community viral abundance using microscopy (for example, Bird et al., 1993; Marchant et al., 2000; Pearce et al., 2007; Yang et al., 2010). The few studies that have experimentally determined microbial mortality as a result of viral infection in the Southern Ocean have demonstrated strong spatial variability in this parameter and indicated that viral lysis is important as a mortality factor as well as a source of organic matter for microorganisms (Guixa-Boixereu et al., 2002; Brussaard et al., 2008; Evans et al., 2009; Evans and Brussaard, 2012; Malits et al., 2014). Further, investigation of Southern Ocean viral genomic diversity has previously been limited to evaluations of viral genome size distribution (Brussaard et al., 2008), single-gene-based examination of algal virus diversity (Short and Suttle, 2002) and identification of viral genes within six bacterial fosmids (Cottrell and Kirchman, 2012). Recently published results from a global-scale analysis of quantitatively produced viromes include two samples from the Southern Ocean and suggest that viral diversity in this region is lower than that observed at lower-latitude locations (Brum et al., 2015). Thus, although viruses are undoubtedly important in Southern Ocean ecosystem function, they remain understudied there as compared with other marine environments.
Viruses in polar systems are also thought to have diverse replication strategies, which may be critical to understanding viral ecology in low-temperature environments (Anesio and Bellas, 2011). Viral infections can be exclusively lytic, wherein viral takeover of cellular machinery results in new viral progeny and lysis of the host, or may involve a lysogenic stage in the case of temperate viruses wherein viral DNA is maintained within the host as a prophage until induced to replicate lytically (reviewed by Fuhrman, 2000; Miller and Day, 2008). The current paradigm, based on cultivated temperate virus–host systems, is that lytic replication dominates when bacterial production and abundance are high, whereas lysogeny is favored when they are low (reviewed by Miller and Day, 2008; Paul, 2008). However, this paradigm remains contentious because of the conflicting evidence obtained from community-level environmental studies (reviewed by Weinbauer, 2004; Paul, 2008). Specifically, community-level temporal studies examining the relationship between lysogeny and bacterial production and/or abundance have shown mixed results including strong negative relationships between these variables (Laybourn-Parry et al., 2007; Payet and Suttle, 2013), weak negative relationships (Williamson et al., 2002), no relationship (Boras et al., 2009) or differing relationships within depth-differentiated data sets from the same study (Thomas et al., 2011). We note that of these studies, the strongest support for the paradigm of negative temporal relationships between lysogeny and bacterial production has been found in polar aquatic regions including the Arctic Ocean (Payet and Suttle, 2013) and Antarctic lakes (Laybourn-Parry et al., 2007). A spatial study including the Southern Ocean also suggests that lysogeny may be more prevalent in regions with lower nutrients and abundance of prokaryotes (Evans and Brussaard, 2012). Further, although no viral metagenomic data exist in Antarctic marine waters, such data from the Arctic Ocean derived from non-quantitatively amplified DNA (Yilmaz et al., 2010) yielded more prophage-related sequences in the Arctic than lower-latitude marine systems (Angly et al., 2006), which is at least qualitatively consistent with polar regions having more temperate viruses. We thus hypothesized that temperate viruses (that is, those capable of both lysogeny and lytic replication) are more abundant in polar regions, and that this may explain the divergent support for this paradigm in aquatic systems across latitudes.
Here we target the WAP region of the Southern Ocean to specifically evaluate the hypotheses that viral replication strategy is determined by bacterial activity, and that temperate viruses dominate polar aquatic regions. To this end, we combined quantitative ecological, metagenomic and bioinformatic analyses across the WAP spring–summer transition to (i) track temporal changes in lysogeny and lytic viral replication in relation to bacterial production and abundance, (ii) evaluate the influence of temperate viruses in this region and (iii) compare WAP dsDNA viral communities with those from lower-latitude marine systems.
Results and Discussion
Temporal changes in viral replication strategies
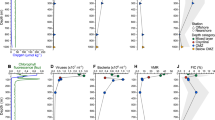
Nearly 3 months of sampling across the spring–summer transition in the WAP revealed contrasting viral and microbial community dynamics (Figure 1), with lysogeny negatively correlated and lytic infections positively correlated with bacterial abundance, bacterial production and viral abundance (Table 1). Springtime’s moderate levels of bacterial abundance, production and chlorophyll a concentration were matched by moderate viral abundance, with few lytic viral infections and a large fraction (5–16%) of lysogens (that is, bacteria that contained inducible prophages). In contrast, summertime had characteristically higher levels of all measured parameters except lysogeny, which was nearly undetectable. Although the virus to bacterium ratio (VBR) varied considerably during the study period, ranging from 6 to 34 (Supplementary Figure S1), there was no significant relationship between VBR and lysogeny or lytic infections (Table 1). Notably, sampling in this study occurred after the sea ice had melted, therefore the observed dynamics cannot be directly attributed to the release of ice-associated viruses and microorganisms into the water column following seasonal ice melt (Paterson and Laybourn-Parry, 2012).
Ecological variables measured in the surface ocean at Palmer LTER Station B from November 2010 through January 2011. Bacteria refers to Bacteria plus Archaea. Lytic viral infections are measured as the frequency of infected cells (FICs) and lysogeny is measured as the percentage of bacteria with inducible prophages. Error bars for FICs are upper and lower 95% confidence intervals. Error bars for bacteria, viruses and lysogeny are s.d. of the means of triplicate samples. 0, below detection; *, statistically significant results for lysogeny; arrows indicate when samples for virome construction were collected.
Comparative analyses of dsDNA viromes constructed from free viruses and induced viruses (that is, temperate viruses chemically induced to switch from lysogeny to lytic replication) at two sampling dates in the spring and summer were conducted using a new shared k-mer-based social network analysis that takes into account the abundance of shared and unique sequences among samples, and does not require assembly or annotation of the virome sequences (Hurwitz et al., 2014). Although these viromes do not include single-stranded DNA or RNA viruses, the <0.2 μm dsDNA viruses that they do include are thought to be numerically dominant in marine viral assemblages (reviewed by Brum and Sullivan, 2015). The spring-free virome was markedly different from both induced viromes, with the majority (59%) of its sequences absent from the induced viromes, suggesting they represented lytic viruses (Figure 2). In contrast, the summer-free virome was highly similar to both induced viromes (sharing 82% of its sequences with the induced viromes) and was therefore assumed to be predominantly comprising temperate viruses capable of utilizing both lysogeny and lytic replication (Figure 2). Given that summer-free viruses were ca fourfold more abundant than spring-free viruses (Figure 1), these results also suggest that temperate viruses dominate the WAP viral assemblage, at least for dsDNA viruses examined here.
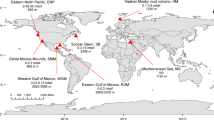
Social network analysis comparing all sequences in the four WAP viromes from the Southern Ocean (SO) with nine lower-latitude, seasonally variable surface samples from the POV data set including five viromes from the LineP transect (LP), three from the Monterey Bay Line 67 transect (MB) and one from Scripps Pier (SP). The network analysis compares viromes based on the abundance of shared k-mers among all sequences in each virome. Small dots represent the probability of the virome position given shared sequence content, labeled white dots represent the mean virome position and dot colors encode proximity of viromes on the plot.
Together, the ecological and metagenomic results suggest that temperate viruses primarily used lysogeny in spring and switched to lytic replication in summer in response to increased bacterial abundance and productivity. This supports the paradigm that viral replication strategy is determined by bacterial physiological status, as evidenced in many cultivated temperate virus–host systems including the model system of Escherichia coli and phage lambda (reviewed by Weinbauer, 2004; Miller and Day, 2008; Paul, 2008). Specifically, in this paradigm host physiological state affects the lytic–lysogenic switch, with poor host physiological conditions (for example, low bacterial production in spring) driving the viruses toward lysogeny, and greater host metabolic health (for example, higher bacterial production in summer) resulting in lytic replication because the viruses have the resources to produce viral progeny (reviewed by Miller and Day, 2008).
Dominance of temperate viruses in polar regions
The hypothesis that temperate viruses are more abundant in polar than lower-latitude aquatic systems was supported by comparative metagenomic analyses in this study. Comparison of WAP dsDNA viromes with nine surface-water lower-latitude Pacific Ocean dsDNA viromes (from the Pacific Ocean virome (POV) data set prepared using the same quantitative methodology; Hurwitz and Sullivan, 2013) showed two clear patterns. First, temperate virus-enriched WAP viromes (that is, summer ‘free’ and both induced viromes) were considerably more divergent from the lower-latitude POV than was the lytic virus-enriched WAP virome (that is, the spring ‘free’ virome; Figure 2). Second, the functions, where annotatable (Supplementary Table S1), of sequences unique to WAP relative to POV paralleled those of unique sequences observed in temperate virus-enriched versus lytic virus-enriched WAP viromes (Table 2). Although the lower-latitude POV samples did not include induced temperate viromes, they did include samples from multiple seasons at the same location. Thus, any significant seasonal changes in the POV-free viromes because of a high portion of temperate viruses should have been evident like they are in the WAP virome data set. Together, this information is consistent with temperate viruses dominating WAP but not lower-latitude dsDNA viral assemblages.
Comparative analysis of viromes also generated new insights regarding differences between temperate and lytic viruses in the WAP. Although the percentage of taxonomically annotated viral sequences was low in WAP viromes (5–7%) as is common in marine viromes (Hurwitz and Sullivan, 2013), notable taxonomic trends emerged. Specifically, myoviruses were more abundant in the lytic-enriched virome (spring-free virome) and podoviruses were more abundant in temperate-enriched viromes (summer-free, spring and summer-induced viromes) as evident in the taxonomic composition of these viromes (Supplementary Figure S2), as well as analysis of the sequences that were unique to temperate and lytic viruses (Figure 3). This contradicted the common view that temperate viruses are mainly siphoviruses, with podoviruses including phage P22 (reviewed by Susskind and Botstein, 1978) as the exception. Of course, as 93–95% of the reads remain taxonomically unassigned, these patterns may or may not hold as more reference genomes become available for refining such analyses.
Functionally, temperate-enriched viromes had more genes encoding for replication and structural functions, whereas metabolic and membrane transport functions were ca five- to ninefold more abundant in the lytic-enriched virome (Supplementary Table S2). Further, metabolic and membrane transport genes comprised only 4% of the functionally annotated unique temperate viral sequences in contrast to 24% of the unique lytic viral sequences, and also drove the difference between the WAP lytic-enriched virome and POV (Table 2). These auxiliary metabolic genes are defined as viral metabolic genes not directly involved in viral replication, and function to modify host metabolic processes for the purpose of enhancing viral replication (reviewed by Breitbart, 2012; Brum and Sullivan, 2015). The enrichment of auxiliary metabolic genes in WAP lytic viruses implies that to survive intense seasonal changes in bacterial production in the WAP, the minority lytic viruses enhance their host’s metabolism through auxiliary metabolic genes, presumably to aid replication when resources are limiting. In contrast, the numerically dominant temperate viruses appear to alternate replication strategies to compensate for seasonally variable host productivity.
Quantifying the importance of temperate viruses in nature has long been desired as they are critical for modeling microbial ecosystems. However, such cultivation-independent quantification has previously not been attainable because of technical limitations that result in non-quantitative viromes (Yilmaz et al., 2010; Duhaime and Sullivan, 2012; Marine et al., 2014; Brum and Sullivan, 2015). Our findings utilizing a combination of experimental methods and quantitative dsDNA virome construction and comparison suggest that temperate viruses dominate polar aquatic systems, probably because the intense seasonal variations in productivity that characterize these regions favor viruses capable of switching replication strategies in response to changes in host productivity. This may explain the lack of consensus among temporal aquatic studies relating lysogeny to bacterial production and abundance at various latitudes (Table 3). Although the range of percent lysogeny detected in these studies shows no relationship with latitude, it is clear from the observed temporal trends that higher-latitude environments display strong seasonality in percent lysogeny, with the highest values in winter and spring, in contrast to lower-latitude environments where lysogeny is sporadically detected. Further, significant correlations between lysogeny and bacterial production and abundance (where available) are weak or absent in lower-latitude environments but strengthen at high latitudes. These trends with latitude may be explained by the relative abundance of temperate viruses. Specifically, if the majority of viruses in lower-latitude aquatic environments are incapable of lysogeny (that is, are not temperate viruses), this could explain the reduced strength of temporal correlations between lysogeny and total community bacterial production and abundance, resulting in the weak relationships or lack of relationship between these variables observed in lower-latitude studies (Williamson et al., 2002; Boras et al., 2009; Thomas et al., 2011). In contrast, the strong seasonal relationships observed between lysogeny and bacterial production in polar aquatic systems (Laybourn-Parry et al., 2007; Payet and Suttle, 2013; this study) most likely resulted from higher relative abundances of temperate viruses.
Effects of temperate viral dominance on polar ecosystem function
Dominance of temperate viruses in the Southern Ocean also has significant implications for interpreting corresponding microbial dynamics. For example, prophage induction alters bacterial community composition through selective mortality (Hewson and Fuhrman, 2007; Supplementary Figure S3) and likely contributes to observed temporal variability in WAP bacterial assemblage composition (Ghiglione and Murray, 2012; Grzymski et al., 2012; Williams et al., 2012; Supplementary Figure S3) and metabolic potential (Grzymski et al., 2012; Williams et al., 2012). Specifically, chemical induction of lysogens in spring resulted in a decrease in Gammaproteobacteria and a concomitant increase in Flavobacteria (Supplementary Figure S3). Although bacterial composition is only available for two sampling dates in this study, the available data are consistent with the hypothesis that the shift from lysogeny to lytic viral replication when bacterial productivity increased (late December; Figure 1) could substantially alter the WAP bacterial community.
In addition, temperate viral dominance may help resolve the decades-long debate regarding whether the relationship between primary and bacterial production in the Southern Ocean is altered in comparison with lower-latitude marine ecosystems (reviewed by Moran et al., 2001; Kirchman et al., 2009; Ducklow et al., 2012a). In the Southern Ocean, increased solar radiation in springtime results in the melting of sea ice followed by generation of phytoplankton blooms and subsequent increases in bacterial production fueled by phytoplankton-derived dissolved organic carbon (Ducklow et al., 2007, 2012a, b). Studies conducted during the onset of Southern Ocean phytoplankton blooms conclude that bacterial production is ‘uncoupled’ from primary production inferred from a greater delay between phytoplankton bloom initiation and increased bacterial production relative to lower latitudes, as well as a reduced ratio of bacterial to primary production (Billen and Becquevort, 1991; Karl et al., 1991; Lochte et al., 1997; Bird and Karl, 1999; Ducklow et al., 2001; Moran and Estrada, 2002; Duarte et al., 2005). In contrast, during full-bloom or non-bloom conditions in the Southern Ocean, bacteria and phytoplankton are described as temporally ‘coupled’, although still with reduced ratios of bacterial to primary production relative to lower latitudes (Moran et al., 2001; Moran and Estrada, 2002; Ducklow et al., 2012b). Although this altered relationship between primary and bacterial production in the Southern Ocean was previously hypothesized to result from lower temperatures (Pomeroy and Wiebe, 2001), further study does not support that hypothesis (Kirchman et al., 2009; Ducklow et al., 2012b).
We hypothesize that temperate virus dominance may help explain these altered relationships between primary and bacterial production in the Southern Ocean as follows (summarized in Figure 4). First, viral gene expression during lysogeny can reduce host metabolism to increase survival when resources are limiting (reviewed by Paul, 2008). Specifically, in cultivated temperate virus–host systems including E. coli and phage lambda, as well as a Pseudoalteromonas phage system from Arctic sea ice, prophage-infected hosts exhibit reduction in substrate usage and growth rate compared with uninfected hosts, thereby increasing the fitness of the infected hosts when resources are scarce (Chen et al., 2005; Paul, 2008; Yu et al., 2015). Thus, lysogeny, which dominated pre-bloom (Figure 1), may decrease the ratio of bacterial to primary production before seasonal phytoplankton blooms. Second, lysogeny sharply declined from 16% (average) to 0% during bloom initiation (in late December), indicating induction of lysogens (Figure 1). Bacterial mortality caused by the induction of these temperate viral ‘time bombs’ (sensu Paul, 2008) likely delays increases in bacterial abundance and production at the onset of phytoplankton blooms. Finally, prophage induction may give viruses a head start on lytic infection dynamics relative to lower latitudes as demonstrated by the increase in viral abundance before bacterial abundance in this study (Figure 1). This would result in reduced bacterial production following phytoplankton blooms because the increased contact rates between viruses and their hosts would contribute to the increase in fatal lytic infections evident in this study (Figure 1). Together, these points suggest temperate viral dominance may help explain the observed temporal uncoupling of bacteria and phytoplankton at the onset of seasonal phytoplankton blooms, as well as reduced responses of bacteria to phytoplankton-derived dissolved organic carbon at all times in the Southern Ocean, relative to lower latitudes (Figure 4).
Generalized illustration showing how the dominance of temperate viruses in the Southern Ocean can delay bacterial response to phytoplankton blooms and reduce the ratio of bacterial production to primary production relative to lower-latitude marine environments. Lysogeny dominates pre-bloom and has been shown to suppress metabolic activity in host bacteria, whereas induction of lysogens at the onset of phytoplankton blooms results in bacterial mortality and production of free viruses, which proceed to cause bacterial mortality via lytic infections.
Conclusions
In summary, this study provides quantitative viral metagenomic data contextualized by viral ecological measurements to demonstrate that temperate viruses dominate the WAP dsDNA viral assemblage, primarily utilizing lysogeny when bacterial production is low and switching to lytic replication when bacterial production increases. Temperate viral dominance in polar regions offers a mechanism for viral ‘survival’ under harsh winter conditions, and helps resolve long-standing questions about the links between lysogeny and lytic viral replication along ecological gradients and altered microbial dynamics observed in the Southern Ocean. These results complement long-term ecological research in the WAP (Ducklow et al., 2012a, 2012b; Saba et al., 2014) and suggest that temperate viruses have substantial roles in modulating microbially driven processes such as carbon and elemental cycling in this region. Given model-predicted increased microbial roles as warming intensifies (Sailley et al., 2013), such fundamental new understanding of virus–host interactions is critical to improving predictions of ecosystem function in polar marine regions, which are currently bearing the brunt of climate change (Doney et al., 2012; Ducklow et al., 2012a).
Materials and methods
Ecological parameters
Surface seawater samples were collected twice weekly, as weather permitted, from the Palmer, Antarctica Long-Term Ecological Research Program (Ducklow, 2008) Station B (64º 46.45' S, 64º 03.27' W), a coastal site near Palmer Station on the WAP, between 8 November 2010 and 31 January 2011. The water samples were collected at the surface by submerging clean bottles into the seawater by hand.
Triplicate samples (4 ml) for viral and bacterial enumeration were preserved with EM-grade glutaraldehyde (2% final concentration), flash-frozen in liquid nitrogen and stored between –72 °C and −80 °C until analysis. Viral and bacterial concentration were determined based on a previously described method (Noble and Fuhrman, 1998) in which thawed samples were filtered onto 0.02-μm-pore-size filters (Anodisc, Whatman, GE Healthcare Life Sciences, Piscataway, NJ, USA), stained with SYBR Gold nucleic acid stain (Invitrogen, Life Technologies, Carlsbad, CA, USA) and enumerated using an epifluorescence microscope (Axio Imager.D2, Zeiss, Jena, Germany).
Chlorophyll a concentration was determined from 1 to 2 liter of seawater (depending on biomass) by fluorescence of extracted samples (Holm-Hansen et al., 1965). Bacterial production rates were measured using the microcentrifuge method (Kirchman et al., 1985; Smith and Azam, 1992; Kirchman, 1993) based on incorporation of 3H-leucine (Perkin-Elmer, Waltham, MA, USA; 144 Ci mmol−1, 20 nM final concentration) in triplicate, 1.5 ml pseudoreplicates incubated for 3 h at in situ temperatures (ca 1 ºC). A blank killed with trichloroacetic acid was also incubated along with the triplicate samples and the 3H-leucine incorporation of the blank was subtracted from the samples. A conversion factor of 1.5 kg C produced per mole of leucine incorporated (Ducklow et al., 2000, 2012b) was used to calculate bacterial production rates.
The percentage of cells with lytic viral infections was determined by transmission electron microscopy to quantify the frequency of visibly infected cells (Proctor and Fuhrman, 1990) using intact cells (Brum et al., 2005). Seawater samples were preserved with EM-grade glutaraldehyde (2% final concentration), flash-frozen in liquid nitrogen and stored between –72 °C and −80 °C until analysis. Samples (26–40 ml, depending on bacterial concentration) were centrifuged for 1 h at 55 000 g using an ultracentrifuge (LM-80, Beckman, Brea, CA, USA) onto grids (200 mesh copper grids with carbon-stabilized formvar support; Ted Pella, Redding, CA, USA) made hydrophilic with 20 s of glow discharge (Hummer 6.2, Anatech, Battle Creek, MI, USA). Grids were then stained with uranyl acetate and analyzed as previously described (Brum et al., 2005) to determine the frequency of visibly infected cells using a transmission electron microscope (CM12, Philips, Eindhoven, The Netherlands). The frequency of infected cells was then calculated from the frequency of visibly infected cells (Binder, 1999). Burst size was also determined from the visibly infected cells as previously described (Brum et al., 2005). Given the low sample size for burst size at most sampling dates (Supplementary Figure S4), this variable was not included in the temporal analysis and instead was used only to calculate percent lysogeny (see below).
The percentage of cells with lysogenic viral infections (lysogeny) was determined using the prophage induction method (Paul and Weinbauer, 2010) in which six replicates (50 ml), three of which were amended with the inducing agent mitomycin C (1 μg ml−1, final concentration), were incubated in the dark at in situ temperatures (ca 1 ºC) for 24 h. Viral concentration was determined at the start and end of the incubation as described above. Increases in viral concentration in the mitomycin C-amended samples versus control samples (no treatment) were evaluated for statistical significance using ANOVA (SigmaPlot version 11.0, Systat Software, Chicago, IL, USA). Lysogeny was calculated using the average burst size of 41 determined for this study, as described above. For samples in which viral concentrations in mitomycin C-amended replicates were slightly (but not significantly) lower than those in control replicates, lysogeny was reported to be below detection.
Virome construction
Viral concentrates for metagenomic analysis were collected on 18 November 2010 and 29 January 2011, at Station B. Surface seawater was pumped into large containers, after which concentrates of free viruses were obtained from 120 liter of 0.2 μm-filtered (Polycap 75 TC, Whatman, GE Healthcare Life Sciences) seawater using iron chloride flocculation (John et al., 2011). Concentrates of induced viruses were also obtained from 120 liter surface samples by first filtering the sample through a 3 μm pore size filter (Isopore, Millipore, Billerica, MA, USA) to reduce the concentration of grazers and large phytoplankton, then concentrating the sample to ca 1.4 liter with a 0.22 μm pore size tangential flow filter (GE Healthcare Life Sciences) to recover bacteria (ca 60% and 63% recovery in November and January, respectively) and remove free viruses (ca 94% and 75% removal in November and January, respectively). This sample was then split into control (ca 200 ml) and mitomycin C-amended (1 μg ml-1 final concentration) treatment (ca 1.2 l) samples and incubated for 24 h in the dark at in situ temperature (ca 1 ºC), resulting in an 2-fold and 1.2-fold increase in viral concentration in the treatment versus control samples for the November (spring) and January (summer) incubations, respectively. Treatment samples were then 0.22 μm filtered (Steripak, Millipore) and viruses were concentrated using iron chloride flocculation (John et al., 2011) and stored at 4 °C until analysis. Approximately 20% of the collected free and induced viral concentrates for each sampling date (equivalent to ca 20 l of starting seawater sample) were then processed to construct viromes as follows, with the remainder set aside for future experiments. Viral concentrates were resuspended with ascorbic acid buffer, treated with DNase and purified using cesium chloride density gradients as previously described (Hurwitz et al., 2013). Viral dsDNA was then extracted and linker amplified as previously described (Duhaime et al., 2012), followed by pyrosequencing (GS-FLX, 454 Life Sciences, Branford, CT, USA) at the Emory Integrated Genomics Core (Emory University, Atlanta, GA, USA).
Virome analysis
Quality-controlled WAP virome reads were compared against the Similarity Matrix of Proteins (Rattei et al., 2006) released 25 June 2011, using BLASTX to assign taxonomy and function (based on Pfam) as previously described (Hurwitz and Sullivan, 2013). Hits were considered significant if they had an E-value <0.001 and only top hits were retained.
Comparisons were made among the four WAP viromes and nine selected 10-m viromes from the POV data set (Hurwitz and Sullivan, 2013) including LineP Station 26 in February, June and August (L.Win.O.10 m, L.Spr.O.10 m and L.Sum.O.10 m, respectively); LineP Station 12 in June (L.Spr.I.10m); LineP Station 4 in June (L.Spr.C.10 m); MBARI Stations H3, 67–70 and 67–155 in October (M.Fall.C.10 m, M.Fall.I.10 m and M.Fall.O.10 m, respectively); and Scripps Pier in April (SFC.Spr.C.5 m). These viromes were compared with one another with a shared k-mer approach using the vmatch package version 2.1.5 (http://www.vmatch.de) with parameters (-pl –allout –v) and k-mer length of 20 to identify unique and shared reads (mode k-mer occurrence >2 and <1000) among the viromes as previously described (Hurwitz et al., 2014). The results of the shared k-mer analysis were then assembled into a matrix consisting of the average of the counts ( ) in common out of the average of the total possible counts (
) in common out of the average of the total possible counts ( ) between virome i and virome j (for i=1,…13) and used as input into a social network analysis for the four WAP viromes and nine POV as previously described (Hurwitz et al., 2014). Briefly, the social network analysis is a robust statistical framework to visually compare samples based on their shared k-mers in k-dimensional latent space wherein viromes that are closer together are more similar, and those that are physically separate are significantly different. The code for analyses is available in the open source github code repository (https://github.com/hurwitzlab/fizkin).
) between virome i and virome j (for i=1,…13) and used as input into a social network analysis for the four WAP viromes and nine POV as previously described (Hurwitz et al., 2014). Briefly, the social network analysis is a robust statistical framework to visually compare samples based on their shared k-mers in k-dimensional latent space wherein viromes that are closer together are more similar, and those that are physically separate are significantly different. The code for analyses is available in the open source github code repository (https://github.com/hurwitzlab/fizkin).
Bacterial community diversity
Bacterial 16S ribosomal RNA gene diversity was obtained from the 0.2 μm pore size filters used to prepare both the free and induced viral concentrates, which were stored between −72 °C and −80 °C until analysis. DNA was extracted based on a filter extraction method (Rich et al., 2008) modified for the capsule filters used in this study (V. Rich, personal communication, May 2012). For the larger filters (Polycap 75 TC, Whatman), 45 ml of lysis buffer (40 mM EDTA, 50 mM Tris pH 8.3, 0.73 M sucrose) was added directly to the frozen filters, which were then sealed and thawed on ice. Lysozyme (13.5 ml of 5 mg ml-1 lysozyme in lysis buffer) was then added and filters were incubated, rotating, at 37 °C for 1 h. Proteinase K (4.5 ml of 10 mg ml−1 proteinase K in lysis buffer) and 6.75 ml of 10% sodium dodecyl sulfate were then added to the filters and incubated, rotating, at 55 °C overnight. The lysate was recovered from the filters, which were then rinsed with an additional 20 ml of lysis buffer and pooled with the recovered lysates. For the smaller filters (Steripak, Millipore), the above volumes were reduced by 75%. Pooled lysates were then extracted twice with phenol–chloroform–isoamyl alcohol (25:24:1, pH 8.0) and once with chloroform–isoamyl alcohol (24:1). The aqueous phases were concentrated to ca 200 μl with centrifugal concentrators (Amicon Ultra-15 with 100 kDa cut-off, Millipore), rinsed twice with 8 ml of TE buffer (Sigma-Aldrich, St Louis, MO, USA) and concentrated to ca 200 μl. Samples were then ethanol extracted, resuspended in TE buffer and stored at −20 ºC. Extracted DNA was pyrosequenced (GS-FLX, 454 Life Sciences) after amplification with 16S ribosomal RNA gene primers (926F and 1492R; Lane, 1991) at the Australian Center for Ecogenomics (University of Queensland, Brisbane, Australia). Taxonomy of the sequence reads was determined using PyroTagger (Kunin and Hugenholtz, 2010) with the Uclust algorithm (http://www.drive5.com/uclust) and a threshold length of 220 bases.
References
Anesio AM, Bellas CM . (2011). Are low temperature habitats hot spots of microbial evolution driven by viruses? Trends Microbiol 19: 52–57.
Angly FE, Felts B, Breitbart M, Salamon P, Edwards RA, Carlson C et al. (2006). The marine viromes of four oceanic regions. PLoS Biol 4: 2121–2131.
Billen G, Becquevort S . (1991). Phytoplankton-bacteria relationship in the Antarctic marine ecosystem. Polar Res 10: 245–254.
Binder B . (1999). Reconsidering the relationship between virally induced bacterial mortality and the frequency of infected cells. Aquat Microb Ecol 18: 207–215.
Bird DF, Karl DM . (1999). Uncoupling of bacteria and phytoplankton during the austral spring bloom in Gerlache Strait, Antarctic Peninsula. Aquat Microb Ecol 19: 13–27.
Bird DF, Maranger R, Karl DM . (1993). Palmer LTER: aquatic virus abundances near the Antarctic Peninsula. Antarct J US 28: 234–235.
Boras JA, Sala MM, Vazquez-Dominguez E, Weinbauer MG, Vaque D . (2009). Annual changes of bacterial mortality due to viruses and protists in an oligotrophic coastal environment (NW Mediterranean). Environ Microbiol 11: 1181–1193.
Breitbart M . (2012). Marine viruses: truth or dare. Ann Rev Mar Sci 4: 425–448.
Brum JR, Ignacio-Espinoza JC, Roux S, Doulcier G, Acinas SG, Alberti A et al. (2015). Patterns and ecological drivers of ocean viral communities. Science 348: 1261498.
Brum JR, Morris JJ, Decima M, Stukel MR . (2014) Mortality in the oceans: causes and consequences. In: Kemp PF (ed) Eco-DAS IX Symposium Proceedings. ASLO: Waco, TX, USA, pp 16–48.
Brum JR, Steward GF, Jiang SC, Jellison R . (2005). Spatial and temporal variability of prokaryotes, viruses, and viral infections of prokaryotes in an alkaline, hypersaline lake. Aquat Microb Ecol 41: 247–260.
Brum JR, Sullivan MB . (2015). Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat Rev Microbiol 13: 147–159.
Brussaard CPD, Timmermans KR, Uitz J, Veldhuis MJW . (2008). Virioplankton dynamics and virally induced phytoplankton lysis versus microzooplankton grazing southeast of the Kerguelen (Southern Ocean). Deep Sea Res Part 2 Top Stud Oceanogr 55: 752–765.
Chen Y, Golding I, Sawai S, Guo L, Cox EC . (2005). Population fitness and the regulation of Escherichia coli genes by bacterial viruses. PLoS Biol 3: e229.
Cottrell MT, Kirchman DL . (2012). Virus genes in Arctic marine bacteria identified by metagenomic analysis. Aquat Microb Ecol 66: 107–116.
Doney SC, Ruckelshaus M, Emmett Duffy J, Barry JP, Chan F, English CA et al. (2012). Climate change impacts on marine ecosystems. Ann Rev Mar Sci 4: 11–37.
Duarte CM, Agusti S, Vaque D, Agawin NSR, Felipe J, Casamayor EO et al. (2005). Experimental test of bacteria-phytoplankton coupling in the Southern Ocean. Limnol Oceanogr 50: 1844–1854.
Ducklow H, Carlson C, Church M, Kirchman D, Smith D, Steward G . (2001). The seasonal development of the bacterioplankton bloom in the Ross Sea, Antarctica, 1994-1997. Deep Sea Res Part 2 Top Stud Oceanogr 48: 4199–4221.
Ducklow H, Clarke A, Dickhut R, Doney SC, Geisz H, Huang K et al. (2012a) The marine system of the western Antarctic Peninsula. In: Rogers AD, Johnston NM, Murphy EJ, Clarke A (ed) Antarctic Ecosystems: An Extreme Environment in a Changing World. Blackwell Publishing Ltd.: Hoboken, NJ, USA, pp 121–159.
Ducklow HW . (2008). Long-term studies of the marine ecosystem along the west Antarctic Peninsula. Deep Sea Res Part 2 Top Stud Oceanogr 55: 1945–1948.
Ducklow HW, Baker K, Martinson DG, Quetin LB, Ross RM, Smith RC et al. (2007). Marine pelagic ecosystems: the West Antarctic Peninsula. Phil Trans R Soc B 362: 67–94.
Ducklow HW, Dickson M-L, Kirchman DL, Steward G, Orchardo J, Marra J et al. (2000). Constraining bacterial production, conversion efficiency and respiration in the Ross Sea, Antarctica, January–February, 1997. Deep Sea Res Part 2 Top Stud Oceanogr 47: 3227–3247.
Ducklow HW, Schofield O, Vernet M, Stammerjohn S, Erickson M . (2012b). Multiscale control of bacterial production by phytoplankton dynamics and sea ice along the western Antarctic Peninsula: a regional and decadal investigation. J Mar Syst 98-99: 26–39.
Duhaime MB, Sullivan MB . (2012). Ocean viruses: rigorously evaluating the metagenomic sample-to-sequence pipeline. Virology 434: 181–186.
Duhaime MBD, Deng L, Poulos BT, Sullivan MB . (2012). Towards quantitative metagenomics of wild viruses and other ultra-low concentration DNA samples: a rigorous assessment and optimization of the linker amplification method. Environ Microbiol 14: 2526–2537.
Evans C, Brussaard CPD . (2012). Regional variation in lytic and lysogenic viral infection in the Southern Ocean and its contribution to biogeochemical cycling. Appl Environ Microbiol 78: 6741–6748.
Evans C, Pearce I, Brussaard CPD . (2009). Viral-mediated lysis of microbes and carbon release in the sub-Antarctic and Polar Frontal zones of the Australian Southern Ocean. Environ Microbiol 11: 2924–2934.
Fuhrman J . (2000) Impact of viruses on bacterial processes. In: Kirchman DL (ed) Microbial Ecology of the Oceans. Wiley-Liss, Inc.: New York, NY, USA, pp 327–350.
Fuhrman JA . (1999). Marine viruses and their biogeochemical and ecological effects. Nature 399: 541–548.
Ghiglione JF, Murray AE . (2012). Pronounced summer to winter differences and higher wintertime richness in coastal Antarctic marine bacterioplankton. Environ Microbiol 14: 617–629.
Grzymski JJ, Riesenfeld CS, Williams TJ, Dussaq AM, Ducklow H, Erickson M et al. (2012). A metagenomic assessment of winter and summer bacterioplankton from Antarctica Peninsula coastal surface waters. ISME J 6: 2901–2915.
Guixa-Boixereu N, Vaque D, Gasol JM, Sanchez-Camara J, Pedro-Alio C . (2002). Viral distribution and activity in Antarctic waters. Deep Sea Res Part 2 Top Stud Oceanogr 49: 827–845.
Hewson I, Fuhrman JA . (2007). Characterization of lysogens in bacterioplankton assemblages of the Southern California Borderland. Microb Ecol 53: 631–638.
Holm-Hansen O, Lorenzen CJ, Holmes RW, Strickland JDH . (1965). Fluorometric determination of chlorophyll. J Cons Int Explor Mer 30: 3–15.
Hurwitz B, Westvald A, Brum J, Sullivan M . (2014). Modeling ecological drivers in marine viral communities using comparative metagenomics and network analyses. Proc Natl Acad Sci USA 111: 10714–10719.
Hurwitz BL, Deng L, Poulos BT, Sullivan MB . (2013). Evaluation of methods to concentrate and purify ocean virus communities through comparative, replicated metagenomics. Environ Microbiol 15: 1428–1440.
Hurwitz BL, Sullivan MB . (2013). The Pacific ocean virome (POV): a marine viral metagenomic dataset and associated protein clusters for quantitative viral ecology. PLoS ONE 8: e57355.
John SG, Mendez CB, Deng L, Poulos B, Kauffman AKM, Kern S et al. (2011). A simple and efficient method for concentration of ocean viruses by chemical flocculation. Environ Microbiol Rep 3: 195–202.
Karl DM, Holm-Hansen O, Taylor GT, Tien G, Bird DF . (1991). Microbial biomass and productivity in the western Bransfield Strait, Antarctica during the 1986-87 austral summer. Deep Sea Res A 38: 1029–1055.
Kirchman D . (1993) Leucine incorporation as a measure of biomass production by heterotrophic bacteria. In: Kemp P (ed) Handbook of Methods in Aquatic Microbial Ecology. Lewis Publishers: Boca Raton, FL, USA, pp 509–512.
Kirchman D, K'nees E, Hodson R . (1985). Leucine incorporation and its potential as a measure of protein synthesis by bacteria in natural aquatic systems. Appl Environ Microbiol 49: 599–607.
Kirchman DL, Moran XAG, Ducklow H . (2009). Microbial growth in the polar oceans - role of temperature and potential impact of climate change. Nat Rev Microbiol 7: 451–459.
Kunin V, Hugenholtz P . (2010). PyroTagger: a fast, accurate pipeline for analysis of rRNA amplicon pyrosequence data. Open J article 1: 1–8.
Lane DJ . (1991) 16S/23S rRNA Sequencing. In: Stackenbrandt E, Goodfellow M (ed) Nucleic Acid Techniques in Bacterial Systematics. John Wiley & Sons: Chichester, NY, USA, pp 115–175.
Laybourn-Parry J, Marshall WA, Madan NJ . (2007). Viral dynamics and patterns of lysogeny in saline Antarctic lakes. Polar Biol 30: 351–358.
Lochte K, Bjornsen PK, Giesenhagen H . (1997). Bacterial standing stock and production and their relation to phytoplankton in the Southern Ocean. Deep Sea Res Part 2 Top Stud Oceanogr 44: 321–340.
Malits A, Christaki U, Obernosterer I, Weinbauer MG . (2014). Enhanced viral production and virus-mediated mortality of bacterioplankton in a natural iron-fertilized bloom event above the Kerguelen Plateau. Biogeosciences 11: 6841–6853.
Marchant H, Davidson A, Wright S, Glazebrook J . (2000). The distribution and abundance of viruses in the Southern Ocean during spring. Antarct Sci 12: 414–417.
Marine R, McCarren C, Vorrasane V, Nasko D, Crowgey E, Polson S et al. (2014). Caught in the middle with multiple displacement amplification: the myth of pooling for avoiding multiple displacement amplification bias in a metagenome. Microbiome 2: 3.
McCalley CK, Woodcroft BJ, Hodgkins SB, Wehr RA, Kim E-H, Mondav R et al. (2014). Methane dynamics regulated by microbial community response to permafrost thaw. Nature 514: 478–481.
Miller RV, Day MJ . (2008) Contribution of lysogeny, pseudolysogeny, and starvation to phage ecology. In: Abedon ST (ed) Bacteriophage Ecology: Population Growth, Evolution, and Impact of Bacterial Viruses. Cambridge University Press: Cambridge, UK, pp 114–143.
Mondav R, Woodcroft BJ, Kim E-H, McCalley CK, Hodgkins SB, Crill PM et al. (2014). Discovery of a novel methanogen prevalent in thawing permafrost. Nat Commun 5: 3212.
Moran XAG, Estrada M . (2002). Phytoplanktonic DOC and POC production in the Bransfield and Gerlache Straits as derived from kinetic experiments of 14C incorporation. Deep Sea Res Part 2 Top Stud Oceanogr 49: 769–786.
Moran XAG, Gasol JM, Pedros-Alio C, Estrada M . (2001). Dissolved and particulate primary production and bacterial production in offshore Antarctic waters during austral summer: coupled or uncoupled? Mar Ecol Prog Ser 222: 25–39.
Noble RT, Fuhrman JA . (1998). Use of SYBR green I for rapid epifluorescence counts of marine viruses and bacteria. Aquat Microb Ecol 14: 113–118.
Paterson H, Laybourn-Parry J . (2012). Antarctic sea ice viral dynamics over an annual cycle. Polar Biol 35: 491–497.
Paul JH . (2008). Prophages in marine bacteria: dangerous molecular time bombs or the key to survival in the seas? ISME J 2: 579–589.
Paul JH, Weinbauer M . (2010) Detection of lysogeny in marine environments. In: Wilhelm SW, Weinbauer MG, Suttle CA (ed) Manual of Aquatic Viral Ecology. ASLO: Waco, TX, USA, pp 30–33.
Payet JP, Suttle CA . (2013). To kill or not to kill: the balance between lytic and lysogenic viral infections is driven by trophic status. Limnol Oceanogr 58: 465–474.
Pearce I, Davidson AT, Bell EM, Wright S . (2007). Seasonal changes in the concentration and metabolic activity of bacteria and viruses at an Antarctic coastal site. Aquat Microb Ecol 47: 11–23.
Pomeroy LR, Wiebe WJ . (2001). Temperature and substrates as interactive limiting factors for marine heterotrophic bacteria. Aquat Microb Ecol 23: 187–204.
Proctor LM, Fuhrman JA . (1990). Viral mortality of marine bacteria and cyanobacteria. Nature 343: 60–62.
Rattei T, Arnold R, Tischler P, Lindner D, Stumpflen V, Mewes HW . (2006). SIMAP: the similarity matrix of proteins. Nucleic Acids Res 34: D252–D256.
Rich VI, Konstantinidis K, DeLong EF . (2008). Design and testing of 'genome-proxy' microarrays to profile marine microbial communities. Environ Microbiol 10: 506–521.
Saba GK, Fraser WR, Saba VS, Iannuzzi RA, Coleman KE, Doney SC et al. (2014). Winter and spring controls on the summer food web of the coastal West Antarctic Peninsula. Nat Commun 5: 4318.
Sailley SF, Ducklow HW, Moeller HV, Fraser WR, Schofield OM, Steinberg DK et al. (2013). Carbon fluxes and pelagic ecosystem dynamics near two western Antarctic Peninsula Adelie pengiun colonies: an inverse model approach. Mar Ecol Prog Ser 492: 253–272.
Short SM, Suttle CA . (2002). Sequence analysis of marine virus communities reveals that groups of related algal viruses are widely distribution in nature. Appl Environ Microbiol 68: 1290–1296.
Smith D, Azam F . (1992). A simple, economical method for measuring bacterial protein synthesis rates in seawater using 3H-leucine. Mar Microb Food Webs 6: 107–114.
Solonenko SA, Sullivan MB . (2013) Preparation of metagenomic libraries from naturally occurring marine viruses. In: DeLong EF (ed) Methods in Enzymology, Vol 531. Academic Press: Waltham, MA, USA, pp 143–165.
Susskind MM, Botstein D . (1978). Molecular genetics of bacteriophage P22. Microbiol Rev 42: 385–413.
Suttle CA . (2007). Marine viruses—major players in the global ecosystem. Nat Rev Microbiol 5: 801–812.
Takahashi T, Sutherland SC, Sweeney C, Poisson A, Metzl N, Tilbrook B et al. (2002). Global sea-air CO2 flux based on climatological surface ocean pCO2, and seasonal biological and temperature effects. Deep Sea Res Part 2 Top Stud Oceanogr 49: 1601–1622.
Thomas R, Berdjeb L, Sime-Ngando T, Jacquet S . (2011). Viral abundance, production, decay rates and life strategies (lysogeny versus lysis) in Lake Bourget (France). Environ Microbiol 13: 616–630.
Weinbauer MG . (2004). Ecology of prokaryotic viruses. FEMS Microbiol Rev 28: 127–181.
Williams TJ, Long E, Evans F, DeMaere MZ, Lauro FM, Raftery MJ et al. (2012). A metaproteomic assessment of winter and summer bacterioplankton from Antarctic Peninsula coastal surface waters. ISME J 6: 1883–1900.
Williamson SJ, Houchin LA, McDaniel L, Paul JH . (2002). Seasonal variation in lysogeny as depicted by prophage induction in Tampa Bay, Florida. Appl Environ Microbiol 68: 4307–4314.
Yang Y, Motegi C, Yokokawa T, Nagata T . (2010). Large-scale distribution patterns of virioplankton in the upper ocean. Aquat Microb Ecol 60: 233–246.
Yilmaz S, Allgaier M, Hugenholtz P . (2010). Multiple displacement amplification compromises quantitative analysis of metagenomes. Nat Methods 7: 943–944.
Yu Z-C, Chen X-L, Shen Q-T, Zhao D-L, Tang B-L, Su H-N et al. (2015). Filamentous phages prevalent in Pseudoalteromonas spp. confer properties advantageous to host survival in Arctic sea ice. ISME J 9: 871–881.
Acknowledgements
This research was supported by the National Science Foundation through a Postdoctoral Fellowship in Polar Regions Research to JRB (award #1019328) and the Palmer LTER project (award #0823101) through the Antarctic Organisms and Ecosystems Program in the Antarctic Sciences Division, and is funded in part by the Gordon and Betty Moore Foundation through grants GBMF2631 and GBMF3790 to MBS. Sequencing was supported through BIO5 funding to MBS. We gratefully acknowledge an allocation of computer time from the UA Research Computing High Performance Computing (HPC) and High Throughput Computing (HPC) at the University of Arizona. We thank A Culley and C Schvarcz for assistance in collecting virus samples; A Alpert and E Woznica for their collection and processing of bacterial production samples; K Coleman, M Garzio, T Miles and C Funkey for assistance with sample collection; the employees of Raytheon Polar Services Company for their support; A Gregory and B Poulos for assistance in conducting linker amplification; B Poulos and V Rich for assistance with bacterial DNA extraction; A Westveld for assistance with social network analysis; and members of the Tucson Marine Phage Lab, E Allers and V Rich for their support and critical review of this research. All metagenomic reads were deposited to iPlant (www.iplantcollaborative.org) under the project name imicrobe/southern_ocean_viromes. Ecological data from the Palmer, Antarctica LTER project (chlorophyll a concentration and bacterial production) are available at: http://oceaninformatics.ucsd.edu/datazoo/data/pallter/datasets.
Author contributions
JRB performed the project planning; JRB performed the experimental work; JRB and BLH performed the data analysis; JRB, BLH, OS, HWD and MBS wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Brum, J., Hurwitz, B., Schofield, O. et al. Seasonal time bombs: dominant temperate viruses affect Southern Ocean microbial dynamics. ISME J 10, 437–449 (2016). https://doi.org/10.1038/ismej.2015.125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.125
This article is cited by
-
The infant gut virome is associated with preschool asthma risk independently of bacteria
Nature Medicine (2024)
-
Presence and role of viruses in anaerobic digestion of food waste under environmental variability
Microbiome (2023)
-
Lysogenic bacteriophages encoding arsenic resistance determinants promote bacterial community adaptation to arsenic toxicity
The ISME Journal (2023)
-
Zea mays genotype influences microbial and viral rhizobiome community structure
ISME Communications (2023)
-
Biogeochemical sulfur cycling of virus auxiliary metabolic genes involved in Napahai plateau wetland
Environmental Science and Pollution Research (2023)