Abstract
Marine phytoplankton can evolve rapidly when confronted with aspects of climate change because of their large population sizes and fast generation times. Despite this, the importance of environment fluctuations, a key feature of climate change, has received little attention—selection experiments with marine phytoplankton are usually carried out in stable environments and use single or few representatives of a species, genus or functional group. Here we investigate whether and by how much environmental fluctuations contribute to changes in ecologically important phytoplankton traits such as C:N ratios and cell size, and test the variability of changes in these traits within the globally distributed species Ostreococcus. We have evolved 16 physiologically distinct lineages of Ostreococcus at stable high CO2 (1031±87 μatm CO2, SH) and fluctuating high CO2 (1012±244 μatm CO2, FH) for 400 generations. We find that although both fluctuation and high CO2 drive evolution, FH-evolved lineages are smaller, have reduced C:N ratios and respond more strongly to further increases in CO2 than do SH-evolved lineages. This indicates that environmental fluctuations are an important factor to consider when predicting how the characteristics of future phytoplankton populations will have an impact on biogeochemical cycles and higher trophic levels in marine food webs.
Similar content being viewed by others
Introduction
The function of marine ecosystems hinges on their smallest primary producers. Marine phytoplankton are responsible for roughly 50% of global primary production (Falkowski et al., 1998), are major drivers of the carbon pump (Beardall and Raven, 2004) and form the base of marine food webs (Rossoll et al., 2012; Thomsen et al., 2013). Marine phytoplankton with large population sizes and short generation times will have ample scope for evolution in a changing ocean (Collins, 2011), where CO2 levels are predicted to have doubled by the end of the century (IPCC report, 2007) and to become less stable (Egleston et al., 2010; Flynn et al., 2012). Thus, phytoplankton are and will be evolving in unstable CO2 environments. Natural fluctuations in CO2 can range from diurnal changes to annual fluctuations and changes on geological timescales, for example, due to microbial activity, severe weather and seasonal vertical mixing (Joint et al., 2011; Gilbert et al., 2008; Li et al., 2009). We have previously shown that average annual fluctuations in CO2 at the sampling location affect the growth responses of marine picoplankton to short-term increases in CO2 (Schaum et al., 2013).
As buffering capacity decreases at lower pH values, any changes in dissolved CO2 are expected to become more pronounced and less predictable in the future (Egleston et al., 2010; Flynn et al., 2012), making it necessary for phytoplankton to not only evolve mechanisms to deal with CO2 levels that are on average higher than today’s but also to cope with larger and more frequent deviations from that average. It has not been tested empirically whether fluctuations modulate responses to further environmental changes in marine phytoplankton, although theory and experiments on other model organisms such as Escherichia coli suggests that it should (Ghalambor et al., 2007; Draghi and Whitlock, 2012; Chevin et al., 2013; Karve et al., 2014). This is particularly important, as even under optimistic scenarios, CO2 levels in the ocean are going to increase long after the 2100 cutoff applied to most climate models (IPCC report, 2007). Yet, despite the likely influence of environmental variability on evolution, selection experiments that aim to characterize phenotypes of marine phytoplankton in a changing ocean are usually carried out in stable environments (for example, Lohbeck et al., 2012; Crawfurd et al., 2011; Tatters et al., 2013; Schlüter et al., 2014).
Most previous studies have used few representatives of a species (or, where species are cryptic, species complex), with no test of whether genotypically or physiologically distinct lineages of the same species differ in the phenotypes they evolve in response to elevated CO2 levels. However, this information is crucial to make predictions about how structures of phytoplankton communities and their function in the ecosystem will be affected by increased CO2 levels or climate change in a broader sense. We need two main pieces of information: who will be present in future phytoplankton communities and what they will be doing. The evolution of lineage growth rates in response to environmental change informs our predictions of how frequencies of lineages within a population are expected to change (Schaum and Collins, 2014), and thus who is likely to be there. The evolution of other aspects of the phenotype, such as size and cellular stoichiometry, allow us to understand what they will be doing and how this will affect higher trophic levels and nutrient cycling. Often, phenotypic characteristics of marine microbial communities to climate change are examined through observation and short-term experiments; however, recent years have seen an increase in studies showing that evolutionary processes need to be considered when predicting effects of climate change on marine microbial communities and the expected phenotypes (see Collins et al., 2013 for a review). In stable environments, these include decreased cell size and a partial restoration of the calcification process in the coccolithophore Emiliania huxleyi (Lohbeck et al., 2012; Schlüter et al., 2014) and lowered C:N ratios in diatoms (Crawfurd et al., 2011). However, it is not known how environmental fluctuations affect phenotypic evolution.
The goal of this study is to measure whether evolving in fluctuating high CO2 (1012±244 μatm CO2, FH) versus stable high CO2 (1031±87 μatm CO2, SH) results in differences in the evolution of key characteristics of a marine phytoplankton. Using 16 genetically distinct lineages of the species complex Ostreococcus (Courties et al., 1994; Subirana et al., 2013) from different habitat types (for example, lineages from coasts and the open ocean, surface waters and near the deep chlorophyll maximum), we can estimate whether trait evolution in the face of elevated CO2 is conserved within a species or genus (Schaum et al., 2013; Zhang et al., 2014). As part of the phenotypic characterization, we investigate how responsive lineages evolved in the SH and FH CO2 environments are to further increases in CO2, that is, how well they are able to adjust their growth rates in response to further CO2 enrichment. The rationale for this is that the ability to adjust growth rates in response to environmental fluctuations is likely an important trait in marine picoplankton when marine environments become more variable. It is not feasible to measure all possible phenotypic traits; hence, here we focus on key traits that determine the role of phytoplankton in biogeochemical cycles, food webs or both. For example, the ratio of oxygen evolution and consumption, combined with size and C:N ratios, data can inform future studies or models on carbon export to the deeper ocean. In addition, changes in size and elemental composition can affect the quality of phytoplankton as a food source for grazers so that changes in the elemental composition of phytoplankton can affect grazer population size and life history (Rossoll et al., 2012; Thomsen et al., 2013).
Materials and methods
Cultures and experimental setup
We used 16 lineages of Ostreococcus from 9 habitat types (see Schaum et al., 2013 and Supplementary Table S1). Lineages were made clonal by dilution at the beginning of the experiment (Schaum and Collins, 2014). Three biological replicates of each lineage were grown in each of the four CO2 selection environments: stable ambient (SA 444±43 μatm CO2), fluctuating ambient (FA 490±97 μatm CO2), SH (1031±87 μatm CO2) and FH (1012±244 μatm CO2). This is a total of 192 populations in the experiment (16 lineages × 3 biological replicates × 4 selection environments; see Figure 1 for experimental setup). Temperature (18 °C), salinity (31) and light intensity (160 μE) were kept stable, while pCO2 was varied. In the fluctuating treatments, pCO2 was set to change every 7 days (in the SA environment, Ostreococcus divide about once a day), to a random value between 430 and 630 μatm CO2 for FA, and 700 and 1300 μatm CO2 for FH environments. Samples were maintained in a semi-continuous batch culture with a transfer approximately every seven generations (see also Schaum and Collins, 2014).
Experimental setup: 3 biological replicates of 16 lineages of the species complex Ostreococcus were evolved in 4 selection environments. These were SA (444±43 μatm CO2), FA (490±97 μatm CO2), SH (1031±87 μatm CO2) and FH (1012±244 μatm CO2). At the beginning of the experiment (t0), as well as at 100 (t100) and 400 (t400) generations into the experiment, phenotypes of all samples were assayed at both 430 and 1000 μatm CO2. We refer to SA- and FA-evolved phenotypes measured at 430 and 1000 μatm CO2 as ambient and plastic responses, respectively. We refer to phenotypes of FH- and SH-evolved lineages measured in the 1000 μatm CO2 assay as evolved responses. FH- and SH-evolved lineages assayed at 430 μatm CO2 are referred to as correlated responses (see Supplementary Figure S1). Shortly after t400, we also assayed all samples at 2000 μatm CO2 and call this a response to further CO2 increase. We discuss all but the correlated responses in detail in the main text. For correlated responses, see Supplementary Information.
Before evolution in any of the selection environments (t0) and also following 100 (t100) and 400 (t400) generations of evolution in the selection environments, populations were grown at 430 and 1000 μatm CO2 for 7–14 generations and then assayed for the investigated phenotypic traits (see Supplementary Figure S1B for FH and SH at 430 μatm CO2). At t600, samples from all selection environments were assayed for growth rates and other phenotypic traits at 2000 μatm CO2 (see Figure 1). To account for the additional generations passed, growth rates, size and chlorophyll content were also re-measured for all populations evolved in SA, FA, SH and FH.
As this study focuses on trait evolution, we report three response types for lineages after they have evolved in their respective selection environments: ambient, plastic and evolved responses. Here the ambient responses are phenotypic traits of lineages evolved at SA or FA that have been measured at 430 μatm CO2. The respective trait values function as a reference to which the plastic and evolved responses are compared. Plastic responses are changes in phenotype that do not require any underlying change in genotype (West-Eberhard, 2003). Plastic responses here are the short-term responses to elevated CO2 levels. To measure the phenotype of the plastic response, lineages evolved at SA or FA are measured at 1000 μatm CO2. These lineages have not evolved in high-CO2 environments; hence, their phenotypes at high CO2 are a measure of the phenotypic change that occurs in the absence of genetic evolution in response to long-term growth in a high-CO2 environment. It is noteworthy that this is sometimes also called an acclimation response in other studies. Here we reserve the word ‘acclimation’ for the period where we allowed populations to acclimate to the assay environment before measuring traits. As our lineages were founded from clonal populations, we argue that any change in average population phenotype observed within 14 generations is unlikely to have arisen through de novo mutation or genotype sorting. We also show that the response is reversible by transferring the samples back to 430 μatm after exposure to elevated CO2 (Supplementary Figure S1A). Likewise, the response of SA-, FA-, SH- and FH-evolved lineages to 2000 μatm CO2 is a plastic response. Evolved responses are changes in phenotype that are due to changes in the genotypic makeup of the population over time. Evolved traits are usually described as changes in phenotype of the evolved populations growing in their selective environment (here, FH or SH growing in high CO2) relative to the phenotype of an ancestral or control population grown in the same environment (here, SA and FA evolved in ambient CO2 environments but were measured in high CO2). We refer to phenotypic traits measured in SH- and FH-evolved populations at 1000 μatm CO2 as evolved responses and describe the correlated responses (SH and FH at 430 μatm CO2) in the Supplementary Information (Supplementary Figure S1B).
The SA environment controlled for any changes in phenotype that might occur due to evolution in a lab environment. The FA environment selected for the maintenance and/or evolution of plasticity (that is, the ability to produce a plastic response) to fluctuations in pCO2 alone. The SH environment selected for growth at high CO2 alone, whereas the FH environment selected for the maintenance and/or evolution of plastic responses due to high as well as fluctuating CO2 environment. For detailed carbonate chemistry tables for the different selection environments, please refer to Supplementary Table S2.
Physiological responses
Samples were acclimated to the appropriate CO2 level as described above before the physiological assays. CO2 levels of 430, 1000 or 2000 μatm CO2 were established through bubbling artificial seawater in the respective CO2 incubator for at least 24 h before any cultures were added. Seawater DIC (dissolved inorganic carbon), alkalinity and pH were measured regularly as described in the Supplementary Information.
Growth rates, oxygen evolution and consumption rates
Growth rates were calculated from the intact, chlorophyll a-positive event counts within an Ostreococcus-specific gate of forward and side scatter on a FACS CANTO (BD Biosciences, Oxford, UK; see Supplementary Figure S2). Oxygen evolution and consumption rates were measured using a Clark-type oxygen electrode under conditions resembling those in the respective incubators (see Supplementary Methods for details).
Cellular stoichiometry
Particular organic carbon and nitrogen samples were taken for eight lineages per selection environment, using three technical replicates of each of three biological replicates. Samples were filtered onto pre-combusted (12 h, 500 °C) GF/F filters (1.2 mm; Whatman, GE Healthcare Life Sciences, Cardiff, UK) and stored in pre-combusted (500 °C) petri dishes. Filters were wrapped in tin foil and particular organic carbon and nitrogen were then measured using an Elementar Vario EL mass spectrometer (Elementar Analysensysteme GmbH, Hanau, Germany).
Cell size, chlorophyll a content and Nile Red fluorescence
Chlorophyll a content, lipid content and relative cell size were determined using a FACS CANTO (for general protocols, see (Marie et al., 2005; Petersen et al., 2012)). The red fluorescence channel was used as a proxy for total chlorophyll a content and was cross-calibrated using chlorophyll a calibration beads as standards (Bangs Laboratories, Inc., Fishers, IN, USA), as well as chlorophyll a extraction (Holm-Hansen and Riemann, 1978). The same channel was used to determine polar lipids after a Nile Red stain (la Jara de et al., 2003; Guzmán et al., 2009). Cell size was inferred from the FSC (forward scatter) and comparison with calibration beads and coulter counter measurements. Owing to the nature of flow cytometry measurements, trait values retrieved from FACS CANTO measurements are approximate (Marie et al., 2005; Petersen et al., 2012), but relative changes between samples are highly informative for comparisons of strains and selection lines.
Statistics and graphical representation
Data were analysed in the R environment, using mixed models within the ‘nlme’ and ‘lme4’ packages and models built based on (Pinheiro and Bates, 2000), to control for temporal effects of CO2 in assay and selection environments, and to account for the same clonal lineages being used across treatments. Owing to the multiple comparisons performed in the omnibus analysis of variance (ANOVA) detecting changes between fluctuating and stable environments across all traits, selection regimes and lineages, P-values were adjusted through a Holm–Bonferroni approach before Tukey’s honest significant difference post-hoc tests performed only on P-values <0.05, that is, all post-hoc results presented throughout are already corrected for multiple comparisons. Smaller ANOVA analyses were performed to gain more detailed information, for example, on the effect of environmental stability on specific traits or groups of lineages. The number of biological replicates per lineages was three. For graphic representation, means and s.e. per lineage, as well as per CO2 treatment, were calculated within the ‘ggplot2’ and ‘plotrix’ package. See Supplementary Information for details.
Results and Discussion
Fluctuating and stable CO2 select for drastically different phenotypes
The plastic responses of the marine picoplankton Ostreococcus to elevated CO2 before the evolution experiment have been described in Schaum et al. (2013), and in line with this we find elevated oxygen evolution and respiration rates, as well as an increase in cell size and C:N ratio in lineages exposed to 1000 μatm CO2 at t0 (before evolution) and in SA- and FA-evolved populations (at t400) assayed at 1000 μatm CO2. Notably, phenotypes characteristic of the plastic response to elevated CO2 are not maintained when populations are allowed enough time to evolve in high-CO2 environments (Figure 2, ANOVA all traits, selection environment × assay environment F3,124=9.69, P<0.05) and begin to reverse after 75–100 generations in the high-CO2 environments (Supplementary Figure S3). This partial reversal of the plastic response in lineages evolving in high-CO2 environments is extremely likely to be due to evolution. This is because first, it occurs while populations are still growing in the high-CO2 environment and, second, it is considerably slower than the reversal of the plastic response seen when we transfer SA or FA populations from 1000 μatm CO2 back to 430 μatm CO2. There, the SA- or FA-evolved phenotype is re-established with 7–14 generations (Supplementary Figure S1A).
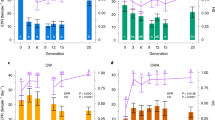
Ambient (left), plastic (middle) and evolved responses (right) to stable (grey) and fluctuating (white) pCO2 of phenotypic traits in Ostreococcus. Selection in SH and FH environments yields different phenotypes, which also differ from plastic response. As a guideline, the dashed line represents mean values of each trait for SA lineages assayed at 430 μatm CO2. Bars are means ±1 s.e. Post-hoc results are summarized in Supplementary Table S3. In the stable environment (grey), high CO2-evolved phenotypes differ from the ambient response with regards to oxygen evolution, oxygen consumption, R:P ratio, C:N ratio and lipid content. The plastic response of SA-evolved lineages differs from the evolved response (SH-evolved lineages) for oxygen evolution, oxygen consumption, R:P ratio, C:N ratio and lipid content. For lineages evolved in a fluctuating environment, there are significant differences between FA- and FH-evolved lineages in oxygen evolution, C:N ratio, chlorophyll a content, size and polar lipid content. The plastic response of FA lineages is different from the evolved response (that is, FH lineages) for C:N ratio, chlorophyll a content, cell size and polar lipid content. Phenotypes of SH lineages are different from those measured in FH lineages for oxygen consumption, C:N ratio chlorophyll a content and size. After evolution in SH or FH, lineages from the stable environment are characterized by lower oxygen consumption rates, higher C:N ratios, higher chlorophyll content per cell volume and larger cells than lineages that had evolved in a fluctuating environment.
In the long-term, elevated CO2 and fluctuations in CO2 both drive evolution of phenotypes in marine phytoplankton and explain ~30% and ~60% of the variation in the evolved phenotypes, respectively (Figure 3). Lineages evolved in FH are different from those evolved in SH (ANOVA all traits, selection CO2 (sco) × environmental fluctuation (ef), F1,118=11.34, P<0.01, Figure 2 and also Supplementary Table S3). For example, FH-evolved lineages respire more than SH-evolved lineages do (post-hoc P<0.05), have lower C:N ratios (post-hoc P<0.01) and are smaller with less chlorophyll a per cell than SH-evolved lineages (both post-hoc P<0.01). Oxygen evolution rates, respiration to photosynthesis ratio (R:P ratio) and lipid content are not significantly different overall, but there are significant differences on the level of the individual lineages for these traits (ANOVA, sco × ef, F15,109=77.43, P<0.05, Supplementary Figure S4). Overall, we cannot use SH-evolved phenotypes to predict FH-evolved ones (see Supplementary Figure S5). Below, we describe the SH- and FH-evolved phenotypes in detail and discuss the potential implications on marine food webs and carbon cycling.
PCA showing that both environmental stability (PC1) and CO2 elevation (PC2) drive phenotypic evolution of lineages. Large symbols, lineages sampled from above 60-m depth; small symbols, lineages sampled from below 60-m depth; circles, evolved in high-CO2 environment; squares, evolved in ambient CO2 environment; grey, evolved in fluctuating environment; white, evolved in stable environment. See Supplementary Information for a color-coded version where each lineage is represented by a unique symbol (Supplementary Figure S7). Together, fluctuation and elevation of pCO2 explain ~90% of the variance in evolved phenotypes and lineages (within CO2 treatments) cluster largely according to sampling depth.
Trait evolution and their implications for productivity and higher trophic levels
Oxygen evolution rates are higher than they are at ambient CO2 levels in both SH-evolved and FH-evolved linages (Figure 2), with rates of 9.07±0.14 and 9.6±1.3 fmol O2 per cell per hour, respectively. The average oxygen consumption rate after selection at SH is 3.2±0.2 fmol O2 per cell and 3.6±0.2 fmol O2 per cell per hour in FH. Mean R:P ratios are 0.35±0.2 and 0.38±0.2 for SH and FH, respectively. Even though there is a general decrease in R:P ratios in both SH- and FH-evolved lineages compared with the ancestral phenotype, there are large differences depending on the lineages’ sampling origin (Figure 3 and Supplementary Figure S6, ANOVA RP ratio, sco × ef, F1,122=59.6, P <0.01).
C:N ratios in SH-evolved lineages are at 7.5±0.15 and at 5.3±0.3 in FH-evolved lineages (Figure 2). Strikingly, this means that SH-evolved lineages evolve a 1.34-fold higher C:N ratio than SA-evolved lineages (post-hoc P<0.01), whereas FH-evolved lineages have lower C:N ratios than FA-evolved lineages (post-hoc P<0.01. C:N ratios in FA are 1.2-fold higher than in FH). The decrease in C:N is particularly marked for Ostreococcus isolated in ‘deep-sea’ locations (Supplementary Figure S6, post-hoc P<0.05).
The average size and chlorophyll content of Ostreococcus cells also changes in response to SH and FH. In SH-evolved lineages, Ostreococcus from the surface and mid-depth locations are slightly larger in size with higher chlorophyll a content per cell than ‘deep sea’ ones (post-hoc P <0.05, Figure 3 and Supplementary Figure S6). On average, SH-evolved cells are 2.19±0.14 μm in size and contain 10.9±1.93 fg chlorophyll a (Figure 2). FH-evolved cells are smaller and contain less chlorophyll a than FA-evolved cells (both post-hoc P <0.01, Figure 2), measuring 1.73±0.03 μm and containing 9.49±0.2 fg per cell. After evolution at SH and FH, intracellular polar lipid content measured as Nile Red fluorescence increases in 6 out of 16 lineages, but no change in polar lipid content was found in the other lineages (Figure 2 and Supplementary Figures S4).
Differences in C:N ratio, cell size and lipid content between lineages evolved in stable and FH environments may affect trophic interactions and biogeochemical cycles. This is particularly the case when we also consider that frequencies of lineages within phytoplankton communities are expected to change. We have previously shown that lineages with larger plastic response to elevated CO2 will increase in frequency in the short term and also have a larger evolutionary response in the long term (Schaum and Collins, 2014). Although between-lineage variance in growth rate is larger before evolution (0.36, see Schaum et al., 2013) than after evolution in SH (0.12) and FH (0.13), our findings show that SH- and FH-evolved lineages have evolved differently.
Ostreococcus evolved in high-CO2 environments show changes in growth rate, size and C:N ratio, indicating that future changes to productivity and carbon export by picoplankton are likely. Our data suggest that the magnitude of this change will vary locally, as there was substantial variation among lineages in some evolved traits (measured in this experiment and in Schaum and Collins, 2014), local variation in both CO2 levels and fluctuations in CO2 levels (Egleston et al., 2010), which will affect how natural selection acts.
Our study was done using many lineages of a single genus. However, differences in evolutionary responses to environmental change between species is also likely to drive changes in primary production and carbon export. For example, mesocosm studies suggest that non-calcifying picoplankton, such as Ostreococcus, may become more frequent in future oceans, as they are less adversely affected by elevated CO2 levels than calcifying phytoplankton (Riebesell et al., 2008; Scheinin et al., 2015). Calcifying phytoplankton (for example, coccolithophores) and non-calcyfying phytoplankton (for example, diatoms and green algae) belong to different functional groups (Falkowski et al., 1998) and have different roles in ecosystem functions and services, which means that the relationship between within-group evolution and ecosystem function is likely complex. For example, picoplankton communities of future oceans may be more productive than picoplankton communities in today’s oceans, but the overall productivity and export of the mixed functional group assemblages might still be lower under CO2-enrichment scenarios that favor an increase in frequency in picoplankon. This is because picoplankton are smaller and sink more slowly than larger plankton. However, even small picoplankton do contribute to carbon export after ingestion by planktonic grazers, or when they become heavier through aggregation (Riebesell and Wolf-Gladrow, 1992).
Size, lipid content and C:N ratio are important aspects of food quality (Thomsen et al., 2013; Rossoll et al., 2012). Organisms that graze on Ostreococcus, mainly heterotrophic nanoflagellates (Christaki et al., 1999; Cuevas, 2006), are size selective and may not be physically able to ingest a food source if it doubles in size, or eat enough of a food source if it decreases substantially in size. Here, FH-evolved lineages were smaller than lineages evolved in the SH environment, with little variation between lineages. Several lineages also evolved phenotypes that stored more lipids at high CO2 than at ambient CO2, which may be part of a general response to elevated CO2, causing a shift from oxidative toward reductive pathways (Rokitta et al., 2012). Overall changes in fatty acid composition of marine plankton can lead to lowered reproductive success in zooplankton grazers (Rossoll et al., 2012). If selection imposed in the FH CO2 environment informs us about selection imposed by global change in oceans, grazers will have to feed on smaller, nitrogen-deplete, lipid-rich cells, and thus will have to ingest more cells than under the present day conditions. If the overall phytoplankton biomass in some lineages decreases, grazers and filter feeders may not be able to fulfil their nutritional requirements, or, alternatively, changes in phytoplankton may drive a co-evolutionary response in grazers. As the plastic response of Ostreococcus to elevated CO2 is different from the evolutionary response, grazers, with longer generation times than phytoplankton, may have to cope with an increasingly variable food source. There are a number of evolutionary responses to changes in their food source that could occur in grazers. For example, grazers may use conservative or diversified bet-hedging strategies to minimize fitness variance in the long term. Here the organisms either use the same, low-risk strategy (conservative) or invest in several strategies at once (diversified) (Beaumont et al., 2009; Ripa et al, 2010). In addition, grazers themselves may respond plastically to changes in their food source by switching the feeding mechanisms depending on the food source present in their habitat (Mariani et al., 2013). Finally, it is also possible that grazers will adjust to more variable food sources through evolution and adaptive tracking (that is, mutants arise and fix to fit the food source when it changes), or use a combination of the responses above.
Exposure to elevated fluctuating CO2 levels broadens the scope to react to further CO2 enrichment
A striking difference between lineages evolved in stable versus fluctuating environments in our experiment is how they respond to further increases in pCO2. Most CO2-enrichment experiments for marine phytoplankton use partial pressures of CO2 of around 700–1000 μatm as their high-CO2 environment. As CO2 levels are expected to rise further and can also transiently rise above these levels today (IPCC report, 2007), it is important to investigate how evolution in moderately high-CO2 environments affects the response to short-term exposure to even higher CO2 levels.
Both 1000 and 2000 μatm CO2 are considered to be ‘high-CO2 conditions’, but the ability of lineages to modulate their growth rates depends on the high-CO2 environment they experienced in their previous selection environment (ANOVA growth, response 2000 μatm × selection environment, F3,105=138.72, P <0.0001). Specifically, only lineages evolved in elevated or fluctuating CO2 environments grow differently at 1000 and 2000 μatm CO2, whereas SA-evolved lineages have the same growth rate in both 1000 and 2000 μatm CO2 (post-hoc P=0.67 for SA-evolved lineages, Figure 4). In FA-evolved lineages, which were transiently exposed to higher CO2 levels during evolution, growth rates are on average 1.22-fold higher at 2000 than at 1000 μatm CO2 (post-hoc P <0.001, Figure 4). In SH- and FH-evolved lineages, growth rates at 2000 μatm CO2 were 1.5- and 1.9-fold higher than at 1000 μatm CO2, respectively (post-hoc difference μ SH and FH at 2000 μatm <0.01). Changes in cell size in response to 2000 μatm CO2 were also larger in lineages from the FH environments than in lineages from the SH environment (ANOVA trait values, response 2000 μatm × selection environment, F3,78=2.89, P <0.05), whereas there was no significant difference in chlorophyll levels.
Selection history affects responses to different levels of elevated pCO2. Here, grey boxes represent pooled lineages from stable environments (SA and SH) and white boxes represent pooled lineages from fluctuating environments (FA and FH). Boxplots are displayed with whiskers extending to the highest and the lowest values that are within 1 interquartile range (IQR) of lower/upper quartile. n=3 per lineage. NS, difference between lineages from stable and fluctuating selection regimes not significant; *P<0.05 and **P<0.01, significant difference between lineages from stable or fluctuating environment, respectively. All values are displayed as fold change compared with the response to 1000 μatm at t400. This is the evolved response for FH and SH, and the plastic response for SA and FA. Any values larger than 1 indicate that the response to pCO2 2000 μatm is stronger than the plastic or evolved (for SH and FH) response to pCO2 1000 μatm.
The differences in lineages evolved in stable and fluctuating environments show that evolving or maintaining the ability to distinguish between different high CO2 levels, even those outside the range of CO2 levels encountered in the past few hundred generations of evolution, only requires occasional exposure to high CO2. In our experiment, even the highest CO2 levels in the FA environment were lower than those in either SH or FH, yet responses of FA to 2000 μatm were comparable to those of lineages evolved in the stable and FH CO2 selection regimes. Although our experiment did not test this directly, we suggest that evolving in an environment where fluctuations are random has a role in lineages evolving the ability to specifically modulate their responses to further changes to the environment. Our reasoning is that an environment switching between two set levels of CO2 could select for generalists displaying the same phenotype across environments or a population composed of specialist subpopulations with distinct phenotypes in high- and low-CO2 environments, but no specific evolution of the ability to deal with or distinguish between different high levels of CO2 (Kassen, 2002; Ketola et al., 2013). One possible reason that lineages evolved in fluctuating environments are able to respond by increasing growth at 2000 μatm CO2 is that they are not operating near limits set by oxidative damage (for example, see Dowling and Simmons, 2009) and are thus metabolically better equipped to deal with further perturbations to their environment. This is consistent with previous work showing the eventual evolution of reduced population growth rates but higher heat shock survival and better competitive abilities against novel conspecific competitors in Ostreococcus lineages evolving in high-CO2 environments (Schaum and Collins, 2014).
Regardless of the underlying cause, we show that evolving in fluctuating environments broadens the scope for using plastic responses in the face of continued environmental change (Figure 4). This is not limited to further increases in pCO2, but also extends to reduced CO2 levels: the correlated responses show that FH-evolved lineages grew more rapidly in their ancestral environment (430 μatm CO2) than did the SH-evolved lineages (Supplementary Figure S1A and B and details in Schaum and Collins, 2014).
These findings indicate that both the average environment and the stability of the environment during evolution affect how populations respond to subsequent environmental fluctuations, and that changes in the stability of marine environments will likely have a key role in driving evolutionary outcomes. Taken together, the differences between lineages evolved in stable and fluctuating environments show that selection experiments carried out under stable conditions will fail to detect key changes in phenotype associated with evolving in fluctuating environments, in particular as we cannot use SH-evolved phenotypes to predict FH-evolved ones.
Conclusions
Phenotypic evolution in high-CO2 environments depends on both deviations from the mean CO2 level and the absolute CO2 levels, and this is consistent over several lineages of a species complex. This study shows that fluctuations in CO2, in addition to differences in average CO2, are likely to be important drivers of evolutionary change in green algae as CO2 levels rise. Although the evolutionary response to long-term CO2 enrichment is to partially revert the plastic (short term) response, persistent changes in cellular size and composition remain. This indicates that evolutionary responses to high and fluctuating CO2 can change the role of marine phytoplankton in food webs and nutrient cycles.
References
Beaumont HJE, Gallie J, Kost C, Ferguson G, Rainey PB . (2009). Experimental evolution of bet hedging. Nature 462: 90–93.
Beardall J, Raven JA . (2004). The potential effects of global climate change on microalgal photosynthesis, growth and ecology. Phycologia 43: 26–40.
Chevin LM, Gallet R, Gomulkiewicz R, Holt RD, Fellous S . (2013). Phenotypic plasticity in evolutionary rescue experiments. Proc R Soc B 368: 20120089.
Christaki U, Jacquet S, Dolan JR, Vaulot D, Rassoulzadegan F . (1999). Growth and grazing on Prochlorococcus and Synechococcus by two marine ciliates. Limnol Oceangr 44: 52–61.
Collins S . (2011). Many possible worlds: expanding the ecological scenarios in experimental evolution. Evol Biol 38: 3–14.
Collins S, Rost B, Rynearson TA . (2013). Evolutionary potential of marine phytoplankton under ocean acidification. Evol Appl 7: 140–155.
Courties C, Vaquer A, Troussellier M, Lautier J, Chrétiennot-Dinet MJ, Neveux J et al. (1994). Smallest eukaryotic organism. Nature 370: 255–255.
Crawfurd KJ, Raven JA, Wheeler GL, Baxter EJ, Joint I . (2011). The response of Thalassiosira pseudonana to long-term exposure to increased CO2 and decreased pH. PLoS One 6: e26695.
Cuevas LA . (2006). Nanoheterotroph grazing on bacteria and cyanobacteria in oxic and suboxic waters in coastal upwelling areas off northern Chile. J Plank Res 28: 385–397.
Dowling DK, Simmons LW . (2009). Reactive oxygen species as universal constraints in life- history evolution. Proc R Soc B 276: 1737–1745.
Draghi JA, Whitlock MC . (2012). Phenotypic plasticity facilitates mutational variance, genetic variance, and evolvability along the major axis of environmental variation. Evolution 66: 2891–2902.
Egleston ES, Sabine CL, Morel FMM . (2010). Revelle revisited: buffer factors that quantify the response of ocean chemistry to changes in DIC and alkalinity. Global Biogeochem Cycles 24: GB1002.
Falkowski PG, Barber R, Smetacek V . (1998). Biogeochemical controls and feedbacks on ocean primary production. Science 281: 200–206.
Flynn KJ, Blackford JC, Baird ME, Raven JA, Clark DR, Beardall J et al. (2012). Changes in pH at the exterior surface of plankton with ocean acidification. Nat Clim Change 2: 510–513.
Ghalambor CK, Mckay JK, Carroll SP, Reznick DN . (2007). Adaptive versus non-adaptive phenotypic plasticity and the potential for contemporary adaptation in new environments. Funct Ecol 21: 394–407.
Gilbert JA, Field D, Huang Y, Edwards R, Li W, Gilna P et al. (2008). Detection of large numbers of novel sequences in the metatranscriptomes of complex marine microbial communities. PLoS One 3: e3042.
Guzmán HM, la Jara Valido de A, Duarte LC, Presmanes KF . (2009). Estimate by means of flow cytometry of variation in composition of fatty acids from Tetraselmis suecica in response to culture conditions. Aquacult Int 18: 189–199.
Holm-Hansen O, Riemann B . (1978). Chlorophyll a determination: improvements in methodology. Oikos 30: 438.
IPCC 2007 Intergovernmental Panel on Climate Change. (2007) Working Group I report ‘The physical science basis’. Available at http://www.ipcc.ch/report/ar5/wg1/.
Joint I, Doney SC, Karl DM . (2011). Will ocean acidification affect marine microbes? ISME J 5: 1–7.
la Jara de A, Mendoza H, Martel A, Molina C, Diaz R . (2003). Flow cytometric determination of lipid content in a marine dinoflagellate, Crypthecodinium cohnii. J Appl Physiol 15: 433–438.
Karve S, Daniel S, Chavhan Y, Anand A, Kharola SS, Dey S . (2014) E. coli populations in unpredictably fluctuating environments evolve to face novel stresses through enhanced efflux activity, bioRxiv, doi. http://dx.doi.org/10.1101/011007.
Kassen R . (2002). The experimental evolution of specialists, generalists, and the maintenance of diversity. J Evol Biol 15: 173–190.
Ketola T, Mikonranta L, Zhang J, Saarinen K, Örmälä A-M, Friman V-P et al. (2013). Fluctuating temperature leads to evolution of thermal generalism and preadaptation to novel environments. Evolution 67: 2936–2944.
Li G, Wu Y, Gao KS . (2009). Effects of Typhoon Kaemi on coastal phytoplankton assemblages in the South China Sea, with special reference to the effects of solar UV radiation. J Geophys Res 114: G04029.
Lohbeck KT, Riebesell U, Reusch TBH . (2012). Adaptive evolution of a key phytoplankton species to ocean acidification. Nat Geosci 5: 346–351.
Mariani P, Andersen KH, Visser AW, Barton AD, Kiørboe T . (2013). Control of plankton seasonal succession by adaptive grazing. Limnol Oceanogr 58: 173–184.
Marie D, Simon N, Vaulot D . (2005) Phytoplankton cell counting by flow cytometry. In Algal Culturing Techniques. Vol 1. Elsevier: Oxford, UK. pp 253–267.
Petersen TW, Brent Harrison C, Horner DN, van den Engh G . (2012). Flow cytometric characterization of marine microbes. Methods 57: 350–358.
Pinheiro JC, Bates DM . (2000) Mixed-effects models in S and S-PLUS. Statistics and Computing. Springer: New York. pp 146–174 283–479.
Riebesell U, Wolf-Gladrow DA . (1992). The relationship between physical aggregation of phytoplankton and particle flux: a numerical model. Deep Sea Res A Oceanogr Res Papers 39: 1085–1102.
Riebesell U, Bellerby RGJ, Grossart HP, Thingstad F . (2008). Mesocosm CO2 perturbation studies: from organism to community level. Biogeosciences 5: 1157–1164.
Ripa J, Olofsson H, Jonzen N . (2010). What is bet-hedging, really? Proc R Soc B. (2010) 277: 1153–1154.
Rokitta SD, John U, Rost B . (2012). Ocean acidification affects redox-balance and ion-homeostasis in the life-cycle stages of Emiliania huxleyi. PLoS One 7: e52212.
Rossoll D, Bermúdez R, Hauss H, Schulz KG, Riebesell U, Sommer U et al. (2012) Ocean acidification-induced food quality deterioration constrains trophic transfer PLoS One 7: e34737.
Schaum CE, Collins S . (2014). Plasticity predicts evolution in a marine alga. Proc Biol Sci 281: 20141486–20141486.
Schaum E, Rost B, Millar AJ, Collins S . (2013). Variation in plastic responses of a globally distributed picoplankton species to ocean acidification. Nat Clim Change 3: 298–302.
Scheinin M, Riebesell U, Rynearson T, Lohbeck K, Collins S . (2015). Experimental evolution gone wild. J R Soc Interface 12: 20150056. http://dx.doi.org/10.1098/rsif.2015.0056.
Schlüter L, Lohbeck KT, Gutowska MA, Gröger JP, Riebesell U, Reusch TBH . (2014). Adaptation of a globally important coccolithophore to ocean warming and acidification. Nat Clim Change 4: 1024–1030.
Subirana L, Pequin B, Michely S, Escande M-L, Meilland J, Derelle E et al. (2013). Morphology, genome plasticity, and phylogeny in the genus Ostreococcus. Ann Anat 164: 643–659.
Tatters AO, Schnetzer A, Fu F, Lie AYA, Caron DA, Hutchins DA . (2013). Short- versus long-term responses to changing CO2 in a coastal dinoflagellate bloom: implications for interspecific competitive interactions and community structure. Evolution 67: 1879–1891.
Thomsen J, Casties I, Pansch C, Körtzinger A, Melzner F . (2013). Food availability outweighs ocean acidification effects in juvenile Mytilus edulis: laboratory and field experiments. Glob Change Biol 19: 1017–1027.
West-Eberhard M. J . (2003) Developmental Plasticity and Evolution. Oxford University Press: New York.
Zhang Y, Klapper R, Lohbeck KT, Bach LT, Schulz KG, Reusch TBH et al. (2014). Between- and within-population variations in thermal reaction norms of the coccolithophore Emiliania huxleyi. Limnol Oceangr 59: 1570–1580.
Acknowledgements
The selection experiment and subsequent assays were conducted at the University of Edinburgh (UK). Intracellular stoichiometry was measured at the Alfred Wegener Institute Bremerhaven (Germany). The research was supported by a Royal Society (UK) University Research Fellowship (SC), by the European Research Council (ERC) under the European Community’s Seventh Framework Programme (FP7/2007-2013), ERC grant agreement 205150 (BR) and a Scottish Universities Life Science Alliance scholarship (C-ES). We thank M Allen and ASSEMBLE (Association of European Marine Biology Laboratories), Roscoff, for providing the Ostreococcus lineages; S Rokitta, K-U Richter and U Richter for assistance in the laboratory at the AWI; M Waterfall (UoE) for assistance with flow cytometry; and JA Raven, A Millar, G Yvon-Durocher and A Buckling for providing feedback on the manuscript.
Author Contributions
ES designed and performed the experiments, analysed data and wrote the manuscript. SC designed the experiments, analysed data, wrote the manuscript and supervised the laboratory work. BR supervised the laboratory work at the Alfred Wegener institute and contributed to the writing of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Schaum, CE., Rost, B. & Collins, S. Environmental stability affects phenotypic evolution in a globally distributed marine picoplankton. ISME J 10, 75–84 (2016). https://doi.org/10.1038/ismej.2015.102
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.102
This article is cited by
-
Multivariate trait analysis reveals diatom plasticity constrained to a reduced set of biological axes
ISME Communications (2021)
-
Transient exposure to novel high temperatures reshapes coastal phytoplankton communities
The ISME Journal (2020)
-
Compensation of ocean acidification effects in Arctic phytoplankton assemblages
Nature Climate Change (2018)
-
Microorganisms and ocean global change
Nature Microbiology (2017)