Abstract
Biogeochemical and microbiological data indicate that the anaerobic oxidation of non-methane hydrocarbons by sulfate-reducing bacteria (SRB) has an important role in carbon and sulfur cycling at marine seeps. Yet, little is known about the bacterial hydrocarbon degraders active in situ. Here, we provide the link between previous biogeochemical measurements and the cultivation of degraders by direct identification of SRB responsible for butane and dodecane degradation in complex on-site microbiota. Two contrasting seep sediments from Mediterranean Amon mud volcano and Guaymas Basin (Gulf of California) were incubated with 13C-labeled butane or dodecane under sulfate-reducing conditions and analyzed via complementary stable isotope probing (SIP) techniques. Using DNA- and rRNA-SIP, we identified four specialized clades of alkane oxidizers within Desulfobacteraceae to be distinctively active in oxidation of short- and long-chain alkanes. All clades belong to the Desulfosarcina/Desulfococcus (DSS) clade, substantiating the crucial role of these bacteria in anaerobic hydrocarbon degradation at marine seeps. The identification of key enzymes of anaerobic alkane degradation, subsequent β-oxidation and the reverse Wood–Ljungdahl pathway for complete substrate oxidation by protein-SIP further corroborated the importance of the DSS clade and indicated that biochemical pathways, analog to those discovered in the laboratory, are of great relevance for natural settings. The high diversity within identified subclades together with their capability to initiate alkane degradation and growth within days to weeks after substrate amendment suggest an overlooked potential of marine benthic microbiota to react to natural changes in seepage, as well as to massive hydrocarbon input, for example, as encountered during anthropogenic oil spills.
Similar content being viewed by others
Introduction
Hydrocarbons are widespread on Earth and can be oxidized by a variety of aerobic and anaerobic microorganisms (e.g. reviewed by Head et al., 2006; Widdel et al., 2010). Natural sources of hydrocarbons in marine systems are gas or oil seeps differing in hydrocarbon concentrations and composition. Marine gas seeps are, in general, dominated by high concentrations of methane (μM–mM range), whereas propane and butane are commonly found only in traces (μM range). In contrast, other marine hydrocarbon seeps contain a broad range of alkanes, alkenes and aromatic compounds (e.g. Anderson et al., 1983; Bazylinski et al., 1988). While degradation of hydrocarbons is well studied under oxic conditions (Head et al., 2006; Kinnaman et al., 2007; Prince et al., 2010; Redmond et al., 2010; Gutierrez et al., 2013) and numerous bacterial and fungal degraders have been described (Shennan, 2006), knowledge about anaerobic hydrocarbon degradation and responsible organisms is still limited. The anaerobic hydrocarbon degradation has been reported to occur under nitrate-reducing, sulfate-reducing, iron-reducing and methanogenic conditions (for a review see Widdel et al., 2010). In marine systems, degradation under sulfate-reducing conditions is of great importance owing to the high availability of sulfate as an electron acceptor. A large fraction of sulfate reduction (SR) at gas and hydrocarbon seeps is fueled by the anaerobic oxidation of methane (AOM; for a review see Reeburgh, 2007). However, the major part of total SR is likely to be fueled by the oxidation of non-methane hydrocarbons (for an overview see Bowles et al., 2011) with a considerable impact on local carbon and sulfur cycling. The oxidation of non-methane alkanes with sulfate has recently been reported to occur across a range of temperatures in diverse marine and terrestrial seeps or vents (e.g. Joye et al., 2004; Kniemeyer et al., 2007; Orcutt et al., 2010; Savage et al., 2010; Quistad and Valentine, 2011; Jaekel et al., 2012; Adams et al., 2013; Bose et al., 2013). Measured rates of alkane degradation and SR closely followed stoichiometric predictions that assume complete oxidation of alkanes to CO2 (e.g. see Quistad and Valentine, 2011; Bose et al., 2013). Reported rates for anaerobic short-chain alkane (SCA) oxidation were high in alkane-amended slurries of Gulf of Mexico or Middle Valley sediments (34–354 nmol C2–C4 ml−1 per day; Adams et al., 2013; Bose et al., 2013). In situ anaerobic propane oxidation rates in fresh sediments from a gas seep located in the Santa Barbara Channel even exceeded rates from the batch cultures with up to 2100 nmol propane cm−3 per day (Quistad and Valentine, 2011). The rates were comparably high to those reported for AOM in similar environments (e.g. Joye et al., 2004, Knittel and Boetius, 2009) suggesting the anaerobic oxidation of SCAs as an important process at marine gas seeps (Quistad and Valentine, 2011; Bose et al., 2013), despite C2–C5 alkanes typically present at much lower concentrations than methane.
Most cultivated or enriched sulfate-reducing hydrocarbon degraders are widespread within the class Deltaproteobacteria (Figure 1). Yet, the only pure culture shown to degrade SCAs is strain BuS5 (Kniemeyer et al., 2007), capable of growth on propane and butane. It is phylogenetically affiliated with the Desulfosarcina/Desulfococcus (DSS) clade belonging to the family Desulfobacteraceae. The DSS clade was defined based on the coverage of probe DSS658 (Manz et al., 1998) that is widely used for in situ detection and quantification of these organisms. Besides strain BuS5, DSS comprises organisms that have been shown to dominate propane- and n-pentane-degrading enrichments from a terrestrial hydrocarbon seep (Savage et al., 2010), or enrichments from Gulf of Mexico or Hydrate Ridge seep sediments degrading C2–C4 alkanes (Jaekel et al., 2012; Bose et al., 2013). In addition to SCA degraders, DSS comprises degraders of mid-chain or long-chain alkanes (LCAs), alkenes or aromatic compounds, such as toluene or naphthalene (e.g. Aeckersberg et al., 1998; Harms et al., 1999; So and Young, 1999; Meckenstock et al., 2000). Further hydrocarbon-degrading sulfate-reducing bacteria (SRB) can be found within non-DSS Desulfobacteraceae, Desulfobulbaceae, Syntrophobacteraceae and Desulfurellales (Widdel et al., 2010; Figure 1). Of these, Desulfurellales might be of particular interest as it comprises a broad diversity of organisms dominating anaerobic batch reactor incubations of sediments from the Middle Valley hydrothermal vent field (Adams et al., 2013), as well as the sulfate-reducing partner bacteria of ANME-1-mediating thermophilic AOM in Guaymas Basin (GB) sediments (Holler et al., 2011). Only few non-deltaproteobacterial SRB capable of degrading hydrocarbons are known and belong to the phylum Firmicutes, that is, Desulfosporosinus sp. strain Y5 (Liu et al., 2004) and Desulfotomaculum sp. strain OX39 (Morasch et al., 2004). All strains available in pure culture can usually degrade only a restricted range of hydrocarbons, for example, few aliphatic hydrocarbons within a narrow range of chain length (e.g. C3–C4 or C6–C10 or C12–C20) or few aromatic compounds.
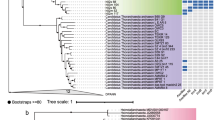
Phylogenetic tree showing the affiliations of 16S rRNA gene sequences retrieved from incubations of AMV and GB sediments with 13C-labeled butane or dodecane with selected reference sequences of the Deltaproteobacteria. A consensus tree based on the obtained maximum-likelihood (PhyML) tree is shown. A selection of gammaproteobacterial sequences was used as outgroup. Partial sequences were inserted into the reconstructed tree by using parsimony criteria. Entries corresponding to T-RFs of identified 13C-labeled alkane degraders are shown in red, and other sequences from the incubations are shown in blue. Sequences of cultivated anaerobic hydrocarbon degraders (black, boldface type) were included as additional reference; their substrate usage is given within parentheses. Probe coverage is indicated by gray-colored boxes. Scale bar represents 10% estimated sequence divergence.
In situ analysis of SRB communities at numerous marine gas and oil seeps indicated a high abundance and diversity of DSS with 10% to >60% of total cells (e.g. Orphan et al., 2001; Orcutt et al., 2010; Schreiber et al., 2010; Kleindienst et al., 2012). Potentially, the DSS clade is one of the most relevant lineages for hydrocarbon degradation in marine sediments, and could thus be of utmost relevance not only at natural hydrocarbon seeps but also upon catastrophic events. Still, the ecology and niche partitioning between these hydrocarbon degraders is quite poorly understood in situ, and studies directly linking functional information from both the lab and the field data are not available. Here, we aim at closing the gap between previous cultivation-dependent studies (i.e. isolates) and biogeochemical rate measurements in situ (i.e. hydrocarbon turnover). We studied two types of anoxic sediments from contrasting natural hydrocarbon seep environments: the GB hydrocarbon seep sediment was selected as a representative for a habitat with changing natural conditions characterized by the presence of steep gradients of temperature and electron acceptors, as well as seepage of a broad spectrum of hydrocarbons including alkanes <C20. In contrast, the Amon mud volcano (AMV) located in the Mediterranean Sea was selected as a representative for a more stable natural seep environment that is characterized by constant temperatures and a narrow range of hydrocarbons (gaseous alkanes C1–C5).
Sediments were incubated under sulfate-reducing conditions with 13C-labeled butane as a model substrate for a gas leakage and 13C-labeled dodecane as a model substrate for an oil spill. We hypothesized (i) that the DSS clade indeed dominates hydrocarbon degradation in situ, and (ii) that niche partitioning of DSS subclades occurs based on alkane chain length. A combination of DNA-, RNA- and protein-based stable isotope probing (SIP) enabled us to identify distinct clades of active SRB, respectively, involved in alkane degradation. These specialized degraders may catalyze a major part of the SR-dependent non-methane hydrocarbon degradation in marine sediments.
Materials and methods
Study sites and sampling
Anoxic sediments were collected from the active AMV and a hydrocarbon seep at GB during the cruises MSM13-3 (RV Maria S. Merian) and AT15-56 (RV Atlantis) in 2009. AMV is located in the Nile deep-sea fan and comprises large subsurface gas reservoirs (Mastalerz et al., 2009). Sampling was carried out with the remotely operated vehicle Quest 4000 during dive 240 at station 929 (PANGAEA database event MSM13/3_929-1; water depth 1122 m, 32°20.1321N, 31°42.6543E). The sediments derived from an area surrounding the central dome and were covered by a microbial mat. For details see Felden et al. (2013). GB is located in the central Gulf of California and harbors petroleum-rich hydrothermal sediments, covered with layers of buried sedimentary organic matter. Hydrothermal fluids contain elevated concentrations of hydrocarbons, including alkanes and aromatic hydrocarbons (Bazylinski et al., 1988; Didyk and Simoneit, 1989). Fine-grained sediments below a white Beggiatoa mat were collected during dive 4573 (water depth 2010 m, 27°0.696N, 111°24.265W) with the submersible Alvin.
Setup of incubations for SIP

Sediments were stored for 4 months at 4 °C. During this time period, preincubations were carried out to identify sediments with highest alkane-degrading microbial activities based on sulfide and 13CO2 production. Slurries were prepared in an anoxic tent (N2:CO2, 9:1 v/v ) by mixing sediments from 2 to 20 cm (AMV) and 0 to 10 cm (GB) depth in a 1:1 ratio with artificial sea water (Widdel and Bak, 1992), using a sterile metal spoon. During further processing and preparation of incubations, samples were always maintained under anoxic conditions at room temperature. Sediment slurries were further distributed in 15 ml aliquots in 22-ml tubes sealed with butyl rubber stoppers, under a headspace of N2:CO2. Either fully 13C-labeled or unlabeled butane were added to the headspace, using N2-purged gastight syringes. 13C-labeled and unlabeled dodecane were injected into the tubes using N2-flushed glass syringes. Hydrocarbon concentrations in our experiments (2.1 mM butane; 1.8 mM dodecane) exceeded the natural concentrations by >100-fold to prevent substrate availability limitations in SIP. Butane is a natural substrate at both seep types occurring in concentrations of 0.5-1.1 μM (AMV) and 6-16 μM (GB, at a site close by; MY Kellermann, personal communication). Dodecane has only been detected at hydrocarbon seeps in the GB (Bazylinski et al., 1988). AMV sediments were incubated at 20 °C, which is slightly above the in situ temperature of 14 °C, whereas for GB sediments, where the temperature range is wider (from 3 °C to 50 °C), 28 °C was chosen. For each site (AMV, GB) and substrate (unlabeled butane, 13C-labeled butane, unlabeled dodecane, 13C-labeled dodecane), six replicate tubes were incubated. Time points for DNA and RNA extraction were chosen based on sulfide production (Figure 2a). For these analyses, two of the replicate tubes were used for each time point. Sediments were centrifuged (10 min, 4000 g) and pellets stored at −80 °C. One tube each was used for nucleic acid-SIP and one for protein-SIP. For catalyzed reporter deposition-fluorescence in situ hybridization (CARD-FISH) analysis, 0.5 ml of incubated sediment slurries were fixed for 1 h at 4 °C with a final concentration of 1% paraformaldehyde, washed with 1 × phosphate-buffered saline (130 mM NaCl, 10 mM sodium phosphate; pH 7.2) and finally stored in phosphate-buffered saline–ethanol (1:1) at −20 °C.
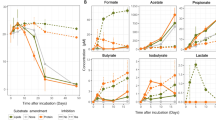
Measurements on incubations of AMV and GB sediments with 13C-labeled butane (SC) or dodecane (LC) under sulfate-reducing conditions. AMV incubations were performed at 20 °C, and GB incubations at 28 °C. (a) Microbial butane- or dodecane-dependent sulfide production. Open symbols (□), incubations with unlabeled alkanes; filled symbols (▪), incubations with 13C-labeled alkanes. Average of two replicate samples is shown. Error bars give the standard deviation for the two replicates. Additionally, consumption of butane () is indicated for AMV-SC and GB-SC 13C incubations. (b) Abundance of 13C-DIC (at%) in incubations with (●) 13C-labeled and (x) -unlabeled substrates in course of the incubation. Average of two replicate samples is shown. Error bars give the standard deviation for the two replicates. (c) Increase of specific cell concentrations of identified alkane degraders of clades SCA1, SCA2, LCA1 and LCA2 over the time course of the four incubations. (d) RIA in peptides (RIA, ) and labeling ratio of 13C-labeled peptides to all peptide forms (LR, ♦).
Chemical analysis
Sulfide was quantified photometrically as colloidal CuS (Cord-Ruwisch, 1985). For each of the two replicate incubations, two technical replicates were measured. Butane concentrations were measured by gas-chromatographic headspace analysis (oven 110 °C, injector 150 °C, detector 280 °C, nitrogen carrier gas) as described before (Musat and Widdel, 2008). The C-isotopic composition of dissolved inorganic carbon (DIC) from sediment slurries was analyzed using gas chromatography combustion isotope ratio mass spectrometry (Fisons VG Optima) according to Assayag et al. (2006). Biological activity of the DIC controls was halted by the addition of 4% paraformaldehyde (final concentration).
CARD-FISH
In situ hybridization with horseradish peroxidase-labeled oligonucleotide probes followed by fluorescently labeled tyramide signal amplification (CARD) and microscopic counting was performed as described previously (Kleindienst et al., 2012). Probes for alkane degraders were developed using the software package ARB and the implemented function ‘probe design’ (Ludwig et al., 2004). The specificity of new probes was evaluated against reference strains having one or more mismatches. For probes SCA1-212a and b, we used strain BuS5 and GMe12, respectively, as a positive control. For probes SCA2, LCA1 and LCA2, stringent hybridization conditions were determined by Clone-FISH (Schramm et al., 2002). Probe sequences and formamide concentrations required for specific hybridization are given in Supplementary Table S1.
DNA/RNA-SIP
Nucleic acids were directly extracted from 3 to 4 g of incubated sediments and stored at −80 °C. From AMV sediment incubations, RNA was extracted according to McIlroy et al. (2008), whereas from GB sediments DNA was extracted according to Zhou et al. (1996), because RNA extraction did not result in the required amounts of high-quality rRNA.
Labeled and unlabeled nucleic acids (5 μg Guaymas-DNA and 750 ng AMV-rRNA) were loaded onto a CsCl gradient (average density 1.71 g ml−1; Calbiochem, Merck, Darmstadt, Germany) or cesium trifluoroacetate (CsTFA; average density 1.795 g ml−1; Amersham Pharmacia Biotech, Freiburg, Germany) in gradient buffer (0.1 M Tris-HCl at pH 8, 0.1 M KCl, 1 mM EDTA). Fractionation of gradients, refractometric measurement of fraction buoyant density and precipitation of nucleic acids was carried out as published before (Lueders, 2010). Distribution of Guaymas-DNA or AMV-RNA within the gradients was analyzed using bacterial 16S rRNA gene-targeted real-time polymerase chain reaction and real-time reverse-transcription polymerase chain reaction in an Mx3000P Cycler (Stratagene, La Jolla, CA, USA) with the primers Ba519f/Ba907r as described in Lueders et al. (2004). FAM-labeled amplicons for terminal restriction fragment length polymorphism fingerprinting of selected density-resolved nucleic acids were generated with bacterial primers 27f/907r as described in detail elsewhere (Lueders et al., 2004; Pilloni et al., 2011). Briefly, amplicons were digested with MspI (Promega, Madison, WI, USA) and size-separated on an ABI 3730 DNA sequencer (Applied Biosystems, Life Technologies, Grand Island, NY, USA). Further analysis was performed with T-REX (Culman et al., 2009). Background noise filtering was on the default factor 1.2; the clustering threshold for aligning peaks across samples was set to 1.5 bp. Relative terminal restriction fragment (T-RF) abundance was inferred from peak heights. For reduction of data complexity, T-RFs that occurred in <10% of the samples were excluded from further analysis.
AMV sediment frozen directly after sampling was used as a control. The resulting T-RF pattern was similar to the one obtained from the SIP inoculum (Supplementary Figure S1), suggesting no pronounced bottle effect on sediment microbiota during the 4-month storage before setting up the SIP incubations.
Pyrotag sequencing was performed on unfractionated total DNA from sediments at the time point of SIP inoculation, as well as on ‘heavy’ fractions of rRNA and DNA from the respective gradients. Amplicon sequencing on a 454 GS FLX pyrosequencer (454 Life Sciences, a Roche company, Branford, CT, USA), data processing and read quality trimming were performed as described (Pilloni et al., 2012). Pyrotag reads were taxonomically classified according to the ARB SILVA taxonomy (SSURef_108, release September 2011) using the SILVA-based next-generation sequencing pipeline (Quast et al., 2013). The taxonomic assignment of sequences to DSS subclades, that is, SEEP-SRB1a, SCA1, SCA2, LCA1 and LCA2 (Table 1), has been carried out using ARB (Ludwig et al., 2004). One representative sequence of all operational taxonomic units (OTUs) (95% criterion) that have been assigned to Desulfobacteraceae based on the SILVA-based next-generation sequencing pipeline has been imported into ARB SILVA database SSURef_111. After manual optimization of alignments sequences have been added to the provided tree under parsimony criteria.
For T-RF prediction, forward and reverse sequences were assembled into contigs (Pilloni et al., 2012). Subsequently, dominating contigs were again taxonomically classified using the SILVA-based next-generation sequencing pipeline and imported into ARB. T-RFs of contigs were predicted using TRiFle (Junier et al., 2008) and search functions implemented in the ARB software package. Given OTU numbers were calculated based on a 95% criterion using the reverse sequences.
masD clone library
The delta subunit of the (1-methylalkyl)succinate synthase gene (masD) was amplified using primers 7757f-1 and 7757f-2/8546r (von Netzer et al., 2013) or ass /bssF and ass/bssR (for GB-SC incubation; Callaghan et al., 2010) as described previously. Cloning and sequencing was performed as described previously (Kleindienst et al., 2012).
Phylogenetic analysis
Phylogenetic 16S rRNA-based trees were calculated by neighbor-joining and maximum-likelihood (RAxML and PhyML) analysis with a filter considering 1244 positions that are conserved in at least 50% of the selected deltaproteobacterial sequences. For tree calculation, only nearly full-length sequences were used. Unstable branching orders are visualized as a multifurcation. Partial sequences from this study were inserted into the reconstructed tree by parsimony criteria. MasD-based trees were calculated from deduced amino acids by maximum-likelihood (RAxML and PhyML) analysis with a filter considering 159-amino-acid positions that are conserved in at least 30% of the selected sequences.
Sequence accession numbers
16S rRNA gene sequences and masD sequences have been submitted to the EMBL/GenBank/DDBJ databases under the accession no. HE797917-HE797929 and HG764655-HG764728, respectively. Pyrotag sequencing raw data have been stored in the sequence read archive under SRA accession number ERP001437.
Protein-SIP
Two grams of incubated sediments were suspended in 5.4 ml extraction buffer (Benndorf et al., 2009) and subjected to three cycles of freeze (in liquid nitrogen) and thaw (water bath at 65 °C) with the addition of 0.6 ml of sodium dodecyl sulfate (10%, w/v) before the last thawing step. After phenol extraction and acetate precipitation of the organic and aqueous phase, extracted proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (Laemmli, 1970). Five sections of each lane were excised from the sodium dodecyl sulfate-polyacrylamide gel electrophoresis and treated according to Jehmlich et al. (2008). Eluted peptides from the aqueous-phase gel slices were combined. Purification was carried out using ZipTipC18 columns (Merck Millipore, Billerica, MA, USA), resulting in six fractions per sample. Mass spectrometric measurement was carried out using the nanoAcquity UPLC (Waters, Milford, MA, USA) coupled LTQ-Orbitrap XL (Thermo Fisher Scientific, Waltham, MA, USA) (Herbst et al., 2013). Trapping was performed for 6 min. To elute the peptides the acetonitrile solvent gradient was set in the first 54 min from 2% to 20% and subsequently 28 min from 20% to 85%.
For peptide identification, first a survey search using Mascot (Perkins et al., 1999) against all prokaryotic entries of NCBInr (28 April 2013) was performed. These unfiltered results and a six-frame translation of the genome of strain BuS5 (Stefan M Sievert, Woods Hole Oceanographic Institution, Woods Whole, MA, USA; JGI project number 403224; http://img.jgi.doe.gov/cgi-bin/w/main.cgi) were used as database for peptide identification. The translation was made with MaxQuant 1.2.0.18 (Cox and Mann, 2008) and a minimum of 50 amino acids per ORF. Furthermore, contaminants and reverse entries as decoys were added to the database. The main search used OMSSA (Geer et al., 2004) in an OpenMS pipeline (Sturm et al., 2008). Carbamidomethylation of cysteins was treated as fixed modification and oxidation of methionines as variable modification. Mass deviation tolerances were 5 p.p.m. for precursor masses and 0.5 Da for product ions. The maximal false discovery rate for peptides was set to 1%.
Protein relative isotope abundance (RIA) and protein labeling ratio
RIA and the labeling ratio were determined using OpenMS and the MetaproSIP tool (http://openms.de/metaprosip) with a minimum correlation of 0.7 and a mass window of 10 p.p.m.
Results and discussion
Bacterial diversity in AMV and GB sediments
We initially characterized the bacterial diversity in contrasting sediments from a gas seep at AMV emitting high fluxes of methane and other SCA and in sediments from a hydrocarbon seep in the GB releasing a complex hydrocarbon mixture similar to crude oil with various n-alkanes and aromatic hydrocarbons. AMV gas seep sediments used as inoculum (t0) for later SIP experiments were clearly dominated by Deltaproteobacteria contributing almost three-quarters of the obtained 5139 pyrosequencing reads (Table 1). The diversity within the Deltaproteobacteria was high with 579 OTUs on the genus level (95% cutoff, Yarza et al., 2010) from overall 1151 bacterial OTUs. About 50% of total AMV sequences (=70% of deltaproteobacterial sequences) were affiliated with the DSS subclade SEEP-SRB1. Of these, 65% affiliated with SEEP-SRB1a, the sulfate-reducing partner of ANME-2a and ANME-2c in AOM habitats (Schreiber et al., 2010). Sequences found at lower frequency were affiliated with Gammaproteobacteria (8%), Epsilonproteobacteria (5%) and candidate division OP9 (3%).
In GB hydrocarbon seep sediments, a total of 546 bacterial OTUs were detected. Of these, 159 OTUs (=29% of bacterial OTUs) have been assigned to Deltaproteobacteria, suggesting a lower diversity in GB sediments compared with AMV. About 14% of total GB sequences (=40% of deltaproteobacterial sequences) were assigned to the deep-branching group Hot-Seep1 that was shown to be involved in thermophilic AOM (Holler et al., 2011; Table 1). Other sequences frequently found in GB sediments were affiliated with Epsilonproteobacteria (12%), Bacteroidetes (8%) and Chloroflexi (5%). The detected phyla match well with the current picture of bacterial diversity in sediments from similar sites at AMV (Omoregie et al., 2009; Pachiadaki et al., 2011) and GB (Teske et al., 2002; Dhillon et al., 2003; Biddle et al., 2011) or more geographic distant seep sites (e.g. Orphan et al., 2001; Knittel et al., 2003; Lloyd et al., 2010; Orcutt et al., 2010; Nunoura et al., 2012; Adams et al., 2013; Rubin-Blum et al., 2014). This congruency also confirms that SIP experiments were carried out with benthic communities that typically occur at these seeps.
Alkane-dependent sulfide production
Incubations with 13C-labeled butane (SC) or dodecane (LC) were started in replicates under sulfate-reducing conditions, resulting in four types of incubations: AMV-SC, AMV-LC, GB-SC and GB-LC.
Alkane-dependent microbial activity differed among the four incubations. The incubations of AMV sediments with butane (AMV-SC) had the fastest response. Butane degradation started without lag phase, as determined based on butane-dependent sulfide production, butane oxidation and production of 13CO2 (Figures 2a and b). After 29 days, the selected end point of AMV-SC incubation, almost complete removal of butane occurred, which coincided with 6 mM sulfide production and a strong labeling of resulting CO2 (15% 13C-DIC). In contrast, a pronounced lag phase of 20 days was observed for incubations of GB sediments with butane (GB-SC). By day 113, butane was consumed resulting in 21% 13C-DIC.
Dodecane-dependent SR was delayed in both incubations (AMV-LC and GB-LC). Significant sulfide production started only after 80 days (Figure 2a) and 6 mM sulfide was produced after 232 days (GB) and 309 days (AMV), leading to an enrichment of 24% and 21% 13C-DIC, respectively (Figure 2b). Our data show that both sediments are able to react within weeks to months on the addition of butane or dodecane.
Four DSS clades are key degraders of butane and dodecane
An early time point and the end point of SIP experiments were selected for DNA and RNA extraction on each of the duplicate 13C-labeled alkane incubations for subsequent ultracentrifugation. Except for GB-LC, no considerable relative increase in total cell counts was observed between t0 and the individual first sampling points (Supplementary Figure S2). The absence of significant cell growth is the prerequisite for a reliable identification of primary degraders by SIP experiments whereas cross-feeding could therefore be neglected at t1. All density-resolved nucleic acid fractions were first subjected to terminal restriction fragment length polymorphism fingerprinting, to identify the T-RFs of 13C-labeled populations, which were subsequently identified by pyrotag sequencing of selected gradient fractions.
Butane degraders
Two subclades of DSS were identified in AMV sediments: after only 9 days, a T-RF of 132 bp length became dominant in the heavy rRNA fraction (Figure 3) and was identified as a close relative of the butane-degrading strain BuS5 (93% 16S rRNA gene sequence similarity; Figure 1). The second subclade was identified after 29 days based on a T-RF of 512 bp length, representing butane degraders closely related (99.6%) to phylotype ‘But12_GMe’ that dominated an enrichment culture on butane (Kniemeyer et al., 2007; Jaekel et al., 2012). A neighbored subclade contains two phylotypes dominant in enrichments with propane (Prop12_GMe) or butane (But12_HR; Jaekel et al., 2012). We name these three neighboring clusters of propane and butane degraders ‘SCA1’. In addition to the identified populations represented by these major labeled T-RFs, several further SCA1-OTUs based on a 95% similarity criterion were identified in AMV sediments, resulting in a total of 5 OTUs for subclade But12_GMe and 14 OTUs for subclade BuS5, suggesting the presence of multiple closely related genera capable of butane degradation. In support of our hypothesis, enrichments with sediments from Garden Banks mud volcano (Northern Gulf of Mexico) supplemented with C2–C4 alkanes were dominated by members of SCA1 (Bose et al., 2013). Furthermore, public database analysis shows that members of SCA1 are indeed globally distributed. They have been retrieved from marine seep sites in the Mediterranean (Pachiadaki et al., 2011), at Hydrate Ridge (Jaekel et al., 2012), Gulf of Mexico or GB (Kniemeyer et al., 2007; Jaekel et al., 2012). With this functional characterization of members of SCA1, we substantiate that this group is likely of relevance in the environment and that the isolate BuS5 is a relevant model organism.
Bacterial community structure as shown by terminal restriction fragment length polymorphism (T-RFLP) fingerprinting of density-resolved rRNA (AMV) and DNA (GB) fractions after density centrifugation of sediments incubated with 13C-labeled butane (SC) or dodecane (LC) under sulfate-reducing conditions. Fingerprints of representative ‘light’, ‘intermediate’ and ‘heavy’ gradient fractions are shown for all gradients of respective 13C-labeled and -unlabeled alkane incubations, for the first and the last sampling time point of each SIP treatment, respectively. The buoyant densities (g ml−1) for each fraction are given in brackets. T-RFs corresponding to taxa identified to be involved in the degradation of butane and dodecane are highlighted with circles. Fingerprints for sediment inocula are shown as reference.
In GB sediments, butane stimulated a different DSS subclade: after 57 days, a 163 bp T-RF (Figure 3) was detected, which we named ‘SCA2’ (Figure 1). SCA2 is phylogenetically closely related to SCA1 with up to 95% sequence similarity, but appears to be restricted to GB. After 113 days, an additional labeled T-RF of 92 bp length emerged and was identified as uncultured Marinilabiaceae of the phylum Bacteroidetes. In general, Bacteroidetes are known to degrade high-molecular-weight compounds (e.g. polysaccharides, proteins) and to have a preference for growth attached to particles (Fernández-Gómez et al., 2013). We therefore assume that members of this clade were secondarily labeled, by using dead biomass of or other trophic interaction with the primary hydrocarbon consumers, rather than degrading butane on its own. Yet, it should be noted that related sequences originate from several seeps in the Mediterranean, Gulf of Mexico and Pacific Ocean (e.g. Omoregie et al., 2008; Ruff et al., 2013). Therefore, it would be worthwhile to investigate the role of this clade in marine hydrocarbon catabolism more specifically.
Dodecane degraders
Another 163 bp T-RF became apparent in heavy rRNA of AMV-LC. It was assigned to organisms of another DSS subclade, which we named ‘LCA1’. LCA1 comprises eight OTUs from AMV sediments and shows an intragroup 16S rRNA similarity of as low as 88.5%. It is phylogenetically clearly distinct from SCA1 and SCA2 (Figure 1). It comprises not only sequences from marine seep sites or hydrocarbon-polluted sediments (Zhang et al., 2010; Elsaied et al., 2011; Acosta-González et al., 2013; Ruff et al., 2013) but also sequences retrieved from the chemocline in the meromictic freshwater lake Cadagno (Halm et al., 2009). Moreover, a second 13C-labeled T-RF (89 bp) was detected in the AMV-LC incubation only for the first time point (Figure 3). However, phylogenetic assignment of this T-RF failed, as no sequences with a matching restriction site were found in the corresponding 454 data set. In GB-LC incubations, a dominating 164 bp T-RF was identified in heavy rRNA after 115 days of incubation with 13C-dodecane. This T-RF represents a novel clade of Desulfobacteraceae that is most closely related to Desulfosalsimonas propionica. We propose to name this clade as ‘LCA2’ and regard it as another subclade of DSS, even though members of clade LCA2 hold one mismatch to probe DSS658 (position 14, C–T). Two and four OTUs of LCA2 were identified in AMV and GB sediments, respectively. Other sequences from marine seeps or oil-contaminated sites (Figure 1; e.g. Lösekann et al., 2007; Abed et al., 2011; Zhang et al., 2011; Ruff et al., 2013) also belong to the LCA2 clade, and could potentially also be involved in dodecane degradation.
A second labeled T-RF (512 bp) was identified in heavy DNA of the GB-LC incubation. The corresponding organisms form a cluster within Desulfobulbaceae, together with sequences retrieved from a mud volcano in the Gulf of Cadiz (e.g. clone GoC_Bac_176, FN820344; Figure 1). If members of this cluster are capable of dodecane oxidation or have been labeled through cross-feeding needs further investigation. Comparative 16S rRNA analysis (Table 1) suggests a low abundance in GB sediments.
Additionally, members of the candidate phylum Omnitrophica (OP3; Rinke et al., 2013), belonging to the Planctomycetes/Verrucomicrobia/Chlamydiae superphylum, were found labeled in GB-LC incubations based on a T-RF of 671 bp length (Figure 3). The OP3 lineage seems to be ubiquitous in anoxic environments, such as marine habitats. Recent studies reported the genomic potential for anaerobic respiration (Glöckner et al., 2010) or the syntrophic involvement of OP3 in anaerobic monoterpene degradation (Rotaru et al., 2012). Further studies are clearly required to unravel whether these microbes were 13C labeled in our incubations owing to syntrophy or cross-feeding. In summary, all identified clades from our study and suggested carbon flow in the distinct sediments are depicted in Figure 4.
Suggested 13C carbon flow for alkane degradation in AMV and GB sediments. The role of active DSS subclades is based on the results of nucleic acid-SIP as visualized by the corresponding T-RFs. Fluorescent images show CARD-FISH signals as obtained after hybridization with oligonucleotide probes specific for SCA1, SCA2, LCA1 and LCA2. Scale bars represent 2 μm.
Detection of SCA- and LCA-degrading DSS subclades in situ
A set of CARD-FISH probes was developed for the specific detection of butane- and dodecane-degrading DSS subclades (Figure 1 and Supplementary Table S1). After careful adjustment of specific hybridization conditions, the new probes were applied on our SIP incubations. Cell size and morphology differed for SCA1 subclades (Figure 4): cells belonging to subclade ‘BuS5’ have an almost coccoid morphology of 1.8 μm × 2 μm, whereas cells of subclade ‘But12_GMe’ are curved rods of 1.5 μm × 2.1 μm. Cells of SCA2 were either short rods (1 μm × 1.3 μm) or vibrio-like (2.5 μm × 1.5 μm). LCA1 and LCA2 cells were both cocci with a diameter of approximately 1 μm.
We observed a strong specific cell increase for all four clades in course of the experiment (Figure 2c). SCA1 and SCA2 cell numbers increased by a factor of approximately 6 within 29 and 113 days, respectively. LCA1 and LCA2 cell numbers increased by a factor of 23 and 59 during incubation. The detected growth of these clades strongly supports the correct taxonomic assignment of the identified T-RFs by polymerase chain reaction-independent means, and thus the capability of these organisms for alkane degradation.
The in situ presence and relevance of SCA1, SCA2 and LCA2 was confirmed by CARD-FISH on nearby sediments with relative abundances between 0.2% and 2% of total single cells (max 4.4 × 107 cells ml−1; Supplementary Table S2). LCA1 cells could not be detected in these samples, possibly because of a too high selectivity of probe LCA1-443, which covers only the specific organisms represented by the 163 bp T-RF, plus one additional sequence.
This evidence for the presence of three out of four of the identified DSS subclades in low abundances at the two investigated seep sites is compelling. From this seed bank, different SC and LC alkane degraders were apparently capable of growing, over relatively short time frames from days to weeks, and initiate alkane degradation.
Diversity of masD genes in SC and LC incubations
MasD gene libraries constructed from the four incubations revealed a high diversity of organisms associated with alkane degradation: of a total of 247 masD sequences, 22 OTUs (based on 93% amino-acid identity) were detected in AMV-SC, 25 OTUs in AMV-LC, 11 OTUs in GB-SC, and 16 OTUs in GB-LC. Several habitat-specific clusters became obvious in the phylogenetic tree comprising sequences either from AMV or from GB (Supplementary Figure S3). Substrate-specific clades were detected only for GB-SC and GB-LC incubations. The high masD diversity supported our findings based on 16S rRNA, that is, the presence of multiple closely related genera capable of alkane degradation. Furthermore, this diversity suggests a great potential of marine benthic communities to respond to hydrocarbon inputs and natural changes in seepage. Gas hydrate decay, gas leakage or oil spills are only some of the possible events able to cause an increased hydrocarbon input on the sediments.
Proteins involved in SC and LC alkane degradation under sulfate-reducing conditions
We directly identified enzymes and catabolic pathways involved in butane and dodecane oxidation in these sediments using protein-SIP. In AMV-SC, AMV-LC, GB-SC and GB-LC incubations, the functional metaproteomic approach identified between 511 (1143 peptide species) and 1493 (3869 peptide species) proteins (see Supplementary Table S3). 13C-labeled peptides showed mostly hits to proteins of the Desulfobacteraceae, thus further substantiating our finding that active alkane degraders in AMV and GB sediments belong to this family.
A crucial step in the anaerobic degradation of alkanes is their activation. A common activation reaction is the addition of alkanes to fumarate yielding alkylsuccinates (Figure 5; Rabus et al., 2001). The genes responsible for the initial alkylsuccinate synthase reaction have been identified and named masD (1-methylalkyl)succinate synthase; Grundmann et al., 2008) or assA (alkylsuccinate synthase; Callaghan et al., 2008). Based on the available BuS5 genome (SCAFFOLD_2:TRUE:69872), 13C-labeled MasD peptides could be identified within the AMV-SC (28 peptide species), AMV-LC (2 peptide species) and GB-SC incubations (2 peptide species; Supplementary Table S3). Fumarate addition is followed by a carbon skeleton rearrangement before β-oxidation. The β-oxidation has been proposed as an analog to the degradation of LC fatty acids (Wilkes et al., 2002). The detection of 13C-labeled peptides in AMV-SC, AMV-LC and GB-SC incubations involved in β-oxidation corroborate this pathway. These peptides were assigned to β-oxidation enzymes responsible for dehydrogenation (i.e. acyl-coenzyme A (CoA) dehydrogenase), hydration (i.e. enoyl-CoA hydratase/isomerase) and oxidation (3-hydroxyl-CoA dehydrogenase) (Supplementary Tables S3 and S4).
Alkane-degrading SRB within the Desulfobacteraceae have the capability for complete substrate oxidation, and thus they further metabolize acetyl-CoA. The most common pathway for acetyl-CoA/acetate dissimilation in SRB is the reverse Wood–Ljungdahl pathway, with carbon monoxide dehydrogenase (reverse acetyl-CoA synthetase) as a key enzyme (Schauder et al., 1986). 13C-labeled peptides corresponding to four enzymes of the Wood–Ljungdahl pathway were indeed identified: carbon monoxide dehydrogenase/acetyl-CoA synthase complex, methylenetetrahydrofolate reductase, formate-tetrahydrofolate ligase (also named formyltetrahydrofolate synthetase) and formate dehydrogenase. These findings strongly indicate that alkane-degrading SRB use the Wood–Ljungdahl pathway for terminal oxidation and that acetyl-CoA is a central metabolite during alkane degradation. The presence of the carbon monoxide dehydrogenase/acetyl-CoA synthase complex suggests that these alkane degraders could, in principle, also grow on acetate as an alternative substrate. The identified labeled peptides allowed the reconstruction of the complete pathway of alkane activation and further degradation to CO2 (Figure 5).
Furthermore, 13C-labeled peptides from AMV incubations could be assigned to the three key enzymes of dissimilatory SR, that is, sulfate adenylyltransferase, adenosine-5′-phosphosulfate reductase and dissimilatory sulfite reductase (DSR, subunits a and b; Supplementary Tables S3 and S4). This corroborates that alkane degradation is directly linked to SR. Labeled peptides involved in SR showed highest BLAST scores to proteins from Desulfobacteraceae supporting that the key butane and dodecane degraders belong to this family. The remaining labeled peptides were mainly derived from highly conserved proteins such as chaperones, ATP synthases and K+-insensitive pyrophosphate-energized proton pumps, again affiliating mainly with Desulfobacteraceae.
Protein-SIP also allowed quantitative interpretation of butane- and dodecane-derived carbon assimilation (Figure 2d). The RIA directly corresponds to the 13C content in newly synthesized proteins. Thus, it reflects the carbon source composition of newly synthesized biomass. As the 13C content of the provided alkane substrates was 100%, but the RIA only approximately 50%, roughly half of the biomass carbon did not derive from alkane degradation. The remaining half could be gained from either other non-labeled substrates or from fixation of inorganic carbon. Particularly in the AMV-SC samples, a good correlation between RIA and 13C-DIC increase could be observed. The increase in the average 13C content corresponded to 50% of the 13C-DIC increase (example shown in Supplementary Figure S4). These SRBs probably use about 50% alkanes and 50% DIC for biomass production. This is in accordance with current knowledge on SRBs, which are known to derive a large fraction of their carbon from CO2 via carboxylation reactions (Rabus et al., 2006). In AMV-SC and AMV-LC incubations, we found several 13C-labeled peptides assigned to pyruvate:ferredoxin/flavodoxin oxidoreductase, an enzyme typically used by SRB for acetyl-CoA assimilation through reductive carboxylation to pyruvate.
Conclusions
The impact of non-methane-dependent SR was thoroughly described for various marine seep sites and has been suggested to be of global significance for the marine carbon and sulfur cycle. However, before our study, this concept has been based mostly on rate measurements (e.g. Joye et al., 2004; Kniemeyer et al., 2007; Orcutt et al., 2010; Bowles et al., 2011; Quistad and Valentine, 2011; Bose et al., 2013) and isolates or enrichment cultures of hydrocarbon-degrading SRB (e.g. Kniemeyer et al., 2007; Jaekel et al., 2012; Adams et al., 2013; Bose et al., 2013). Here, we provide the direct link between biogeochemical patterns and apparently specialized DSS subclades of key alkane degraders in marine sediments. Nucleic-acid-SIP and protein-SIP, which has been combined for the first time on a marine sediment sample, proved as powerful approach for the investigation of the microbial community associated with alkane degradation.
Complementary to aerobic hydrocarbon-degrading bacterial populations in the marine realm (Hazen et al., 2010; Abbriano et al., 2011; Kostka et al., 2011; Bælum et al., 2012; Redmond and Valentine, 2012; Gutierrez et al., 2013), this study corroborates the environmental importance of the DSS clade for the anaerobic degradation of alkanes at marine seeps. Moreover, the identification of diverse and substrate-specialized clades of sulfate-reducing bacteria initiating alkane degradation within days to weeks suggests an overlooked potential of natural marine benthic microbiota to react to massive hydrocarbon input, for example, as encountered during anthropogenic oil spills. Finally, we provided evidence that available isolates and their hydrocarbon activation mechanisms are of great relevance for natural settings.
Accession codes
References
Abbriano RM, Carranza MM, Hogle SL, Levin RA, Netburn AN, Seto KL et al (2011). Deepwater Horizon oil spill: a review of the planktonic response. Oceanography 24: 294–301.
Abed RMM, Musat N, Musat F, Mußmann M . (2011). Structure of microbial communities and hydrocarbon-dependent sulfate reduction in the anoxic layer of a polluted microbial mat. Mar Pollut Bull 62: 539–546.
Acosta-González A, Rosselló-Móra R, Marqués S . (2013). Characterization of the anaerobic microbial community in oil-polluted subtidal sediments: aromatic biodegradation potential after the Prestige oil spill. Environ Microbiol 15: 77–92.
Adams MM, Hoarfrost AL, Bose A, Joye SB, Girguis PR . (2013). Anaerobic oxidation of short-chain alkanes in hydrothermal sediments: potential influences on sulfur cycling and microbial diversity. Frontiers in Microbiology 4: 110.
Aeckersberg F, Rainey FA, Widdel F . (1998). Growth, natural relationships, cellular fatty acids and metabolic adaptation of sulfate-reducing bacteria that utilize long-chain alkanes under anoxic conditions. Arch Microbiol 170: 361–369.
Anderson RK, Scalan RS, Parker PL, Behrens EW . (1983). Seep oil and gas in Gulf of Mexico slope sediment. Science 222: 619–621.
Assayag N, Rivé K, Ader M, Jézéquel D, Agrinier P . (2006). Improved method for isotopic and quantitative analysis of dissolved inorganic carbon in natural water samples. Rapid Commun Mass Spectrom 20: 2243–2251.
Bælum J, Borglin S, Chakraborty R, Fortney JL, Lamendella R, Mason OU et al (2012). Deep-sea bacteria enriched by oil and dispersant from the Deepwater Horizon spill. Environ Microbiol 14: 2405–2416.
Bazylinski DA, Farrington JW, Jannasch HW . (1988). Hydrocarbons in surface sediments from a Guaymas Basin hydrothermal vent site. Org Geochem 12: 547–558.
Benndorf D, Vogt C, Jehmlich N, Schmidt Y, Thomas H, Woffendin G et al (2009). Improving protein extraction and separation methods for investigating the metaproteome of anaerobic benzene communities within sediments. Biodegradation 20: 737–750.
Biddle JF, Cardman Z, Mendlovitz H, Albert DB, Lloyd KG, Boetius A et al (2011). Anaerobic oxidation of methane at different temperature regimes in Guaymas Basin hydrothermal sediments. ISME J 6: 1018–1031.
Bose A, Rogers DR, Adams MM, Joye SB, Girguis PR . (2013). Geomicrobiological linkages between short-chain alkane consumption and sulfate reduction rates in seep sediments. Front Microbiol 4: 386.
Bowles MW, Samarkin VA, Bowles KM, Joye SB . (2011). Weak coupling between sulfate reduction and the anaerobic oxidation of methane in methane-rich seafloor sediments during ex situ incubation. Geochim Cosmochim Acta 75: 500–519.
Callaghan AV, Wawrik B, Ní Chadhain SM, Young LY, Zylstra GJ . (2008). Anaerobic alkane-degrading strain AK-01 contains two alkylsuccinate synthase genes. Biochem Biophys Res Commun 366: 142–148.
Callaghan AV, Davidova IA, Savage-Ashlock K, Parisi VA, Gieg LM, Suflita JM et al (2010). Diversity of benzyl- and alkylsuccinate synthase genes in hydrocarbon-impacted environments and enrichment cultures. Environ Sci Technol 44: 7287–7294.
Cord-Ruwisch R . (1985). A quick method for the determination of dissolved and precipitated sulfides in cultures of sulfate-reducing bacteria. J Microbiol Methods 4: 33–36.
Cox J, Mann M . (2008). MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26: 1367–1372.
Culman S, Bukowski R, Gauch H, Cadillo-Quiroz H, Buckley D . (2009). T-REX: software for the processing and analysis of T-RFLP data. BMC Bioinform 10: 171.
Dhillon A, Teske A, Dillon J, Stahl DA, Sogin ML . (2003). Molecular characterization of sulfate-reducing bacteria in the Guaymas Basin. Appl Environ Microbiol 69: 2765–2772.
Didyk BM, Simoneit BRT . (1989). Hydrothermal oil of Guaymas Basin and implications for petroleum formation mechanisms. Nature 342: 65–69.
Elsaied H, Stokes HW, Kitamura K, Kurusu Y, Kamagata Y, Maruyama A . (2011). Marine integrons containing novel integrase genes, attachment sites, attI, and associated gene cassettes in polluted sediments from Suez and Tokyo Bays. ISME J 5: 1162–1177.
Felden J, Lichtschlag A, Wenzhöfer F, de Beer D, Feseker T, Pop Ristova P et al (2013). Limitations of microbial hydrocarbon degradation at the Amon mud volcano (Nile deep-sea fan). Biogeosciences 10: 3269–3283.
Fernández-Gómez B, Richter M, Schüler M, Pinhassi J, Acinas SG, González JM et al (2013). Ecology of marine Bacteroidetes: a comparative genomics approach. ISME J 7: 1026–1037.
Geer LY, Markey SP, Kowalak JA, Wagner L, Xu M, Maynard DM et al (2004). Open mass spectrometry search algorithm. J Proteome Res 3: 958–964.
Glöckner J, Kube M, Shrestha PM, Weber M, Glöckner FO, Reinhardt R et al (2010). Phylogenetic diversity and metagenomics of candidate division OP3. Environ Microbiol 12: 1218–1229.
Grundmann O, Behrends A, Rabus R, Amann J, Halder T, Heider J et al (2008). Genes encoding the candidate enzyme for anaerobic activation of n-alkanes in the denitrifying bacterium, strain HxN1. Environ Microbiol 10: 376–385.
Gutierrez T, Singleton DR, Berry D, Yang T, Aitken MD, Teske A . (2013). Hydrocarbon-degrading bacteria enriched by the Deepwater Horizon oil spill identified by cultivation and DNA-SIP. ISME J 7: 2091–2104.
Halm H, Musat N, Lam P, Langlois R, Musat F, Peduzzi S et al (2009). Co-occurrence of denitrification and nitrogen fixation in a meromictic lake, Lake Cadagno (Switzerland). Environ Microbiol 11: 1945–1958.
Harms G, Zengler K, Rabus R, Aeckersberg F, Minz D, Rosselló-Móra R et al (1999). Anaerobic oxidation of o-xylene, m-xylene, and homologous alkylbenzenes by new types of sulfate-reducing bacteria. Appl Environ Microbiol 65: 999–1004.
Hazen TC, Dubinsky EA, DeSantis TZ, Andersen GL, Piceno YM, Singh N et al (2010). Deep-sea oil plume enriches indigenous oil-degrading bacteria. Science 330: 204–208.
Head IM, Jones DM, Roling WFM . (2006). Marine microorganisms make a meal of oil. Nat Rev Microbiol 4: 173–182.
Herbst F-A, Bahr A, Duarte M, Pieper DH, Richnow H-H, von Bergen M et al (2013). Elucidation of in situ polycyclic aromatic hydrocarbon degradation by functional metaproteomics (protein-SIP). Proteomics 13: 2910–2920.
Holler T, Widdel F, Knittel K, Amann R, Kellermann MY, Hinrichs K-U et al (2011). Thermophilic anaerobic oxidation of methane by marine microbial consortia. ISME J 5: 1946–1956.
Jaekel U, Musat N, Adam B, Kuypers M, Grundmann O, Musat F . (2012). Anaerobic degradation of propane and butane by sulfate-reducing bacteria enriched from marine hydrocarbon cold seeps. ISME J 7: 885–895.
Jehmlich N, Schmidt F, Hartwich M, von Bergen M, Richnow H-H, Vogt C . (2008). Incorporation of carbon and nitrogen atoms into proteins measured by protein-based stable isotope probing (Protein-SIP). Rapid Commun Mass Spectrom 22: 2889–2897.
Joye SB, Boetius A, Orcutt BN, Montoya JP, Schulz HN, Erickson MJ et al (2004). The anaerobic oxidation of methane and sulfate reduction in sediments from Gulf of Mexico cold seeps. Chem Geol 205: 219–238.
Junier P, Junier T, Witzel K-P . (2008). TRiFLe: a program for in silico terminal restriction fragment length polymorphism analysis with user-defined sequence sets. Appl Environ Microbiol 74: 6452–6456.
Kinnaman FS, Valentine DL, Tyler SC . (2007). Carbon and hydrogen isotope fractionation associated with the aerobic microbial oxidation of methane, ethane, propane and butane. Geochim Cosmochim Acta 71: 271–283.
Kleindienst S, Ramette A, Amann R, Knittel K . (2012). Distribution and in situ abundance of sulfate-reducing bacteria in diverse marine hydrocarbon seep sediments. Environ Microbiol 14: 2689–2710.
Kniemeyer O, Musat F, Sievert SM, Knittel K, Wilkes H, Blumenberg M et al (2007). Anaerobic oxidation of short-chain hydrocarbons by marine sulphate-reducing bacteria. Nature 449: 898–910.
Knittel K, Boetius A . (2009). Anaerobic oxidation of methane: progress with an unknown process. Annu Rev Microbiol 63: 311–334.
Knittel K, Boetius A, Lemke A, Eilers H, Lochte K, Pfannkuche O et al (2003). Activity, distribution, and diversity of sulfate reducers and other bacteria in sediments above gas hydrate (Cascadia margin, Oregon). Geomicrobiol J 20: 269–294.
Kostka JE, Prakash O, Overholt WA, Green SJ, Freyer G, Canion A et al (2011). Hydrocarbon-degrading bacteria and the bacterial community response in Gulf of Mexico beach sands impacted by the Deepwater Horizon oil spill. Appl Environ Microbiol 77: 7962–7974.
Laemmli UK . (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Liu A, Garcia-Dominguez E, Rhine ED, Young LY . (2004). A novel arsenate respiring isolate that can utilize aromatic substrates. FEMS Microbiol Ecol 48: 323–332.
Lloyd KG, Albert DB, Biddle JF, Chanton JP, Pizarro O, Teske A . (2010). Spatial structure and activity of sedimentary microbial communities underlying a Beggiatoa spp. mat in a Gulf of Mexico hydrocarbon seep. PLoS One 5: e8738.
Lösekann T, Knittel K, Nadalig T, Fuchs B, Niemann H, Boetius A et al (2007) Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby Mud Volcano, Barents Sea. Appl Environ Microbiol 73: 3348–3362.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Lueders T, Manefield M, Friedrich MW . (2004). Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol 6: 73–78.
Lueders T, Wagner B, Claus P, Friedrich MW . (2004). Stable isotope probing of rRNA and DNA reveals a dynamic methylotroph community and trophic interactions with fungi and protozoa in oxic rice field soil. Environ Microbiol 6: 60–72.
Lueders T . (2010). Stable isotope probing of hydrocarbon-degraders. In: Timmis KN, (ed) Handbook of Hydrocarbon and Lipid Microbiology vol. 5 Springer: Berlin, Heidelberg, Germany, pp 4011–4026.
Manz W, Eisenbrecher M, Neu TR, Szewzyk U . (1998). Abundance and spatial organization of gram-negative sulfate-reducing bacteria in activated sludge investigated by in situ probing with specific 16S rRNA targeted oligonucleotides. FEMS Microbiol Ecol 25: 43–61.
Mastalerz V, de Lange GJ, Dählmann A . (2009). Differential aerobic and anaerobic oxidation of hydrocarbon gases discharged at mud volcanoes in the Nile deep-sea fan. Geochim Cosmochim Acta 73: 3849–3863.
McIlroy S, Porter K, Seviour RJ, Tillett D . (2008). A simple and safe method for the simultaneous isolation of microbial RNA and DNA from problematic populations. Appl Environ Microbiol 74: 6806–6807.
Meckenstock RU, Annweiler E, Michaelis W, Richnow HH, Schink B . (2000). Anaerobic naphthalene degradation by a sulfate-reducing enrichment culture. Appl Environ Microbiol 66: 2743–2747.
Morasch B, Schink B, Tebbe CC, Meckenstock RU . (2004). Degradation of o-xylene and m-xylene by a novel sulfate-reducer belonging to the genus Desulfotomaculum. Arch Microbiol 181: 407–417.
Musat F, Widdel F . (2008). Anaerobic degradation of benzene by a marine sulfate-reducing enrichment culture, and cell hybridization of the dominant phylotype. Environ Microbiol 10: 10–19.
Nunoura T, Takaki Y, Kazama H, Hirai M, Ashi J, Imachi H et al (2012). Microbial diversity in deep-sea methane seep sediments presented by SSU rRNA gene tag sequencing. Microbes Environment 27: 382–390.
Omoregie EO, Mastalerz V, de Lange G, Straub KL, Kappler A, Røy H et al (2008). Biogeochemistry and community composition of iron- and sulfur-precipitating microbial mats at the Chefren mud volcano (Nile Deep Sea fan, Eastern Mediterranean). Appl Environ Microbiol 74: 3198–3215.
Omoregie EO, Niemann H, Mastalerz V, de Lange GJ, Stadnitskaia A, Mascle J et al (2009). Microbial methane oxidation and sulfate reduction at cold seeps of the deep Eastern Mediterranean Sea. Mar Geol 261: 114–127.
Orcutt BN, Joye SB, Kleindienst S, Knittel K, Ramette A, Reitz A et al (2010). Impact of natural oil and higher hydrocarbons on microbial diversity, distribution, and activity in Gulf of Mexico cold-seep sediments. Deep-Sea Res Part II 57: 2008–2021.
Orphan VJ, Hinrichs K-U, Ussler W, Paull CK, Taylor LT, Sylva SP et al (2001). Comparative analysis of methane-oxidizing archaea and sulfate-reducing bacteria in anoxic marine sediments. Appl Environ Microbiol 67: 1922–1934.
Pachiadaki M, Kallionaki A, Dählmann A, De Lange G, Kormas K . (2011). Diversity and spatial distribution of prokaryotic communities along a sediment vertical profile of a deep-sea mud volcano. Microb Ecol 62: 655–668.
Perkins DN, Pappin DJC, Creasy DM, Cottrell JS . (1999). Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20: 3551–3567.
Pilloni G, von Netzer F, Engel M, Lueders T . (2011). Electron acceptor-dependent identification of key anaerobic toluene degraders at a tar-oil-contaminated aquifer by Pyro-SIP. FEMS Microbiol Ecol 78: 165–175.
Pilloni G, Granitsiotis MS, Engel M, Lueders T . (2012). Testing the limits of 454 pyrotag sequencing: reproducibility, quantitative assessment and comparison to T-RFLP fingerprinting of aquifer microbes. PLoS One 7: e40467.
Prince RC, Gramain A, McGenity TJ . (2010). Prokaryotic hydrocarbon degraders. In: Timmis KN, (ed) Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin Heidelberg, Germany, pp 1669–1692.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41: D590–D596.
Quistad SD, Valentine DL . (2011). Anaerobic propane oxidation in marine hydrocarbon seep sediments. Geochim Cosmochim Acta 75: 2159–2169.
Rabus R, Wilkes H, Behrends A, Armstroff A, Fischer T, Pierik AJ et al (2001). Anaerobic initial reaction of n-alkanes in a denitrifying bacterium: evidence for (1-methylpentyl)succinate as initial product and for involvement of an organic radical in n-hexane metabolism. J Bacteriol 183: 1707–1715.
Rabus R, Hansen TA, Widdel F . (2006). Dissimilatory sulfate- and sulfur-reducing prokaryotes. In: Dworkin M, Falkow S, Rosenberg E, (eds) The Prokaryotes. A Handbook on the Biology of Bacteria. Springer: Berlin, Germany, pp 659–768.
Redmond MC, Valentine DL, Sessions AL . (2010). Identification of novel methane-, ethane-, and propane-oxidizing bacteria at marine hydrocarbon seeps by stable isotope probing. Appl Environ Microbiol 76: 6412–6422.
Redmond MC, Valentine DL . (2012). Natural gas and temperature structured a microbial community response to the Deepwater Horizon oil spill. Proc Natl Acad Sci USA 109: 20292–20297.
Reeburgh WS . (2007). Oceanic methane biogeochemistry. Chem Rev 107: 486–513.
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng J-F et al (2013). Insights into the phylogeny and coding potential of microbial dark matter. Nature 499: 431–437.
Rotaru A-E, Schauer R, Probian C, Mußmann M, Harder J . (2012). Visualization of candidate division OP3 cocci in limonene-degrading methanogenic cultures. J Microbiol Biotechnol 22: 457–461.
Rubin-Blum M, Antler G, Turchyn AV, Tsadok R, Goodman-Tchernov BN, Shemesh E et al (2014). Hydrocarbon-related microbial processes in the deep sedimentsof the Eastern Mediterranean Levantine Basin. FEMS Microbiol Ecol 3: 780–796.
Ruff SE, Arnds J, Knittel K, Amann R, Wegener G, Ramette A et al (2013). Microbial communities of deep-sea methane seeps at Hikurangi continental margin (New Zealand). PLoS One 8: e72627.
Savage KN, Krumholz LR, Gieg LM, Parisi VA, Suflita JM, Allen J et al (2010). Biodegradation of low-molecular-weight alkanes under mesophilic, sulfate-reducing conditions: metabolic intermediates and community patterns. FEMS Microbiol Ecol 72: 485–495.
Schauder R, Eikmanns B, Thauer RK, Widdel F, Fuchs G . (1986). Acetate oxidation to CO2 in anaerobic bacteria via a novel pathway not involving reactions of the citric acid cycle. Arch Microbiol 145: 162–172.
Schramm A, Fuchs BM, Nielsen JL, Tonolla M, Stahl DA . (2002). Fluorescence in situ hybridization of 16S rRNA gene clones (Clone-FISH) for probe validation and screening of clone libraries. Environ Microbiol 4: 713–720.
Schreiber L, Holler T, Knittel K, Meyerdierks A, Amann R . (2010). Identification of the dominant sulfate-reducing bacterial partner of anaerobic methanotrophs of the ANME-2 clade. Environ Microbiol 12: 2327–2340.
Shennan JL . (2006). Utilisation of C2–C4 gaseous hydrocarbons and isoprene by microorganisms. J Chem Technol Biotechnol 81: 237–256.
So CM, Young LY . (1999). Initial reactions in anaerobic alkane degradation by a sulfate reducer, strain AK-01. Appl Environ Microbiol 65: 5532–5540.
Sturm M, Bertsch A, Gropl C, Hildebrandt A, Hussong R, Lange E et al (2008). An open-source software framework for mass spectrometry. BMC Bioinformatics 9: 163.
Teske A, Hinrichs K-U, Edgcomb V, de Vera Gomez A, Kysela D, Sylva SP et al (2002). Microbial diversity of hydrothermal sediments in the Guaymas Basin: evidence for anaerobic methanotrophic communities. Appl Environ Microbiol 68: 1994–2007.
von Netzer F, Pilloni G, Kleindienst S, Krüger M, Knittel K, Gründger F et al (2013). Enhanced gene detection assays for fumarate-adding enzymes allow uncovering of anaerobic hydrocarbon degraders in terrestrial and marine systems. Appl Environ Microbiol 79: 543–552.
Widdel F, Bak F . (1992). Gram-negative mesophilic sulfate-reducing bacteria. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer KH, (eds) The Prokaryotes 2nd edn. Springer-Verlag: New York, NY, USA, pp 3352–3378.
Widdel F, Knittel K, Galushko A . (2010). Anaerobic hydrocarbon-degrading microorganisms: an overview. in: Timmis KN, McGenity T, van der Meer JR, de Lorenzo V, (eds) Handbook of Hydrocarbon and Lipid Microbiology. Springer: Berlin, Heidelberg, Germany, pp 1997–2021.
Wilkes H, Rabus R, Fischer T, Armstroff A, Behrends A, Widdel F . (2002). Anaerobic degradation of n-hexane in a denitrifying bacterium: further degradation of the initial intermediate (1-methylpentyl)succinate via C-skeleton rearrangement. Arch Microbiol 177: 235–243.
Winderl C, Penning H, Netzer FV, Meckenstock RU, Lueders T . (2010). DNA-SIP identifies sulfate-reducing Clostridia as important toluene degraders in tar-oil-contaminated aquifer sediment. ISME J 4: 1314–1325.
Yarza P, Ludwig W, Euzéby J, Amann R, Schleifer K-H, Glöckner FO et al (2010). Update of the all-species living tree project based on 16S and 23S rRNA sequence analyses. Syst Appl Microbiol 33: 291–299.
Zhang F, She Y, Zheng Y, Zhou Z, Kong S, Hou D . (2010). Molecular biologic techniques applied to the microbial prospecting of oil and gas in the Ban 876 gas and oil field in China. Appl Microbiol Biotechnol 86: 1183–1194.
Zhang Y, Maignien L, Zhao X, Wang F, Boon N . (2011). Enrichment of a microbial community performing anaerobic oxidation of methane in a continuous high-pressure bioreactor. BMC Microbiol 11: 137.
Zhou J, Bruns MA, Tiedje JM . (1996). DNA recovery from soils of diverse composition. Appl Environ Microbiol 62: 316–322.
Acknowledgements
We acknowledge the officers, crews and shipboard scientific parties of the cruises MSM13-3 and AT15-56. Sampling was funded by the EU project HERMIONE (FP6; contract number 226354) and the DFG (METEOR/MERIAN SPP, MSM13-3 awarded to Antje Boetius). We especially acknowledge Antje Boetius, Janine Felden, Andreas Teske and Gunter Wegener for sampling, Gaute Lavik, Tim Kalvelage, Katrin Hörmann, Giovanni Pilloni and Marion Engel for analytical support and Barbara MacGregor for sharing RNA extraction protocols. This study was funded by grants to KK, TL and MvB within the Priority Program SPP1319 ‘Transformation of hydrocarbons under anoxic conditions: from molecular to global scale’ of the DFG. Further support for this work was provided by the EU-MAGICPAH (FP7-KBBE-2009-245226) project, the Helmholtz Society and the Max Planck Society.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Kleindienst, S., Herbst, FA., Stagars, M. et al. Diverse sulfate-reducing bacteria of the Desulfosarcina/Desulfococcus clade are the key alkane degraders at marine seeps. ISME J 8, 2029–2044 (2014). https://doi.org/10.1038/ismej.2014.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.51
Keywords
This article is cited by
-
Microbial remediation of oil-contaminated shorelines: a review
Environmental Science and Pollution Research (2023)
-
Microbiome and metagenomic analysis of Lake Hillier Australia reveals pigment-rich polyextremophiles and wide-ranging metabolic adaptations
Environmental Microbiome (2022)
-
Subsurface hydrocarbon degradation strategies in low- and high-sulfate coal seam communities identified with activity-based metagenomics
npj Biofilms and Microbiomes (2022)
-
Phylogenetically and catabolically diverse diazotrophs reside in deep-sea cold seep sediments
Nature Communications (2022)
-
Comparative study of gut microbiota from decomposer fauna in household composter using metataxonomic approach
Archives of Microbiology (2022)