Abstract
Dysregulated immune responses to gut microbes are central to inflammatory bowel disease (IBD), and gut microbial activity can fuel chronic inflammation. Examining how IBD-directed therapies influence gut microbiomes may identify microbial community features integral to mitigating disease and maintaining health. However, IBD patients often receive multiple treatments during disease flares, confounding such analyses. Preclinical models of IBD with well-defined disease courses and opportunities for controlled treatment exposures provide a valuable solution. Here, we surveyed the gut microbiome of the T-bet−/− Rag2−/− mouse model of colitis during active disease and treatment-induced remission. Microbial features modified among these conditions included altered potential for carbohydrate and energy metabolism and bacterial pathogenesis, specifically cell motility and signal transduction pathways. We also observed an increased capacity for xenobiotics metabolism, including benzoate degradation, a pathway linking host adrenergic stress with enhanced bacterial virulence, and found decreased levels of fecal dopamine in active colitis. When transferred to gnotobiotic mice, gut microbiomes from mice with active disease versus treatment-induced remission elicited varying degrees of colitis. Thus, our study provides insight into specific microbial clades and pathways associated with health, active disease and treatment interventions in a mouse model of colitis.
Similar content being viewed by others
Introduction
Inflammatory bowel disease (IBD) is linked to alterations in gut microbial communities and dysregulated mucosal immune responses (Ott et al., 2004; Frank et al., 2007; Sokol et al., 2008; Packey and Sartor, 2009; Frank et al., 2011; Erickson et al., 2012; Morgan et al., 2012; Presley et al., 2012). Management of IBD has relied on nonspecific immunosuppressive therapies, agents targeting pro-inflammatory pathways, antibiotics and more recently, probiotics (Dethlefsen et al., 2008; Engel and Neurath, 2010; Hill et al., 2010; Jakobsson et al., 2010; Manichanh et al., 2010; Pineton de Chambrun and Sandborn, 2012; Ubeda and Pamer, 2012). However, many IBD-directed therapies are not effective in all patients and some carry a high risk of complications and side effects. How these therapies perturb the aggregate ecology and biomolecular environment of the gut microbiome is poorly understood. Thus, determining what aspects of the gut microbiome structurally and functionally change in active colitis and treatment-induced remission may provide improved therapeutic targets.
Mouse models of IBD provide an opportunity to identify microbes and microbial pathways involved in IBD and host–microbiota responses to therapies, which can be difficult to discern in humans given their genetic diversity and variability in environmental and treatment exposures. Deciphering which gut microbial community members and functions are similarly or differentially modulated by therapeutic interventions has important implications for IBD management and may facilitate customization of existing and future therapies.
TRUC (T-bet−/− Rag2−/− ulcerative colitis) mice develop an early onset, spontaneous ulcerative colitis (UC) due to genetic defects in innate and adaptive immunity (Garrett et al., 2007). TRUC pathogenesis is driven, in part, by tumor necrosis factor-alpha (TNF-α) (Garrett et al., 2007) and is dependent on the gut microbiome, as germ-free (GF) TRUC mice do not develop disease (Garrett et al., 2010). In the presence of an endogenous microbiota, specific gut microbes have been shown to trigger TRUC colitis, including Klebsiella pneumoniae and Proteus mirabilis (Garrett et al., 2010) – and more recently, Helicobacter typhlonius (Powell et al., 2012). Like human IBD, TRUC colitis is responsive to immunomodulatories that dampen pro-inflammatory responses to gut microbes (Garrett et al., 2007) and to oral antibiotics (Garrett et al., 2010). Daily consumption of a fermented milk product (FMP) has also been shown to mitigate TRUC colitis (Veiga et al., 2010). Provided that antibiotics, immunomodulatories and dietary interventions mechanistically act on different aspects of the host–microbiota interface to ameliorate TRUC colitis, they likely differentially modify the gut microbiome. Thus, TRUC mice offer a tractable model for evaluating gut microbiome contributions to colonic inflammatory pathogenesis and for characterizing gut microbiome responses to therapeutic interventions.
Here, we investigated the effects of diverse treatment interventions on host disease status and on gut microbiome structure and function in TRUC mice. Using 16S ribosomal RNA (rRNA) gene surveys, we analyzed gut microbial communities following treatment with: oral antibiotics (gentamicin, metronidazole or vancomycin), immunomodulatories (TNF-α neutralizing antibodies (anti-TNF-α) or infusion of regulatory T cells (TRegs)) and dietary interventions (FMP or non-fermented milk control (MC)). In addition to examining taxonomic shifts associated with treatment exposure, we used gnotobiotic TRUC mice to test the inflammatory phenotypes of gut microbiomes from treatment groups in vivo. To functionally interrogate the metabolic potential of gut microbiomes associated with active colitis versus treatment-induced remission, we performed in silico analysis of 16S rRNA gene sequences coupled with reference genomes to infer microbial community function. We found that TRUC gut microbiomes with active colitis had a reduced potential for both carbohydrate and energy metabolism and an enhanced potential for flagellar assembly, tetrathionate respiration and benzoate degradation. Collectively, our study identified microbes and microbial functions underlying colitis-associated dysbiosis that were similarly or differentially modulated by host- and microbiota-targeted therapies, illustrating the potential for therapeutic manipulation of the gut microbiome in colitis.
Materials and methods
Animal husbandry
Specified-pathogen-free (SPF) BALB/c T-bet−/− Rag2−/− mice have been described (Garrett et al., 2007), see Supplementary Methods.
Treatment interventions
Antibiotics
Mice were treated with antibiotics dissolved in their autoclaved drinking water: gentamicin (2 g l−1; Cellgro, Manassas, VA, USA), metronidazole (1 g l−1; Sigma-Aldrich, St Louis, MO, USA), vancomycin (500 mg l−1; Sigma-Aldrich) (Garrett et al., 2010).
Anti-TNF-α injections
Mice were injected (15 mg kg−1) with a hamster anti-mouse TNF-α neutralizing antibody (clone TN4-19.12) (Bio X Cell, West Lebanon, NH, USA) weekly for 4 weeks starting at 4 weeks of age (Garrett et al., 2007).
TReg cell infusion
FACS-sorted peripheral lymph node CD4+CD62LhiCD25+ cells (75 000 cells per mouse) were intravenously injected at 4 weeks of age (Garrett et al., 2007).
Dietary interventions
For details on the FMP and MC, see Supplementary Methods.
Histology
Upon sacrifice, colons were resected, fixed in 4% paraformaldehyde (Sigma-Aldrich) and embedded in paraffin. Five-micron sections were H&E-stained and evaluated in a blinded fashion for epithelial hyperplasia (0–3), epithelial injury (0–3), polymorphonuclear infiltration (0–3) and mononuclear infiltration (0–3), these indices were summed to generate the histologic colitis score (Garrett et al., 2007).
Microbial DNA preparation from stool and mesenteric lymph nodes (MLNs)
Samples were collected and homogenized in RNAlater (Ambion, Grand Island, NY, USA), held at 4 °C overnight and stored at −80 °C before processing. For details on DNA extraction, see Supplementary Methods.
16S rRNA gene survey analysis of gut microbial communities
For details on 16S rRNA gene amplification, 454 pyrosequencing, sequence processing, operational taxonomic unit (OTU) selection, microbial composition and community structure analyses, metagenome inference and metabolic pathway reconstruction, and microbial biomarker discovery, see Supplementary Methods.
Whole metagenome shotgun (WMS) sequence analysis
For details on Illumina shotgun sequencing, sequence processing and microbial gene and pathway abundance analyses, see Supplementary Methods.
Dopamine ELISA
To measure dopamine levels, the DOP Research ELISA (Labor Diagnostika Nord, Nordhorn, Germany) was employed following the manufacturer’s instructions. For details, see Supplementary Methods.
Gnotobiotic mouse experiments
GF Balb/c T-bet−/− Rag2−/− mice were maintained on autoclavable Mouse Breeder Diet 5021 (LabDiet, St Louis, MO, USA) and housed in gnotobiotic isolators in the Harvard Digestive Disease Center Facility. For details, see Supplementary Methods.
RT-qPCR for FMP strains
For detailed RT-qPCR (real-time quantitative PCR) methods and primers, see Supplementary Methods.
Statistical analysis
Significant P-values associated with microbial clades and functions identified by LEfSe (Linear Discriminant Analysis with Effect Size) were corrected for multiple hypothesis testing using the Benjamini and Hochberg false discovery rate correction (Benjamini and Hochberg, 1995). All P-values and q-values are in Supplementary Table S3. Other statistical tests for significance were performed in GraphPad Prism version 5.0b for Mac OS X (GraphPad Software, La Jolla, CA, USA). All averages are mean±SEM.
Accession number
Sequences have been deposited on MG-RAST under project ID 6698.
Results
Assessing gut microbiome structure during active disease and treatment-induced remission with 16S rRNA gene surveys
TRUC mice manifest signs of colitis prior to 3 weeks of age, which increase in severity over time (Garrett et al., 2007). At 3 weeks of age, mice were randomized to the following groups: antibiotics (gentamicin, metronidazole or vancomycin), immunomodulatories (anti-TNF-α or TRegs), dietary interventions (FMP or MC), or untreated (sham) control, and began treatment 1 week later. Stool samples were collected from mice at 4 weeks of age (baseline/pre-intervention) and directly upon completion of the intervention at 8 weeks of age (post-intervention). The study design and histology-based colitis scores to relate gut microbiome changes to host disease status are shown (Figures 1a and b).
Experimental design and influence of interventions on the gut microbiome. (a) Study experimental schema. (b) Histologic colitis scores. Symbols represent individual mice. Error bars indicate mean±SEM. Colitis scores >2 indicate active colitis and scores ⩽2 remission. Sham, untreated, handling control; Gent, gentamicin; Metro, metronidazole; Vanco, vancomycin; Immunomods, anti-TNF-α or TRegs; MC, non-fermented milk control; FMP, fermented milk product; Diet, dietary intervention with FMP or MC in addition to ad libitum chow. (c) PCoA using unweighted UniFrac distances of gut microbial communities obtained from stool samples collected at baseline (pre-intervention) and upon treatment completion (post-intervention). The first two principal coordinates (PC) from the PCoA are plotted. Symbols represent data from individual mice, color-coded by the indicated metadata. (d) Gut microbiomes were clustered by similarity using the UPGMA clustering algorithm on the unweighted UniFrac distances. Samples from individual mice were clustered by the indicated intervention class (outside ring) or by the specific treatment (inside ring). (e) Phylum-level phylogenetic classification of 16S rRNA gene sequences. Bars represent mean relative abundances for each pre- or post-intervention group.
In total, 1 014 181 quality-filtered 16S rRNA gene sequences were obtained from 152 stool samples (6672±335 reads per sample; Supplementary Table S1). Reads were binned de novo into approximately species-level OTUs at ⩾97% sequence similarity (586±30 OTUs per sample; Supplementary Table S1). Microbiome analysis tools included: QIIME (Quantitative Insights Into Microbial Ecology) for sequence processing (Caporaso et al., 2010), PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) for metagenome inference (Langille et al., 2013), HUMAnN (The HMP Unified Metabolic Analysis Network) for functional profiling (Abubucker et al., 2012), and LEfSe for univariate contrasts (Segata et al., 2011) (Supplementary Figure S1).
Effects of treatment interventions on gut microbiome composition
Principal coordinate analysis (PCoA) of the unweighted UniFrac distances (Lozupone et al., 2011) revealed that baseline (pre-intervention) communities clustered together (Figure 1c, far left panel). Comparing post-treatment stool, samples tended to separate by type of intervention, with distinct clusters observed for gut microbiomes exposed to antibiotics, immunomodulatories and dietary interventions (Figure 1c, middle right panel). PCoA of the weighted UniFrac distances demonstrated similar trends (Supplementary Figure S2), as did a single-dimensional UPGMA (unweighted pair group method with arithmetic mean) hierarchical clustering of these samples (Figure 1d).
The 16S rRNA gene survey data revealed the greatest variation in gut microbial community composition with antibiotic exposure, particularly for gentamicin and vancomycin (Figure 1c, far right panel). In contrast, immunomodulatories and dietary interventions (Figure 1c, right panels) induced changes of smaller effect size. Shifts in phylum-level relative abundances of samples collected at baseline and upon intervention completion also confirmed these high taxonomic level gut microbial response patterns (Figure 1e). Both in terms of microbial presence/absence and in overall clade abundances, immunomodulatories, dietary interventions and individual antibiotics each perturbed the gut microbiome in distinctive ways and to varying extents, with antibiotics having the most substantial effect on microbial community structure.
Antibiotic-driven microbial community shifts may be influenced by early-life exposures and are associated with specific clade responses
Despite mitigating some IBD flares (Wang et al., 2012), antibiotics disrupt gut microbial community structure and diversity, pushing the microbiome to an alternate state and potentially prolonging susceptibility to pathogens (Dethlefsen et al., 2008; Hill et al., 2010; Jakobsson et al., 2010; Manichanh et al., 2010; Ubeda and Pamer, 2012). We investigated the extent to which three oral antibiotics perturb the gut microbiomes of TRUC mice: metronidazole, gentamicin and vancomycin. Gentamicin and metronidazole (the latter used clinically to treat IBD and Clostridium difficile colitis) both ameliorated TRUC colitis (Figure 1b). However, vancomycin, also employed clinically to treat C. difficile colitis, did not attenuate TRUC colitis (Figure 1b). Given that gentamicin, metronidazole and vancomycin have disparate modes of action and effects on TRUC colitis, we characterized shifts in community composition and diversity following treatment exposure and identified which members of the gut microbiome were similarly or differentially modulated by each antibiotic.
In contrast to baseline microbial communities, which consisted mostly of Firmicutes followed in decreasing order of relative abundance by Bacteroidetes, Proteobacteria, Actinobacteria, Deferribacteres and TM7, each antibiotic had marked effects on gut microbiome composition (Figure 2a). Gentamicin exposure led to a dominance of Bacteroidetes (99.6±0.1%, relative abundance), particularly Bacteroidaceae, whereas vancomycin promoted an expansion of Proteobacteria – levels increased from 3.4±1.8% to 47.0±10.8%. Responses within the metronidazole and vancomycin treatment groups were distinct in terms of overall community composition (Figure 1c, far right panel) and by relative abundance analysis (Figure 2a). To address if host or environmental factors affect gut microbiome responses to these antibiotics, we evaluated whether treatment, caging or legacy effects (influence of parental transmission on microbial composition) corresponded to the greatest degree of microbial variation. The greatest source of variation was not cage effects – observed in some mouse gut microbiome studies and attributed to cohabitation and coprophagia (Hildebrand et al., 2013). Rather, microbial communities segregated first by parental origin and second by the antibiotic administered (metronidazole versus vancomycin) (Figures 2b and c; Supplementary Figure S3a), suggesting that community structure may be influenced by early-life exposures and by post-weaning interactions with other environmental exposures, respectively.
Antibiotic-driven microbial community shifts may be influenced by early-life exposures and are associated with specific clade responses. (a) Family-level phylogenetic classification of 16S rRNA gene sequences from stool samples collected pre- and post-intervention. Bars represent relative abundances of samples from individual mice. Labels indicate families with average relative abundances ⩾1% in at least one pre- or post-intervention group. Remaining families and reads assigned to higher level taxonomies were binned together in their associated phylum as ‘other’ or ‘unclassified’ (uncl.), respectively. (b) PCoA plots of the unweighted UniFrac distances of post-intervention stool samples from metronidazole- (n=10) and vancomycin (n=10)-treated mice. The first two PCs from the PCoA are plotted. Symbols represent data from individual mice, color-coded by the indicated metadata. For caging, M, metronidazole-treated; V, vancomycin-treated. (c) UPGMA clustering algorithm on the unweighted UniFrac distances of samples, color-coded by breeding pair. The letter and number assigned to the sire and dam, respectively, correspond to the PCoA plots in b. (d) Differentially abundant microbial clades in stool from mice with active colitis versus remission. Symbols represent data from individual mice from three independent experiments. Error bars indicate±SEM. Kruskal–Wallis test with Dunn’s multiple comparison test: * or +P<0.05; ** or ++P<0.01; and *** or +++P<0.001.
To identify gut microbiome responses associated with antibiotics and legacy effects, we determined pre- and post-treatment microbial clade differences in progeny of breeding pairs using LEfSe. Mice enriched in Actinobacteria and Bacteroides (BP-II) (Supplementary Figure S3b) at baseline became further enriched in these microbial clades with metronidazole treatment (Figure 2a, Metro-6-8,10). Because metronidazole selectively targets anaerobic bacteria, it may enable aerotolerant members of the Actinobacteria and Bacteroidetes phyla to bloom. Mice enriched in δ-proteobacteria (BP-III) (Supplementary Figure S3c) at baseline became further enriched in Proteobacteria with vancomycin treatment (Figure 2a, Vanco-1-5). This vancomycin-induced expansion of Proteobacteria has been observed previously (Carvalho et al., 2012; Ubeda and Pamer, 2012). Thus, identifying microbial clades enriched at baseline may predict gut microbial responses to antibiotics and may be useful for tailoring antibiotic treatment.
Alterations of gut microbial communities following gentamicin, metronidazole and vancomycin treatment coupled with their disparate effects on colitis provided an opportunity to identify which gut microbiome members are preferentially targeted by a given antibiotic and probe for members that are consistently associated with disease pathogenesis. Using LEfSe, we performed an all-against-all multi-class comparison of gut microbiome samples from antibiotic-treated mice to identify clades specifically modulated by each antibiotic. Although gentamicin-treated communities were completely dominated by Bacteroidetes, the remaining Firmicutes were enriched in Erysipelotrichi (Supplementary Figure S4a). Metronidazole treatment was associated with enrichments in Firmicutes and unclassified bacteria (Supplementary Figure S4a). In contrast with gentamicin and metronidazole, vancomycin treatment led to a significant expansion of γ-proteobacteria and ε-proteobacteria, including Escherichia and Helicobacter (Supplementary Figure S4a), which have been associated with intestinal inflammatory pathogenesis (Mukhopadhya et al., 2012).
To explore microbial biomarkers of active colitis, we looked for clades consistently reduced in gentamicin- and metronidazole-treated communities but augmented in vancomycin and sham communities. There were significant enrichments across three microbial lineages in colitogenic gut microbiomes: Mucispirillum, Desulfovibrio and Helicobacteraceae (Figure 2d; Supplementary Figure S4b), all of which discriminated between active colitis versus remission. Other studies have pointed to their opportunistic nature given their putative capacity to degrade mucin (Mucispirillum) (Robertson et al., 2005; Berry et al., 2012) and during active inflammation to produce high levels of hydrogen sulfide (Desulfovibrio) (Carbonero et al., 2012) and ammonia (Helicobacteraceae) (Hansen et al., 2011), which may further fuel inflammation.
Immunomodulatories alter low abundance communities of the gut microbiome and drive distinct clade responses
Use of immunomodulatories in IBD has increased given their longer disease-remission periods and fewer side effects as compared with corticosteroids (Yanai and Hanauer, 2011; Pineton de Chambrun and Sandborn, 2012). However, knowledge of the effects of specific immunomodulatories on gut microbiomes is limited (Engel and Neurath, 2010). Because TRUC colitis is driven, in part, by dysregulated colonic TNF-α production, which can be ameliorated with TNF-α neutralizing antibodies or infusion of immunosuppressive TRegs (Garrett et al., 2007), we utilized TRUC mice to unravel the effects of TNF-α-directed therapies on a colitogenic gut microbiome.
To determine the extent to which gut microbiomes are affected by immunomodulatories, we inspected exclusive and shared species-level phylotypes (OTUs) among anti-TNF-α injected, TReg infused and untreated mice. In all, 1 611 out of 4 420 total OTUs (36.5%) identified in immunomodulatory-treated and untreated mice were shared (Figure 3a, left panel). Shared OTUs were mostly more abundant species – 240 306 out of 265 344 (90.6%) of all the sequences present across samples – whereas OTUs unique to each group were mostly low abundance species (Figure 3a, right panel). TReg-infused mice had the largest number of unique species (17.3%) compared to anti-TNF-α treated (10.3%) and untreated (10.4%) mice (Figure 3a, left panel), suggesting that immunomodulatories, particularly TRegs, modulate low abundance gut microbiome members as compared to antibiotics, in agreement with the smaller effect size of shifts in community structure (Figure 1c, right panels).
Immunomodulatories influence low abundance gut microbial community members and drive distinct microbial responses. (a) Left panel: Venn diagram of exclusive and shared species-level phylotypes (non-singleton OTUs, at ⩾97% sequence identity) in anti-TNF-α injected (n=10), TReg infused (n=10) or sham (untreated; n=12) mice. In total, 4420 OTUs were present across samples. Right panel: number of 16S rRNA gene sequences in each of the indicated segments of the Venn diagram. In total, 265 344 sequences were present across samples. (b) Differentially abundant microbial clades in stool from immunomodulatory-treated (anti-TNF-α or TRegs; n=20) versus sham (n=12) mice. (c) Differentially abundant microbial clades in stool from anti-TNF-α-(n=10) versus TReg (n=10)-treated mice. For cladograms, white circles represent non-significant microbial clades.
As this argued for a focus on clade abundances rather than microbial presence/absence, we identified which microbial clades were consistently modified by immunomodulatories using LEfSe. With immunomodulatories, gut microbiomes were most significantly enriched in Actinobacteria, including Corynebacterium, as well as, Bacillales – specifically the genus Staphylococcus (Figure 3b; Supplementary Figure S5a). The gut microbiomes of anti-TNF-α-treated mice were enriched in Staphylococcus, in contrast with that of TReg-infused mice, which were most significantly enriched in Aerococcaceae (Figure 3c; Supplementary Figure S5b). Examining shared and differential taxonomic responses illustrated that Staphylococcus expanded with both immunomodulatories, however, to a greater extent following treatment with anti-TNF-α. Helicobacter spp. were also enriched in immunomodulatory-treated versus sham mice and reflect the presence of Helicobacter ganmani – a species found in mice and humans but not associated with colitis (Robertson et al., 2001; Tolia et al., 2004). Although treatment with anti-TNF-α and TRegs resulted in colitis remission, these immunomodulatories drove specific microbial responses. These findings suggest that gut microbiomes may distinguish and differentially respond to distinct host-directed therapies.
FMP consumption has subtle but distinct effects on the gut microbiome
Lactic acid-producing bacteria (LAB) within FMPs can improve gut homeostasis by providing microbes with beneficial functions to the host (Nanau and Neuman, 2012; Veerappan et al., 2012). LAB have shown promise in IBD management, given their ability to promote anti-inflammatory immune responses without inducing severe side effects in IBD patients (Veerappan et al., 2012). However, little is known of how LAB modify the gut microbiome in IBD in relation to other therapies (Rijkers et al., 2010). Previous studies in TRUC mice have shown that colitis can be ameliorated with daily consumption of a LAB-containing FMP consisting of Bifidobacterium animalis subsp. lactis (CNCM I-2494), Streptococcus thermophilus (CNCM I-1630), two strains of Lactobacillus delbrueckii subsp. bulgaricus (CNCM I-1632 and CNCM I-1519) and Lactococcus lactis subsp. cremoris (CNCM I-1631) (Veiga et al., 2010) (Figure 1b). In contrast, MC administration was less effective in attenuating colitis (Veiga et al., 2010) (Figure 1b). To determine the differential effects of the FMP and MC on the gut microbiome, we orally instilled either product daily for 4 weeks.
Bifidobacteriaceae and Coriobacteriaceae have been associated with human colonic health and IBD remission (Morgan et al., 2012; Papa et al., 2012) and were enriched with FMP treatment (Figure 4a; Supplementary Figure S6a). The observed increase in Bifidobacteriaceae may reflect its presence in the FMP or an FMP-mediated expansion of endogenous bifidobacteria. Similar to earlier studies, we found an increase in lactate-consuming bacteria with FMP, such as Desulfovibrionaceae, that may be associated with elevated levels of the electron donor lactate produced by the dietary LAB in the FMP (Veiga et al., 2010). Differential clade responses between FMP- and MC-fed mice included enrichments of Lactobacillus and Streptococcus in the MC group, which may have been secondary to the presence of lactose and absence of LAB in the MC product (Figure 4b; Supplementary Figures S6b and c). Proteobacteria were enriched with FMP – this finding reflects the presence of Helicobacter spp., specifically H. ganmani, as was observed with immunomodulatories (Figures 3b and 4b, Supplementary Figure S6c). These results point to administration of FMP having subtle but distinctive effects on the gut microbiome, which facilitate gut microbial community changes that directly or indirectly ameliorate colitis.
A FMP influences the microbial communities of the gut and MLNs. (a) Differentially abundant microbial clades in stool collected before and after FMP (n=10). (b) Differentially abundant microbial clades in stool after FMP (n=10) versus MC (n=10). (c) PCoA plots of the unweighted UniFrac distances of post-intervention stool samples (FMP, n=10; MC, n=10) and MLNs (FMP, n=21; MC, n=16; 5 MLNs/mouse). The first two PCs from the PCoA are plotted. Symbols represent data from individual mice, color-coded by the indicated metadata. (d) Phylum-level phylogenetic classification of 16S rRNA gene sequences from pre-intervention (n=20) and post-intervention stool samples (FMP, n=10; MC, n=10) and post-intervention MLNs (FMP, n=21; MC, n=16; 5 MLNs/mouse). Pie charts represent the mean relative abundances of phyla across mice from each group. (e) Differentially abundant microbial clades in post-intervention samples from stool (FMP, n=10; MC, n=10) versus MLNs (FMP, n=21; MC, n=16; 5 MLNs/mouse). (f) Differentially abundant microbial clades in post-intervention MLNs of FMP- (n=21) versus MC (n=16)-fed mice. *, aerotolerant genera;+, genera shared between MLNs and deep colonic crypt communities (Pédron et al., 2012). For cladograms, white circles represent non-significant microbial clades.
FMP administration influences the mucosal immune system by modulating microbial communities trafficked to the MLNs
Given the subtle clade differences in stool observed with the FMP, we questioned whether there were greater changes in other gut-associated microbial communities, like the mesenteric lymph nodes (MLNs). MLNs function as a ‘firewall’ that prevents live gut microbes from reaching the systemic immune system (Macpherson and Smith, 2006; Eberl and Lochner, 2009). Microbial DNA was sequenced from pooled MLNs (n=5 MLNs per mouse) of individual FMP- and MC-fed mice. In total, 288 500 quality-filtered 16S rRNA gene sequences were obtained with an average of 7797±5728 reads per sample. Comparing stool and MLN microbial communities, we observed that the major variation is explained by sampling site (biogeography) (Figure 4c; Supplementary Figure S7).
To identify which microbes are preferentially sampled by the host mucosal immune system or otherwise trafficked through the lymphatics to the MLNs, we examined taxonomic differences between microbiomes from stool and MLNs. We observed greater differences between MLN microbial communities than between stool communities with administration of FMP versus MC, particularly for the ratio of Firmicutes to Bacteroidetes and the proportion of total Proteobacteria (Figure 4d). This suggests that FMP exposure might affect which gut microbes reach the MLNs and consequently shape host immune responses. Microbial communities of MLNs were highly enriched in aerobes and facultative anaerobes (Figure 4e). TRUC MLNs compositionally resembled deep colonic crypt communities (Pédron et al., 2012) with both being enriched in aerotolerant clades that are rare in stool (Figure 4e; Supplementary Figure S8). Based on evidence for a dominant presence of aerotolerant genera at the oxygen-rich mucosal surfaces of intestinal epithelial cells (Engel and Neurath, 2010; Marteyn et al., 2010; Pineton de Chambrun and Sandborn, 2012), our findings suggest that aerotolerant microbes may have greater access to the MLNs.
MLNs of MC-fed mice, which tended to have more severe colitis compared with FMP-fed mice (Figure 1b), were highly enriched in Proteobacteria, including an increased relative abundance of Enterobacteriaceae. Gut microbiome studies of IBD patients have demonstrated expansions of Proteobacteria, particularly Enterobacteriaceae (Garrett et al., 2007; Mukhopadhya et al., 2012). We observed increased levels of Klebsiella in the MLNs of MC-fed mice (Figure 4f; Supplementary Figure S9), which have been implicated as opportunistic drivers of inflammation in TRUC mice (Garrett et al., 2007). In contrast, MLNs of FMP-fed mice were enriched in Firmicutes – including Lactobacillales, Clostridiales and Coriobacteriales (Figure 4f; Supplementary Figure S9). Gut microbiome status (homeostasis versus dysbiosis) can influence transport of commensal and pathogenic bacterial antigens from the lumen to the MLNs (Diehl et al., 2013). Our analysis of stool and MLN microbial communities support these findings. Furthermore, increased Proteobacteria in stool and MLNs of mice with active colitis aligns with the association between Proteobacteria and IBD-associated dysbiosis.
Microbial community perturbations in active disease versus treatment-induced remission and their functional validation in gnotobiotic mice
Discriminatory microbial lineages for active colitis included the Deferribacteres, Mucispirillum; Anaerotruncas; and Proteobacteria, particularly Enterobacteriaceae, Helicobacteraceae, Desulfovibrio and Sutterella (Figure 5a; Supplementary Figure S10). In contrast, clades associated with remission included Actinobacteria; the Bacteroidetes, S24-7; and the Firmicutes, Staphylococcaceae and Erysipelotrichales (Figure 5a; Supplementary Figure S10).
Gut microbiome composition in active colitis and treatment-induced remission with in vivo functional validation of pro-inflammatory activity in gnotobiotic TRUC mice. (a) Differentially abundant microbial clades in stool from mice with active colitis (n=31) versus remission (n=51) upon intervention completion. For cladogram, white circles represent non-significant microbial clades. (b) Experimental schema of 8-week gnotobiotic TRUC association with conventionally raised, SPF TRUC donor stool. SPF donors were treated for 4 weeks prior to stool collection. Stool from the indicated number of donors was pooled and transplanted into gnotobiotic TRUC recipients. (c) Histologic colitis scores of donors and recipients. Symbols represent data from individual mice. Error bars indicate±SEM. Mann–Whitney test: *P<0.05. (d) B. lactis and L. lactis levels quantified by RT-qPCR in stool from fermented milk (FMP)-treated donors (pooled; n=4) and their corresponding GF recipients (n=5).
Gnotobiotic mice represent a tractable system for testing the function or inflammatory capacity of gut microbiomes (Tremaroli and Bäckhed, 2012). Disease phenotypes, for example, diabetes and obesity, can be transferred to gnotobiotic mice by inoculating them with gut microbiomes of afflicted mice or human donors (Turnbaugh et al., 2006; Wen et al., 2008). To assess the inflammatory potential of gut microbiomes exposed to antibiotics, immunomodulatories or FMP compared with an untreated (sham) control, we performed fecal transfers from treated or untreated SPF TRUC donors to GF TRUC recipients (Figure 5b experimental schema). Pro-inflammatory input communities from untreated donors, on average, induced colitis over the duration of an 8-week association, whereas anti-inflammatory input communities from gentamicin and anti-TNF-α-treated donors did not induce colitis (Figure 5c). Despite FMP ameliorating colitis in SPF donors, gnotobiotic recipients tended to develop mild to moderate colitis (Figure 5c). To follow up on this finding, we measured levels of two FMP bacterial strains, B. lactis and L. lactis, in donor and recipient stool samples using RT-qPCR. FMP strains were detected in donor but not recipient stool (Figure 5d). These findings suggest inefficient intestinal colonization of FMP strains from the donor stool samples and support studies showing that maintaining the benefits of FMPs requires routine administration (Veiga et al., 2010; Sanders, 2011). In contrast with antibiotics and immunomodulatories, these experiments suggest a transient protective effect of FMP on the gut microbiome. Together, these experiments demonstrate that inflammatory phenotypes of gut microbiomes are capable of being transmitted and tested in vivo. Moreover, our results point to differences in the durability of gut microbiomes and their associated disease phenotypes with treatment.
Microbial metabolic functions associated with active colitis versus treatment-induced remission
To investigate the gut microbiome functions associated with active colitis versus remission in TRUC mice following treatment, we used PICRUSt to infer putative metagenomes from our 16S rRNA gene profiles (Langille et al., 2013). Reads were binned into OTUs at ⩾97% sequence identity using a closed reference-based strategy that searches against the available collection of Greengenes reference OTUs. PICRUSt transformed counts of reference-based OTUs into metagenome prediction counts of functional genes on a per-sample basis and evaluated prediction accuracy by calculating the extent to which microbes in a sample are related to sequenced reference genomes using the weighted Nearest Sequenced Taxon Index (Supplementary Table S2). Identified microbial gene families (specified by KEGG Orthology groups) were grouped into metabolic pathways and broader functional categories based on the BRITE hierarchy. We used LEfSe to identify significant, differentially abundant microbially relevant functions associated with active colitis versus remission following treatment (Supplementary Figure S11). Categories associated with remission included carbohydrate metabolism and energy metabolism, as well as the biosynthesis of secondary metabolites. In contrast, gut microbiomes with active colitis were enriched in categories associated with cell motility, signal transduction and xenobiotics biodegradation and metabolism, as well as lipid metabolism (Figure 6a). Thus, gut microbiomes associated with active colitis may have a reduced capacity for energy harvest and dysregulated microbial signaling and cellular processing pathways.
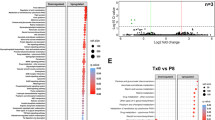
Inferred gut microbiome functions associated with active colitis and treatment-induced remission. Relative abundances of KO gene families grouped into BRITE functional hierarchies, as inferred by PICRUSt from 16S rRNA gene sequences. Differentially abundant microbial functions associated with active colitis (n=31) versus remission (n=51) upon intervention completion organized by KEGG BRITE categories (a) and pathways (b–d). Boxplots denote top quartile, median and bottom quartile. Whiskers and outliers are plotted by the Tukey method. (e) ELISA-based determinations of fecal dopamine levels. Symbols represent data from individual mice from three independent experiments. Mann–Whitney test: *P<0.05; **P<0.01 and ***P<0.001.
Within the cell motility category, we observed an increased capacity for bacterial motility proteins, including genes for flagellar assembly (Figure 6b). Flagellar bacterial antigens have been implicated as disease drivers in both mouse models of colitis and human IBD (Steiner, 2007). Within the signal transduction category, we detected the most significant gene abundances within two-component regulatory systems (Figure 6c). Data suggest that opportunistic microbes have an ability to utilize substrates generated under inflammatory conditions. We examined whether there were differential gene abundances for tetrathionate respiration, a metabolic pathway underlying the fitness advantage of Salmonella enterica subsp. Typhimurium in an inflamed gut (Winter et al., 2010), and observed enhanced potential for tetrathionate respiration with active disease in TRUC mice (K0s: 08357,08358,08539,13040,13041; Figure 6c, inset). Within the xenobiotics biodegradation and metabolism category, we found an enhanced potential for benzoate degradation (Figure 6d). Catechols (1,2-dihydroxybenzene) – intermediaries of benzoate metabolism – have the ability to promote Enterobacteriaceae growth and virulence (Freestone et al., 2007; Lyte et al., 2011). Catecholamines, which are host-derived catechols with a side-chain amine, are detectable in stool (Asano et al., 2012). K. pneumoniae has been implicated in TRUC pathogenesis (Garrett et al., 2010) and members of the Klebsiella genus are one of the few Enterobacteriaceae with the genomic potential to fully metabolize catecholamines (see benzoate degradation pathway kpn00362). Thus, active colitis, with its Enterobacteriaceae enrichment, may result in decreased levels of fecal catecholamines. To test this hypothesis, we measured dopamine, the most abundant catecholamine in the colonic lumen (Asano et al., 2012), in stool collected from mice with active colitis and treatment-induced remission. We examined samples from mice treated with vancomycin and gentamicin, which increases or decreases Enterobacteriaceae, respectively. Dopamine levels were significantly decreased in stool of sham- and vancomycin-treated mice with active colitis as compared with gentamicin-treated mice in remission. In addition, there was a trend, although not statistically significant, toward lower levels of dopamine in mice treated with vancomycin versus sham (Figure 6e).
To validate the inferred functions determined by PICRUSt, we performed WMS sequencing – the conventional means of assessing microbiome functional potential – on a subset of banked stool samples from anti-TNF-α-treated mice (n=6). A total of 419 659 443 quality-filtered shotgun sequences were obtained with an average of 69 943 241±31 774 397 reads per sample. Microbial gene abundances estimated from WMS and 16S rRNA gene sequence data were correlated (Spearman correlation, r=0.6835). This was consistent with correlations in WMS and 16S functional data for human stool samples (Morgan et al., 2012).
Discussion
A goal of microbiome research is defining the structure and function of the gut microbiome in health and active disease states (Lemon et al., 2012). Our study identified shifts in microbial clades and inferred functions associated with active colitis and remission following treatment with antibiotics, immunomodulatories or a fermented milk–dietary intervention in an experimental colitis mouse model. Using gnotobiotic fecal transplant experiments, we found that treatments’ effect on the durability of gut microbiome inflammatory phenotypes varies. Collectively, we identified microbial biomarkers (both clades and functions) that may have clinical relevance for tracking disease when it is asymptomatic and utility as therapeutic targets for managing IBD.
An emerging concept in dysbiosis and bacterial pathogenesis is that certain bacteria have the ability to utilize host substrates to gain a fitness advantage during inflammation (Fleckenstein et al., 2010; Buffie and Pamer, 2013; Winter et al., 2013). The ability to respire tetrathionate and nitrate is central to the fitness of several Enterobacteriaceae (Winter et al., 2013), as these metabolites are readily available in an inflamed gut and can be used as electron acceptors to generate ATP. We observed that tetrathionate utilization was associated with active colitis, which supports a link between enhanced oxidative stress and Enterobacteriaceae-mediated dysbiosis previously described in the TRUC model (Garrett et al., 2010). Dampening the redox stress associated with intestinal inflammation may reduce the abundance of these electron acceptors and eliminate the fitness advantage of colitogenic bacteria, thus restoring intestinal homeostasis.
Enrichment of genes for microbial benzoate degradation in active colitis was unexpected. Catecholamines have garnered interest as communication molecules between host and microbes (Lyte et al., 2011). Enterobacteriaceae can degrade catecholamines and catecholamines can promote Enterobacteriaceae growth and expression of bacterial virulence factors (Eloe-Fadrosh and Rasko, 2013). The histidine sensor kinase QseC is necessary for bacterial responses to host catecholamines and a compound that inhibits QseC, LED209, has been shown to inhibit pathogen virulence in vivo and in vitro (Rasko et al., 2008). Given that LED209 selectively interferes with bacterial virulence and colonization without affecting bacterial growth, typical antibiotic resistance patterns that plague traditional antimicrobials are unlikely to develop (Rasko and Sperandio, 2010). Our observations on tetrathionate respiration and benzoate degradation highlight how gut microbiome studies in mouse models of disease are useful for identifying novel microbial therapeutic targets.
Fecal transplantation represents a long-standing treatment with the potential to address IBD dysbioses and its practice is resurging. However, its use requires substantial consideration from a safety and regulatory perspective (Buffie and Pamer, 2013). We observed that the health status of a host and its gut microbiome could be transient in some cases. Despite a host being in a state of health, as confirmed by histology, transferring its gut microbiome to a GF recipient resulted in colonic inflammation. As applications for fecal transplantation develop for humans, gnotobiotic mouse models may prove useful for evaluating whether microbiomes selected for transplant will confer the intended health outcomes for the recipient.
In summary, our analyses point to features of microbiome dysbiosis and dysfunction in experimental colitis. Improvements in animal model systems and opportunities for translational medical research warrant future studies that incorporate 16S rRNA gene surveys with other ‘omic’ approaches to recognize the gut microbiome’s full potential and ultimately guide therapeutic strategies for manipulating the microbiome to manage disease.
References
Abubucker S, Segata N, Goll J, Schubert AM, Izard J, Cantarel BL et al (2012). Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput Biol 8: e1002358.
Asano Y, Hiramoto T, Nishino R, Aiba Y, Kimura T, Yoshihara K et al (2012). Critical role of gut microbiota in the production of biologically active, free catecholamines in the gut lumen of mice. Am J Physiol Gastrointest Liver Physiol 303: G1288–G1295.
Benjamini Y, Hochberg Y . (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57: 289–300.
Berry D, Schwab C, Milinovich G, Reichert J, Ben Mahfoudh K, Decker T et al (2012). Phylotype-level 16S rRNA analysis reveals new bacterial indicators of health state in acute murine colitis. ISME J 6: 2091–2106.
Buffie CG, Pamer EG . (2013). Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol 13: 790–801.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Carbonero F, Benefiel AC, Gaskins HR . (2012). Contributions of the microbial hydrogen economy to colonic homeostasis. Nat Rev Gastroenterol Hepatol 9: 504–518.
Carvalho FA, Koren O, Goodrich JK, Johansson MEV, Nalbantoglu I, Aitken JD et al (2012). Transient inability to manage proteobacteria promotes chronic gut inflammation in TLR5-deficient mice. Cell Host Microbe 12: 139–152.
Dethlefsen L, Huse S, Sogin ML, Relman DA . (2008). The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol 6: e280.
Diehl GE, Longman RS, Zhang J-X, Breart B, Galan C, Cuesta A et al (2013). Microbiota restricts trafficking of bacteria to mesenteric lymph nodes by CX(3)CR1(hi) cells. Nature 494: 116–120.
Eberl G, Lochner M . (2009). The development of intestinal lymphoid tissues at the interface of self and microbiota. Mucosal Immunol 2: 478–485.
Eloe-Fadrosh EA, Rasko DA . (2013). The human microbiome: from symbiosis to pathogenesis. Annu Rev Med 64: 145–163.
Engel MA, Neurath MF . (2010). New pathophysiological insights and modern treatment of IBD. J Gastroenterol 45: 571–583.
Erickson AR, Cantarel BL, Lamendella R, Darzi Y, Mongodin EF, Pan C et al (2012). Integrated metagenomics/metaproteomics reveals human host-microbiota signatures of Crohn's disease. PLoS One 7: e49138.
Fleckenstein JM, Hardwidge PR, Munson GP, Rasko DA, Sommerfelt H, Steinsland H . (2010). Molecular mechanisms of enterotoxigenic Escherichia coli infection. Microbes Infect 12: 89–98.
Frank DN, Robertson CE, Hamm CM, Kpadeh Z, Zhang T, Chen H et al (2011). Disease phenotype and genotype are associated with shifts in intestinal-associated microbiota in inflammatory bowel diseases. Inflamm Bowel Dis 17: 179–184.
Frank DN St, Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR . (2007). Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104: 13780–13785.
Freestone PPE, Walton NJ, Haigh RD, Lyte M . (2007). Influence of dietary catechols on the growth of enteropathogenic bacteria. Int J Food Microbiol 119: 159–169.
Garrett WS, Gallini CA, Yatsunenko T, Michaud M, DuBois A, Delaney ML et al (2010). Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 8: 292–300.
Garrett WS, Lord GM, Punit S, Lugo-Villarino G, Mazmanian SK, Ito S et al (2007). Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 131: 33–45.
Hansen R, Thomson JM, Fox JG, El-Omar EM, Hold GL . (2011). Could Helicobacter organisms cause inflammatory bowel disease? FEMS Immunol Med Microbiol 61: 1–14.
Hildebrand F, Nguyen ATL, Brinkman B, Yunta RG, Cauwe B, Vandenabeele P et al (2013). Inflammation-associated enterotypes, host genotype, cage and inter-individual effects drive gut microbiota variation in common laboratory mice. Genome Biol 14: R4.
Hill DA, Hoffmann C, Abt MC, Du Y, Kobuley D, Kirn TJ et al (2010). Metagenomic analyses reveal antibiotic-induced temporal and spatial changes in intestinal microbiota with associated alterations in immune cell homeostasis. Mucosal Immunol 3: 148–158.
Jakobsson HE, Jernberg C, Andersson AF, Sjölund-Karlsson M, Jansson JK, Engstrand L . (2010). Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS One 5: e9836.
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA et al (2013). Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 319: 814–821.
Lemon KP, Armitage GC, Relman DA, Fischbach MA . (2012). Microbiota-targeted therapies: an ecological perspective. Sci Transl Med 4: 137rv5.
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R . (2011). UniFrac: an effective distance metric for microbial community comparison. ISME J 5: 169–172.
Lyte M, Vulchanova L, Brown DR . (2011). Stress at the intestinal surface: catecholamines and mucosa-bacteria interactions. Cell Tissue Res 343: 23–32.
Macpherson AJ, Smith K . (2006). Mesenteric lymph nodes at the center of immune anatomy. J Exp Med 203: 497–500.
Manichanh C, Reeder J, Gibert P, Varela E, Llopis M, Antolin M et al (2010). Reshaping the gut microbiome with bacterial transplantation and antibiotic intake. Genome Res 20: 1411–1419.
Marteyn B, West NP, Browning DF, Cole JA, Shaw JG, Palm F et al (2010). Modulation of Shigella virulence in response to available oxygen in vivo. Nature 465: 355–358.
Morgan XC, Tickle TL, Sokol H, Gevers D, Devaney KL, Ward DV et al (2012). Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol 13: R79.
Mukhopadhya I, Hansen R, El-Omar EM, Hold GL . (2012). IBD-what role do Proteobacteria play? Nat Rev Gastroenterol Hepatol 9: 219–230.
Nanau RM, Neuman MG . (2012). Nutritional and probiotic supplementation in colitis models. Dig Dis Sci 57: 2786–2810.
Ott SJ, Musfeldt M, Wenderoth DF, Hampe J, Brant O, Fölsch UR et al (2004). Reduction in diversity of the colonic mucosa associated bacterial microflora in patients with active inflammatory bowel disease. Gut 53: 685–693.
Packey CD, Sartor RB . (2009). Commensal bacteria, traditional and opportunistic pathogens, dysbiosis and bacterial killing in inflammatory bowel diseases. Curr Opin Infect Dis 22: 292–301.
Papa E, Docktor M, Smillie C, Weber S, Preheim SP, Gevers D et al (2012). Non-invasive mapping of the gastrointestinal microbiota identifies children with inflammatory bowel disease. PLoS One 7: e39242.
Pédron T, Mulet C, Dauga C, Frangeul L, Chervaux C, Grompone G et al (2012). A crypt-specific core microbiota resides in the mouse colon. MBio 3: e00116–12.
Pineton de Chambrun GP, Sandborn WJ . (2012). IBD in 2011: advances in IBD management—towards a tailored approach. Nat Rev Gastroenterol Hepatol 9: 70–72.
Powell N, Walker AW, Stolarczyk E, Canavan JB, Gökmen MR, Marks E et al (2012). The transcription factor T-bet regulates intestinal inflammation mediated by interleukin-7 receptor+ innate lymphoid cells. Immunity 37: 674–684.
Presley LL, Ye J, Li X, Leblanc J, Zhang Z, Ruegger PM et al (2012). Host-microbe relationships in inflammatory bowel disease detected by bacterial and metaproteomic analysis of the mucosal-luminal interface. Inflamm Bowel Dis 18: 409–417.
Rasko DA, Moreira CG, Li DR, Reading NC, Ritchie JM, Waldor MK et al (2008). Targeting QseC signaling and virulence for antibiotic development. Science 321: 1078–1080.
Rasko DA, Sperandio V . (2010). Anti-virulence strategies to combat bacteria-mediated disease. Nat Rev Drug Discov 9: 117–128.
Rijkers GT, Bengmark S, Enck P, Haller D, Herz U, Kalliomaki M et al (2010). Guidance for substantiating the evidence for beneficial effects of probiotics: current status and recommendations for future research. J Nutr 140: 671S–676S.
Robertson BR, O'Rourke JL, Neilan BA, Vandamme P, On SLW, Fox JG et al (2005). Mucispirillum schaedleri gen. nov., sp. nov., a spiral-shaped bacterium colonizing the mucus layer of the gastrointestinal tract of laboratory rodents. Int J Syst Evol Microbiol 55: 1199–1204.
Robertson BR, O'Rourke JL, Vandamme P, On SL, Lee A . (2001). Helicobacter ganmani sp. nov., a urease-negative anaerobe isolated from the intestines of laboratory mice. Int J Syst Evol Microbiol 51: 1881–1889.
Sanders ME . (2011). Impact of probiotics on colonizing microbiota of the gut. J Clin Gastroenterol 45 Suppl: S115–S119.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS et al (2011). Metagenomic biomarker discovery and explanation. Genome Biol 12: R60.
Sokol H, Lay C, Seksik P, Tannock GW . (2008). Analysis of bacterial bowel communities of IBD patients: what has it revealed? Inflamm Bowel Dis 14: 858–867.
Steiner TS . (2007). How flagellin and toll-like receptor 5 contribute to enteric infection. Infect Immun 75: 545–552.
Tolia V, Nilsson HO, Boyer K, Wuerth A, Al-Soud WA, Rabah R et al (2004). Detection of helicobacter ganmani-like 16S rDNA in pediatric liver tissue. Helicobacter 9: 460–468.
Tremaroli V, Bäckhed F . (2012). Functional interactions between the gut microbiota and host metabolism. Nature 489: 242–249.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031.
Ubeda C, Pamer EG . (2012). Antibiotics, microbiota, and immune defense. Trends Immunol 33: 459–466.
Veerappan GR, Betteridge J, Young PE . (2012). Probiotics for the treatment of inflammatory bowel disease. Curr Gastroenterol Rep 14: 324–333.
Veiga P, Gallini CA, Beal C, Michaud M, Delaney ML, DuBois A et al (2010). Bifidobacterium animalis subsp. lactis fermented milk product reduces inflammation by altering a niche for colitogenic microbes. Proc Natl Acad Sci USA 107: 18132–18137.
Wang S-L, Wang Z-R, Yang C-Q . (2012). Meta-analysis of broad-spectrum antibiotic therapy in patients with active inflammatory bowel disease. Exp Ther Med 4: 1051–1056.
Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC et al (2008). Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature 455: 1109–1113.
Winter SE, Lopez CA, Bäumler AJ . (2013). The dynamics of gut-associated microbial communities during inflammation. EMBO Rep 14: 319–327.
Winter SE, Thiennimitr P, Winter MG, Butler BP, Huseby DL, Crawford RW et al (2010). Gut inflammation provides a respiratory electron acceptor for Salmonella. Nature 467: 426–429.
Yanai H, Hanauer SB . (2011). Assessing response and loss of response to biological therapies in IBD. Am J Gastroenterol 106: 685–698.
Acknowledgements
We thank members of the Garrett and Huttenhower Labs for discussion. The following grants supported this study: to MGR, NIAID 5T32AI007638; to CH, NIH R01HG005969, NSF DBI-1053486 and ARO W911NF-11-1-0473; to WSG, NIH (R01CA154426, K08AI078942), Burroughs Wellcome CAMS, Searle Scholars, Cancer Research Institute and Danone Research Awards.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Rooks, M., Veiga, P., Wardwell-Scott, L. et al. Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME J 8, 1403–1417 (2014). https://doi.org/10.1038/ismej.2014.3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.3