Abstract
Although the importance of trophic linkages, including ‘top-down forcing’, on energy flow and ecosystem productivity is recognized, the influence of metazoan grazing on Archaea and the biogeochemical processes that they mediate is unknown. Here, we test if: (1) Archaea provide a food source sufficient to allow metazoan fauna to complete their life cycle; (2) neutral lipid biomarkers (including crocetane) can be used to identify Archaea consumers; and (3) archaeal aggregates are a dietary source for methane seep metazoans. In the laboratory, we demonstrated that a dorvilleid polychaete, Ophryotrocha labronica, can complete its life cycle on two strains of Euryarchaeota with the same growth rate as when fed bacterial and eukaryotic food. Archaea were therefore confirmed as a digestible and nutritious food source sufficient to sustain metazoan populations. Both strains of Euryarchaeota used as food sources had unique lipids that were not incorporated into O. labronica tissues. At methane seeps, sulfate-reducing bacteria that form aggregations and live syntrophically with anaerobic-methane oxidizing Archaea contain isotopically and structurally unique fatty acids (FAs). These biomarkers were incorporated into tissues of an endolithofaunal dorvilleid polychaete species from Costa Rica (mean bulk δ13C=−92±4‰; polar lipids −116‰) documenting consumption of archaeal-bacterial aggregates. FA composition of additional soft-sediment methane seep species from Oregon and California provided evidence that consumption of archaeal-bacterial aggregates is widespread at methane seeps. This work is the first to show that Archaea are consumed by heterotrophic metazoans, a trophic process we coin as ‘archivory’.
Similar content being viewed by others
Introduction
Trophic interactions represent the most important class of feedback phenomena on Earth (Worm and Duffy, 2003), but we have yet to include one of the most abundant forms of life in the oceans, Archaea (Karner et al., 2001; Francis et al., 2005; Lipp et al., 2008), into our understanding of these trophic relationships. Archaea perform key ecosystem services, including nitrification (Beman and Francis, 2006), anaerobic methane oxidation (Treude et al., 2003) and chemosynthetic production (Wuchter et al., 2003; Boetius and Suess, 2004; Herndl et al., 2005). These services are likely impacted by grazing pressure, a ‘top-down’ force that can cause changes in productivity and nutrient cycling (McNaughton, 1985; Belovsky and Slade, 2000). Integrating Archaea into our understanding of marine–trophic relationships may provide fundamental information about mechanisms that control archaeal-driven biogeochemical processes. Yet, Archaea are known to prevail in low-energy systems and grow slowly (Valentine, 2007); thus, it is unclear if Archaea can provide a food source that supports persistent metazoan populations. In addition, highly-stable cell membranes of Archaea (van de Vossenberg et al., 1998) may provide protection from digestion while failing to provide essential fatty acids (FAs) that most Metazoa need for growth and reproduction. Here, we demonstrate that (1) Archaea is a food source that can provide the nutrients necessary for a metazoan to complete its life cycle and (2) archaeal–bacterial syntrophic aggregates are a primary food source for a family of methane-seep annelids, thus identifying a novel relationship between Archaea and Metazoa.
Methane-seep ecosystems contain abundant methanotrophic Archaea that have a unique carbon isotopic and lipid signature. Aggregates of anaerobic-methane oxidizing Archaea (ANMEs) and sulfate-reducing δ-proteobacteria (SRBs) consume the majority of methane released from deep-sea reservoirs through the anaerobic oxidation of methane (AOM; Boetius et al., 2000; Orphan et al., 2001b, 2002; Reeburg, 2007). Methane often has a distinct 13C to 12C ratio that is depleted in 13C relative to 12C (Whiticar, 1998—δ13C notation is explained within). In turn, ANME consortia are defined by similar highly negative δ13C values, ranging between −30‰ and −100‰ for archaeal cells and −15‰ to −70‰ for associated symbiotic SRB (Orphan et al., 2002; House et al., 2009). These values are often distinct from those of photosynthetic production, which normally has a δ13C value of −15‰ to −25‰ (Fry and Sherr, 1994). In addition to their unique cell carbon-isotopic composition, both ANMEs and SRBs possess unique membrane lipids, which also exhibit a highly negative carbon-isotopic composition. ANME archaeal lipids have δ13C values between −128‰ to −61‰ and are composed of repeating isoprene units with or without an ether-linked glycerol head (Elvert et al., 1999; Boetius et al., 2000; Hinrichs et al., 2000; Niemann and Elvert, 2008). SRB within these consortia have FAs largely composed of 16:1(n−5), cyc17:0w5,6, and/or a15:0 and 18:1(n−7) FAs with δ13C values between −112‰ and −65‰ (Elvert et al., 2003; Blumenberg et al., 2004; Niemann and Elvert, 2008). These aggregates can make up to 80% of the microbial biomass in methane seep sediments (Boetius et al., 2000) and act as deep-sea engineers by creating hard substrate in the form of authigenic carbonates (Greinert et al., 2001; Luff et al., 2004). In addition, ANME aggregates routinely occur in the top four centimeters of sediment (Boetius et al., 2000; Elvert et al., 2003; Knittel et al., 2003; House et al., 2009), placing them in contact with metazoan grazers, including polychaetes of the family Dorvilleidae (Levin et al., 2003).
The 13C-depleted Archaea at methane seeps provide a model system to explore archivory, a term we define here to mean the consumption of Archaea. The carbon-isotopic signature of heterotrophic organisms is derived from the isotopic signature of the primary producer they or their prey feed upon (DeNeiro and Epstien, 1978). Within methane seeps, dorvilleids have been reported to exhibit extremely negative δ13C values (for example, −90.6‰ in the Gulf of Alaska; Levin and Mendoza, 2007). These isotopic values potentially reflect the consumption of ANME aggregates (Levin and Michener, 2002). Yet, bacterial methanotrophs also have a depleted 13C, methane-derived isotopic composition (Elvert et al., 2000; Werne et al., 2002) and chemoautotrophic producers, including sulfide-oxidizing bacteria, can incorporate a 13C-depleted isotopic signature from a 13C-depleted DIC pool (Fisher, 1990). Thus, a consumer with an extreme 13C depletion need not directly consume methanotrophic Archaea.
Grazer lipids are partially derived from their diet (Dalsgaard et al., 2003), providing a tool for tracking microbial consumption and potentially archivory. This phenomenon is best known for FAs, a component of both eukaryotic and bacterial cell membranes that can provide a quantitative measure of food web linkages (Iverson, 2009). In seep settings, key FA biomarkers include 16:1(n−7) and 18:1(n−7) that are abundant in sulfide-oxidizing bacteria (McCaffrey et al., 1989), the aforementioned 16:1(n−5) and cyc17w5,6 that are abundant in sulfate-reducing bacteria (Elvert et al., 2003; Blumenberg et al., 2004), and 16:1(n−6) and 16:1(n−8) that are indicative of type I aerobic bacterial methanotrophs (Bowman et al., 1991). Composition of photosynthetic production can be identified by polyunsaturated fatty acids, including 22:6(n−3) and 20:5(n−3); (reviewed in Dalsgaard et al., 2003). Archaeal lipid biomarkers are diagnostic yet their incorporation into metazoan tissues has yet to be documented, limiting their utility in food-web studies.
Owing to the ubiquity of dorvilleids in Archaea-fueled methane seep sediments and their 13C-depleted carbon-isotopic composition (Levin and Michener, 2002; Levin et al., 2003; Levin, 2005; Levin and Mendoza, 2007; Menot et al., 2010; Ritt et al., 2010), we chose to use this polychaete family to study archivory. Dorvilleids are tolerant to sulfide stress allowing them to numerically dominate the macrofauna in microbial-mat covered sediments at seeps (Levin et al., 2003; Levin, 2005), a habitat that has high ANME abundance (Boetius and Suess, 2004). To begin testing if Archaea are a viable and utilized food source, we combined laboratory-based feeding assays with trophic studies in soft-sediment and authigenic-carbonate seep habitats to address the following questions: (1) Can metazoans survive and reproduce on a diet exclusively of Archaea? (2) Does an archaeal diet manifest itself in consumer lipid signatures? (3) Is there evidence for archaeal consumption by metazoans in natural populations?
Materials and methods
Sample collection and preparation
To test if archaeal biomass provides sufficient nutrition for sustaining metazoan populations and results in a unique lipid pattern within archaeal consumers, we raised a species of dorvilleid polychaete on Archaea within the laboratory. The shallow water dorvilleid, Ophryotrocha labronica, was raised on monocultures of two types of halophilic Euryarchaea, Halobacterium salinarium and Haloferax volcanii, and its growth rate over a 44–48 day period was compared with the same species fed monocultures of bacterial (Bacillus subtilis, a gram-positive Firmicute, or Photobacterium profundum, a gram-negative γ-proteobacterium) or eukaryotic food sources (Oryza sp., rice, or Spinacia oleracea, spinach). Individuals of O. labronica were raised on either one of the two-archaeal sources or the bacterium B. subtilis for neutral lipid comparison. Between 11 and 14 feeding trials were run with each food source during which egg mass deposition was noted if it occurred (see Supplementary Materials for additional information).
In situ archivory was studied using specimens and substrate from three deep-sea, methane-rich locations with high abundances of Archaea: authigenic carbonates precipitated on Mound 12, Costa Rica (8°55.8′N 84°18.8′W; 1000 m) and soft-sediment seep habitats at Eel River, California (40°47.1′N 124°35.68′W; 490–520 m) and at Hydrate Ridge, Oregon (44°40.1′N 125°05.8′W; 580–890 m). Authigenic carbonate rocks were collected from Costa Rica during RV Atlantis cruise 15–44 (rock L2; 21 February – 8 March, 2009) and 15–59 (rock E3; 6–12 January, 2010). The rocks were recovered from active seep areas by the submersible DSRV Alvin and placed into insulated boxes for transport to the ship. Carbonates were broken open using a chisel and endolithofauna were removed. We focused on a single undescribed dorvilleid polychaete species within the genus Dorvillea, which was found living within the rocks. Soft-sediment seep habitats off California and Oregon containing a high abundance of methanotrophic Archaea (Orphan et al., 2001b; Elvert et al., 2003; Knittel et al., 2003; Boetius and Suess, 2004) were sampled during R/V Atlantis cruises 15–7 (13–27 July, 2006) and 15–11 (26 September–10 October, 2006) using the DSRV Alvin. Sediment was collected using push cores or ‘scoops’ from clam beds and microbial mats. Sediment samples were sieved with a 300-μm sieve and infauna were sorted live, yielding five species of dorvilleid polychaetes for biomarker analysis (Ophryotrocha maciolekae, Ophryotrocha platykephale, Parougia oregonensis, and an undescribed species of each Parougia and Exallopus). All individuals analyzed were allowed to evacuate their guts overnight in 25-μm filtered sea water and frozen at −80 °C. To document the availability of ANME aggregates to rock-dwelling dorvilleids, the microbial community present within a subsample of rock E3 was identified using FA profile analysis (modified from Lewis et al., 2000; see Supplementary Material for additional methods), 16S rRNA analysis and epifluorescence microscopy protocols described in Orphan et al. (2001a).
Lipid analyses
Lipid profiles of archaeal food sources, O. labronica fed those food sources, and dorvilleids collected in the field were measured to assess whether archaeal lipids can function as archivory biomarkers. Key archaeal lipids, including hydroxyarchaeol, archaeol, crocetane and pentamethylicosane (PMI) are present in the neutral lipid fraction (Elvert et al. 1999; Hinrichs et al. 1999; Thiel et al., 1999; Oba et al., 2006) and if present within the tissue of metazoans that consume Archaea, would provide a powerful tool to identify Archaea. FAs of field-collected individuals and rock E3 were extracted using a one-step extraction–transesterification. This technique was chosen due to its effectiveness at extracting low-biomass (<0.2 mg) samples. This allowed us to avoid pooling individuals across samples and thus have true replication in all but one of the species collected. Owing to the small biomass of most dorvilleids, gas chromatography-mass spectrometry (GC–MS) analysis, which is highly sensitive but less quantitative than GC–FID, was employed. Thus, whereas internal comparisons within this study are valid, absolute concentrations are not quantitatively comparable to values generated by other instruments.
Carbon isotopic analyses
In all, three types of isotopic analysis were performed on field-collected samples to track methane-derived carbon, potentially indicative of archivory: ‘δ13C Bulk’, ‘δ13C Bulk-Lipid’, and ‘δ13C Compound-Specific’. ‘δ13C Bulk’ is the carbon-isotopic composition of the non-lipid extracted dorvilleid biomass. ‘δ13C Bulk-Lipid’ involved separate analyses of polar and neutral lipids within Dorvillea sp. from rock L2. Biomarkers that are directly incorporated from methanotrophic Archaea should have a very 13C-depleted carbon isotopic signature, thus this analysis identifies whether the polar or neutral fraction contains Archaea-derived carbon. ‘δ13C Compound-Specific’ was used to identify the isotopic composition of each of the FAs present within both rock E3 microorganisms and the Dorvillea sp. that inhabited it. This allowed fine-scale identification of which FAs were present within the food source (the rock) and were incorporated into Dorvillea sp. tissues.
Results
Archaea as a food source
Laboratory growth and fecundity experiments with O. labronica demonstrated that Archaea provide sufficient nutrition for this species to close its life cycle. Ophryotrocha labronica grew from 0.2 mm (post hatching) to ∼1.11±0.4 mm (adult size) in as little as 12 days on monospecific archaeal diets of both H. salinarium (n=13, separate egg broods, which we heretofore refer to as a separate cohorts) and H. volcanii (n=11, separate cohorts). Mean growth rate of O. labronica over a 44–47-day period did not vary as a function of food source (Figure 1; F5,29=1.36, P=0.27). After 45–58 days, three cohorts fed H. salinarium produced egg masses and one cohort fed H. volcanii produced an egg mass. This reproductive success was similar to the cohorts fed the other two domains of life over a similar 45–48-day period. The two bacterial food sources, B. subtilis and P. profundum supported production of three and one egg mass by 14 and 12 separate cohorts, respectively. The 12 cohorts of O. labronica fed Oryza sp. or S. oleracea, both resulted in deposition of a single egg mass.
Neutral lipid biosignatures in laboratory-reared specimens
Archaeal food sources did not generate a distinctive neutral lipid pattern in O. labronica. Both strains of Euryarchaeota had a neutral lipid composition that included squalene, dihydrosqualene and tetrahydrosqualene in addition to a few other isoprene-based molecules (Table 1). The worms fed these two archaeal species did not incorporate these molecules, but instead had a neutral lipid profile consisting largely of cholesterol and other sterols. With the exception of β-sitosterol, these same compounds were found in O. labronica fed the bacterium B. subtilis (Table 1). None of the neutral lipids in Archaea-fed O. labronica tissues were archaeal-specific, despite the fact that this species could grow to reproduction on an Archaea monoculture. We note that although other researchers have found archaeol in both of the Archaea that we have used here (Qiu et al., 2000; Stiehl et al., 2005), we did not resolve these compounds within our cultures and thus our findings do not include intact polar lipids.
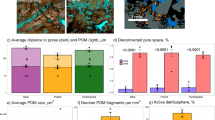
ANME aggregates in authigenic carbonates
Archaea available to dorvilleids at methane seeps often occur as ANME aggregates. In the interior of carbonate rock E3, microbial aggregates resembling previously described consortia of ANME and SRB were observed (Figure 2; Orphan et al., 2002). PCR-based 16S rRNA gene analysis of DNA recovered from rock E3 revealed an archaeal assemblage that was dominated by methanotrophic ANME-2 (68%), with a lower percentage of ANME-1 (10%) phylotypes (following the classification of Orphan et al., 2001b). Rock E3 was largely composed of two FAs, cyc17:0 and 16:1(n−5), each comprised more than 20% percent of the total FA composition of the rock (Figure 3). The isotopic signatures of these two FAs were extremely depleted in 13C, with δ13C values of −115‰ and −110‰, respectively, providing further evidence that they are derived from SRBs that participate in AOM (sensu Elvert et al., 2003). In addition, branched 15-carbon FAs and 18:1(n−7), which are common in ANME-1/SRB aggregates (Elvert et al., 2003; Blumenberg et al., 2004), formed a combined 23.8% of the FAs present within the rock. This FA profile supports the findings of the 16S rRNA gene sequences that indicated that the majority of the microbial community within the rock was comprised of ANME-2 consortia with a smaller subset belonging to ANME-1-associated SRB. Furthermore, crocetane, an archaeal lipid, had a δ13C value of −122‰ that clearly indicated a methane-derived energy source for the Archaea present. Both 16:0 and 18:0 were more enriched in 13C than the other FAs present. Although these data were corrected for concurrently run blanks, trace contaminants from the sample processing were apparent in both of these saturated FAs and thus their isotopic values are unlikely to represent the isotopic composition of those FAs from the sample. There were no contaminats in the other FAs analyzed. Diagnostic FA biomarkers for aerobic bacterial methanotrophs 16:1(n−6) and 16:1(n−8) (Bowman et al., 1991) were not recovered from carbonate rock E3.
Carbon isotopic compositions of fatty acids (FAs) and FA distribution within (upper panel) carbonate rock E3 and (lower panel) Dorvillea sp. from Costa Rica. Left y axis and bars are percentage of FA and right y axis and points are isotopic composition. Asterisks indicate that isotopic composition was potentially largely impacted by contaminants (see text for more details). Value for this omitted FA ranged between −104‰ to −47‰. 18:2 and 20:1(n−11) were left off this figure as they were <2% in only one individual of Dorvillea sp. See text for explanation of de novo synthesis identification.
SRB/ANMEs in the diet of dorvilleids
A combination of stable-carbon isotope and FA analysis indicate that the carbonate-associated Dorvillea sp. at the Costa Rica seeps consumed methanotrophic ANME/SRB aggregates as their main food source. Among the most 13C depleted metazoans on record, the ‘δ13C bulk’ signatures of Dorvillea sp. were as low as −101‰ (from authigenic carbonate rock L2) and within rock E3 this species had a mean δ13C of −91.7±3.5 (s.e.) ‰ (n=10). The Dorvillea sp. lipid profile included FAs likely derived from ANME-associated SRBs (Figure 3). These FAs included 16:1(n−5), which composed 1.4–1.8% of the total FAs present within Dorvillea sp. from rocks E3 and L2. The FA 18:1(n−5) also formed between 3.3% and 5.3% of the Dorvillea's FA profile. This FA can be synthesized by eukaryotes if they are provided with 16:1(n−5), but was not present within rock E3. Both of these (n−5) FAs had isotopic signatures within 3‰ of that of the sulfate-reducing bacteria (n−5) FA (Figure 3). This indicates that these FAs were derived from the carbonate AOM consortia.
To test if consumption of the AOM consortia is an evolutionary oddity limited to Costa Rica carbonates or a common occurrence within methane-seep environments, we then examined five additional dorvilleid polychaete species from NE Pacific methane seep sediments. The ‘δ13C bulk’ composition of these species varied between −19.5‰ and −57.9‰ indicating use of a diversity of food sources, including potentially photosynthetic and AOM-derived organic matter. To understand the relationship between carbon isotopic signature and FA composition of the worms, a regression of percent (n−5) FA on δ13C was performed. As with the carbonate endolithofauna, a subset of these dorvilleids possessed (n−5) FAs (Table 2). The distribution of these FAs was not uniform among the species; instead the greater the composition of (n−5) FAs the more negative the δ13C value (Figure 4). This simple relationship between percent (n−5) and δ13C explained 50% of the variance observed in the carbon isotopic composition of these species (F1,20=20.4, P<0.01; R2=50.5):

Relationship between carbon isotopic composition and sum of 16:1(n−5) and 18:1(n−5) FAs present within five species of polychaete from Eel River and Hydrate Ridge. Solid line is a linear regression, including all dorvilleid polychaetes in which both isotopic and FA data are available within a single sample (R2=50.5). Dotted line indicates linear regression if the point to the far right is removed (R2=47.4). Symbols indicate species.
Removal of one outlier appeared to have a large influence on this regression (FA composition of 10.1% (n−5) FAs) and increased the negative slope of the regression from −3.5 to −5.6 but had little effect on the variance explained (F1,19=17.1, p<0.01; R2=47.4). Thus, we chose to retain all data. Inclusion of the (n−7) FAs, common in sulfide-oxidizing bacteria (McCaffrey et al., 1989), improved the fit of the model only by 1.5%. A regression of isotope signature against solely percent (n−7) FAs, explained only 20% of the variance (F1,20=5.30, P=0.03; R2=20.9). There was a small percentage of 16:1(n−6) FA present in three of the dorvilleid species, which are FAs indicative of aerobic methanotrophy, yet these were a minor component of the lipid profile (Table 2).
Neutral lipids of species that consume SRB/ANME aggregates
The neutral lipids within the Costa Rica seep dorvilleid species were isotopically distinct from archaeal lipids and had a sterol composition that was uniform among the species analyzed from Costa Rica, NE Pacific and the laboratory (Table 1). The carbonate-associated individual with a bulk δ13C of −101‰, had a polar-lipid δ13C of −116‰ and a neutral lipid δ13C of −68‰. This indicated that the most 13C-depleted carbon, and thus biomarkers from methanotroph biomass, was incorporated into the polar-lipid fraction rather than the neutral lipids when viewed in bulk. Although the neutral-lipid fraction also showed clear incorporation of methane-derived carbon, this isotopic composition is far removed from the lipid signature of the SRB (here, δ13C=−110‰; Figure 3), the archaeal lipid composition (δ13C=−122‰) or the dorvilleid isotopic composition (‘δ13C Bulk’=−92±4‰) recovered from rock E3. In addition, the neutral-lipid composition of this species, Dorvillea sp., and all of the other dorvilleid species did not include crocetane.
Discussion
Ingestion or digestion
The FAs present within the Dorvillea sp. had isotopic signatures that were <−100‰ (except for a 20:0 FA that was −95‰), clearly indicating incorporation of methane-derived carbon (Figure 3). Of special note, two polyunsaturated fatty acids, 20:5(n−3) and 20:4(n−6), commonly used as indicators of phytoplanktonic production (Dalsgaard et al., 2003), had δ13C values between −103‰ and −109‰, indicating that they were formed in situ by consumed methane-fueled bacteria or synthesized de novo by the dorvilleids. The possibility that annelids may be able to synthesize these FAs has been previously considered; three species that belong to the family Siboglinidae, which lack both mouth and anus and live off chemoautotrophic energy from endosymbionts, all possessed these FAs, and in one instance they had polyunsaturated fatty acids with a δ13C of −72‰ (Pond et al., 2002; Lösekann et al., 2008).
Dorvilleids possess chitonized jaws that provide a mechanism to harvest ANME aggregates off of carbonate rocks. The effectiveness of this feeding strategy is likely aided by the large size of ANME aggregations hosted in carbonate rocks (up to 15 μm in diameter; Figure 2, compared with average diameters of 3–7 μm for ANME aggregations in sediments; House et al., 2009; Orphan et al., 2009) and is analogous to gastropod use of their radula to harvest epilithic algae in intertidal habitats. However, the fate of ANME/SRB aggregates after the aggregate is consumed may be explained by two potential hypotheses: (1) dorvilleids may consume the aggregate but only digest the sulfate-reducing bacteria and excrete the archaeal component, potentially alive or (2) dorvilleids digest the aggregates en masse and thus their diet is composed of both Archaea and SRB. Either of these scenarios would impact the symbiotic relationship of the aggregate and the rate of AOM but to different extents. Previous studies have not found δ13C SRB biomass to be more 13C depleted than −70‰ (Orphan et al., 2002; House et al., 2009), thus the incredibly 13C-depleted isotopic composition of the endolithofaunal Dorvillea sp. supports the idea that archaeal biomass is indeed digested, as was observed in the laboratory feeding trials.
More information can be gained about these two hypotheses from the FA type and isotopic composition. Eukaryotes have a suite of enzymes that allow them to form or modify dietary-derived FAs. In addition to being able to elongate FAs, eukaryotes can also synthesize a diversity of unsaturated (n−7) and (n−9) FAs and a variety of 20:1 FAs (Kattner and Hagen, 1995). Yet, knowledge to date suggests that (n−5) FAs are not synthesized by eukaryotes (MacAvoy et al., 2003). Thus, the FA profile of Dorvillea sp. is as expected from a species that synthesizes almost all of its FAs except for the (n−5) FAs provided by SRB. Elongation of dietary lipid occurs through the addition of acetate. Assuming Dorvillea sp. forms 18:1(n−5) from 16:1(n−5), we can apply a carbon mass-balance approach to calculate the δ13C of acetate used by Dorvillea sp. to gain insight into its carbon source. This results in an estimate of δ13Cacetate=−109‰ and −128‰ used by Dorvillea sp. from rocks L2 and E3, respectively. Within rock E3, the only compound that had an isotopic composition that was close to −128‰ was the archaeal lipid crocetane (δ13C=−122‰). This suggests that Dorvillea sp. at Costa Rica was elongating its FAs using archaeal-lipid derived carbon. As no 18:1(n−5) was found in the rocks and two-carbon additions to dietary-derived FAs follows established metazoan FA biosynthesis pathways, the general assumptions behind this mass-balance calculation are likely appropriate. However, this does assume that there is no enzymatic fractionation, or specifically selection of the more 13C depleted 16:1(n-5) FAs that are then converted to 18:1(n-5) or selection of the most 13C depleted acetate molecules used for this elongation; enzymatic carbon selectivity could preferentially select 12C molecules for use during the elongation process. Furthermore, a pool of available acetate occurs within the sediment and can also be quite 13C depleted. Few seep samples have had δ13Cacetate analyzed, but within Black Sea seep sediments this source of acetate was found to be as negative as δ13Cacetate=−85‰ (Heuer et al., 2006). Thus, the acetate used by Dorvillea sp. to elongate its FAs may be a combination of acetate derived from its diet and acetate present within the dissolved organic carbon pool within the rock.
Archaeal biomarkers
None of the archaeal lipids that we identified in this study were preserved unmodified in those taxa that consumed Archaea, and neutral lipid composition appeared independent of diet. These later findings do not appear to be entirely unique to dorvilleids; Capitella sp.I also augments its dietary-derived sterols through biosynthesis (Marsh et al., 1990). Although beyond the scope of this study, whose aim was to identify if Archaea biomass as a whole was sufficient to sustain metazoans populations, we can gain some insight into the catabolic pathway of Archaea based on our isotopic results. Squalene, a lipid provided by both archaeal food sources from the laboratory study, is a de novo formed precursor to most sterols synthesized by metazoans (Kanazawa, 2001). Pentamethylicosane and Crocetane, two archaeal hydrocarbons, have a similar structure to squalene yet differ by saturation state. The incorporation of any of these hydrocarbons into sterol synthetic pathways by archaeal consumers may benefit species that live in low-energy systems. However, this pathway seems unlikely as the bulk neutral-lipid isotopic composition for Dorvillea sp. was −68‰, a value far removed from crocetane's δ13C of −122‰. Although we did not measure the isotopic composition of other archaeal lipids from the rock, we can use the isotopic values of Stadnitskaia et al. (2008) who measured the δ13C of archaeal lipids as=−97‰ to −98‰ within the authigenic carbonates at Mound 11, a nearby Costa Rican methane seeps, as a potential isotopic value for larger archaeal lipids. Alternatively, we can use crocetane's isotopic composition as a proxy for the isotopic composition of archaeal lipids in rock E3. Previous research has found that archaeal lipids are at most 12‰ enriched in 13C compared with crocetane, if not more depleted (Niemann and Elvert, 2008). Therefore, we can estimate that the δ13C value of the archaeal lipids in this rock are <−110‰, a value far removed from the neutral lipid isotopic signature measured for Dorvillea sp. As such, it is unlikely that the neutral lipid value reflects direct incorporation of archaeal lipids.
A caveat for the extraction technique used to analyze neutral lipids is that it may have co-extracted carbohydrates and/or proteins, contaminants that may mask archaeal input into the neutral lipid class. Any contaminants extracted in this lipid class, if 13C enriched, would impact our conclusion that the neutral lipids do not contain archaeal-derived lipids beyond potential, and unresolved, trace amounts. The bulk isotopic composition of the carbon within Dorvillea sp. was also much more 13C depleted than the neutral lipids that were extracted from it. As proteins and to a lesser extent carbohydrates are often much more abundant in annelids than lipids (Blackstock et al. 1982) this bulk isotopic composition likely reflects the carbon isotopic composition of the potential contaminants. Therefore, even if we did co-extract non-neutral lipids, potentially masking 13C-depleted sterols or unresolved compounds, these contaminants would likely skew our neutral lipid signature more negative rather than positive.
The δ13C value of the neutral lipids may be explained by the worm deriving this carbon from non-lipid sources of either SRB or ANME aggregates, which are enriched in 13C compared with their lipids, or by the uptake of sterols from the environment, potentially reflecting recalcitrant pools of carbon. As sediments containing active AOM aggregates also have 13C-enriched sterols that are far removed from the 13C-depleated values of the AOM aggregates (Niemann and Elvert, 2008), uptake of these 13C-enriched sterol pools could impact the sterol neutral lipid isotopic composition. What is clear is that when analyzing the neutral lipids of these species, no clear biomarkers are apparent. Further analysis should examine larger compounds (including intact polar lipids, and specifically targeting Archaeol and sn-2-hydroxyarchaeol as well at tetraethers), although the neutral lipid isotopic composition measured here suggests they are unlikely to be abundant.
Archaea in food webs
Archaea are a ubiquitous domain of life, and in addition to their previously recognized importance, we have shown that they are capable of supporting growth and reproduction of a heterotrophic metazoan. Symbiotic relationships are known between Archaea and ciliates (van Hoek et al., 2000), sponges (Preston et al., 1996) and even terrestrial ruminants (Joblin, 2005), defining a key role for Archaea in supporting eukaryotes through symbioses. It is unlikely that archaeal symbioses were present in Dorvillea sp. as attempts to PCR amplify archaeal 16S rRNA from this species were unsuccessful (S Goffredi pers. com.), and, as with all the dorvilleids within this study, Dorvillea sp. had a well-developed digestive tract complete with mouth and anus. Thus, this family of polychaete provides the first evidence for the ecological role of free-living Archaea as a food source for heterotrophic metazoans. Further research may identify the specific catabolic pathways by which the different archaeal cellular constituents are digested and used by consumers.
ANME are thought to be the terminal sink of methane throughout the world's oceans (Reeburg, 2007) yet the impact of metazoan grazing on ANMEs is unknown. Because of aerobic methanotrophic bacteria and ANME's metabolic activities, a majority of methane emitted at seeps is consumed within the sediment before release into the water column (Sommer et al., 2006; Reeburg, 2007). Here, we demonstrate that the anaerobic part of this sediment filter is subject to grazing by metazoans at three separate seep locations, with unknown ramifications for the biogeochemical cycling of methane within these seeps. Observation of this phenomenon within authigenic carbonates, habitats that are widespread on margins, opens a new avenue of C cycling investigation. The incredibly 13C-depleted carbon isotopic composition of the endolithofaunal polychaete within this study highlights the unknown but clearly active role of carbonate-associated biota in the global methane cycle.
References
Belovsky GE, Slade JB . (2000). Insect herbivory accelerates nutrient cycling and increases plant production. Proc Natl Acad Sci USA 97: 14412–14417.
Beman JM, Francis CA . (2006). Diversity of ammonia-oxidizing Archaea and bacteria in the sediments of a hypernutrified subtropical estuary: Bahia del Tobari, Mexico. Appl Environ Microbiol 72: 7767–7777.
Blackstock J, Barnes M, Barnes the late H . (1982). The loch eil project: biochemical composition of the polychaete, Glycera alba (Müller), from Loch EIL. J Exp Marine Biol Ecol 57: 85–92.
Blumenberg M, Seifert R, Reitner J, Pape T, Michaelis W . (2004). Membrane lipid patterns typify distinct anaerobic methanotrophic consortia. Proc Nat 30: 11111–11116.
Boetius A, Ravenschlag K, Schubert CJ, Richert D, Widdle F, Gieske A et al. (2000). A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407: 623–626.
Boetius A, Suess E . (2004). Hydrate Ridge: a natural laboratory for the study of microbial life fueled by methane from near-surface gas hydrates. Chem Geol 205: 291–310.
Bowman JP, Skerratt JH, Nichols PD, Sly LI . (1991). Phospholipid fatty acid and lipopolysaccharide fatty acid signature lipids in methane-utilizing bacteria. FEMS Microbiol Ecol 85: 15–22.
Dalsgaard J, St John M, Kattner G, Müller-Navarra D, Hagen W . (2003). Fatty acid trophic markers in the pelagic marine environment. Adv Mar Biol 46: 225–340.
DeNeiro MJ, Epstien S . (1978). Influence of diet on the distribution of carbon isotopes in animals. Geochim Cosmochim Acta 42: 495–506.
Elvert M, Boetius A, Knittel K, Jorgensen BB . (2003). Characterization of specific membrane fatty acids as chemotaxonomic markers for sulfate-reducing bacteria involved in anaerobic oxidation of methane. Geomocrbiol J 20: 403–419.
Elvert M, Greinert J, Suess E, Whiticar MJ . (2000). Archaea mediating anaerobic methane oxidation in deep-sea sediments at cold seeps of the eastern aleutian subduction zone. Organic Geochem 31: 1175–1187.
Elvert M, Suess E, Whiticar MJ . (1999). Anaerobic methane oxidation associated with marine gas hydrates: superlight c-isotopes from saturated and unsaturated C20 and C25 irregular isoprenoids. Naturwissenchaften 86: 295–300.
Fisher CR . (1990). Chemoautotrophic and methanotrophic symbioses in marine invertebrates. Rev Aquat Sci 2: 399–436.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB . (2005). Ubiquity and diversity of ammonia-oxidizing Archaea in water columns and sediments of the oceans. Proc Natl Acad Sci 102: 14683–14688.
Fry B, Sherr EB . (1994). δ13C measurements as indicators of carbon flow in marine and freshwater systems. Contrib Mar Sci 27: 13–46.
Greinert J, Bohrmann G, Suess E . (2001). Gas hydrate-associated carbonates and methane-venting at Hydrate Ridge classification, distribution, and origin of authigenic lithologies. Geophys Monogr 124: 99–113.
Herndl GJ, Reinthaler T, Tiera E, van Aken H, Veth C, Pernthaler A et al. (2005). Contribution of Archaea to total prokaryotic production in the deep Atlantic Ocean. App Environ Microbiol 71: 2303–2309.
Heuer V, Elvert M, Tille S, Krummen M, Mollar XP, Hmelo LR et al. (2006). Online δ13C analysis of volatile fatty acids in sediment/porewater systems by liquid chromatography-isotope ratio mass spectrometry. Limnol Oceanogr Meth 4: 346–357.
Hinrichs KU, Hayes JM, Sylva SP, Brewer PG, DeLong EF . (1999). Methane-consuming archaebacteria in marine sediments. Nature 398: 802–805.
Hinrichs K-U, Summons RE, Orphan V, Sylva SP, Hayes JM . (2000). Molecular and isotopic analysis of anaerobic methane-oxidizing communities in marine sediments. Org Geochem 31: 1685–1701.
House CH, Orphan VJ, Turk KA, Thomas B, Pernthaler A, Vrentas JM et al. (2009). Extensive carbon isotopic heterogeneity among methane seep microbiota. Envirion Microbiol 11: 2207–2215.
Iverson SJ . (2009). Tracing aquatic food webs using fatty acids: from qualitative indicators to quantitative determination. In: Arts M, Brett M, Kainz M (eds). Lipids in Aquatic Ecosystems. Springer: New York, pp 281–307.
Joblin K . (2005). Methanogenic archaea. In: Makkear HPS, McSweeny CS (eds). Methods in Gut Microbial Ecology for Ruminants. Springer: New York, pp 47–53.
Kanazawa A . (2001). Sterols in marine invertebrates. Fisheries Sci 67: 997–1007.
Karner MB, Delong EF, Karl DM . (2001). Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature 409: 507–510.
Kattner G, Hagen W . (1995). Lipid metabolism of the Antarctic euphausiid Euphausia crystallorophias and its ecological implications. Mar Ecol Prog Ser 170: 203–213.
Knittel K, Boetius A, Lemke A, Eilers H, Lochte K, Pfannkucke O et al. (2003). Activity, distribution, and diversity of sulfate reducers and other bacteria in sediments above gas hydrate (Cascadia Margin, Oregon). Geomicrobiol J 20: 269–294.
Levin LA . (2005). Ecology of cold seep sediments: interactions of fauna with flow, chemistry and microbes. Oceanogr Mar Biol Ann Rev 43: 1–46.
Levin LA, Mendoza GF . (2007). Community structure and nutrition of deep methane-seep macrobenthos from the North Pacific (Aleutian) Margin and the Gulf of Mexico (Florida Escarpment). Mar Ecol 28: 131–151.
Levin LA, Michener R . (2002). Isotopic evidence of chemosynthesis-based nutrition of macrobenthos: the lightness of being at Pacific methane seeps. Limnol Oceanog 47: 1336–1345.
Levin LA, Ziebis W, Mendoza G, Growney V, Tryon M, Brown K et al. (2003). Spatial heterogeneity of macrofauna at northern California methane seeps: the influence of sulfide concentration and fluid flow. Mar Ecol Prog Ser 265: 123–139.
Lewis T, Nichols PD, McMeekin TA . (2000). Evaluation of extraction methods for recovery of fatty acids from lipid-producing microheterotrophs. J Microbiol Methods 43: 107–116.
Lipp JS, Morono Y, Inagaki F, Hinrichs K-U . (2008). Significant contribution of Archaea to extant biomass in marine subsurface sediments. Nature 424: 991–994.
Lösekann T, Robador A, Niemann H, Knittel K, Boetius A, Dubilier N . (2008). Endosymbioses between bacteria and deep-sea siboglinid tubeworms from an Arctic Cold Seep (Haakon Mosby Mud Volcano, Barents Sea). Environ Microbiol 10: 3237–3254.
Luff R, Wallmann K, Aloisi G . (2004). Numerical modeling of carbonate crust formation at cold vent sites: significance for fluid and methane budgets and chemosynthetic biological communities. Eart Planet Sci Lett 22: 337–353.
MacAvoy SE, Macko SA, Carney RS . (2003). Links between chemosynthetic production and mobile predators on the Louisiana continental slope: stable carbon isotopes of specific fatty acids. Chem Geol 201: 229–237.
Marsh AG, Harvey HR, Gremaré A, Tenore KR . (1990). Dietary effects on oocyte yolk-composition in Capitella sp. I (Annelida: Polychaeta): fatty acids and sterols. Mar Biol 106: 369–374.
McCaffrey MA, Farrington JW, Repeta DJ . (1989). Geochemical implications of the lipid composition of Thioploca spp. from the Peru upwelling region—15 S. Org Geochem 14: 61–68.
McNaughton SJ . (1985). Ecology of a grazing ecosystem: the Serengeti. Ecol Monogr 55: 259–294.
Menot L, Galéron J, Olu K, Caprais J-C, Crassous P, Khripounoff A et al. (2010). Spatial heterogeneity of macrofaunal communities in and near a giant pockmark area in the deep Gulf of Guinea. Mar Ecol 31: 78–93.
Niemann H, Elvert M . (2008). Diagnostic lipid biomarker and stable carbon isotope signatures of microbial communities mediating the anaerobic oxidation of methane with sulphate. Org Geochem 39: 1668–1677.
Oba M, Sakata S, Tsunogai U . (2006). Polar and neutral isopranyl glycerol ether lipids as biomarkers of Archaea in near-surface sediments from the Nankai trough. Org Geochem 37: 1643–1654.
Orphan VJ, Hinrichs K-U, Ussler III W, Paull CK, Taylor LT, Sylva SP et al. (2001a). Comparative analysis of methane-oxidizing Archaea and sulfate-reducing bacteria in anoxic marine sediments. Appl Environ Microbiol 67: 1922–1934.
Orphan VJ, House CH, Hinrichs KU, Mckeegan KD, Delong EF . (2001b). Methane-consuming Archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 293: 484–487.
Orphan VJ, House CH, Hinrichs KU, McKeegan KD, DeLong EF . (2002). Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc Natl Acad Sci USA 99: 7663–7668.
Orphan VJ, Turk KA, Green AM, House CH . (2009). Patterns of 15N assimilation and growth of methanotrophic ANME-2 Archaea and sulfate–reducing bacteria within structured syntrophic consortia revealed by FISH-SIMS. Environ Microbiol 11: 1777–1791.
Pond DW, Allen CE, Bell MV, Van Dover CL, Fallick AE, Dixon DR et al. (2002). Origins of long-chain polyunsaturated fatty acids in the hydrothermal vent worms Ridgea piscesae and Protis hydrothermica. Mar Ecol Prog Ser 225: 219–216.
Preston CM, Ying Wu K, Molinski TF, DeLong EF . (1996). A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov. Proc Natl Acad Sci 93: 6241–6246.
Qiu D, Games M, Xiao X, Games D, Walton T . (2000). Characterisation of membrane phospholipids and glycolipids from a halophilic archaebacterium by high-performance liquid chromatography/electrospray mass spectrometry. Rapid Commun Mass Spectrom 14: 1586–1591.
Reeburg WS . (2007). Oceanic methane biogeochemistry. Chem Rev 107: 486–513.
Ritt B, Sarrazin J, Caprais J-C, Noël P, Gauthier O, Pierre C et al. (2010). First insights into the structure and environmental setting of cold-seep communities in the Marmara Sea. Deep-Sea Res I 57: 1120–1136.
Sommer S, Pfannkucke O, Linke P, Luff R, Greinert J, Drews M et al. (2006). Efficiency of the benthic filter: biological control of emission of dissolved methane from sediments containing shallow gas hydrates at Hydrate Ridge. Global Biogeochem Cycles 20: 1–14.
Stadnitskaia A, Bouloubassi I, Elvert M, Hinrichs KU, Sinninghe Damsté J . (2008). Extended hydroxyarchaeol, a novel lipid biomarker for anaerobic methanotrophy in cold seepage habitats. Organic Geochem 39: 1007–1014.
Stiehl T, Rullkotter J, Nissenbaum A . (2005). Molecular and isotopic characterization of lipids in cultured halophilic microorganisms from the Dead Sea and comparison with the sediment record of this hypersaline lake. Organic geochem 36: 1242–1251.
Thiel V, Peckmann J, Seifert R, Wehrung P, Reitner J, Michaelis W . (1999). Highly isotopically depleted isoprenoids: molecular markers for ancient methane venting. Geochimica et Cosmochimica Acta 63: 3959–3966.
Treude T, Boetius A, Knittel K, Wallmann K, Jørgensen BB . (2003). Anaerobic oxidation of methane above gas hydrates at Hydrate Ridge, NE Pacific Ocean. Mar Ecol Prog Ser 264: 1–14.
Valentine DL . (2007). Adaptations to energy stress dictate the ecology and evolution of the Archaea. Nat Rev Microbiol 5: 316–323.
van Hoek AHAM, van Alen TA, Sprakel VSI, Leunissen JAM, Brigge T, Bogels GD et al. (2000). Multiple acquisition of methanogenic archaeal symbionts by anaerobic ciliates. Mol Biol Evol 17: 251–258.
Van de Vossenberg JLCM, Driessen AJM, Konings WN . (1998). The essence of being extremophilic: the role of the unique archaeal membrane lipids. Extremophiles 2: 163–170.
Werne JP, Baas M, Sinninghe Damste JS . (2002). Molecular isotopic tracing of carbon flow and trophic relationships in a methane-supported benthic microbial community. Limnol Oceanogr 47: 1694–1701.
Whiticar MJ . (1998). Carbon and hydrogen isotope systematic of bacterial formation and oxidation of methane. Chem Geol 161: 291–314.
Worm B, Duffy JE . (2003). Biodiversity, productivity, and stability in real food webs. Trends Ecol Evo 18: 628–632.
Wuchter C, Schouten S, Boshker HTS, Sinninghe Damste JS . (2003). Bicarbonate uptake by marine Crenarchaeota. FEMS Microbiol Lett 219: 203–220.
Acknowledgements
We are indebted to the Captains, Crew, Alvin group and Science parties from RV Atlantis legs 15-9; 15-11; 15-44; and 15-59. Guillermo Mendoza, Jennifer Gonzalez and Drs Brigitte Ebbe, Ken Halanych, Ray Lee and Greg Rouse all helped tremendously at sea with sorting and identification of the dorvilleid species. Dr Shana Goffredi kindly checked for archaeal symbionts within the Costa Rican Dorvilleid. Dr William Gerwick provided access to the analytical instrumentation necessary to carry out this research and Cameron Coates, Jo Nunnery and Tak Suyama were always present to help trouble shoot analytical problems. Two anonymous reviewers, and Drs G Rouse and Lihini Aluwihare provided helpful comments on an earlier version of this manuscript. This research was supported by NSF Grants OCE 0425317, OCE 0826254 and OCE-0939557, and grant UAF-050141 from the West Coast National Undersea Research Center to LA Levin, OCE-0939559 and OCE-0825791 grants to VJ Orphan and a UC Marine Council award to W Gerwick and LA Levin, in addition to the Michael M Mullin Memorial fellowship, Sidney E Frank Foundation Fellowship, UC Marine Council CEQI Fellowship and the graduate office of Scripps Institution of Oceanography's support of AR Thurber.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Thurber, A., Levin, L., Orphan, V. et al. Archaea in metazoan diets: implications for food webs and biogeochemical cycling. ISME J 6, 1602–1612 (2012). https://doi.org/10.1038/ismej.2012.16
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.16
Keywords
This article is cited by
-
Symbiotic Interactions of Archaea in Animal and Human Microbiomes
Current Clinical Microbiology Reports (2023)
-
Labelling halophilic Archaea using 13C and 15N stable isotopes: a potential tool to investigate haloarchaea consumption by metazoans
Extremophiles (2019)
-
Comparison of Archaeal and Bacterial Diversity in Methane Seep Carbonate Nodules and Host Sediments, Eel River Basin and Hydrate Ridge, USA
Microbial Ecology (2015)