Abstract
Magnetosomes are prokaryotic organelles produced by magnetotactic bacteria that consist of nanometer-sized magnetite (Fe3O4) or/and greigite (Fe3S4) magnetic crystals enveloped by a lipid bilayer membrane. In magnetite-producing magnetotactic bacteria, proteins present in the magnetosome membrane modulate biomineralization of the magnetite crystal. In these microorganisms, genes that encode for magnetosome membrane proteins as well as genes involved in the construction of the magnetite magnetosome chain, the mam and mms genes, are organized within a genomic island. However, partially because there are presently no greigite-producing magnetotactic bacteria in pure culture, little is known regarding the greigite biomineralization process in these organisms including whether similar genes are involved in the process. Here using culture-independent techniques, we now show that mam genes involved in the production of magnetite magnetosomes are also present in greigite-producing magnetotactic bacteria. This finding suggest that the biomineralization of magnetite and greigite did not have evolve independently (that is, magnetotaxis is polyphyletic) as once suggested. Instead, results presented here are consistent with a model in which the ability to biomineralize magnetosomes and the possession of the mam genes was acquired by bacteria from a common ancestor, that is, the magnetotactic trait is monophyletic.
Similar content being viewed by others
Introduction
Magnetite-producing magnetotactic bacteria (MTB) are phylogenetically affiliated with the Alphaproteobacteria, Gammaproteobacteria and Deltaproteobacteria classes of the Proteobacteria and the Nitrospirae phylum (Bazylinski and Frankel, 2004; Lefèvre et al., 2010a). Greigite-producing MTB have not been cultured and include a group of morphologically similar multicellular magnetotactic prokaryotes affiliated with Deltaproteobacteria (Abreu et al., 2007; Simmons and Edwards, 2007) and large rod-shaped bacteria (Pósfai et al., 1998a), whose phylogeny has not been studied in detail. One report suggests that at least one type of greigite-producing MTB is affiliated with the Gammaproteobacteria (Simmons et al., 2004). However, some doubt has been raised regarding the true phylogenetic relationship of this organism (Amann et al., 2006) and the recent isolation of two new gammaproteobacterial rod-shaped MTB (Lefèvre et al., 2010b) suggests that the organism described by Simmons et al. (2004) biomineralizes magnetite. Confirmed greigite-producing MTB whose 16S rRNA gene has been sequenced include only the multicellular magnetotactic prokaryote Candidatus Magnetoglobus multicellularis and bacteria morphologically similar to it (Abreu et al., 2007; Simmons and Edwards, 2007). This microorganism consists of an assemblage of genetically identical, Gram-negative bacteria that are capable of collectively migrating along magnetic field lines because of the coordinated rotation of flagella that cover each cell on one side. Individual cells of this microorganism do not move or respond in a magnetic field. Ca. M. multicellularis is characterized by a multicellular life cycle with no apparent cell differentiation and intercellular communication between cells (Abreu et al., 2007). The first report about greigite-producing MTB and its affiliation to Deltaproteobacteria suggest, but do not confirm, that magnetotaxis based on iron oxide and iron sulfide magnetosomes evolved independently and that the trait was polyphyletic (DeLong et al., 1993).
Magnetite magnetosome formation in the Alphaproteobacteria has been extensively characterized in some cultured species (Jogler and Schüler, 2009) and briefly studied in uncultured organisms from environmental samples (Jogler et al., 2009b). The genes responsible for magnetite biomineralization, the mam and mms genes, make up the mamAB, mamGFDC, mamXY and mms6 operons (Jogler and Schüler, 2009) in a genomic island known as the magnetosome island (MAI; Schüler, 2004). These genes are responsible for controlling the size and morphology of magnetite crystals in MTB, as well as magnetosome chain organization (Schüler, 2004). Comparisons between the MAIs of different cultured magnetite-producing MTB show that gene content and organization differ among them and are thought to be responsible for differences in magnetosome crystal morphology and size and magnetosome organization (Jogler et al., 2009a).
The ability of magnetosome synthesis is thought to have been distributed between these organisms by horizontal gene transfer of the MAI (Jogler and Schüler, 2009). According to a current model (Jogler et al., 2009a), magnetite magnetosome genes were acquired by the magnetospirilla, the coccus Candidatus Magnetococcus marinus strain MC-1 (Schübbe et al., 2009) and the vibrio Candidatus Magnetovibrio blakemorei strain MV-1 (Schübbe et al., 2009) through independent events of horizontal gene transfer from an unknown ancestor. The close phylogenetic affinity of the magnetospirilla to a genus of photosynthetic bacteria, Phaeospirillum, suggests that the ancestor of these two groups of prokaryotes might have been a phototrophic organism. The absence of selective pressure for magnetotaxis and the loss of the MAI might have led to the occurrence of non-magnetotactic representatives within the MTB (Jogler et al., 2009a). This model, however, describes magnetotaxis evolution only within the Alphaproteobacteria and considers only genes related to magnetite biomineralization. Genes for greigite magnetosome formation have not yet been found until now, so they have not been considered in this evolution model of magnetotaxis.
In this work, we show that genes related to magnetite biomineralization are conserved in greigite-producing MTB, suggesting that similar genes are involved in the biomineralization of both magnetite and greigite, but also that biomineralization of magnetite and greigite might not have evolved independently (that is, magnetotaxis is polyphyletic) as once suggested (DeLong et al., 1993). Results presented here are consistent with a model in which the ability to biomineralize magnetosomes and the possession of the mam genes were possibly acquired by bacteria of the Nitrospirae and certainly acquired by Proteobacteria from an ancient common ancestor in independents horizontal gene transfer events, that is, the magnetotactic trait is monophyletic. This hypothesis is supported by the fact that Ca. M. multicellularis mam genes, when analyzed individually, are more related to orthologues found in both Deltaproteobacteria and also Alphaproteobacteria MTB and, when analyzed together, they are recovered as a deep branching lineage within bacteria.
Materials and methods
Sampling and sequencing
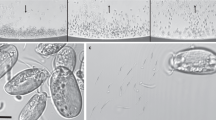
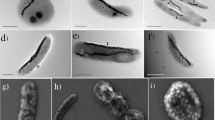
Because Ca. M. multicellularis has not been cultured, DNA samples were prepared according to a modified magnetic enrichment procedure (Lins et al., 2003) which is based on capillary racetrack principle (Wolfe et al., 1987). Water and sediment were collected from Araruama Lagoon, Rio de Janeiro, Brazil (22° 50′S, 42° 13′W) and magnetic enrichment was carried out with additional washing steps using lagoon sterile autoclaved water. This procedure yielded a highly concentrated sample of Ca. M. multicellularis. DNA extraction was performed (Chen and Kuo, 1993) and genomic DNA was amplified using REPLI-g mini kit (Qiagen, Hilden, Germany). Ca. M. multicellularis DNA was sequenced on 454 GS FLX System sequencer (Roche Diagnostics GmbH/454 Life Sciences Corporation, Branford, CT, USA). The purity of the Ca. M. multicellularis cell sample for DNA extraction was checked using light and electron microscopy on Zeiss Axiostar Plus microscope (Carl Zeiss, Oberkochen, Germany) and FEI Morgagni transmission electron microscope (FEI Company, Eindhoven, The Netherlands). The detection of Ca. M. multicellularis was greatly simplified by its unique morphology and the presence of magnetosomes (Supplementary Figure 1). No unicellular bacterium with or without magnetosomes was observed. The purity of the samples was also checked by amplification and sequencing of 16S rRNA genes according to Abreu et al. (2007). The only 16S rRNA gene obtained from MTB was that of Ca. M. multicellularis.
Comparative analysis of magnetosome genes
The system for automated bacterial integrated annotation platform (Almeida et al., 2004) was used to predict open reading frames (ORFs) position and sequence analysis using a tblastx tool (Altschul et al., 1997). MAI genes of M. magneticum AMB-1 (AP007255), M. gryphiswaldense MRS-1 (AM085146), M. magnetotacticum MS-1 (NZ_AAAP01003731), Ca. Magnetococcus marinus strain MC-1 (NC_008576), Ca. Magnetovibrio blakemorei strain MV-1 (FP102531) and Desulfovibrio magneticus (AP010904) were compared with Ca. M. multicellularis sequences using tblastx for identity, positives and e-value analysis (Altschul et al., 1997). Contigs having valid similarity values (e-value <1e-05) with mam genes were completely analyzed. Similar sequences of each MTB were compared between them for e-value determination. These sequences have been submitted to the GenBank databases under accession numbers HQ336745 and HQ336746.
Phylogenetic and bioinformatic analyses
Maximum likelihood trees based on 16S rRNA gene sequences and amino acid composition of mam genes were constructed although, for this study, phylogenetic analysis of Mam protein amino acid sequences was favored because sequences from distantly related taxa were analyzed (Opperdoes, 2009). Sequence alignment was performed with Muscle (version 3.6; Edgar, 2004). The general time-reversible model (GTR; Yang, 1994; Zharkikh, 1994) with gamma-distributed substitution rates was selected as the DNA substitution model by Modelgenerator (version 0.84; Keane et al. 2006). The Bayesian information criterion was used for the phylogenetic reconstruction. A maximum likelihood tree was then reconstructed using PhyML (version 3.0; Guindon and Gascuel, 2003). The tree topology space was explored using the nearest neighbor interchange and subtree pruning and regrafting algorithms starting from five random starting trees generated by BioNJ (Guindon and Gascuel, 2003; Guindon et al., 2010). Branch support was calculated using the approximate likelihood ratio test (aLRT) with SH-like interpretation. This approach is as conservative and accurate as bootstrapping but is less computationally intensive (Anisimova and Gascuel, 2006; Guindon et al., 2010).
Results
Partial genome sequencing of Ca. M. multicellularis and comparative analysis based on known mam and mms genes revealed the following orthologues (Table 1): mamA, mamB, mamE, mamK, mamM, mamO, mamP, mamQ and mamT. These genes were found in two segments of DNA (contigs) shown in Figure 1. Contig 1 (GenBank accession number HQ336746) is 15 950 bp long and consists of 754 reads (20 × coverage) whereas contig 2 (GenBank accession number HQ336745) is 9131 bp long and consists of 618 reads (30 × coverage), confirming data reliability and accuracy. Besides putative mam genes, contig 1 contains ORFs encoding for other genes possibly involved in magnetosome synthesis including an iron-dependent repressor, a putative ferrous iron transporter and several hypothetical proteins. The tblastx analysis of the first two sequences showed that they are similar to a putative iron repressor (e-value 7e-49; coverage 86%; identity 55%; positives 74%) and ferrous iron transporter found in the MAI (e-value 6e-110; coverage 85%; identity 52%; positives 73%) of the magnetite-producing deltaproteobacterium Desulfovibrio magneticus (Nakazawa et al., 2009). The position of these genes is also similar to D. magneticus as they are located upstream of mam genes. ORFs similar to an ATPase domain protein (2e-103; coverage 96%; identity 42%; positives 60%) and a hypothetical protein (e-value 8e-71; coverage 73%; identity 50%; positives 68%) found in D. magneticus were also present.
Organization of open reading frames in two segments of DNA (contigs) from Candidatus Magnetoglobus multicellularis containing putative magnetosome-related genes compared to the organization of magnetosome genes within the magnetosome gene island of several cultivated magnetotactic bacteria. Scale bar indicates 1000 bp. Hypothetical proteins are represented as unlabeled arrows.
The G+C content of the two contigs containing mam genes is higher than the contig containing the rrs gene of Ca. M. multicellularis and putative genes involving metabolic pathways, possibly indicating that this genomic region is part of a MAI acquired by horizontal gene transfer from other organisms as described for magnetite-producing MTB (Jogler et al., 2009a, 2009b). The conserved genes mamA, mamB, mamE, mamO and mamP, mamQ and mamT appear to be organized in an operon; these genes are most likely essential for greigite magnetosome formation, as they may participate in the assembly of multiprotein complexes for magnetosome formation, iron transport and magnetosome organization (Jogler and Schüler, 2009). Interestingly, Ca. M. multicellularis contains two ORFs similar to the mamP gene. However, these sequences are more similar to each other (e-value 7e-19; coverage 61%; identities 34%; positives 59%) than to mamP from other MTB.
The mms6, mamD, mamC, mamF, mamG, mamJ, mamX and mamY genes, strongly conserved in most magnetite-producing magnetotactic Alphaproteobacteria (Jogler et al., 2009a), were not found in our sequences or in the genome of D. magneticus (Nakazawa et al., 2009). These genes are considered to be of great importance in controlling magnetosome size and morphology in MTB of the Alphaproteobacteria. Nakazawa et al. (2009) suggested that their absence in D. magneticus might explain the presence of irregular, bullet-shaped magnetosomes in this species rather than the consistent hexa- and octahedra produced by Alphaproteobacteria MTB that possess these genes. Greigite magnetosome crystals, such as the bullet-shaped magnetite crystals, are pleomorphic (Abreu et al., 2007), which might be an indication that orthologues of these genes are not present in MTB that biomineralize greigite and bullet-shaped magnetite crystals. Moreover, some greigite-producing MTB biomineralize bullet-shaped magnetite crystals in addition to greigite (Bazylinski et al., 1993). Our results suggest that the minimum magnetosome-related genes required for the synthesis of magnetite or greigite magnetosomes include mamA, mamB, mamE, mamK, mamM, mamO, mamP, mamQ and mamT, which are shared among MTB of both the Alphaproteobacteria and Deltaproteobacteria.
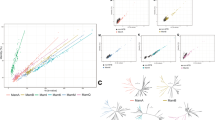
The highest similarity values according to tblastx analysis for ORFs from the contigs of Ca. M. multicellularis were for mamA (e-value 2e-25), mamB (e-value 3e-62), mamK (e-value 8e-137), mamO (e-value 1e-49), mamP* (e-value 7e-11), mamP (e-value 3e-19) and mamT (e-value 8e-08), all from D. magneticus, whereas mamE (e-value 1e-50) and mamQ (e-value 2e-18) were more similar to those of Ca. Magnetococcus marinus and mamM (e-value 1e-14) to that of Ca. Magnetovibrio blakemorei strain MV-1. Ca. M. multicellularis Mam amino acid sequences, when analyzed together, are more similar to those of D. magneticus, which is in accordance with the phylogenetic analysis based on 16S rRNA gene sequences (Figures 1 and 2). Phylogenetic analysis based on Mam amino acids sequences suggests that a more recent horizontal gene transfer occurred from Ca. M. multicellularis and D. magneticus than the other species, but they are still distantly related (Figure 2).
Phylogenetic analyses of Candidatus Magnetoglobus multicellularis: (a) maximum likelihood tree based on 16S rRNA gene sequences using the GTR+Γ DNA substitution model. Numbers at nodes represent bootstrap values for 1000 replicates. Numbers in parentheses are GenBank accession numbers. (b) Amino acid composition of conserved mam genes using the WAG+I+Γ+F matrix. Sequences of the uncultured MTB C. Thermomagnetovibrio paiutensis HSMV-1 (Lefèvre et al., 2010a) and Ca. Magnetobacterium bavaricum (Spring et al., 1993), both phylogenetically affiliated with the Nitrospirae phylum, are included as outgroups in this analysis. Numbers between branches are aLRT values.
Discussion
The relatively high similarity values found between the mam orthologues in Ca. M. multicellularis and the mam genes of MTB of both the Alphaproteobacteria (mamE, mamM and mamQ) and Deltaproteobacteria (mamA, mamB, mamK, mamO, mamP and mamT) suggests that these genes evolved from a common ancestor. Moreover, phylogenetic relationships of MTB based on Mam amino acid sequences and on 16S rRNA gene sequences suggest that magnetite and greigite biomineralization did not evolve independently, at least in the Proteobacteria. Phylogenetic analysis based on Mam protein amino acids sequences and mam genes content suggest that D. magneticus and Ca. M. multicellularis might share an unknown magnetotactic deltaproteobacterial ancestor. According to the prokaryotic evolution point of view (Gogarten et al., 2002), frequency of successful exchange between taxa will depend on specific factors, including propinquity, metabolic compatibility, adaptation to the abiotic environment, gene expression systems and gene-transfer mechanisms. Based on this, acquisition of mam genes by horizontal gene transfer between D. magneticus and Ca. M. multicellularis might have a high probability as they both inhabit anoxic sulfide-rich environments below the oxic–anoxic transition zone, unlike almost all other magnetite-producing MTB.
A number of magnetosome-related genes, including mamA, mamB, mamE, mamI, mamM, mamP and mamQ, were recently found to be present in the genome of Candidatus Magnetobacterium bavaricum, a magnetite-producing MTB phylogenetically affiliated with the Nitrospirae phylum (Jogler et al., 2011). These genes not only show similar sequence homologies to others of their type, the organization of these genes show similar short intergenic distances between them and an identical direction of transcription, providing evidence that they are organized as an operon as described for other MTB (Schübbe et al., 2006). The presence of mam genes and their organization in the Nitrospirae and various classes of the Proteobacteria strongly supports a monophyletic origin for magnetite-based magnetotaxis. Because both the greigite-producing Ca. M. multicellularis and the magnetite-producing MTB share similar mam genes, the capability of magnetosome, regardless of whether they contain iron oxide or iron sulfide crystals, synthesis in all currently known MTB appears to be a result of the acquisition of mam genes by independent horizontal gene transfer events from a common ancestor during evolution.
This scenario raises a number of interesting and important questions. What was the first magnetic mineral biomineralized by the MTB? It seems from the information presented here that it was likely magnetite, as MTB from the most deeply branching groups of Bacteria (Emerson et al., 2007) that contain them, the Nitrospirae and the Deltaproteobacteria produce this mineral. Moreover, MTB from these groups are known to biomineralize only bullet-shaped magnetite crystals suggesting that this crystal morphology is the earliest form of magnetosome magnetite. If true, this finding has important implications in the finding and interpretation of magnetofossils, the putative remains of MTB magnetite crystals in ancient and recent sediments (Jimenez-Lopez et al., 2010). It is unclear if any of the mam genes are involved in the selective precipitation of magnetite or greigite in magnetosomes. Greigite formation may have been a modification of magnetite biomineralization in some sulfate-reducing MTB such as Ca. M. multicellularis under certain conditions. This possibility is supported by the fact that the positioning of Ca. M. multicellularis in sediment is influenced by the physicochemical properties of the microenvironment, for example, the redox potential (Eh) and iron/sulfur availability (Sobrinho et al., 2011). According to Sobrinho et al. (2011) Ca. M. multicellularis is positioned where the Eh and pH facilitates the formation of iron monosulfides, a condition necessary for the formation of greigite in magnetosomes (Pósfai et al., 1998b). In addition, environmental conditions appear to influence the magnetosome mineral composition in some greigite-producing rod-shaped bacteria that also produce magnetite (Bazylinski et al., 1995).
More recently diverging groups of the Proteobacteria (Emerson et al., 2007), the Alpha- and Gammaproteobacteria, that contain MTB that biomineralize cubooctahedral and elongated prismatic crystal morphologies of magnetite (Bazylinski and Frankel, 2004; Lefèvre et al., 2010b) and possess more mam genes, might have acquired this ability through gene mutation and gene duplication or acquisition of additional mam-like genes that were later distributed between species of these groups by horizontal gene transfer. The presence of mam genes may not be sufficient to explain all the differences in morphology and mineral content of magnetosomes; the presence of a membrane enveloping the crystals appears to be essential to the biomineralization of the consistent, regularly-shaped magnetosome morphologies (for example, hexahedral prisms) observed in some species. Our data show that the mam gene content of D. magneticus and Ca. M. multicellularis is similar but these microorganisms appear to be distinct in the presence/absence of a membrane enveloping the crystals and in their magnetosome crystal morphologies. The absence of a magnetosome membrane in D. magneticus (Byrne et al., 2010) may explain the irregular morphologies of magnetite formed by this microorganism despite the presence of mam genes in its genome (Nakazawa et al., 2009). Ca. M. multicellularis magnetosomes are enveloped by a membrane (Abreu et al., 2008) similar to those present in some Nitrospirae (Lefèvre et al., 2011) and in magnetotactic multicellular prokaryotes capable of simultaneous biomineralization of magnetite and greigite magnetosomes (Lins et al., 2007); these uncultured microorganisms produce regularly shaped magnetosomes. It is also possible that the process of magnetite biomineralization used by D. magneticus is different under the culture conditions used to grow this organism compared with what occurs in nature. Further investigation of the physiology of divergent MTB species is needed to achieve a deeper understanding of the differences between magnetite and greigite biomineralization in MTB.
References
Abreu F, Martins JL, Silveira TS, Keim CN, Lins de Barros HG P, Gueiros-Filho F et al. (2007). ‘Candidatus Magnetoglobus multicellularis’ a multicellular magnetotactic prokaryote from a hypersaline environment. Int J Syst Bacteriol 57: 1318–1322.
Abreu F, Silva KT, Farina M, Keim C, Lins U . (2008). Greigite magnetosome membrane ultrastructure in Candidatus Magnetoglobus multicellularis. Int Microbiol 11: 75–80.
Almeida LG, Paixão R, Souza RC, Costa GC, Barrientos FJ, Santos MT et al. (2004). A system for automated bacterial (genome) integrated annotation—SABIA. Bioinformatics 20: 2832–2833.
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 389–3402.
Amann R, Peplies J, Schüler D . (2006). Diversity and taxonomy of magnetotactic bacteria. In: Schüler D (ed). Magnetoreception and Magnetosomes in Bacteria. Springer: Berlin, Germany, pp 25–36.
Anisimova M, Gascuel O . (2006). Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol 55: 539–552.
Bazylinski DA, Frankel RB . (2004). Magnetosome formation in prokaryotes. Nat Rev Microbiol 2: 217–230.
Bazylinski DA, Frankel RB, Heywood BR, Mann S, King JW, Donaghay PL et al. (1995). Controlled biomineralization of magnetite (Fe3O4) and greigite (Fe3S4) in a magnetotactic bacterium. Appl Environ Microbiol 61: 3232–3239.
Bazylinski DA, Heywood BR, Mann S, Frankel RB . (1993). Fe3O4 and Fe3S4 in a bacterium. Nature 366: 218.
Byrne ME, Ball DA, Guerquin-Kern JL, Rouiller I, Wu TD, Downing KH et al. (2010). Desulfovibrio magneticus RS-1 contains an iron- and phosphorus-rich organelle distinct from its bullet-shaped magnetosomes. Proc Natl Acad Sci USA 107: 12263–12268.
Chen W, Kuo T . (1993). A simple and rapid method for the preparation of Gram-negative bacterial genomic DNA. Nucleic Acids Res 21: 2260.
DeLong EF, Frankel RB, Bazylinski DA . (1993). Multiple evolutionary origins of magnetotaxis in bacteria. Science 259: 803–806.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Emerson D, Rentz JA, Lilburn TG, Davis RE, Aldrich H, Chan C et al. (2007). A novel lineage of proteobacteria involved in formation of marine Fe-oxidizing microbial mat communities. PLoS ONE 8: e667.
Gogarten JP, Doolittle WF, Lawrence JG . (2002). Prokaryotic evolution in light of gene transfer. Mol Biol Evol 19: 2226–2238.
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O . (2010). New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59: 307–321.
Guindon S, Gascuel O . (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52: 696–704.
Jimenez-Lopez C, Romanek CS, Bazylinski DA . (2010). Magnetite as a prokaryotic biomarker: a review. J Geophys Res-Biogeo 115: G00G03.
Jogler C, Kube M, Schübbe S, Ullrich S, Teeling H, Bazylinski DA et al. (2009a). Comparative analysis of magnetosome gene clusters in magnetotactic bacteria provides further evidence for horizontal gene transfer. Environ Microbiol 15: 1267–1277.
Jogler C, Lin W, Meyerdierks A, Kube M, Katzmann E, Flies C et al. (2009b). Toward cloning of the magnetotactic metagenome: identification of magnetosome island gene clusters in uncultivated magnetotactic bacteria from different aquatic sediments. Appl Environ Microbiol 75: 3972–3979.
Jogler C, Schüler D . (2009). Genomics, genetics, and cell biology of magnetosome formation. Annu Rev Microbiol 63: 501–521.
Jogler C, Wanner G, Kolinko S, Nieber M, Amann R, Petersen N et al. (2011). Conservation of proteobacterial magnetosome genes and structures in an uncultivated member of the deep-branching Nitrospira phylum. Proc Natl Acad Sci USA 108: 1134–1139.
Keane TM, Creevey CJ, Pentony MM, Naughton TJ, McLnerney JO . (2006). Assessment of methods for amino acid matrix selection and their use on empirical data shows that ad hoc assumptions for choice of matrix are not justified. BMC Evol Biol 6: 29.
Lefèvre CT, Abreu F, Schmidt ML, Lins U, Frankel RB, Hedlund BP et al. (2010a). Moderately thermophilic magnetotactic bacteria from hot springs in Nevada. Appl Environ Microbiol 76: 3740–3743.
Lefèvre CT, Frankel RB, Abreu F, Lins U, Bazylinski DA . (2011). Culture-independent characterization of a novel, uncultivated magnetotactic member of the Nitrospirae phylum. Environ Microbiol 13: 538–549.
Lefèvre CT, Viloria N, Abreu F, Pósfai M, Prozorov T, Lins U et al. (2010b). Phylogenetic diversity and ecophysiology of new cultured and uncultured magnetite-producing magnetotactic bacteria from aquatic environments in the desert. Abstract, 2nd International Symposium on Magnetotactic Bacteria and Biomineralization, Beijing, China. pp 34–36.
Lins U, Freitas F, Keim CN, Lins de Barros HGP, Esquivel DSM, Farina M . (2003). Simple homemade apparatus for harvesting uncultured magnetotactic microorganisms. Braz J Microbiol 34: 111–116.
Lins U, Keim CN, Evans FF, Farina M, Buseck PR . (2007). Magnetite (Fe3O4) and greigite (Fe3S4) crystals in multicellular magnetotactic prokaryotes. Geomicrobiol J 24: 43–50.
Nakazawa H, Arakaki A, Narita-Yamada S, Yashiro I, Jinno K, Aoki N et al. (2009). Whole genome sequence of Desulfovibrio magneticus strain RS-1 revealed common gene clusters in magnetotactic bacteria. Genome Res 19: 1801–1808.
Opperdoes FR . (2009). Phylogenetic analysis using protein sequences. In: Lemey P, Salemi M, Vandamme A-M (eds). The Phylogenetic Handbook: A Practical Approach to Phylogenetic Analysis and Hypothesis Testing. Cambridge University Press: Cambridge, pp 313–331.
Pósfai M, Buseck PR, Bazylinski DA, Frankel RB . (1998a). Iron sulfides from magnetotactic bacteria: structure, composition, and phase transitions. Am Mineral 83: 1469–1481.
Pósfai M, Buseck PR, Bazylinski DA, Frankel RB . (1998b). Reaction sequence of iron sulfide minerals in bacteria and their use as biomarkers. Science 280: 880–883.
Schübbe S, Williams TJ, Xie G, Kiss HE, Brettin TS, Martinez D et al. (2009). Complete genome sequence of the chemolithoautotrophic marine magnetotactic coccus strain MC-1. Appl Environ Microbiol 75: 4835–4852.
Schübbe S, Würdemann C, Peplies J, Heyen U, Wawer C, Glöckner FO et al. (2006). Transcriptional organization and regulation of magnetosome operons in Magnetospirillum gryphiswaldense. Appl Environ Microbiol 72: 5757–5765.
Schüler D . (2004). Molecular analysis of a subcellular compartment: the magnetosome membrane in Magnetospirillum gryphiswaldense. Arch Microbiol 181: 1–7.
Simmons SL, Bazylinski DA, Edwards KJ . (2004). Spatiotemporal distribution of marine magnetotactic bacteria in a seasonally stratified coastal salt pond. Appl Environ Microbiol 70: 6230–6239.
Simmons SL, Edwards KJ . (2007). Unexpected diversity in populations of the many-celled magnetotactic prokaryote. Environ Microbiol 9: 206–215.
Sobrinho R, Lins U, Bernardes M . (2011). Geochemical characteristics related to the greigite-producing multicellular magnetotactic prokaryote ‘Candidatus Magnetoglobus multicellularis’ in a hypersaline lagoon. Geomicrobiol J (in press).
Spring S, Amann R, Ludwig W, Schleifer KH, van Gemerden H, Petersen N . (1993). Dominating role of an unusual magnetotactic bacterium in the microaerobic zone of freshwater sediment. Appl Environ Microbiol 59: 2397–2403.
Wolfe RS, Thauer RK, Pfennig N . (1987). A ‘capillary racetrack’ method for isolation of magnetotactic bacteria. FEMS Microbiol Ecol 45: 31–35.
Yang Z . (1994). Estimating the pattern of nucleotide substitution. J Mol Evol 39: 105–111.
Zharkikh A . (1994). Estimation of evolutionary distances between nucleotide sequences. J Mol Evol 39: 315–329.
Acknowledgements
The financial support from the Brazilian agencies, CNPq, CAPES and FAPERJ is acknowledged. CTL and DAB were supported by the US National Science Foundation grant EAR-0920718.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Abreu, F., Cantão, M., Nicolás, M. et al. Common ancestry of iron oxide- and iron-sulfide-based biomineralization in magnetotactic bacteria. ISME J 5, 1634–1640 (2011). https://doi.org/10.1038/ismej.2011.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.35
Keywords
This article is cited by
-
Renaissance for magnetotactic bacteria in astrobiology
The ISME Journal (2023)
-
Magnetotactic bacteria and magnetofossils: ecology, evolution and environmental implications
npj Biofilms and Microbiomes (2022)
-
Occurrence of south- and north-seeking multicellular magnetotactic prokaryotes in a coastal lagoon in the South Hemisphere
International Microbiology (2022)
-
Magnetotactic bacteria: concepts, conundrums, and insights from a novel in situ approach using digital holographic microscopy (DHM)
Journal of Comparative Physiology A (2022)
-
Expanding magnetic organelle biogenesis in the domain Bacteria
Microbiome (2020)