Abstract
Terephthalate (TA) is one of the top 50 chemicals produced worldwide. Its production results in a TA-containing wastewater that is treated by anaerobic processes through a poorly understood methanogenic syntrophy. Using metagenomics, we characterized the methanogenic consortium inside a hyper-mesophilic (that is, between mesophilic and thermophilic), TA-degrading bioreactor. We identified genes belonging to dominant Pelotomaculum species presumably involved in TA degradation through decarboxylation, dearomatization, and modified β-oxidation to H2/CO2 and acetate. These intermediates are converted to CH4/CO2 by three novel hyper-mesophilic methanogens. Additional secondary syntrophic interactions were predicted in Thermotogae, Syntrophus and candidate phyla OP5 and WWE1 populations. The OP5 encodes genes capable of anaerobic autotrophic butyrate production and Thermotogae, Syntrophus and WWE1 have the genetic potential to oxidize butyrate to CO2/H2 and acetate. These observations suggest that the TA-degrading consortium consists of additional syntrophic interactions beyond the standard H2-producing syntroph–methanogen partnership that may serve to improve community stability.
Similar content being viewed by others
Introduction
Terephthalate (TA) is used as the raw material for the manufacture of numerous plastic products (for example, polyethylene TA bottles and textile fibers). During its production, TA-containing wastewater is discharged in large volumes (as high as 300 million m3 per year) and high concentration (up to 20 kg COD (chemical oxygen demand) m−3) (Razo-Flores et al., 2006). This wastewater is generally treated by anaerobic biological processes under mesophilic conditions (∼35 °C). However, anaerobic processes operated at hyper-mesophilic (46–50 °C) and thermophilic (∼55 °C) temperatures may be preferable because of the ability to achieve higher loading rate (van Lier et al., 1997; Chen et al., 2004), which reduces the reactor volume. Moreover, TA wastewater is usually generated at 54–60 °C, and does not require additional energy input for maintaining reactor temperature (Chen et al., 2004). The microbial biomass usually occurs in the form of granules or biofilms attaching on the surface of porous media. Under such environments, TA degradation has been hypothesized (Kleerebezem et al., 1999) to be based on a syntrophic microbial relationship whereby fermentative H2-producing bacteria (syntrophs) convert TA through benzoate to acetate and H2/CO2, and acetoclastic and hydrogenotrophic methanogens further convert the intermediates to methane by physically positioning themselves close to the syntrophs to overcome the thermodynamic barrier (Stams, 1994; Conrad, 1999; Dolfing, 2001).
In practice, the complexities of TA-degrading communities are not as well known. The communities require a long maturation phase (>200–300 days), are difficult to maintain, and do not always result in a successful syntrophic interaction. If the syntrophic interaction is disturbed and the treatment rendered ineffective, the resulting high-concentration effluent must be treated with a more energy-intensive down-stream aerobic biological process. These factors can significantly increase the operational cost of the process and limit its application on a wider scale.
Several studies have investigated the microbial populations present in methanogenic TA degradation, often using laboratory-scale reactors operated at various temperatures (Kleerebezem et al., 1999; Wu et al., 2001; Chen et al., 2004). Using ribosomal RNA (rRNA)-based molecular methods, these studies have found that TA-degrading consortia in bioreactors are dominated by two to three bacterial populations and two types of methanogens (Wu et al., 2001; Chen et al., 2004). The methanogens are relatively straightforward to classify being mainly acetoclastic Methanosaeta-related species and a novel hydrogenotrophic methanogenic species in the family Methanomicrobiales. The syntrophic bacteria, however, are difficult to identify based on phylogenetic classification, and extremely difficult to obtain in pure culture. In the last decade, only three bacterial species that can degrade TA and its isomers have been successfully co-cultured with methanogens under mesophilic conditions (Qiu et al., 2006, 2008), and these isolates are different from those found under thermophilic conditions (Chen et al., 2004). This greatly limits the effort to understand the microbial interaction and function in the TA-degrading consortia.
A metagenome analysis was chosen for this study since it has been proven as an effective method for retrieving nearly complete microbial genomes of dominant populations in relatively simple microbial ecosystems (Tyson et al., 2004; Martin et al., 2006). In particular, this study aims to elucidate the microbiology underpinning anaerobic TA-degrading processes, including improved knowledge of the diversity and physiology of participating syntrophs and methanogens, and the mechanism behind the establishment and maintenance of the partnership. This knowledge may lead to the generation of principles, which could be applied to establish different consortia for treating other chemicals discharged from industrial production lines, or to treat contaminants in other environments.
Materials and methods
The anaerobic microbial consortium that degrades TA was selectively enriched using a 1-l laboratory-scale hybrid bioreactor (Figure 1a) as described previously (Angelidaki et al., 1990; Chen et al., 2004) (see Supplementary Text). Biomass was collected from only porous packing filters on days 221 and 280, and from both filters and sludge bed on days 346 and 430 for further analyses. These biomass samples were used for genomic DNA extraction, library construction and sequencing according to standard protocols (http://www.jgi.doe.gov/sequencing/protocols/prots_production.html) (see Supplementary Text). Detailed metagenome analysis methods are described in the Supplementary Text. Data can be accessed through the Integrated Microbial Genome/Microbiome (http://img.jgi.doe.gov/m/) system.
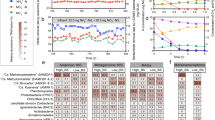
TA-degrading laboratory-scale anaerobic hybrid bioreactor. (a) Schematic of the laboratory-scale anaerobic TA-degrading hybrid reactor operated with a temperature gradient from ∼46 °C at the bottom to 50 °C in the upper zone. Inserts illustrate freshly grown biofilm biomass on the surface of the media after 2 and 11months of enrichment; and (b) performance of the reactor over 480-day operation. Under the initial operational conditions (that is, TA-loading rate of 0.70–0.78 gTA/d.l, and hydraulic retention time (HRT) of 4 days), TA removal efficiency was gradually improved to 72.7% by day 124. By shortening the HRT (3 d on day 127, 2 d on day 168 and then to 1.5 d on day 182) and increasing the TA loading concentration (to 3.2 g on day 364), the TA loading rate was increased to 2.13 gTA/d.l by day 364. Concurrently, the TA removal efficiency increased over the operation period reaching a 99% removal efficiency by day 308. During the entire operation no sulfate reduction activity was detected. Samples were removed at the indicated time points (arrows) and the genomic DNA was extracted for 16S rRNA clone library construction and metagenomics analysis.
Results and discussion
Bioreactor operation and performance
An anaerobic hybrid reactor was successfully operated with TA as the only carbon and energy source for 480 days. This reactor was constructed with an upper section filled with ring-shape porous filters to support the growth of microbial biofilms, and a lower section for the development of anaerobic granular sludge (Figure 1a). This reactor is unusual and novel, in that it was the first methanogenic reactor operated in the hyper-mesophilic temperature zone (46–50 °C), whereas previously published studies of TA-degrading communities were at mesophilic (∼35 °C) or thermophilic (∼55 °C) temperatures (Kleerebezem et al., 1999; Wu et al., 2001; Chen et al., 2004). After achieving a good TA removal efficiency (Figure 1b), sludge samples were taken from the surface of the filter media at days 221, 280, 346 and 430. Samples were also taken from the sludge bed at days 346 and 430. These samples taken were used for 16S rRNA and metagenome analysis.
16S rRNA-based community profiling
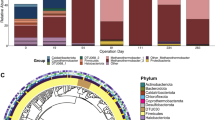
The rarefaction curve analysis indicates that the bacterial population diversity is much higher in terms of operational taxonomic unit number than the archaeal population diversity in any given sample taken from the TA reactor (Supplementary Figure 1). The phylogenetic distribution of bacterial 16S rRNA clones (Figure 2) indicates that Peptococcaceaea (mostly Pelotomaculum), Thermotogae, Syntrophaceae, and candidate phyla OP5 and WWE1 were the dominant bacterial lineages present. Between the biofilm samples (rings 1, 2, 3 and 5) and sludge bed samples (rings 4 and 6) taken, differences in the abundances of major phyla including Firmicutes, Thermotogae, Proteobacteria and OP5 were observed. These differences are likely attributed to the differences in growth temperature (46 °C vs 50 °C) and form (biofilms vs granules). Archaeal representatives were less diverse and consisted of two major types of methanogens belonging to the orders Methanomicrobiales and Methanosarcinales (Supplementary Figure 2). Differences in growth temperature may further explain the variations observed in the microbial populations enriched in previous studies (Supplementary Figure 3). Using the 16S rRNA gene and McrA gene as biomarkers, temperature-dependent variations were also observed with acetoclastic methanogen populations found in the order Methanosarcinales (Supplementary Figure 4). The hydrogenotrophic methanogens identified here are closely related to methanogens found in mesophilic and thermophilic TA-degrading reactors (Wu et al., 2001; Chen et al., 2004), and together with Methanolinea tarda NOBI-1 recently isolated from anaerobic digestion processes (Imachi et al., 2008), form a novel cluster separate from other known hydrogenotrophic methanogens. The comparison of McrA genes further suggests that the methanogens found in the new cluster are likely different from M. tarda NOBI-1 (Supplementary Figure 4B).
Bacterial population dynamics of the TA-degrading bioreactor as revealed by 16S rRNA clone library. Samples (number of 16S rRNA sequences) from inner to outer of the ring chart were day 221 biofilms (287), day 280 biofilms (254), day 346 biofilms (337), day 346 sludge bed (289), day 430 biofilms (352) and day 430 sludge bed (287).
Shotgun sequencing
The assembled sequence data contained 37 818 singlets and 14 526 contiguous fragments of intermediate length (the largest fragment was approximately 240 kb (Supplementary Figure 5) and 45 fragments between 24 and 167 kb). Gene prediction on the entire data set using Genemark resulted in the prediction of 93 104 protein-coding genes. A composition-based classifier, PhyloPythia (McHardy et al., 2007), was used to assign those contigs and singlets into major phylogenetic groups (Table 1), including Pelotomaculum species, candidate phylum OP5 species, Methanolinea species and Methanosaeta species. The highest coverage of an isolate reference genome was observed for M. thermophila (∼80%) followed by Pelotomaculum thermopropionicum (∼60%) (Supplementary Figure 6). However, the Methanolinea population appears to be the best covered one as the average read depth is 5.3 × with many contigs having 10 × read depth (Supplementary Figure 5). In the case of OP5, in which a closely related microbial genome was not present in the database, the occurrence was calculated with the phylogenetic marker clusters of orthologous genes in the OP5 bin (1.41 Mb; G+C content, 28%). Approximately 50% of the OP5 genome is estimated to be covered by the metagenomic data (Table 1).
Pelotomaculum
As a known catabolic-degrading organism abundant in the reactor, Pelotomaculum is assumed to be largely responsible for catabolic degradation of TA to CO2, H2 and acetate. With an average read depth of 3.2 × , 1083 contigs were assigned to the Pelotomaculum population, comprising 4.3 Mb in total (Table 1). We first searched for genes with known decarboxylase functions that are responsible for the first decarboxylation step of TA degradation (Supplementary Figure 7). Two gene sets (tadcc27178-79-80 and tadcc16349) from the Pelotomaculum bin were identified to have high sequence similarity and a subunit complement with a known 4-hydroxybenzoate decarboxylase, EC 4.1.1.61, from Sedimentibacter hydroxybenzoicum that consists of three subunits (AAD50377, AAY67850 and AAY67851) and belongs to the UbiD family of proteins (Lupa et al., 2005). Two mechanisms have been described for the subsequent fermentation of benzoate to acetate and CO2: the well-known benzoyl-CoA reductase (BCR, EC 1.3.99.15) route used by Thauera aromatica (Boll and Fuchs, 1995) and the less understood BCR-independent mechanism for reductive dearomatization (Wischgoll et al., 2005). Examining the TA decarboxylase data set did not show any clear homologs of the Thauera BCRs. Instead, homologs are found in the alternative BCR-independent mechanism within a set of 44 genes that have been postulated to operate in Geobacter metallireducens and ‘Syntrophus aciditrophicus’ (Butler et al., 2007; McInerney et al., 2007; Peters et al., 2007). The key BCR enzyme in G. metallireducens was successfully characterized in vitro (Kung et al., 2009). The metagenome analysis further predicted the pathways that are used for conversion of benzoate to hydroxypimelyl-CoA and subsequent conversion of 3-hydroxypimelyl-CoA via β-oxidation to acetyl-CoA, which in turn gives rise to acetate through substrate-level phosphorylation (Supplementary Figure 7; Figure 3). The Pelotomaculum bin also contains genes and pathways for the production of butyrate (Supplementary Figure 7). These observations indicate that TA fermentation by Pelotomaculum may lead to the formation of butyrate in addition to acetate. Two genes assigned to Pelotomaculum (tadcc25255 and tadcc12813) belong to the Fe-only hydrogenase protein family and are potentially involved in hydrogen generation.
Methanogens
Three major groups or bins of methanogens belonging to the genera Methanosaeta and Methanolinea were identified and are known to be syntrophic partners of Pelotomaculum (Table 1). Complete pathways for both acetoclastic and hydrogenotrophic methanogenesis were identified (Supplementary Table 1). The first step in acetoclastic methanogenesis is the formation of acetyl-CoA from acetate. It has been proposed (Smith and Ingram-Smith, 2007) that acetoclastic methanogenesis in Methanosaeta proceeds with a modified version of the pathway compared with Methanosarcina, which utilize the acetate kinase/phosphotransacetylase pathway to convert acetate to acetyl-CoA. In contrast, the M. thermophila genome does not include a readily identifiable acetate kinase and it has been proposed that this species utilizes an acetate transporter coupled with acetyl-CoA synthetases to convert acetate to acetyl-CoA (Smith and Ingram-Smith, 2007). Analysis of the TA data set indicates the presence of acetate transporters (tadcc8417) and acetyl-CoA synthetases, EC 6.2.1.1, (tadcc27524, tadcc27520, tadcc27522, tadcc21744, tadcc21743) in the Methanosaeta bin. A complete set of the five acetyl-CoA decarbonylase subunits (EC 1.2.99.2) was identified as well as genes for the remaining steps of methanogenesis (Supplementary Table 1).
Hydrogenotrophic methanogenesis in the TA community is performed by the Methanolinea group. A complete set of Eha hydrogenase enzyme subunits is found in the Methanolinea bin. This set is adjacent to formyl-methanofuran dehydrogenase, which reduces CO2 to formyl-methanofuran (tadcc39592–39604), suggesting that it may be the enzyme reducing the ferredoxin used by the dehydrogenase (Anderson et al., 2008). In addition, complete sets of ech (tadcc3040–tadcc3045) and mbh (tadcc17854–tadcc17865) hydrogenases can be found in the Methanolinea bin. These hydrogenases are proposed to provide H2 for the reduction of heterodisulfide (CoM-S-S-CoB) by the heterodisulfide reductase in the absence of MvhADG hydrogenase (Anderson et al., 2008; Thauer et al., 2008). In this way they link the regeneration of CoM to the reduction of ferredoxin. No homologs to the MvhADG hydrogenase were identified in the Methanolinea bin, suggesting that this organism couples ferredoxin and CoB-S-S-CoM reduction to hydrogen.
OP5
Analysis of the gene content in the OP5 bin (Table 1) revealed the existence of a gene fragment (tadcc9232) related to the Archaeoglobus type III RuBisCO. This fragment contains 181 amino acids and exhibits 60% identity to the N-terminus of the large subunit of the Archaeoglobus ribulose 1,5-bisphosphate carboxylase, EC 4.1.1.39, raising a link between OP5 and autotrophic CO2 fixation through the Calvin–Benson–Bassham cycle. Previous work has established that type III RuBisCOs are functional enzymes in vitro and also complement RuBisCO deletion in photosynthetic organisms indicating their functionality in vivo (Tabita et al., 2007). However, other experiments have shown that type III RuBisCO enzymes are involved in adenosine monophosphate metabolism (Sato et al., 2007). Thus, future experiments are required to validate whether OP5 species can use type III RuBisCO enzymes for autotrophic CO2 fixation. Although organisms that contain type III RuBisCOs usually lack recognizable phosphoribulokinases (as is the case for Archaea), the OP5 bin contains a gene (tadcc30466) that belongs to the phosphoribulokinase protein family (Pfam domain 00485), which provides the second substrate for the RuBisCO reaction, ribulose 1,5-bisphosphate. These are the two unique enzymatic activities required for CO2 assimilation. The OP5 bin also contains genes encoding phosphoglycerate kinase, EC 2.7.2.3, (tadcc33464), glyceraldehyde-3-phosphate dehydrogenase, EC 2.7.1.12, (tadcc33465), and phosphoglycerate mutase, EC 5.4.2.1, (tadcc16672) although no representatives of the remaining Calvin–Benson–Bassham cycle genes are readily recognizable in the OP5 bin.
OP5 also contains phosphate butyryltransferase, EC 2.3.1.19, (tadcc17546) and two copies of butyrate kinase, EC 2.7.2.7, (tadcc17547 and tadcc17544) indicating its ability to produce butyrate and gain energy by substrate-level phosphorylation. No acetate kinases or adenosine diphosphate-forming acyl-CoA synthetases were detected in the OP5 genes binned by PhyloPythia. However, inspecting unassigned contigs with GC content <31% identified an acetate kinase (EC 2.7.2.1) gene (tadcc15543 on contig taComm3_C5047) that may originate from OP5. On the basis of these observations, it is proposed that the OP5 populations within the TA community participate in the syntrophic interactions by removing CO2 and H2 and producing butyrate and potentially acetate. An operon on contig C11376 binned in OP5 was found to contain a system of Ni-hydrogenases (tadcc33916 and tadcc3391) potentially involved in hydrogen utilization.
Syntrophaceae, Thermotogae and WWE1
Syntrophaceae are members of syntrophic communities and are a minor component of the TA-degrading community (Figure 2). To our knowledge, no known Syntrophaceae isolates have been reported to degrade TA and most of the isolates utilize propionate, long-chain fatty acids and benzoate. The Thermotogae and WWE1 groups were estimated to constitute a significant proportion of the community based on the 16S rRNA analysis. Both for WWE1 and Thermotogae, the respective sample populations could not be modeled directly in composition-based binning, because of a lack of sample-specific training data, for WWE1 there was also not sufficient data to directly model the clade (Table 1). A protein-similarity comparison with sequenced members of the phylum Thermotogae (utilizing the distribution of BLAST (Basic Local Alignment Search Tool) matches for protein-coding genes in the data set) resulted in 1066 genes with a BLAST matches >60%, with additional 1646 genes having BLAST matches >30%. Among these, acetate kinase (tadcc6136) (Supplementary Figure 8) and a phosphotransacetylase (tadcc64919) were identified, suggesting that members of the Thermotogae in the TA community may participate in the syntrophic interactions by producing acetate from an intermediate molecule. This intermediate molecule may be the butyrate produced by the OP5 population. A fragment encoding butyryl-CoA dehydrogenase, EC 1.3.99.2, (tadcc28367) further suggests the existence of the butyrate utilizing pathway, and a contig encoding a Fe-only hydrogenase (tadcc1433) suggests the ability of this population to generate H2. On the basis of these observations, we hypothesize that the Thermotogae species may oxidize butyrate to acetate and H2.
The sequence similarity-based phylogenetic profiler tool of IMG identified a set of genes from the TA-degrading community with high similarity to Candidatus Cloacamonas acidaminovorans, which presents the only sequenced bacterial genome of the WWE1 candidate phylum through genome sequence reconstruction and is predicted as a syntrophic bacterium in anaerobic digesters (Pelletier et al., 2008). Comparing the common genes between the TA community data set and the C. acidaminovorans genome identified 1607 and 3228 genes with sequence identity greater than 80% or 60%, respectively. These genes are likely to originate from the WWE1 population in the TA-degrading community. Among them, acetate kinases (tadcc38857) and acetyl phosphotransferases, EC 2.3.1.8, (tadcc25853, tadcc25854) were identified, suggesting an oxidative pathway generating acetate and energy through substrate-level phosphorylation and Fe-only hydrogenases (tadcc1522, tadcc13376 and tadcc38378), which presumably produce hydrogen. The substrate for this oxidative pathway may be butyrate because members of the butyrate-oxidizing pathway can be identified in the data set.
Methanogenic syntrophy
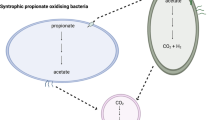
Methanogenic syntrophy has a critical role in the complete degradation of TA to methane (Figure 3). Thermodynamic considerations suggest a low and narrow-range hydrogen concentration as the essential regulator to establish the syntrophic association between the H2-producing bacteria and the H2-consuming methanogens (Schink, 1997; Conrad, 1999). This is because the first reaction from TA to acetate and CO2/H2 (TA + 8H2O → 3 Acetate + 3H+ + 2HCO3− + 3H2, ΔGo′=43.2 kJ mol–1) can occur only at a low pH2 (<5 Pa, 1 a.t.m.=10 1325 Pa) by coupled with methanogenesis (4TA+35 H2O → 17HCO3− + 9H+ + 15CH4, ΔGo′ <−151.9) (Schink, 1997). Also, a minimal threshold pH2 is required for the H2-dependent methanogenesis step to produce the minimum amount of energy required for cell maintenance (Conrad, 1999; Dolfing, 2001). When H2 concentration is higher than this threshold level, H2-dependent methanogenesis ceases. Such a low, narrow-range pH2 is thought to be maintained by ‘interspecies hydrogen transfer’ (Stams, 1994), in which H2-producing syntrophs and H2-consuming methanogens cooperate intimately by arranging themselves in close physical proximity in flocs or in a biofilm with short diffusion distances to facilitate hydrogen transfer. TA community metagenomic data revealed a set of hydrogenases, which generate hydrogen in Pelotomaculum and consume hydrogen in methanogens.
Using fluorescence in situ hybridization analysis (Supplementary Figure 9), we observed not only close physical proximity among methanogens and Pelotomaculum but also the presence of other microbes (that is, OP5, WWE1, Thermotogae and Syntrophus), as illustrated in Figure 3, associating with syntrophs and methanogens. Metagenomic analysis suggests that these populations can actively participate in the syntrophic interactions to tightly regulate pH2. OP5 are likely to consume CO2 and H2 that are produced by Pelotomaculum through the degradation of TA, and produce butyrate. This concept is supported by the ΔGo′ value (−198.05 kJ mol–1) for the conversion of CO2+H2 to butyrate (10H2 + 4CO2 → C4H8O2 + 6H20), which is even more favorable under hyper-mesophilic conditions than mesophilic conditions. OP5 and methanogens can compete for H2 but the competition is likely to be pH2 dependent.
Populations of Syntrophus, Thermotogae and WWE1 may be involved in utilizing and recycling butyrate produced by OP5 probably through the secondary β-oxidation pathway. The presence of hydrogenases indicates that both Thermotogae and WWE1 gain energy through substrate-level phosphorylation. Although there is no clear evidence for the carbon source that these populations utilize, butyrate (produced by OP5) may serve as a key carbon source. This would suggest that, like Syntrophus, some members of Themotogae and WWE1 are possibly syntrophs. However, they are not persistently dominant populations and their abundance varies throughout the reactor operation (Figure 2). TA metagenomics data indicate the presence of butyrate kinases and phosphotransacetylases in the Pelotomaculum bin, suggesting that this population may ferment TA not only to acetate but also butyrate. Our previous study also observed a detectable level of butyrate by using 2-bromoethanesulfonate to inhibit the methanogenesis step in a mesophilic TA-degrading consortium (Wu et al., 2001). It is possible that this type of fermentation results in the production of a second end product (butyrate in addition to acetate) and triggers a secondary syntrophic interaction involving butyrate-oxidizing organisms.
Several studies (Chan, 2000; Qiu et al., 2006; Imachi et al., 2008) have observed the existence of multiple bacterial populations in highly enriched methanogenic cultures degrading carbon substrates like formate, acetate, propionate and phthalate isomers. These observations were shown through a defined mixed culture (Dolfing et al., 2008), suggesting that syntrophic interactions in methanogenic enrichments are more complex than simple pairwise syntroph–methanogen relationships. Rather, they include other members that maintain and regulate the interspecies hydrogen transfer, which is the cornerstone of syntrophy. In conclusion, our overall observations imply that degradation of organic carbon is not simply a syntrophic interaction between H2-producing syntrophs and methanogenic archaea. They further support the hypothesis that additional secondary interactions take place to maintain the stability of the TA-degrading community. (Supplementary Table 2).
References
Anderson I, Rodriguez J, Susanti D, Porat I, Reich C, Ulrich LE et al. (2008). Genome sequence of Thermofilum pendens reveals an exceptional loss of biosynthetic pathways without genome reduction. J Bacteriol 190: 2957–2965.
Angelidaki I, Petersen SP, Ahring BK . (1990). Effects of lipids on thermophilic anaerobic-digestion and reduction of lipid inhibition upon addition of bentonite. Appl Microbiol Biotechnol 33: 469–472.
Boll M, Fuchs G . (1995). Benzoyl-coenzyme A reductase (dearomatizing), a key enzyme of anaerobic aromatic metabolism—ATP dependence of the reaction, purification and some properties of the enzyme from Thauera aromatica strain K172. Eur J Biochem 234: 921–933.
Butler JE, He Q, Nevin KP, He ZL, Zhou JZ, Lovley DR . (2007). Genomic and microarray analysis of aromatics degradation in Geobacter metallireducens and comparison to a Geobacter isolate from a contaminated field site. BMC Genomics 8: 180.
Chan OC . (2000). Characterization of microbial consortia in anaerobic granular sludge—a ribosomal RNA-based molecular approach. In: Civil and Environmental Engineering. University of Hong Kong: Hong Kong, p. 221.
Chen CL, Macarie H, Ramirez I, Olmos A, Ong SL, Monroy O et al. (2004). Microbial community structure in a thermophilic anaerobic hybrid reactor degrading terephthalate. Microbiology 150: 3429–3440.
Conrad R . (1999). Contribution of hydrogen to methane production and control of hydrogen concentrations in methanogenic soils and sediments. FEMS Microbiol Ecol 28: 193–202.
Dolfing J . (2001). The microbial logic behind the prevalence of incomplete oxidation of organic compounds by acetogenic bacteria in methanogenic environments. Microbial Ecol 41: 83–89.
Dolfing J, Jiang B, Henstra AM, Stams AJM, Plugge CM . (2008). Syntrophic growth on formate: a new microbial niche in anoxic environments. Appl Environ Microbiol 74: 6126–6131.
Imachi H, Sakai S, Sekiguchi Y, Hanada S, Kamagata Y, Ohashi A et al. (2008). Methanolinea tarda gen. nov., sp nov., a methane-producing archaeon isolated from a methanogenic digester sludge. Int J Syst Evol Microbiol 58: 294–301.
Kleerebezem R, Pol LWH, Lettinga G . (1999). Anaerobic degradation of phthalate isomers by methanogenic consortia. Appl Environ Microbiol 65: 1152–1160.
Kung JW, Loffler C, Dorner K, Heintz D, Gallien S, Van Dorsselaer A et al. (2009). Identification and characterization of the tungsten-containing class of benzoyl-coenzyme A reductases. Proc Natl Acad Sci USA 106: 17687–17692.
Lupa B, Lyon D, Gibbs MD, Reeves RA, Wiegel J . (2005). Distribution of genes encoding the microbial non-oxidative reversible hydroxyarylic acid decarboxylases/phenol carboxylases. Genomics 86: 342–351.
Martin HG, Ivanova N, Kunin V, Warnecke F, Barry KW, McHardy AC et al. (2006). Metagenomic analysis of two enhanced biological phosphorus removal (EBPR) sludge communities. Nat Biotechnol 24: 1263–1269.
McHardy AC, Martin HG, Tsirigos A, Hugenholtz P, Rigoutsos I . (2007). Accurate phylogenetic classification of variable-length DNA fragments. Nat Methods 4: 63–72.
McInerney MJ, Rohlin L, Mouttaki H, Kim U, Krupp RS, Rios-Hernandez L et al. (2007). The genome of Syntrophus aciditrophicus: life at the thermodynamic limit of microbial growth. Proc Natl Acad Sci USA 104: 7600–7605.
Pelletier E, Kreimeyer A, Bocs S, Rouy Z, Gyapay G, Chouari R et al. (2008). ‘Candidatus Cloacamonas acidaminovorans’: genome sequence reconstruction provides a first glimpse of a new bacterial division. J Bacteriol 190: 2572–2579.
Peters F, Shinoda Y, McInerney MJ, Boll M . (2007). Cyclohexa-1,5-diene-1-carbonyl-coenzyme A (CoA) hydratases of Geobacter metallireducens and Syntrophys aciditrophicus: evidence for a common benzoyl-CoA degradation pathway in facultative and strict anaerobes. J Bacteriol 189: 1055–1060.
Qiu YL, Hanada S, Ohashi A, Harada H, Kamagata Y, Sekiguchi Y . (2008). Syntrophorhabdus aromaticivorans gen. nov., sp. nov., the first cultured anaerobe capable of degrading phenol to acetate in obligate syntrophic associations with a hydrogenotrophic methanogen. Appl Environ Microbiol 74: 2051–2058.
Qiu YL, Sekiguchi Y, Hanada S, Imachi H, Tseng IC, Cheng SS et al. (2006). Pelotomaculum terephthalicum sp. nov. and Pelotomaculum isophthalicum sp. nov.: two anaerobic bacteria that degrade phthalate isomers in syntrophic association with hydrogenotrophic methanogens. Arch Microbiol 185: 172–182.
Razo-Flores E, Macarie H, Morier F . (2006). Application of biological treatment systems for chemical and petrochemical wastewaters. In: Cervantes FJ, Pavlostathis SG, van Haandel AC (eds). Advanced Biological Treatment Processes for Industrial Wastewaters. IWA Publishing: London, UK pp. 267–297.
Sato T, Atomi H, Imanaka T . (2007). Archaeal type IIIRuBisCOs function in a pathway for AMP metabolism. Science 315: 1003–1006.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Smith KS, Ingram-Smith C . (2007). Methanosaeta, the forgotten methanogen? Trends Microbiol 15: 150–155.
Stams AJM . (1994). Metabolic interactions between anaerobic bacteria in methanogenic environments. Antonie Van Leeuwenhoek Int J Gen Mol Microbiol 66: 271–294.
Tabita FR, Hanson TE, Li HY, Satagopan S, Singh J, Chan S . (2007). Function, structure, and evolution of the RuBisCO-like proteins and their RubisCO homologs. Microbiol Mol Biol Rev 71: 576–599.
Thauer RK, Kaster AK, Seedorf H, Buckel W, Hedderich R . (2008). Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol 6: 579–591.
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM et al. (2004). Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428: 37–43.
van Lier JB, Rebac S, Lettinga G . (1997). High-rate anaerobic wastewater treatment under psychrophilic and thermophilic conditions. Water Sci Technol 35: 199–206.
Wischgoll S, Heintz D, Peters F, Erxleben A, Sarnighausen E, Reski R et al. (2005). Gene clusters involved in anaerobic benzoate degradation of Geobacter metallireducens. Mol Microbiol 58: 1238–1252.
Wu JH, Liu WT, Tseng IC, Cheng SS . (2001). Characterization of microbial consortia in a terephthalate-degrading anaerobic granular sludge system. Microbiology 147: 373–382.
Acknowledgements
This work was performed under the auspices of the US Department of Energy's Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344 and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396.
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper was corrected on 21st December 2010 to include contributing authors inadvertently ommitted from the original version of the paper
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Lykidis, A., Chen, CL., Tringe, S. et al. Multiple syntrophic interactions in a terephthalate-degrading methanogenic consortium. ISME J 5, 122–130 (2011). https://doi.org/10.1038/ismej.2010.125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.125
Keywords
This article is cited by
-
Isophthalate:coenzyme A ligase initiates anaerobic degradation of xenobiotic isophthalate
BMC Microbiology (2022)
-
Microbial degradation and valorization of poly(ethylene terephthalate) (PET) monomers
World Journal of Microbiology and Biotechnology (2022)
-
Emergence of a Synergistic Diversity as a Response to Competition in Pseudomonas putida Biofilms
Microbial Ecology (2020)
-
Anaerobic degradation of xenobiotic isophthalate by the fermenting bacterium Syntrophorhabdus aromaticivorans
The ISME Journal (2019)
-
Identification of naphthalene carboxylase subunits of the sulfate-reducing culture N47
Biodegradation (2019)