Abstract
Members of the Burkholderia cepacia complex (Bcc), such as B. ambifaria, are effective biocontrol strains, for instance, as plant growth-promoting bacteria; however, Bcc isolates can also cause severe respiratory infections in people suffering from cystic fibrosis (CF). No distinction is known between isolates from environmental and human origins, suggesting that the natural environment is a potential source of infectious Bcc species. While investigating the presence and role of phase variation in B. ambifaria HSJ1, an isolate recovered from a CF patient, we identified stable variants that arose spontaneously irrespective of the culture conditions. Phenotypic and proteomic approaches revealed that the transition from wild-type to variant types affects the expression of several putative virulence factors. By using four different infection models (Drosophila melanogaster, Galleria mellonella, macrophages and Dictyostelium discoideum), we showed that the wild-type was more virulent than the variant. It may be noted that the variant showed reduced replication in a human monocyte cell line when compared with the wild-type. On the other hand, the variant of isolate HSJ1 was more competitive in colonizing plant roots than the wild-type. Furthermore, we observed that only clinical B. ambifaria isolates generated phase variants, and that these variants showed the same phenotypes as observed with the HSJ1 variant. Finally, we determined that environmental B. ambifaria isolates showed traits that were characteristic of variants derived from clinical isolates. Our study therefore suggest that B. ambifaria uses phase variation to adapt to drastically different environments: the lung of patients with CF or the rhizosphere.
Similar content being viewed by others
Introduction
The Burkholderia genus comprises more than 40 species that inhabit remarkably diverse ecological niches, as they have been isolated from soil, rhizosphere, insects, fungi and infected humans. Many Burkholderia have developed beneficial interactions with their plant hosts and have considerable ecological significance: several species of Burkholderia have proven to be very efficient biocontrol and bioremediation agents (Chiarini et al., 2006). This is especially true for members of the Burkholderia cepacia complex (Bcc), which currently comprises 17 closely related genomovars (Vanlaere et al., 2009). For instance, Burkholderia ambifaria J82 was registered by the US Environmental Protection Agency for use as a biospecticide and was commercially used for a few years (Parke and Gurian-Sherman, 2001). However, Bcc isolates are now increasingly identified as opportunistic pathogens, notably infecting people suffering from cystic fibrosis (CF) or chronic granulomatous disease (Govan et al., 2007). For this reason, their use as biological control agents is not allowed anymore (Chiarini et al., 2006).
Within the Bcc, B. ambifaria (genomovar VII) typically illustrates the duality between isolates with biological control purposes and those that cause infections in humans (Coenye et al., 2001). This dichotomy is indeed underlined by the choice of the name ambifaria from the latin name ambifarius (having two sides; ambiguous) (Coenye et al., 2001). One of the most investigated biocontrol agents, the sequenced strain B. ambifaria AMMD (LMG19182), isolated from the rhizosphere of the pea, shows well-demonstrated antagonistic activity against a number of plant pathogenic fungi (Parke et al., 1991). On the other hand, several B. ambifaria isolates have been isolated from CF patients (Coenye et al., 2001). B. ambifaria is among the most common Bcc species in the plant rhizosphere; however, a minority of Bcc-infected CF patients harbor this genomovar (Ramette et al., 2005; Reik et al., 2005).
No clear distinction is known between Bcc isolates from environmental and human origins, and they seem to be clonally related. For instance, the biocontrol strain B. ambifaria AMMD and clinical isolate B. ambifaria AU0212 are clonal (Payne et al., 2005). A B. cepacia genomovar III isolate recovered from agricultural soil in the mid-Atlantic region of the United States is indistinguishable from an isolate frequently recovered from CF patients of the same region using the fingerprinting method (LiPuma et al., 2002).
Phase variation, or phenotypic switching, is an adaptive process conferring to bacteria the capacity to adapt to fluctuating environmental conditions. Phase variation implies phenotypic changes resulting from genetic or epigenetic alterations at specific genetic loci (Villemur and Déziel, 2005; Wisniewski-Dyé and Vial, 2008). In contrast to spontaneous mutations that occur at a frequency of approximately 10−8 mutations per growing cell per generation, phase variation occurs at frequencies higher than 10−5 switches per cell per generation and always affects the same phenotype(s) (van der Woude and Baumler, 2004). This phenomenon results in an increased diversity within a bacterial population and favors bacterial fitness. Phase variation has an important adaptive role in the capacity of bacteria to colonize different host anatomical niches and escape host defense mechanisms.
While investigating the presence and role of phase variation in B. ambifaria isolates recovered from CF patients, we identified stable, smooth-colony variants showing numerous phenotypic differences when compared with their rough-colony parent. Strikingly, we found that clinically rough isolates are more virulent than their variants, whereas these phase variants are more competitive in rhizosphere colonization. Our data suggest that, by phase variation, B. ambifaria adopts two different phenotypic states: the first one is better adapted to the lungs of CF patients and the second is more adapted to plant roots environment.
Materials and methods
Bacterial strains and growth conditions
The B. ambifaria strains used in this study are listed in Supplementary Table S1. Unless otherwise stated, strains were grown in tryptic soy broth (TSB, Difco, Detroit, MI, USA) at 37 °C with shaking (240 r.p.m.), or on TSB agar plates containing 0.1% (w/v) Congo red (CRTSB). Specific culture conditions can be found in the Supplementary data.
Phenotypic characterization
Several enzymatic activities were evaluated using API-ZYM strips (bioMerieux, Lyon, France). Siderophore, proteolytic and antifungal activities were measured as described earlier (Vial et al., 2008). Hemolytic activity was assessed on TSB agar supplemented with 5% (v/v) sheep blood. TSB agar containing tributyrin 1% v/v was used to detect lipolytic activity. Cholesterol oxidase activity was assessed using TSB agar plates containing 0.34% Triton X-100 and 0.9 mM cholesterol (Sigma-Aldrich, Oakville, Ontario, Canada) (Doukyu and Aono, 2001). Specific details can be found in the Supplementary data.
The presence of elastase was detected using elastin-Congo red (Sigma) as substrate (Pearson et al., 1997) and phospholipase C activity was detected as described earlier (Esselmann and Liu, 1961). The capacity of B. ambifaria to produce exopolysaccharides (EPSs) was determined as described by Zlosnik et al. (2008).
Disc diffusion assay was used to test the susceptibility of B. ambifaria to H2O2. In brief, logarithmic-phase cells (OD600 of 3.0) or stationary-phase cells (OD600 of 6.0) were spread as a lawn on TSB agar plates and disks with 10 μl of 30% H2O2 were applied to the surface. The plates were incubated for 16 h at 37 °C, and then the diameter of the zone of growth inhibition around each disk was measured.
Biofilm formation and abiotic adherence assay
The biofilm formation protocol was performed as before (Déziel et al., 2001), with the following modification: 12 × 75-mm2 polystyrene tubes containing TSB were inoculated from overnight TSB cultures and incubated at 37 °C without agitation for 24 h.
The abiotic adherence assay was performed as described by Aubert et al. (2008), with the following modifications: approximately 105 colony-forming units (CFUs) were added to polystyrene tubes containing 1 ml of NaCl 0.8% and then incubated at 37 °C for 20 min. After four washes with NaCl 0.8%, the bound bacteria were detached from the surface by adding 500 μl of 10 mM EDTA and 1.5 ml of 1% Triton X-100. The adherence of the wild-type and variant was calculated by dividing the output number by the input number. Experiments were performed in triplicate and independently repeated three times.
Sensitivity to antimicrobial peptides
For sensitivity to antimicrobial peptides, bacteria were grown on TSB agar plate for 24 h at 37 °C, and then for 24 h at room temperature. Cells (4 × 105 CFU) were resuspended in 200 μl of 0.1 × phosphate buffered saline containing various concentrations of antimicrobial peptides: 50 or 100 μg ml−1 melittin (Sigma), 200 μg ml−1 LL-37 (AnaSpec, San Jose, CA, USA) or 1000 μg ml−1 polymyxin B (Sigma). Bacteria were then incubated without agitation at 37 °C for 8 h. After treatment, viable cells were enumerated on TSB agar plates. All assays were conducted in triplicate and repeated independently at least twice. The survival (%) was defined as follows: (CFU of peptide-exposed culture/CFU of nonexposed culture) × 100.
Lipopolysaccharide (LPS) analysis
LPSs were prepared for gel analysis as described earlier (Apicella et al., 1994), with slight modifications as outlined in the Supplementary data.
Quantifications of 4-hydroxy-2-alkylquinolines (HAQs) and N-acyl-L-homoserine lactones (AHLs)
Quantifications of HAQs and AHLs were performed as described earlier (Lépine et al., 2003; Vial et al., 2008).
Bacterial survival/proliferation and competitive index in human macrophages
Macrophage infection assays were performed using the human acute monocytic leukemia cell line THP-1 as described in Forest et al. (2007), with the following modifications: bacteria grown overnight at 37 °C in static conditions in TSB were added to a cell monolayer at a multiplicity of infection of 10. After incubation for 20 min at 37 °C (time 0), the infected cells were washed three times with prewarmed phosphate buffered saline. In order to kill extracellular bacteria, fresh medium containing 125 μg ml−1 tetracycline and 1 mg ml−1 ceftazidime was added, and cultures were incubated for another 2 h at 37 °C. The number of surviving bacteria (intracellular bacteria) was determined by bacterial plate counting (CFU) on CRTSB at time 0 and at 2, 4, 6, and 8 h after initial inoculation. At 8 h after initial inoculation, the final wash was plated for determining the number of viable extracellular bacteria. The level of phagocytosis was expressed as a percentage of the initial inoculum. All assays were conducted in duplicate and repeated independently at least three times.
For competitive index (CI) in macrophages, a 1:1 mixture of two bacterial strains was added to THP-1-cultured cells at a multiplicity of infection of 10 as described above. The numbers of viable intracellular bacteria were determined 2 and 8 h after infection. Infections were performed in triplicate. The CI in proliferation is defined as the ratio between the two strains in the output (intracellular bacteria recovered 8 h after infection) divided by their ratio in the input (intracellular bacteria after the 2-h treatment with antibiotics) (Segura et al., 2004).
Dictyostelium discoideum plaque assay
D. discoideum strain DH1-10 was grown in a Petri dish at 21 °C in HL-5 liquid medium containing 15 μg ml−1 tetracycline (Cornillon et al., 2000; Mercanti et al., 2006). Amoebae were collected by centrifugation, washed and resuspended in HL-5 at the concentration of 2 × 105 per 5 μl. Subsequent serial dilutions were carried out in HL-5 to obtain the following concentrations (per 5 μl): 1 × 105, 5 × 104, 2.5 × 104, 1.25 × 104, 6.25 × 103, 3.13 × 103, 1.56 × 103 and 7.8 × 102. B. ambifaria strains were grown on CRTSB for 24 h at 37 °C and then for 24 h at 25 °C. Suspensions of bacteria for each strain were plated on HL-5 agar and the bacterial lawns were spotted with 5-μl droplets of the D. discoideum serial dilutions. Plates were incubated at 23 °C for 24 h and examined for plaque formation. This experiment was performed four times with HSJ1 and its variant (HSJ1v), and at least two times with all other strains. Similar results were obtained each time.
Infection of Galleria mellonella larvae
G. mellonella killing assays were performed as described previously (Seed and Dennis, 2008). In competition experiments, larvae were injected with wild-type and variant in a 1:1 ratio (1 × 104 CFU of wild-type and variant), and the CI was evaluated 48 h after co-inoculation, just before the death of the larvae. To estimate B. ambifaria concentration inside the larvae, each of them was surface sterilized (by dipping in ethanol and passing the larvae through a Bunsen flame), crushed and homogenized in 1 ml 0.8% NaCl. The CI is defined as the CFU output (48 h after infection) ratio of HSJ1v in comparison with HSJ1, divided by their ratio in the input (inoculum). This experiment was repeated twice.
Rhizosphere colonization
Pisum sativum seeds were sterilized as described earlier (O'Sullivan et al., 2007). After 3 days, germinated seeds were sown in tubes containing Perlite as the solid substrate and 8 mM KNO3−-supplemented Fahraeus liquid medium (Fahraeus, 1957). Bacterial isolates were inoculated in a 1:1 ratio (105 CFU of wild-type and variant). Plants were maintained in a controlled-condition incubator: 12 h of light at 25 °C with 65% relative humidity. Bacteria were recovered from the rhizosphere by vortexing the roots for 2 min in a tube containing 10 ml of 0.8% NaCl with glass beads and by plating on CRTSB with antibiotics (25 μg ml−1 gentamycin, 15 μg ml−1 polymyxin B, 20 μg ml−1 cycloheximide). Roots were sampled 2, 5, 10 and 15 days after inoculation. The CI for colonization was defined as the CFU output (day 15 after inoculation) ratio of HSJ1v in comparison with HSJ1, divided by the CFU input (inoculum) ratio of variant to wild-type.
Results
Phase variation in B. ambifaria HSJ1
Phase variation was observed in the B. ambifaria strain HSJ1 isolated at the Sainte-Justine Hospital (Montréal, Canada) from a patient suffering from CF. When a single colony of strain HSJ1 was plated on CRTSB agar, two colonial forms developed (Figure 1). Most colonies (wild-type phenotype) were rough, often wrinkled and bound Congo red, giving them a red appearance. The other form of colony (variant phenotype) was translucent, smoother and unable to bind Congo red. To verify the stability of the two phenotypes, single colonies were individually grown overnight in TSB and plated on CRTSB. The wild-type (HSJ1) continued to produce variant colonies, whereas the variant (HSJ1v) remained stable: no reversion to the wild-type or appearance of another phenotype was detected. Variant colonies arose spontaneously at low frequencies, irrespective of the culture conditions used (that is, growth medium and carbon source). The frequency of occurrence of variant colonies was 1.37 × 10−4 (±0.85 × 10−4) per cell per generation in the conditions tested. Identical profiles were obtained using polymerase chain reaction/restriction-fragment length polymorphism of the 16S ribosomal DNA region. This result confirms that HSJ1v is not a co-cultivated bacterium, but is instead derived from the wild-type strain HSJ1.
Phenotypic comparison between B. ambifaria wild-type (W) and variant (V). (a) Colony distinction between B. ambifaria HSJ1 wild-type and variant on Congo red tryptic soy broth (CRTSB) medium. Plates were incubated for 24 h at 37 °C and then kept for 72 h at room temperature. (b) Siderophore production on chrome azurol S (CAS) agar. The orange halo surrounding the colony indicates the release of siderophores. (c) Proteolytic activity on agar plate containing 1% skim milk powder. (d) Cholesterol oxidase activity on TSB agar plate containing 0.9 mM cholesterol. The presence of a turbid halo around the colony indicates cholesterol oxidase activity. (e) Biofilm assay. Biofilms developed in polystyrene tubes were stained with crystal violet and quantified by spectrophotometry (A600) after solubilization of the dye in ethanol. Error bars represent the standard deviations. Abbreviations: WT, B. ambifaria HSJ1 wild-type; V, B. ambifaria HSJ1 variant. A full colour version of this figure is available at The ISME Journal online.
Pleiotropic differences of the B. ambifaria HSJ1 variant
The growth rates of the wild-type and of the variant were determined to be the same in TSB and in M63, with the following carbon sources: dextrose, mannitol, fructose and galactose (Supplementary Figure S1a and b and data not shown). Both types were unable to use lactose or maltose. Interestingly, inferior growth of HSJ1 was observed on saccharose, xylose, sorbitol and glycerol when compared with HSJ1v (Supplementary Figure S1).
Many physiological and biochemical characteristics of the variant HSJ1v were different from those of the wild-type HSJ1 (Table 1 and Figure 1). The variant type showed pleiotropic phenotypic losses, such as the absence of hemolytic and antifungal activities. On the other hand, HSJ1v type was frankly mucoid (+++) on yeast extract-mannitol agar, whereas HSJ1 was only partially mucoid (+) (Table 1). Furthermore, the variant produced more siderophores than the wild-type, suggesting that HSJ1v possesses a greater ability for iron scavenging than HSJ1. Indeed, superior growth of the variant was observed in a minimal medium supplemented with an iron-chelating agent (Supplementary Figure S1). Moreover, an increase in the proportion of variant was observed when iron was limiting (Supplementary Figure S2).
Secretomic studies on the two phenotypic types revealed three major secreted proteins that were absent from HSJ1v (Supplementary Figure S3). Two spots (WT1 and WT2) were identified as zinc metallopeptidases ZmpB and ZmpA (Bamb_4475 and Bamb_3836, according to the annotation of sequenced B. ambifaria strain AMMD) associated with the protease activity of B. cenocepacia (Kooi et al., 2006). Accordingly, HSJ1v showed significantly less proteolytic activity on skim milk than the wild-type (Table 1 and Figure 1c). Spot WT3 was identified as an FAD-dependent cholesterol oxidase (Bamb_6465) (Doukyu and Aono, 2001). Absence of the corresponding activity in HSJ1v was confirmed on cholesterol-containing agar plates (Figure 1d).
When inoculated into TSB and incubated at 37 °C without shaking, HSJ1v grew suspended in the broth, whereas the wild-type formed aggregates, mostly at the air–liquid interface. Accordingly, the wild-type formed more biofilm than the variant (Figure 1e). Moreover, HSJ1 was 1.6-fold more adherent to abiotic surfaces than HSJ1v. Bacterial motility mechanisms are known to have an important role in biofilm formation. However, swimming activity was similar for HSJ1 and HSJ1v on tryptone swim plate and we were unable to detect any swarming motility for both types (data not shown).
Eight wild-type colonies and eight variant colonies were randomly chosen from several CRTSB plates and screened for antifungal, proteolytic and hemolytic activities. No variability between the different wild-type colonies was observed and each variant colony showed the same phenotypes, suggesting that all the wild-type and variant colonies show a consistent phenotype pattern. Moreover, in a wide range of conditions tested (temperature, carbon source and oxidative stress), no reversion from variant to wild-type was observed.
The variant type is unable to produce 4-hydroxy-3-methyl-2-alkylquinolines (HMAQs)
The production of extracellular proteases, siderophores and of an unknown antifungal compound is under quorum-sensing control in B. ambifaria (Zhou et al., 2003; Wopperer et al., 2006). As these quorum-sensing-regulated phenotypes are altered in the variant, we hypothesized that quorum-sensing might be affected. We thus quantified the production of octanoyl-homoserine lactone, C8-HSL, the predominant quorum-sensing signal produced by B. ambifaria (Zhou et al., 2003; Vial et al., 2008). However, no significant difference was observed between the wild-type and the variant: for instance, during exponential phase (OD600=4.00), concentrations were 1.43 mg ml−1 for HSJ1 versus 1.29 mg ml−1 for HSJ1v.
Recently, we have reported that the wild-type HSJ1 strain produces HAQ analogues bearing a methyl group at the third position: HMAQs (Vial et al., 2008). HAQs, such as the Pseudomonas quinolone signal, are implicated in the regulation of virulence genes in P. aeruginosa (Dubern and Diggle, 2008). We thus analyzed several randomly chosen variant colonies and found that none were producing HMAQs. We therefore compared the phenotypes of an HMAQ-negative hmqA− mutant with those of HSJ1 and HSJ1v. However, despite the absence of HMAQ production in HSJ1v, an hmqA− mutant showed mostly wild-type features (data not shown). Furthermore, we have recently described that the hmqA− mutant shows greater antifungal and proteolytic activity than the wild-type (and consequently more than the variant) (Vial et al., 2008). Taken together, these results indicate an absence of correlation between loss of HMAQ production and the variant phenotype.
Sensitivity to antimicrobial peptides and modification of the LPS profile
By disk diffusion assay, we observed that the wild-type and its variant present essentially the same antibiogram (Supplementary Table S2). However, as Bcc isolates are highly resistant to antimicrobial peptides, we challenged the HSJ1 wild-type and variant with antimicrobial peptide in suspension, followed by enumeration of viable cells. Using this precise method, we analyzed further the response to three antimicrobial peptides. Figure 2a shows that the survival of the wild-type is not affected by the presence of 1000 μg ml−1 polymyxin B or 200 μg ml−1 LL-37, a human cathelicidin. In contrast, HSJ1v type shows only 60% of survival in the presence of these compounds, and this increased sensitivity of the variant is even more pronounced with the honeybee polypeptide mellitin (Figure 2b).
Antimicrobial peptides assay and analysis of lipopolysaccharides (LPSs). (a) The graph shows antimicrobial peptide survival of B. ambifaria wild-type (hatched bars) and variant (white bars). Antimicrobial peptide survival was calculated from the ratio of exposed cells to nonexposed cells. (b) Analysis of LPSs by tricine SDS-polyacrylamide gel electrophoresis (SDS-PAGE). Abbreviations: WT, B. ambifaria wild-type; V, B. ambifaria variant.
The altered appearance of variant colonies suggests changes in the cell wall, possibly in LPS structure. Such differences are known to affect the response to antimicrobial peptides (Rosenfeld and Shai, 2006). As shown in Figure 2c, tricine SDS-polyacrylamide gel electrophoresis analysis revealed modifications in the O-antigen and lipid A-core regions.
The wild-type is more virulent in several infection models
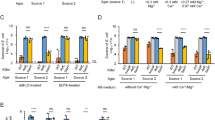
Certain Bcc strains can invade and survive within respiratory epithelial cells and macrophages (Valvano et al., 2005). The levels of phagocytosis and intracellular replication after infection with the wild-type and the variant were thus evaluated using THP-1 cells. Although HSJ1v is more amenable to phagocytosis, with a twofold higher level (30.7±4.1%) compared with HSJ1 (15.9±2.8%), it shows a much reduced ability to replicate in macrophages (Figure 3a). Importantly, during these experiments, no difference in the frequency of phase variation between the wild-type inoculum and the population recovered from the macrophages 8 h after infection was observed.
Comparison of the virulence of B. ambifaria wild-type HSJ1 and variant. (a) Quantification of B. ambifaria HSJ1 wild-type (▪) and variant (○) within human acute monocytic leukemia cell line (THP-1) macrophages. The results are expressed as the means±s.d. of the replicate experiments. (b) G. mellonella larvae survival over time infected with B. ambifaria HSJ1 wild-type and variant. The larvae were injected with bacteria (1 × 106 or 1 × 105) and monitored for their survival. Each data set is representative of a single trial with the specified strain (N=10). (c) Competitive index (CI) analysis of B. ambifaria wild type and variant. CI in proliferation in THP-1 macrophages is defined as the ratio between the two strains in the output (intracellular bacteria recovered 8 h after infection) divided by their ratio in the input (intracellular bacteria after the 2-h treatment with antibiotics). CI in G. mellonella is defined as the colony-forming unit (CFU) output (48 h after infection) ratio of the variant in comparison with the wild-type strain, divided by the CFU input ratio (inoculum) of mutant to wild-type; CI for colonization of the rhizosphere of P. sativum is defined as the CFU output (day 15 after inoculation) ratio of the variant in comparison with the wild-type strain, divided by the CFU input (inoculum) ratio of variant to wild-type. The wild-type is more competitive than the variant when the CI is greater than 1.
The wax moth, G. mellonella, was recently reported as a useful surrogate host for determining the virulence of the Bcc, notably B. ambifaria (Seed and Dennis, 2008). When compared with wild-type, HSJ1v shows reproductively less virulence towards G. mellonella larvae (Figure 3b). This result was also confirmed using the Drosophila pricking model (Supplementary Figure S4).
Finally, the virulence of B. ambifaria wild type and variant was quantitatively characterized by spotting serial dilutions of D. discoideum on bacterial lawns (Figure 4). The social amoeba D. discoideum is a suitable host for studying the virulence of bacterial pathogens, including the Bcc (Aubert et al., 2008). D. discoideum forms clear plaques on HSJ1v variant lawns at dilutions containing at least 25 000 amoebae. However, on wild-type lawns, D. dictyostelium is unable to produce clear plaques, even at the highest concentration of amoebae. This result suggests that the wild-type, in contrast with the variant, is able to resist D. discoideum predation.
Plaque assay with D. discoideum. The bacterial lawn was spotted with 5-μl droplets of the serial dilutions of D. discoideum, and the ability of D. discoideum to produce clearing plaques in the bacterial lawn was recorded after 24 h at 23 °C. Small holes, such as the one pointed by the arrow, are not clearing plaques but only scratches made when amoebae were deposited on the bacterial lawn.
To better define the difference of the wild-type and the variant in intracellular replication abilities within THP-1 cell lines, we produced a co-infection to generate a CI in proliferation (Segura et al., 2004). A mean value of 2.2 for the CI confirmed the better survival and replication of the wild-type within THP-1 cells (Figure 3d). A CI performed during G. mellonella larvae infection further established that the wild-type is also better adapted to survive and replicate in this host (Figure 3d).
Several G. mellonella larvae and Drosophila melanogaster flies were infected with B. ambifaria variant, and at different times after the inoculation, we evaluated the presence of revertant or occurrence of another phenotype on CRTSB. In these conditions, neither reversion to wild-type nor appearance of new phenotypes was detected. These results showed that the variant type was stable in all conditions tested.
The variant of B. ambifaria HSJ1 is more competitive in rhizosphere colonization
As several B. ambifaria environmental strains have been isolated from the rhizosphere of the pea (Pisum sativum) (Coenye et al., 2001), this plant was used as our model to investigate the rhizosphere-colonizing competence of HSJ1 wild-type and variant. Our results showed that HSJ1v was more competitive in colonizing the rhizosphere (Figures 3c and 5). At 15 days after co-inoculation, the proportion of variants in the rhizospheric population nearly reached 80%. In the control medium, no significant difference in the proportion of the two populations was observed. These results highlight a central difference in competitiveness between the wild-type and the variant in colonizing the rhizosphere because when the wild-type was inoculated alone no increase in the variant proportion was observed (data not shown). The difference observed in competitiveness during colonization did not seem to be the consequence of a better root attachment of the variant (Supplementary Figure S5).
Competitive root colonization by B. ambifaria wild type and variant. Plants were inoculated 1:1 with the wild-type and the mutant. Results are shown as means±s.d. Hatched bars and white bars represent the percentage of colonies recovered from the wild-type strain and the variant strain, respectively. Medium represents the percentage of colonies recovered from Fahraeus (FP) mineral medium inoculated and stored 15 days in the same condition as the roots.
Occurrence of phase variation in the B. ambifaria genomovar
To ascertain that B. ambifaria phase variation is not solely restricted to isolate HSJ1, B. ambifaria isolates recovered from CF patients (n=7) and from environmental sources (n=6) were grown overnight in TSB medium and plated on CRTSB (Supplementary Table S1). We found that clinical isolates formed rough, wrinkled colonies binding Congo red, the phenotype typical of HSJ1 wild-type. Moreover, all clinical isolates generated variants at a frequency of 10−4 variants per cell per generation (Supplementary Table S3). These variants were translucent and unable to bind Congo red, similar to HSJ1v. Moreover, all the isolates recovered from CF patients produced variant colonies with the same phenotypes observed with HSJ1v: absence of hemolytic, antifungal or cholesterol oxidase activities (Table 2). Only clinical isolates were able to produce HMAQs, with the exception of CEP0516. Virulence of the clinical isolates and their variants was investigated using the D. discoideum plaque assay. We observed, such as with HSJ1, that the amoebae were unable to form clear plaques on the wild-type clinical isolates lawns, in contrast with their variants (Table 2).
On the other hand, all isolates recovered from environmental sources failed to bind Congo red, showed only one type of colony on CRTSB, even after several subcultures, and showed common features with the variants derived from the clinical isolates (Table 2). Notably, environmental isolates were frankly mucoid on yeast extract-mannitol agar and did not produce HMAQs. With one exception, they did not show antifungal and cholesterol oxidase activities, similar to the variants derived from clinical isolates. Finally, most environmental isolates were susceptible to predation by the amoebae, similar to the variants (Table 2).
Discussion
B. ambifaria uses phase variation for niche adaptation
Bcc isolates that occur in the environment are also capable of infecting humans and it is widely accepted that differentiation between the environmental and clinical isolates of Bcc is unrealistic (Govan et al., 2000). To the best of our knowledge, this is the first report that distinguish between two distinct phenotypic states inside a genomovar of the B. cepacia complex. The first phenotypic type of B. ambifaria is composed of isolates adapted to clinical infections or interactions with eukaryotic hosts and are recovered from patients suffering from CF. The second type is associated with environmental isolates and is better adapted to the rhizosphere. We suggest that this distinction mediated by phase variation allows the adaptation of a B. ambifaria strain to drastically different environments: the lungs of patients with CF or the plant roots environment.
A striking phenomenon observed with B. ambifaria is that only clinical isolates generate phase variants, under laboratory conditions. Most importantly, these variants show features typical of isolates recovered from the environment, suggesting that phase variants from clinical isolates are in fact rhizosphere colonizers. An extensive number of B. ambifaria isolates would be required to confirm this hypothesis.
Features of B. ambifaria clinical isolates
As the virulence of Bcc strains can vary depending on the model host used, it was appropriate to test the virulence of HSJ1 and HSJ1v using several models. A number of infection models have earlier been tested with Bcc members, including alfalfa seedlings, G. mellonella and Caenorhabditis elegans (Bernier et al., 2003; Cardona et al., 2005; Seed and Dennis, 2008). By using four infection models (D. melanogaster, G. mellonella, macrophages and D. discoideum), we clearly showed that the wild-type is more virulent than the variant. Interestingly, our study using D. discoideum uncovers an intriguing distinction between the two phenotypic states in B. ambifaria: amoebae are unable to form clear plaques in lawns of clinical isolates, whereas most environmental isolates and all the variants are permissive to D. discoideum. Further investigations are required for a better understanding of clinical isolate resistance to amoebae.
Direct evidence of intracellular survival of the Bcc species has been shown in airway epithelial cells, in murine macrophages, in human monocytic cells and in amoebae (Marolda et al., 1999; Saini et al., 1999; Savoia and Zucca, 2007). The capacity of Bcc strains to survive and replicate within eukaryotic cells may explain their persistence in the respiratory system (Valvano et al., 2005). However, the replication ability of Burkholderia inside cell lines is known to vary between isolates or genomovars (Saini et al., 1999; Sajjan et al., 2006). In this study we showed that both B. ambifaria HSJ1 wild-type and variant can replicate within phagosomes, but HSJ1v clearly displayed reduced replication in the human monocyte cell line compared with HSJ1. Our results are consistent with those of Savoia and Zucca (2007), who observed replication of a B. ambifaria clinical strain in murine macrophages but no significant replication for an environmental strain.
Numerous factors involved in the virulence of Bcc strains have been identified, especially in B. cenocepacia (Kooi et al., 2006; Aubert et al., 2008; Sajjan et al., 2008). In our study, phenotypic and proteomic approaches reveal that transition from wild-type to variant affects the expression of several putative virulence factors, for instance, biofilm formation or protease activity. Indeed, HSJ1 shows greater protease activity than HSJ1v, probably because of low ZmpA and ZmpB expression in HSJ1v. In B. cenocepacia, zmpA− zmpB− mutants were less virulent in a rat agar bead chronic infection model (Kooi et al., 2006). These proteins are probably also involved in the virulence of B. ambifaria, as they have a very high degree of homology inside the Bcc (for example, 93% identity between B. ambifaria and B. cenocepacia). In the same way, cholesterol oxidation, which is detected only in the wild-type, could be involved in the virulence of HSJ1. Indeed, cholesterol oxidase is required for mice infection by Mycobacterium tuberculosis (Brzostek et al., 2007).
A remarkable difference between the two subgroups of B. ambifaria is the production of HMAQs, which is synthesized by clinical isolates. These compounds are also produced by the human pathogen B. pseudomallei (Vial et al., 2008). Their role in B. ambifaria is still under investigation. In P. aeruginosa, some of the HAQs are implicated in the regulation of multiple virulence factors (Dubern and Diggle, 2008). Therefore, HMAQs could also be involved in the virulence of clinical isolates, especially during infections in CF patients.
Bcc isolates are resistant to a wide variety of antimicrobial compounds, including antimicrobial peptides (Mahenthiralingam et al., 2005; Loutet et al., 2006). In the Bcc, the complete lipid A-core oligosaccharide is required for resisting polymyxin B and melittin (Loutet et al., 2006). The modification of the LPS profile that we observed between the wild-type and the variant might explain their difference in sensitivity to antimicrobial peptides.
All these differences between the two phenotypic states could explain that clinical isolates are more virulent to animal hosts, and possibly better adapted to the lungs of CF patients.
Adaptation of the variant to the rhizosphere
Our data show that the variant is more competitive in colonizing the rhizosphere than the wild-type. Interestingly, in P. fluorescens F113, phenotypic variation occurs during rhizosphere colonization (Martinez-Granero et al., 2006). For B. ambifaria HSJ1, an increase in the proportion of variants generated by the wild-type during the colonization of the pea rhizosphere was not observed. However, it is conceivable that the rhizosphere of other plants could stimulate the frequency of phase variation.
Using API-ZYM strips, we detected that HSJ1v possesses a stronger β-glucosidase activity when compared with HSJ1 (Table 1). β-Glucosidases belong to a heterogeneous group of enzymes that catalyze the hydrolysis of cellobiose and related β-glucosides. Furthermore, only the variant can grow with saccharose and xylose as carbon sources. Finally, HSJ1v grows faster than HSJ1 when polyols (glycerol and sorbitol) are used as carbon sources. The HSJ1 variant is nutritionally more versatile than the clinical wild-type and consequently probably better adapted to the rhizosphere. Indeed, these sugars recovered from plant root exudates could be used as carbon source.
The variants and the environmental isolates of B. ambifaria produce more EPSs than the clinical isolates, a result that is in agreement with the data from Zlosnik et al. (2008). In that study, 92.5% of the B. ambifaria environmental strains were frankly mucoid, whereas the clinical strain CEP0996 was not. Interestingly, B. cenocepacia strains, the most virulent species of the Bcc, are generally non-mucoid (Zlosnik et al., 2008). This would suggest that EPS production is not a feature of CF lungs colonization, in contrast with P. aeruginosa. The role of EPSs in plant–Burkholderia interaction is not well documented. However, onion tissues induce EPS production from Burkholderia, probably through the carbohydrates produced by the onion (Bartholdson et al., 2008). In some plant–bacterial associations (for example, Rhizobium), EPSs are involved in the attachment of bacteria to the roots. EPS from the Bcc and notably from B. ambifaria could be involved in bacterial–plant interactions.
B. ambifaria, and the other Bcc members, exert beneficial effects on plant growth by preventing the deleterious functions of pathogenic microorganisms, generally by producing antibiotics or by competing for nutrients such as iron (Compant et al., 2008). The variant HSJ1v expresses a much greater ability for iron scavenging than HSJ1. In contrast, only the wild-type HSJ1 produces an antifungal compound in vitro. Surprisingly, this activity is absent from variants and from most environmental strains. Indeed, B. ambifaria is well characterized as a biocontrol agent: B. ambifaria AMMD is known to show antagonistic activity against the plant pathogens Pythium and Aphanomyces euteiches (Parke et al., 1991). We have confirmed that B. ambifaria AMMD possesses the capacity to inhibit the growth of Pythium (data not shown), suggesting the presence of several compounds responsible for the antifungal activity in B. ambifaria.
Phase variation in B. ambifaria
Phase variants of B. ambifaria arise at frequencies of 10−4 per cell per generation, which is similar to those reported for variants in many bacteria (Wisniewski-Dyé and Vial, 2008). However, in most reports, the variants are unstable and revert to the original phenotype. In our case, variants of all the clinical B. ambifaria isolates showed stable phenotypes over multiple subculturing or culture conditions in the laboratory. Notably, even after interactions with eukaryotic hosts (for instance, in the G. mellonella or macrophage cell line) no reversion could be detected. Remarkably, for the phytopathogen Ralstonia solanacearum, a phenotypic conversion from mucoid to non-mucoid state was evident in vitro, but revertants could only be detected ex planta or in the presence of tomato root exudates (Poussier et al., 2003). In the case of B. ambifaria, reversion to the wild-type might be restricted to very specific niche conditions, as, for instance, typically obtained in CF lungs. Thus, it is conceivable that environmental isolates of B. ambifaria would represent the primary source of infection of CF patients.
Phase variation mechanisms can involve a genome modification (for example, gene conversion and site-specific inversion) or epigenetic mechanisms such as differential methylation. In B. cenocepacia strain K56-2, a putative LysR transcriptional regulator (BCAS0225) was responsible for the shiny colony morphology and several shiny colony variants harbor spontaneous mutation in this gene. Interestingly, a highly conserved homologue of this gene is present in B. ambifaria (Bamb_6152 for B. ambifaria AMMD, >90% identity in amino acid sequences with BCAS0225). However, and in contrast with B. cenocepacia, no difference was detected between the gene sequence for the wild-type and for the variant, suggesting that this LysR regulator is not involved in the regulation of phase variation in B. ambifaria (data not shown).
Finally, it must be mentioned that until now, B. ambifaria has been essentially isolated from the plant rhizosphere and from CF lung infections. From the perspective we used here, the wild-type phenotype is the clinical form of the bacterium. Alternatively, because the variant type we identified is apparently the regular state in the environment, the latter could be considered closer to the actual wild type from the environmental point of view.
Conclusion
Currently, phase variation is viewed as a mechanism to evade immune system or for adaptation to different sites during host interaction (van der Woude and Baumler, 2004). The case of B. ambifaria appears unprecedented as it suggests that an opportunistic pathogen is able to generate a variant form that is less virulent to animal hosts but better adapted to the rhizosphere. In other words, a phase variation phenomenon occurs in B. ambifaria for niche adaptation.
Phase variation is likely not limited to B. ambifaria within the Bcc. For instance, shiny colony variants, isolated from B. cenocepacia strain K56-2, are less virulent in a chronic lung infection system and in the alfalfa model (Bernier et al., 2008). These variants, as is the case for B. ambifaria, could be preferentially adapted to the rhizosphere and show beneficial properties for plant growth.
Further studies are needed to decipher the mechanism used by B. ambifaria for phase variation, notably to understand the phenotypical stability of the variants. This could allow the development of genetically stable variants, which would combine desirable biocontrol properties with a reduced risk to cause human infections.
References
Apicella MA, Griffiss JM, Schneider H . (1994). Isolation and characterization of lipopolysaccharides, lipooligosaccharides, and lipid A. Methods Enzymol 235: 242–252.
Aubert DF, Flannagan RS, Valvano MA . (2008). A novel sensor kinase-response regulator hybrid controls biofilm formation and type VI secretion system activity in Burkholderia cenocepacia. Infect Immun 76: 1979–1991.
Bartholdson SJ, Brown AR, Mewburn BR, Clarke DJ, Fry SC, Campopiano DJ et al. (2008). Plant host and sugar alcohol induced exopolysaccharide biosynthesis in the Burkholderia cepacia complex. Microbiology 154: 2513–2521.
Bernier SP, Nguyen DT, Sokol PA . (2008). A LysR-type transcriptional regulator in Burkholderia cenocepacia influences colony morphology and virulence. Infect Immun 76: 38–47.
Bernier SP, Silo-Suh L, Woods DE, Ohman DE, Sokol PA . (2003). Comparative analysis of plant and animal models for characterization of Burkholderia cepacia virulence. Infect Immun 71: 5306–5313.
Brzostek A, Dziadek B, Rumijowska-Galewicz A, Pawelczyk J, Dziadek J . (2007). Cholesterol oxidase is required for virulence of Mycobacterium tuberculosis. FEMS Microbiol Lett 275: 106–112.
Cardona ST, Wopperer J, Eberl L, Valvano MA . (2005). Diverse pathogenicity of Burkholderia cepacia complex strains in the Caenorhabditis elegans host model. FEMS Microbiol Lett 250: 97–104.
Chiarini L, Bevivino A, Dalmastri C, Tabacchioni S, Visca P . (2006). Burkholderia cepacia complex species: health hazards and biotechnological potential. Trends Microbiol 14: 277–286.
Coenye T, Mahenthiralingam E, Henry D, LiPuma JJ, Laevens S, Gillis M et al. (2001). Burkholderia ambifaria sp. nov., a novel member of the Burkholderia cepacia complex including biocontrol and cystic fibrosis-related isolates. Int J Syst Evol Microbiol 51: 1481–1490.
Compant S, Nowak J, Coenye T, Clement C, Ait Barka E . (2008). Diversity and occurrence of Burkholderia spp. in the natural environment. FEMS Microbiol Rev 32: 607–626.
Cornillon S, Pech E, Benghezal M, Ravanel K, Gaynor E, Letourneur F et al. (2000). Phg1p is a nine-transmembrane protein superfamily member involved in Dictyostelium adhesion and phagocytosis. J Biol Chem 275: 34287–34292.
Déziel E, Comeau Y, Villemur R . (2001). Initiation of biofilm formation by Pseudomonas aeruginosa 57RP correlates with emergence of hyperpiliated and highly adherent phenotypic variants deficient in swimming, swarming, and twitching motilities. J Bacteriol 183: 1195–1204.
Doukyu N, Aono R . (2001). Cloning, sequence analysis and expression of a gene encoding an organic solvent- and detergent-tolerant cholesterol oxidase of Burkholderia cepacia strain ST-200. Appl Microbiol Biotechnol 57: 146–152.
Dubern JF, Diggle SP . (2008). Quorum sensing by 2-alkyl-4-quinolones in Pseudomonas aeruginosa and other bacterial species. Mol Biosyst 4: 882–888.
Esselmann MT, Liu PV . (1961). Lecithinase production by gram-negative bacteria. J Bacteriol 81: 939–945.
Fahraeus G . (1957). The infection of clover root hairs by nodule bacteria studied by a simple glass slide technique. J Gen Microbiol 16: 374–381.
Forest C, Faucher SP, Poirier K, Houle S, Dozois CM, Daigle F . (2007). Contribution of the stg fimbrial operon of Salmonella enterica serovar Typhi during interaction with human cells. Infect Immun 75: 5264–5271.
Govan JR, Brown AR, Jones AM . (2007). Evolving epidemiology of Pseudomonas aeruginosa and the Burkholderia cepacia complex in cystic fibrosis lung infection. Future Microbiol 2: 153–164.
Govan JRW, Balendreau J, Vandamme P . (2000). Burkholderia cepacia—friend and foe. ASM News 66: 124–125.
Kooi C, Subsin B, Chen R, Pohorelic B, Sokol PA . (2006). Burkholderia cenocepacia ZmpB is a broad-specificity zinc metalloprotease involved in virulence. Infect Immun 74: 4083–4093.
Lépine F, Déziel E, Milot S, Rahme LG . (2003). A stable isotope dilution assay for the quantification of the Pseudomonas quinolone signal in Pseudomonas aeruginosa cultures. Biochim Biophys Acta 1622: 36–41.
LiPuma JJ, Spilker T, Coenye T, Gonzalez CF . (2002). An epidemic Burkholderia cepacia complex strain identified in soil. Lancet 359: 2002–2003.
Loutet SA, Flannagan RS, Kooi C, Sokol PA, Valvano MA . (2006). A complete lipopolysaccharide inner core oligosaccharide is required for resistance of Burkholderia cenocepacia to antimicrobial peptides and bacterial survival in vivo. J Bacteriol 188: 2073–2080.
Mahenthiralingam E, Urban TA, Goldberg JB . (2005). The multifarious, multireplicon Burkholderia cepacia complex. Nat Rev Microbiol 3: 144–156.
Marolda CL, Hauroder B, John MA, Michel R, Valvano MA . (1999). Intracellular survival and saprophytic growth of isolates from the Burkholderia cepacia complex in free-living amoebae. Microbiology 145 (Pt 7): 1509–1517.
Martinez-Granero F, Rivilla R, Martin M . (2006). Rhizosphere selection of highly motile phenotypic variants of Pseudomonas fluorescens with enhanced competitive colonization ability. Appl Environ Microbiol 72: 3429–3434.
Mercanti V, Charette SJ, Bennett N, Ryckewaert JJ, Letourneur F, Cosson P . (2006). Selective membrane exclusion in phagocytic and macropinocytic cups. J Cell Sci 119: 4079–4087.
O'Sullivan LA, Weightman AJ, Jones TH, Marchbank AM, Tiedje JM, Mahenthiralingam E . (2007). Identifying the genetic basis of ecologically and biotechnologically useful functions of the bacterium Burkholderia vietnamiensis. Environ Microbiol 9: 1017–1034.
Parke JL, Gurian-Sherman D . (2001). Diversity of the Burkholderia cepacia complex and implications for risk assessment of biological control strains. Annu Rev Phytopathol 39: 225–258.
Parke JL, Rand RE, Joy AE, King EB . (1991). Biological control of Aphanomyces root rot and Pythium damping-off of peas by application of Pseudomonas cepacia or Pseudomonas fluorescens applied to seed. Plant Dis 75: 987–992.
Payne GW, Vandamme P, Morgan SH, Lipuma JJ, Coenye T, Weightman AJ et al. (2005). Development of a recA gene-based identification approach for the entire Burkholderia genus. Appl Environ Microbiol 71: 3917–3927.
Pearson JP, Pesci EC, Iglewski BH . (1997). Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J Bacteriol 179: 5756–5767.
Poussier S, Thoquet P, Trigalet-Demery D, Barthet S, Meyer D, Arlat M et al. (2003). Host plant-dependent phenotypic reversion of Ralstonia solanacearum from non-pathogenic to pathogenic forms via alterations in the phcA gene. Mol Microbiol 49: 991–1003.
Ramette A, LiPuma JJ, Tiedje JM . (2005). Species abundance and diversity of Burkholderia cepacia complex in the environment. Appl Environ Microbiol 71: 1193–1201.
Reik R, Spilker T, Lipuma JJ . (2005). Distribution of Burkholderia cepacia complex species among isolates recovered from persons with or without cystic fibrosis. J Clin Microbiol 43: 2926–2928.
Rosenfeld Y, Shai Y . (2006). Lipopolysaccharide (Endotoxin)-host defense antibacterial peptides interactions: role in bacterial resistance and prevention of sepsis. Biochim Biophys Acta 1758: 1513–1522.
Saini LS, Galsworthy SB, John MA, Valvano MA . (1999). Intracellular survival of Burkholderia cepacia complex isolates in the presence of macrophage cell activation. Microbiology 145 (Pt 12): 3465–3475.
Sajjan SU, Carmody LA, Gonzalez CF, LiPuma JJ . (2008). A type IV secretion system contributes to intracellular survival and replication of Burkholderia cenocepacia. Infect Immun 76: 5447–5455.
Sajjan US, Yang JH, Hershenson MB, LiPuma JJ . (2006). Intracellular trafficking and replication of Burkholderia cenocepacia in human cystic fibrosis airway epithelial cells. Cell Microbiol 8: 1456–1466.
Savoia D, Zucca M . (2007). Clinical and environmental Burkholderia strains: biofilm production and intracellular survival. Curr Microbiol 54: 440–444.
Seed KD, Dennis JJ . (2008). Development of Galleria mellonella as an alternative infection model for the Burkholderia cepacia complex. Infect Immun 76: 1267–1275.
Segura I, Casadesus J, Ramos-Morales F . (2004). Use of mixed infections to study cell invasion and intracellular proliferation of Salmonella enterica in eukaryotic cell cultures. J Microbiol Methods 56: 83–91.
Valvano MA, Keith KE, Cardona ST . (2005). Survival and persistence of opportunistic Burkholderia species in host cells. Curr Opin Microbiol 8: 99–105.
van der Woude MW, Baumler AJ . (2004). Phase and antigenic variation in bacteria. Clin Microbiol Rev 17: 581–611.
Vanlaere E, Baldwin A, Gevers D, Henry D, De Brandt E, LiPuma JJ et al. (2009). Taxon K, a complex within the Burkholderia cepacia complex, comprises at least two novel species, Burkholderia contaminans sp. nov. and Burkholderia lata sp. nov. Int J Syst Evol Microbiol 59: 102–111.
Vial L, Lépine F, Milot S, Groleau MC, Dekimpe V, Woods DE et al. (2008). Burkholderia pseudomallei, B. thailandensis, and B. ambifaria produce 4-hydroxy-2-alkylquinoline analogues with a methyl group at the 3 position that is required for quorum-sensing regulation. J Bacteriol 190: 5339–5352.
Villemur R, Déziel E . (2005). Phase variation and antigenic variation. In: Mullany P (ed) The Dynamic Bacterial Genome. Cambridge University Press: New York. pp 277–322.
Wisniewski-Dyé F, Vial L . (2008). Phase and antigenic variation mediated by genome modifications. Antonie Van Leeuwenhoek 94: 493–515.
Wopperer J, Cardona ST, Huber B, Jacobi CA, Valvano MA, Eberl L . (2006). A quorum-quenching approach to investigate the conservation of quorum-sensing-regulated functions within the Burkholderia cepacia complex. Appl Environ Microbiol 72: 1579–1587.
Zhou H, Yao F, Roberts DP, Lessie TG . (2003). AHL-deficient mutants of Burkholderia ambifaria BC-F have decreased antifungal activity. Curr Microbiol 47: 174–179.
Zlosnik JE, Hird TJ, Fraenkel MC, Moreira LM, Henry DA, Speert DP . (2008). Differential mucoid exopolysaccharide production by members of the Burkholderia cepacia complex. J Clin Microbiol 46: 1470–1473.
Acknowledgements
We are grateful to Arnaud Graindorge (University of Lyon I), Deborah Henry (University of British Columbia), John J LiPuma (University of Michigan Medical School), Jim Tiedje (Michigan State University) and Céline Laferrière (Sainte-Justine Hospital, Montréal) for providing Burkholderia strains. We are also grateful to Claude Guertin for his expertise with G. mellonella. This study was supported by a Canadian Institutes of Health Research (CIHR) Operating grant to ED. LV and MGL were both recipients of postdoctoral fellowships from the Fondation Armand-Frappier, and VD of an FQRNT PhD fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Vial, L., Groleau, MC., Lamarche, M. et al. Phase variation has a role in Burkholderia ambifaria niche adaptation. ISME J 4, 49–60 (2010). https://doi.org/10.1038/ismej.2009.95
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.95
Keywords
This article is cited by
-
Molecular changes associated with spontaneous phenotypic variation of Paenibacillus polymyxa, a commonly used biocontrol agent, and temperature-dependent control of variation
Scientific Reports (2020)
-
Characterization of in vitro phenotypes of Burkholderia pseudomallei and Burkholderia mallei strains potentially associated with persistent infection in mice
Archives of Microbiology (2017)
-
Within-host evolution decreases virulence in an opportunistic bacterial pathogen
BMC Evolutionary Biology (2015)
-
Phenotypic characterization of colony morphological mutants of Burkholderia glumae that emerged during subculture
Journal of General Plant Pathology (2013)
-
Diversity of potential pathogenicity and biofilm formation among Burkholderia cepacia complex water, clinical, and agricultural isolates in China
World Journal of Microbiology and Biotechnology (2012)