Abstract
Background/Objectives:
Inter-individual variability in weight loss during obesity treatment is complex and poorly understood. Here we use whole body and tissue approaches to investigate fuel oxidation characteristics in skeletal muscle fibers, cells and distinct circulating protein biomarkers before and after a high fat meal (HFM) challenge in those who lost the most (obese diet-sensitive; ODS) vs the least (obese diet-resistant; ODR) amount of weight in a highly controlled weight management program.
Subjects/Methods:
In 20 weight stable-matched ODS and ODR women who previously completed a standardized clinical weight loss program, we analyzed whole-body energetics and metabolic parameters in vastus lateralis biopsies and plasma samples that were obtained in the fasting state and 6 h after a defined HFM, equivalent to 35% of total daily energy requirements.
Results:
At baseline (fasting) and post-HFM, muscle fatty acid oxidation and maximal oxidative phosphorylation were significantly greater in ODS vs ODR, as was reactive oxygen species emission. Plasma proteomics of 1130 proteins pre and 1, 2, 5 and 6 h after the HFM demonstrated distinct group and interaction differences. Group differences identified S-formyl glutathione hydratase, heat shock 70 kDA protein 1A/B (HSP72), and eukaryotic translation initiation factor 5 (eIF5) to be higher in ODS vs ODR. Group-time differences included aryl hydrocarbon interacting protein (AIP), peptidylpropyl isomerase D (PPID) and tyrosine protein-kinase Fgr, which increased in ODR vs ODS over time. HSP72 levels correlated with muscle oxidation and citrate synthase activity. These proteins circulate in exosomes; exosomes isolated from ODS plasma increased resting, leak and maximal respiration rates in C2C12 myotubes by 58%, 21% and 51%, respectively, vs those isolated from ODR plasma.
Conclusions:
Findings demonstrate distinct muscle metabolism and plasma proteomics in fasting and post-HFM states corresponding in diet-sensitive vs diet-resistant obese women.
Similar content being viewed by others
Introduction
Weight loss response in clinical obesity treatment programs is highly variable1, 2 and there is growing evidence of important biological factors that contribute to inherent differences in oxidative metabolism, and/or adaptation to energy restriction.3, 4, 5, 6, 7 In previous analyses of over 3400 patient records in the Ottawa Hospital Weight Management Program (OHWMP), we documented a two to threefold difference in weight loss response in diet compliant individuals.1, 2, 7, 8
Here, age-, sex- and initial body weight-matched adherent patients in the upper versus lower quintiles for weight loss in the first 6 weeks of meal-replacement are defined as obese diet-sensitive (ODS) and obese diet-resistant (ODR), respectively.
Using these criteria, ODS patients were found to have a higher proportion of type I muscle fibers; muscle fiber hypertrophy; increased expression of genes involved in oxidative metabolism; higher proton leak in isolated mitochondria and primary myotubes2, 8, 9 and greater expression of OXPHOS genes in whole blood prior to diet intervention.10 Moreover, patients defined as ODR in the meal-replacement program demonstrate impaired weight loss in response to bariatric surgery.11 These findings are consistent with the hypothesis that capacity for weight loss has an important biological basis.
With the goal of elucidating possible mechanisms, we have interrogated metabolic responses in a newly recruited group of closely matched ODS and ODR individuals under resting fasting conditions and in response to a defined high fat meal (HFM). We report distinct differences in muscle fiber fatty acid-supported respiration, maximal oxidative phosphorylation and circulating protein biomarkers.
Materials/subjects and methods
Participants
This study was approved by the Ottawa Health Science Network Research Ethics Board and participants provided informed written consent. These included weight stable, sedentary, non-smoking, non-diabetic females, with BMI >30 kg m−2. During the first 12 weeks of the 26 week program, participants consume 900 kcal day−1 of Optifast liquid meals (Nestlé Health Sciences). Rate of weight loss, corrected for age, initial body mass and BMI is calculated on the basis of serial measures in the first 6 weeks.2, 12 On the basis of comprehensive compliance measures as described previously2, 8 only diet-adherent women who had previously completed the OHWMP and were strictly identified as ODS or ODR.
Patients in the OHWMP were excluded from data analysis and further metabolic studies if: they did not meet the adherence criteria; they were absent for more than two visits during the initial 6 weeks on meal-replacement; physician notes expressing reservations about self-reported compliance; and/or there was inadequate completion of the laboratory testing protocol. Patients were also excluded from metabolic and clinical research if they had any medical conditions or were taking any medications that may affect rate of weight loss including prior bariatric surgery, use of any medication known to affect rate of weight loss or glucose homeostasis, abnormal thyroid indices, congestive heart failure, malignancy or obstructive sleep apnea (during the program or at the time of biopsy). Ten pairs of ODS and ODR subjects not previously studied were matched for age (±10 years), body mass (±10 kg) and BMI (±2 kg m−2) at program enrollment and at the time of study (±10 years).
Study design
Meeting 1
Participants maintained normal daily activities and physical activity during waking hours for the preceding 7 days was determined using Actical accelerometers (Bend, OR, USA) and written records. After an overnight fast, anthropometric measures including DEXA scan were obtained.
Meeting 2: Trial day
Participants consumed a standardized meal and arrived by taxi at 0700 hours after a 12 h fast and were met with a wheelchair, to minimize exertion.
Indirect calorimetry
Participants rested supine in the dark for 20 min. Expired gases were collected for 30 min using a hood (Vmax Encore 29N, SensorMedics, Yorba Linda, CA, USA). Following the HFM, gases were collected for 30 min at 1, 2, 3, 4 and 5 h. Fuel utilization was determined.13 Throughout the remainder of the day, when participants were not undergoing expired gas collection, they remained in a resting seated or reclined position. Urine was quantitatively collected for urea determination to assess protein oxidation.
Blood sampling
Samples were at 30 min intervals from 0 h (baseline) to 6 h via a forearm catheter.
Muscle biopsies
Pre and 6 h post-HFM, vastus lateralis biopsies were obtained using a Bergstrom needle in alternate legs. Muscle was placed immediately in ice-cold saline: 20–30 mg was transferred to ice-cold BIOPS14 for respiration and reactive oxygen species (ROS) measurements; the remaining tissue was flash frozen.
High fat meal
Following baseline sampling, participants consumed within 5 min a liquid HFM, modified from15 equivalent to 35% of total daily energy requirements (TDE). TDE was determined from the sum of resting metabolic rate (RMR; determined from indirect calorimetry), average daily energy expended from physical activity and the thermic effect of food (assumed 10% TDE).
Analyses of tissues, plasma and primary cells
High resolution respirometry
Permeabilized muscle fibers (1–4 mg) were prepared and analyzed in duplicate, at 37 °C, with 200–400 nmol O2/l14 in an Oxygraph O2K (Oroboros, Innsbruck, Austria).
Protocol 1
Routine leak in the absence of adenylates was measured in the presence of 1.5 mm malate (M) and 200 μm octanoylcarnitine (OC), followed by 5 mm ADP to assess state 3 FA-supported respiration. Maximal OXPHOS was measured with complex I substrates (5 mm pyruvate (P) and 10 mm glutamate (G)) followed by succinate (10 mm; (S)), and additional ADP. State 4 was assessed with oligomycin (Oligo; 2.5 μm). Non-mitochondrial respiration was assessed with antimycin A (AA; 2.5 μm). Cytochrome c oxidase (COX) activity was measured with tetramethylphosphodinitrate (TMPD; 5 mm), and ascorbate (5 mm). Autoxidation of TMPD was inhibited with sodium azide (100 mm).
Protocol 2
Complex I and downstream complexes were assessed with M and P, followed by ADP and G. Maximal OXPHOS was measured with S and ADP reinjections. Cytochrome C (Cyt C; 10 μm) was injected to confirm integrity of the mitochondria; respiration rates did not increase >5% of OXPHOS. Maximal uncoupled respiration was measured with FCCP (0.25–1 μm). Non-mitochondrial respiration was that remaining in the presence of AA. Maximal capacity of COX was measured with TMPD, as above.
H2O2 emission
H2O2 emission was measured in ~10 mg of permeabilized muscle prepared using the Oxygraph O2K fluorimeter, as described.16
Western blots
Proteins were blotted for: total OXPHOS (1:1000, ab-22604, Abcam, Cambridge, UK), and HSP72 (1:5000; C92F3A-S, Enzo Life Sciences, Farmingdale, NY, USA) and analyzed using enhanced chemiluminescence, and Image J software with vinculin (1:1000, ab129002, Abcam) or Ponceau as loading controls.
Enzyme activities
Citrate synthase and β-HAD activities were determined as described.17, 18
Plasma analytes
Standard kits were used: glucose (Sigma Aldrich, St Louis, MO, USA); free fatty acids and triglycerides (WAKO Diagnostics, Richmond, VA, USA); insulin (Sigma Aldrich).
Primary myotubes
Muscle satellite cells were isolated from vastus lateralis of ODR (42±5 year; 96.7±4.8 kg; BMI: 34.0±1.5 kg m−2, n=5) and ODS (42±3 year; 97.7±4.8 kg; BMI: 35.8±0.5 kg m−2, n=5).9, 19 On day 6, cells were treated with differentiation media containing 2% BSA and 1 mm carnitine with or without 500 μm palmitate for 24 h.
Reactive oxygen species
Cellular ROS emission was measured in a 96-well plate using 2′,7′–dichlorofluorescin diacetate.9
Fatty acid oxidation
Myotubes were grown to confluence in a 12-well plate and induced to differentiate for 7 days. FA oxidation was measured as described.20
Plasma acylcarnitine levels
Methods were as previously described21 with minor modifications,16 using baseline, 0.25, 0.5, and 1- 6 hourly samples.
Plasma proteomics
Analyses were performed on plasma collected at baseline and at 1, 2, 5, 6 h following the HFM using SOMAscan technology (Somalogics, Boulder, CO, USA). Each of the 1130 plasma proteins in the SOMAscan assay has a targeted SOMAmer (Slow Off-rate Modified Aptamer) reagent, which is used as an affinity-binding reagent and quantified on a custom Agilent hybridization chip. ODS and ODR samples were randomly assigned to plates within each assay and run along with a set of calibration and normalization samples. Intra-run normalization and inter-run calibration were performed according to SOMAscan v3 quality-control procedures.22
Isolation and analysis of plasma exosomes, and effects on muscle cells
Exosomes were isolated from 1.0 ml of plasma using differential ultracentrifugation and analyzed to confirm quality and quantity using Zetaview, as previously described.23 C2C12 myoblasts were seeded at 10 000 cells per well in 96-well Seahorse cell culture plates and differentiated 7 days. Myotubes were treated in pentuplicate with isolated exosomes in a substrate-limited media (DMEM with 0.5 mm glucose, 1 mm glutamine, 0.5 mm carnitine, 1% FBS) for 24 h. There was no significant difference in the absolute concentration of exosomes across patient groups, and the exosome final concentrations in the medium approximated that in plasma in vivo. A standard FA oxidation assay (Seahorse XFe96, Santa Clara, CA, USA) was performed in the presence of palmitate-conjugated BSA or a BSA control. Data were normalized to protein concentration.
Statistical analysis
Two-way ANOVA with Bonferroni post hoc analyses were used to assess meal and group effects. One-way ANOVA was used to assess differences in body mass, BMI and age. For plasma proteomics, repeated measures ANOVA was used for group effects, time effects and group × time effects. Biomarkers remaining significant after correction for multiple testing (n=1130) including Bonferroni-corrected P<0.05, Benjamini–Hochberg FDR q<0.05 and local false discovery rates (LFDR) <0.05 were selected as significant hits. All statistical analyses were conducted in R (version 3.0.1, Vienna, Austria) and Prism (Version 7.0; GraphPad; La Jolla, CA, USA).
Results
Anthropometry and weight loss characteristics
Prior to enrollment in the weight loss program, there were no differences in age, body mass or BMI. Following 6 week of dietary restriction, ODS women lost 44% more weight compared to ODR (ODR vs ODS; 6.9±0.5 kg, vs 11.5±0.6 kg, P<0.001; Table 1). At the time of the HFM challenge, age, body mass, BMI, waist circumference, body fat mass and lean mass were similar between ODS and ODR (Table 1). As in a previous study,8 there were no differences between groups in daily physical activity (Table 1).
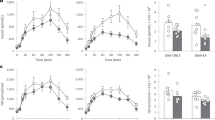
Following the HFM, whole-body O2 consumption, CO2 production and TEE increased over time to a similar extent in the two groups (Figures 1a, b and e; P<0.0001, effect of time, e; effect of time vs time 0). As expected, lipid utilization increased over time, whereas reliance on carbohydrate decreased (Figures 1c and d; P<0.01).
Whole-body response to the high fat meal in ODS and ODR individuals. Whole-body VO2 (a), VCO2 (b), carbohydrate utilization (c), lipid utilization (d) and total energy expenditure (e) at baseline and in response to the HFM. Plasma fatty acids (f), triglyceride (TG; (g, h) glucose and (i) insulin at baseline (0 min) and following a high fat meal in ODR and ODS individuals. Two-way ANOVA, significant effect of time, f–i: P<0.0001; a–d: P<0.01. (e) Total energy expenditure, Significant effect of time P<0.05, post hoc significantly different from baseline within group. Data are mean±s.e.m., n=9–10.
Plasma FA, TAG, glucose and insulin all increased over time, but were not different between groups. Plasma FA initially dropped (0–2 h) after the HFM then levelled-off, and began to increase at 4 h (Figure 1f; P<0.0001), whereas TG, glucose and insulin increased in both groups and began to decline at ~4 h, coinciding with the rise in FA (Figures 1g, i and d; P<0.0001).
Metabolic characteristics of skeletal muscle fibers
For the first time, we have investigated oxidative processes in isolated muscle fibers of ODS and ODR patients, and compared group responses to a HFM. Following the HFM, FA-supported respiration was significantly increased in both groups (Figure 2a; P<0.001) and tended to be higher in ODS vs ODR fibers (P=0.09). We also probed for differences in activity of the electron transport complexes and observed a post-meal increase in complex I (PMPG), that occurred in both groups (Figure 2b; P<0.01). Importantly, the maximal oxidative phosphorylation rate through complex I, II and ETF (PMOPGS) was greater in ODS vs ODR (Figure 2c; P<0.05) with a group difference following the meal. When we used traditional OXPHOS substrates, we also found higher respiration in ODS (Figure 2d; P=0.06). Moreover, maximal OXPHOS capacity was greater in ODS vs ODR fibers (Figure 2e; P<0.05). Non-phosphorylating respiration measured in the presence of malate and pyruvate, and in the presence of oligomycin did not differ between groups.
Skeletal muscle mitochondrial respiration but not content is increased in ODS compared to ODR prior to and in response to the high fat meal. Mitochondrial respiration was measured in permeabilized muscle fibers of ODS and ODR individuals pre and post the high fat meal. (a) State 3 fatty acid-supported respiration with 2 mm malate (M), 200 μm octanoylcarnitine and 5 mm ADP (PMO3). Two-way ANOVA, P=0.09, effect of group, *P<0.05 significant effect of meal. (b) State 3 complex I respiration with electron supply from 2 mm malate, 5 mm pyruvate (P), 10 mm glutamate (g) (PMPG3). *P<0.05, significant effect of meal. (c) Maximal oxidative phosphorylation with electron supply to complex I and II with substrate M, OC, P, G and 10 mm succinate (S)+ADP (PMOPGS3). *P<0.05 two-way ANOVA, significant effect of group, a post hoc significantly different between groups. (d) Maximal oxidative phosphorylation with electron supply to complex I and II with substrate M, P,G, S (PMPGS3), P=0.06, two-way ANOVA, effect of group. (e) Maximal capacity of the electron transfer system, as measured with the chemical uncoupler FCCP. *P<0.05 significant effect of group. (f) Expression of electron transfer system proteins (complex I, II, III, IV, V), (g) citrate synthase activity and (h) beta HAD activity in skeletal muscle of ODR and ODS at baseline. (i) COX activity in permeabilized muscle fibers of ODS and ODR was measured with 5 mm TMPD maintained in a reduced state by 5 mm ascorbate and reducing cytochrome C as the substrate. (j) FA-supported respiration expressed per COX activity. *Two-way ANOVA, significant effect of group. **P<0.01, significant effect of meal, post hoc significantly different post between ODS and ODR. (k) Maximal oxidative phosphorylation (PMOPGS3) expressed per COX activity. *P<0.05 two-way ANOVA significant effect of group. (l) Maximal oxidative phosphorylation (PMPGS3) expressed per COX activity. *P<0.05 two-way ANOVA significant effect of group and meal. Data are mean±s.e.m.
We then sought to determine if differences in fiber energetics were related to mitochondrial content. However there was no difference in OXPHOS protein levels (Figure 2f), citrate synthase activity (Figure 2g) or muscle fiber COX activity (Figure 2i) between ODS and ODR. We also measured β-HAD activity, as a marker of FA oxidative capacity, and there were no difference between groups (Figure 2h). Moreover, when we normalized FA-supported maximal OXPHOS and uncoupled respiration to COX activity, rates remained higher in ODS vs ODR (Figures 2j–l, P<0.05). Altogether, these findings demonstrate greater FA-driven respiration and OXPHOS system activity in fibers of ODS vs ODR that are not due to differences in mitochondrial content.
As the formation of mitochondrial supercomplexes promotes electron flow through the ETS,24 we measured supercomplexes in skeletal muscle. There were no differences between groups or in response to the HFM (Supplementary Figure 1). Thus, differences in muscle energetics between groups and following the HFM are not due to changes in ETS supercomplexes.
FA oxidation
As circulating acylcarnitines can be indicative of disordered FA oxidation,25 we measured plasma acylcarnitines at baseline and after the HFM. Short (C2–C4), medium (C6–C10) and long chain (C12-C18) acylcarnitines changed over time (Figures 3a–c; P<0.01, effect of time), with an initial increase at 15 min. Medium chain acylcarnitines were significantly affected by the HFM (Figure 3b; ODR 15 min vs 2, 3, 4, 5 h, P<0.05; ODS 15 min vs 3 h, 5 h, P<0.05) and peaked again at 6 h in ODR (6 h vs 3 h, P<0.05) but not in ODS. The AUC for long chain acylcarnitines was lower in ODS (P=0.06) suggesting that FA oxidation is more complete in ODS.
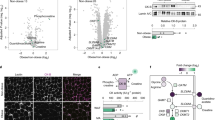
FA metabolism and oxidative stress in ODS and ODR. Plasma (a) short, (b) medium and (c) long chain acylcarnitine content and area under the curve (AUC) in ODS and ODR at baseline and in response to the HFM. Two-way ANOVA, Bonferroni post hoc, significant effect of time **P<0.01, ***P<0.001. Data are mean±s.e.m. (n=9). (d–h) FA oxidation in myotubes derived from ODR and ODS±glucose pre-treatment to test metabolic flexibility. (N=5) Data are mean±s.e.m. Two-way ANOVA, line above graph indicates significant effect of glucose *P<0.05, **P<0.01, ***P<0.001. (d) ASP in the media, post hoc effect of glucose ODR **P<0.01, ODS ***P<0.001 (e) ASP in the cell, post hoc effect of glucose, ODS *P<0.05; (f) Incomplete FA oxidation, post hoc effect of glucose, ODS P<0.01; (g) Complete FA oxidation to CO2 production, post hoc effect of glucose ODR, *P<0.05, ODS ***P<0.001; and (h) total FA oxidation, post hoc effect of glucose ODR *P<0.05, ODS **P<0.01. (i) Oligomycin-induced H2O2 emission in skeletal muscle of ODS and ODR pre and post-HFM. *P<0.05 Two-way ANOVA, significant effect of group (N=7–8). (j) ROS emission in myotubes derived from ODS and ODR with and without 24 h 500 μm palmitate treatment. Two-way ANOVA **P<0.01 significant effect of treatment. A P<0.05 post hoc significant difference from control condition within ODS n=5.
We then assessed FA oxidation in cultured primary myotubes from ODS and ODR individuals with or without glucose pre-treatment (Figures 3d–h). As expected, glucose pre-incubation decreased FA oxidation (Figures 3d–h). Complete, incomplete (acid soluble product (ASP) in the media+ASP in the cells) and total FA oxidation were similar between groups. Interestingly, in ODR cellular ASP and incomplete FA oxidation did not differ in the presence of glucose (Figures 3e and f) indicative of reduced metabolic flexibility.
Oxidative stress
A similar HFM was reported to increase muscle ROS.15 Interestingly, we found that ROS emissions were higher in ODS vs ODR, with no HFM effect (Figure 3i). We also assessed ROS in primary myotubes following 24 h of 500 μm palmitate exposure. Palmitate increased ROS (overall effect), and in ODS specifically (Figure 3j), consistent with observed increases in ROS in ODS ex vivo muscle (above).
As a measure of oxidative damage, protein carbonyls did not differ in muscle (group or HFM effects), but the HFM increased carbonylation in adipose in both groups (Supplementary Figure 2; P<0.05).
Plasma proteomic biomarkers
To potentially identify biomarkers of diet responsiveness, we conducted proteomic analyses of 1130 proteins in plasma collected at baseline, 1, 2, 5 and 6 h after the HFM. QQ plots of P-values for the group effects, and group-time interaction effects are shown in Figures 4a and e, respectively. Local false discovery rate and Q-values are available in Supplementary Table 1. After correction for multiple testing, proteins differing between groups (regardless of the fasted or fed states) included S-formyl glutathione hydratase (esterase D), heat shock 70 kDA protein 1 A/B (heat shock protein 72; HSP72), and eukaryotic translation initiation factor 5 (group effect, Figures 4b–d). Moreover, plasma aryl hydrocarbon receptor-interacting protein (AIP; also known as ARA9, part of the acryl hydrocarbon receptor protein complex), peptidylprolyl isomerase D (PPID; also known as cyclophilin D) and tyrosine protein-kinase Fgr (FGR) increased in ODR following the HFM (group × time effect, Figures 4f–h). Additional proteins of interest that were nominally significant include glutathione-S transferase P (Figure 4i group effect; P<0.01), tyrosine protein phosphatase non-receptor 11 (SHP2, Figure 4j, group effect; P<0.01) higher in ODS and brain derived neurotrophic factor (BDNF; Figure 4k group × time effect, P<0.05) and Peptide YY (PYY Figure 4l, group × time, P<0.01).
Plasma proteins at baseline and in response to a high fat meal in ODR and ODS individuals. (a) QQ plot—group analysis, (b) Esterase D, (c) heat shock protein 70 1 A/1B (HSP72) and (d) eukaryotic translation initiation factor 5 (eIF5); significant effect of group. (e) QQ plot—group × time analysis (f) aryl hydrocarbon interacting protein (AIP), (g) PPID and (h) tyrosine protein-kinase Fgr (FGR); significant different group × time, two-way repeated measures ANOVA, Bonferroni-corrected P-value, Q-value and LFDR<0.05 **P<0.01, *P<0.05). (i–l) Nominally significant plasma proteins at baseline and in response to the high fat meal. Group effect: (i) Glutathione-S transferase and (j) tyrosine protein phosphatase non-receptor type 11 (SHP2). Group × time effect: (k) brain derived neurotropic factor (BDNF) and (l) Protein YY (PYY). Two-way repeated measures ANOVA, **P<0.01, *P<0.01. HSP72 is positively correlated to (m) FA-supported respiration in muscle fibers (PMO3), (n) OXPHOS activity in muscle fibers (PMOPGS3), (o) complex I respiration in muscle fibers (PMP3) and (p) skeletal muscle citrate synthase activity. Data are mean±s.e.m., n=9–10.
Plasma HSP72 levels were higher in ODS versus ODR in both fasted and fed states (group effect, Bonferroni-corrected, P<0.05; Figure 4c). HSP72 has a role in reducing inflammation and insulin resistance in obesity and high fat feeding,26 and as we hypothesized, baseline HSP72 was positively correlated with FA-supported respiration (PMO3, P<0.05 Figure 4m), OXPHOS (PMOPGS3, P<0.05, Figure 4n), complex I respiration (PMP3, P=0.05, Figure 4o) and citrate synthase activity (P=0.05, Figure 4p). Muscle HSP72 protein expression was not different between ODS and ODR (Supplementary Figure 3).
Effects of plasma exosomes on muscle cell energetics
As HSP72 and the other identified proteins circulate in exosomes, which are biologically potent,27, 28, 29, 30 and our results demonstrated a FA oxidation defect in ODR vs ODS muscle (Figures 2a and 3), we investigated the ability of plasma exosomes from ODS and ODR to elicit effects on FA metabolism in a well characterized muscle cell line.
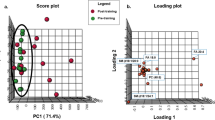
Exosome size distribution and the median size analyses demonstrated successful isolation of exosomes31, 32, 33 and no differences in size or plasma concentration between groups (Figures 5a and b; Supplementary Figure 4). Differentiated C2C12 myotubes were treated with exosomes from 1.0 ml of fasting (0 h) or fed (5 h post-HFM) plasma. Exosomes from ODS fasting plasma elicited resting, leak, and maximal oxygen consumption rates (OCRs) that were 58%, 21%, and 51% higher, respectively, as compared to exosomes from fasting ODR plasma (Figure 5c). In contrast, resting glycolysis rates were higher in ODR than ODS by 67% (Figure 5d). When OCRs were plotted against extracellular acidification rates (ECARs) to delineate the metabolic poise of cells, myotubes treated with fasting ODS exosomes are clearly less glycolytic and have a greater oxidative flexibility than the cells treated with the fasting ODR exosomes (Figure 5e). Interestingly, although exosomes isolated from the plasma taken 5 h after the HFM caused some stimulation of oxidative and glycolytic activities in the C2C12 cells, there were no differences between ODS and ODR groups (Figures 5f–h).
Differential metabolic effects of ODS and ODR plasma-derived exosomes on differentiated myotubes. Cellular respiration and extracellular acidification was measured in 7 day differentiated C2C12 myotubes in response to exosomes isolated from plasma samples pre (ODS n=9, ODR n=9) and 5 h post (ODS n=9, ODR n=8) HFM. (a) Representative Zetaview analysis of the size distribution of the vesicles. (b) Median sizes of the exosomes for ODS and ODR groups. (c, f) Cellular oxygen consumption rates (OCR) at rest, following oligomycin injection (3 μm; proton leak), and FCCP injection (2 μm; maximal respiration) with exosomes isolated from fasted (0 h) or post-HFM (5 h) respectively. (d, g) Extracellular acidification rates (ECAR) at rest, and following oligomycin and FCCP injection (maximal glycolysis) with exosomes isolated from fasted (0 h) or post-HFM (5 h) respectively. (e, h) Graph illustrating the metabolic flexibility from resting OCR and ECAR (left data points) and following oligomycin and FCCP injection (right data points). Data are mean±s.e.m., n=8–9, *P<0.05, **P<0.01.
Discussion
Here we have further interrogated the complex biology underlying weight loss variability in obesity. We focused on mechanisms in plasma and skeletal muscle related to fuel oxidation and metabolic flexibility in vivo and in vitro. We demonstrate that both FA-supported muscle respiration and maximal oxidative phosphorylation are greater in ODS vs ODR individuals under fasting, resting conditions, and following a high fat meal. These differences are independent of muscle mitochondrial content and oxidative phosphorylation supercomplex assembly. Seeking possible plasma markers of diet sensitivity vs resistance we surveyed 1130 circulating proteins before and at several time points after the HFM and identified several proteins that were either different between groups or exhibited different post-meal trajectories. HSP72 was positively correlated with muscle fiber respiration and citrate synthase activity. Given that all of the identified proteins circulate in exosomes, another key finding is that plasma exosomes from ODS stimulate oxidative metabolism in muscle cells to a greater extent than those from ODR.
In permeabilized muscle fibers in which mitochondrial structures remain intact and highly functional, we demonstrate that FA-supported respiration, oxidative phosphorylation with NADH- and FADH2-mediated electron supply through complexes I and II and the maximal capacity of the ETS were higher in ODS muscle. In addition, NADH-driven respiration through complex I is acutely increased following a HFM, consistent with the control of mitochondrial oxidative capacity through acute processes, such as post-translational modifications. Differences in mitochondrial respiration are often ascribed to differences in mitochondrial content.14, 34, 35, 36 Despite the use of various well recognized techniques, we observed no difference in mitochondrial content.
Mitochondrial ETS proteins interact to form supercomplexes, and dysfunctional supercomplexes are associated with aging,37 cardiovascular disease38 and type 2 diabetes,39 which may be related to oxidative stress.40 In the current study, muscle mitochondrial supercomplex were not different between ODS and ODR. Thus the observed differences in mitochondrial respiration (group and meal effects) may be a result of post-translational modification of ETS proteins, which will require further study.
Consistent with a lower capacity for FA oxidation in ODR muscle, plasma long chain acylcarnitines and peak medium chain acylcarnitines (15 min, 6 h) were higher in ODR than ODS. Acylcarnitines are intermediary metabolites of FA oxidation. Circulating acylcarnitines can be indicative of FA oxidation dysfunction,25, 41, 42 and plasma and skeletal muscle acylcarnitine profiles can be abnormal in obesity.41, 43, 44 Owing to the tissue constraints, we were unable to measure muscle acylcarnitines. We did however observe metabolic inflexibility in primary myotubes of ODR individuals. Indeed, FA-supported respiration in permeabilized muscle fibers was also lower in ODR. Taken together, findings indicate that FA metabolism is impaired in skeletal muscle of ODR individuals.
Plasma proteomics revealed that FGR, PPID and AIP increased in ODR following the HFM. FGR, a Src kinase, interacts with succinate dehydrogenase and aconitase.45 Redox sensitive Src kinases and tyrosine kinases have been found in mitochondria and can be activated by H2O2 or oxidative stress.46, 47 Src kinase is decreased in cardiac tissue in sepsis and associated with decreased tyrosine phosphorylation and activity of OXPHOS proteins.48 PPID (cyclophilin D) is an important regulator of mitochondrial physiology and interacts with proteins including glycogen synthase kinase 3, adenine nucleotide translocator, F1FO ATP synthase and the mitochondrial permeability transition pore (MPTP).49 MPTP has a key role in apoptosis as well as calcium handling and metabolism50 and the PPID locus is linked to serum metabolite levels.51 Finally, AIP (ARA9) is part of the aryl hydrocarbon receptor protein complex (AHR), which influences cell cycle, death, xenobiotic metabolism, and mitochondrial function, potentially through its interaction with ATP5α1. The unique increases in the levels of these circulating proteins in ODS, and the absence of responses in ODR may contribute at least in part to the observed differences in muscle mitochondrial metabolism following the HFM.
Proteomics also revealed that levels of esterase D, eIF5 and HSP72 are higher in ODS vs ODR. Esterase D is also referred to as S-formyl glutathione hydrolase, and is thought to hydrolyze a formyl group from S-formyl glutathione yielding glutathione.52 This may be relevant given that glutathione is the most important non-protein antioxidant in cells and also has a role in controlling mitochondrial protein activity through glutathionylation.53, 54 Glutathione redox is impaired in aging and many metabolic disorders55 and we previously reported that glutathione redox (GSH:GSSG) is impaired in primary myotubes of ODR compared to ODS.9 Further research is required to elucidate how glutathione metabolism becomes disordered in muscle of diet-resistant individuals. eIF5 is a protein translation initiation factor and catalyzes GTP hydrolysis from the 40S ribosomal initiation complex. It is possible that there are differences in protein translation in diet-resistant individuals, and this would require additional study.
Of particular interest, we found that plasma HSP72 was higher in ODS, and positively correlated with muscle FA-supported respiration, OXPHOS activity, citrate synthase activity and complex I respiration. HSPs mitigate oxidative stress and inflammation by acting as chaperones for naïve, aberrantly folded or mutated proteins.56 Muscle HSP72 expression is decreased in obesity26 and type 2 diabetes57 and correlates with muscle oxidative capacity in humans.57 HSP72 overexpression in muscle reduces high fat diet-induced inflammation, weight gain, and increases whole-body energy metabolism and mitochondrial content.26, 58 Interestingly, we did not find any difference in skeletal muscle HSP72 expression between ODS and ODR, despite the profound difference observed in plasma and its correlation with skeletal muscle oxidative capacity. Previous studies have found skeletal muscle HSP72 levels to be low in physiological states characterized by mitochondrial dysfunction (that is, type 2 diabetes). Some studies that have shown skeletal muscle HSP72 to be low in physiological states characterized by mitochondrial dysfunction, measured mRNA but not protein levels. As HSP72 is a stress-induced protein that controls protein turnover and chaperones proteins within cells and out of cells, HSP72 turnover is high, and this is controlled through HSP72 ubiquitination.59 Thus differences may be more apparent at the mRNA level, than the protein level. It has also been proposed that HSP72 may act as a myokine, thereby supporting the possibility of its active excretion from muscle.60 Unfortunately, due to tissue constraints we were unable to measure HSP72 mRNA expression. On the basis of the high proportion of total body mass that is skeletal muscle (~40%), and on prior research that links oxidative metabolism and skeletal muscle HSP72 levels,26, 57, 58 we feel that skeletal muscle is a likely source of plasma HSP72 in the current study. Further studies will need to be conducted to decipher the main source of plasma HSP72.
As proof-of-concept we then explored how these proteins (which circulate in exosomes) might affect muscle metabolism, and found that exosomes from ODS vs ODR groups differentially altered myotube FA metabolism. Fasting plasma exosomes from ODS enhanced FA oxidation, whereas ODR exosomes enhanced glycolysis, despite excess palmitate in the medium. ODS exosomes stimulated proton leak uncoupling; proton leak was previously found to be higher in isolated muscle mitochondria and in primary myotubes of ODS vs ODR.2, 9 However, we did not detect increased leak on ODS vs ODR permeabilized myofibers herein, an observation possibly due to differences in tissue preparation and incubation conditions.
Treatment of C2C12 cells with exosomes from the 5 h post-HFM time point resulted in a similar metabolic phenotype between groups; the effect of the group seems to be overpowered by the effect of the meal. Given the distinct differences in the circulating proteome following the meal, it is quite likely that different mechanisms are involved. Future studies should focus on the proteome, microRNA and mRNA content of the ODS and ODR exosomes, in fasted and fed states.
Skeletal muscle accounts for about 20% of total resting energy expenditure,61 which may explain in part why no measurable difference was observed in resting and postprandial whole-body energetics between ODR and ODS individuals despite marked differences in skeletal muscle metabolism observed.2, 9, 12 Undoubtedly, detection of differences in whole-body energy expenditure between ODS and ODR require longer time frames than the 30 min used here. It may be that differences in whole-body energy metabolism could be detected during exercise; this will be pursued in an upcoming study.
We also sought to investigate ROS and oxidative stress and observed higher ROS emission and glutathione-S transferase in ODS and increased ROS emission response to palmitate in primary myotubes from ODS, suggesting that ODS may have a greater capacity to respond to ROS compared to ODR. Contrary to Anderson et al.15 who found increased ROS emission 4 h following a HFM, we did not find increased oxidative stress following the HFM in either group. However, at 6 h we may have missed the maximal effect of the HFM; indeed, plasma TG and insulin began to decline at 4 h.
In summary, we provide novel insights into the biology underlying weight loss resistance, including lower muscle fiber oxidative characteristics independent of mitochondrial content or supercomplex assembly and metabolic inflexibility (cellular FA oxidation, plasma acylcarnitines). Multiple differences in the plasma proteome provide exciting clues relevant to diet resistance, and provide potentially useful biomarkers to predict weight loss success. In particular, greater HSP72 correlated with muscle respiration. Finally, the differential effects of plasma exosomes from ODS vs ODR on muscle cell metabolism are consistent with the possibility that factors circulating in exosomes could contribute to weight loss variability and lead to novel therapeutic approaches.
References
Dent R, McPherson R, Harper ME . Variability in weight loss in highly compliant women on a controlled dietary regimen. Obes Res 1999; 7 (Suppl. 1): 98S.
Harper ME, Dent R, Monemdjou S, Bezaire V, Van Wyck L, Wells G et al. Decreased mitochondrial proton leak and reduced expression of uncoupling protein 3 in skeletal muscle of obese diet-resistant women. Diabetes 2002; 51: 2459–2466.
Tremblay A, Lepage C, Panahi S, Couture C, Drapeau V . Adaptations to a diet-based weight-reducing programme in obese women resistant to weight loss. Clin Obes 2015; 5: 145–153.
Doucet E, St-Pierre S, Almeras N, Despres JP, Bouchard C, Tremblay A . Evidence for the existence of adaptive thermogenesis during weight loss. Br J Nutr 2001; 85: 715–723.
Leibel RL, Rosenbaum M, Hirsch J . Changes in energy expenditure resulting from altered body weight. N Engl J Med 1995; 332: 621–628.
Rosenbaum M, Leibel RL . Adaptive thermogenesis in humans. Int J Obes 2010; 34 (Suppl 1): S47–S55.
Thrush AB, Dent R, McPherson R, Harper ME . Implications of mitochondrial uncoupling in skeletal muscle in the development and treatment of obesity. FEBS J 2013; 280: 5015–5029.
Gerrits MF, Ghosh S, Kavaslar N, Hill B, Tour A, Seifert EL et al. Distinct skeletal muscle fiber characteristics and gene expression in diet-sensitive versus diet-resistant obesity. J Lipid Res 2010; 51: 2394–2404.
Thrush AB, Zhang R, Chen W, Seifert EL, Quizi JK, McPherson R et al. Lower mitochondrial proton leak and decreased glutathione redox in primary muscle cells of obese diet-resistant versus diet-sensitive humans. J Clin Endocrinol Metab 2014; 99: 4223–4230.
Ghosh S, Dent R, Harper ME, Stuart J, McPherson R . Blood gene expression reveal pathway differences between diet-sensitive and resistant obese subjects prior to caloric restriction. Obesity 2011; 19: 457–463.
Azar M, Nikpay M, Harper ME, McPherson R, Dent R . Can response to dietary restriction predict weight loss after Roux-en-Y gastroplasty? Obesity 2016; 24: 805–811.
Gerrits M, Ghosh S, Kavaslar N, Hill B, Tour A, Seifert E et al. Distinct skeletal muscle fiber characteristics and gene expression in diet-sensitive versus diet-resistant obesity. J Lipid Res 2010; 51: 2394–2404.
Blondin DP, Depault I, Imbeault P, Peronnet F, Imbeault MA, Haman F . Effects of two glucose ingestion rates on substrate utilization during moderate-intensity shivering. Eur J Appl Physiol 2010; 108: 289–300.
Boushel R, Gnaiger E, Schjerling P, Skovbro M, Kraunsoe R, Dela F . Patients with type 2 diabetes have normal mitochondrial function in skeletal muscle. Diabetologia 2007; 50: 790–796.
Anderson EJ, Lustig ME, Boyle KE, Woodlief TL, Kane DA, Lin CT et al. Mitochondrial H2O2 emission and cellular redox state link excess fat intake to insulin resistance in both rodents and humans. J Clin Invest 2009; 119: 573–581.
Beauchamp B, Thrush AB, Quizi J, Antoun G, McIntosh N, Al-Dirbashi OY et al. Undernutrition during pregnancy in mice leads to dysfunctional cardiac muscle respiration in adult offspring. Biosci Rep 2015; 35 (pii): e00200.
Srere PA . Citrate synthase. Methods Enzymol 1969; 13: 3–11.
Bruce CR, Thrush AB, Mertz VA, Bezaire V, Chabowski A, Heigenhauser GJ et al. Endurance training in obese humans improves glucose tolerance and mitochondrial fatty acid oxidation and alters muscle lipid content. Am J Physiol Endocrinol Metab 2006; 291: E99–E107.
Costford SR, Crawford SA, Dent R, McPherson R, Harper ME . Increased susceptibility to oxidative damage in post-diabetic human myotubes. Diabetologia 2009; 52: 2405–2415.
Aguer C, Fiehn O, Seifert EL, Bezaire V, Meissen JK, Daniels A et al. Muscle uncoupling protein 3 overexpression mimics endurance training and reduces circulating biomarkers of incomplete beta-oxidation. FASEB J 2013; 27: 4213–4225.
Turgeon C, Magera MJ, Allard P, Tortorelli S, Gavrilov D, Oglesbee D et al. Combined newborn screening for succinylacetone, amino acids, and acylcarnitines in dried blood spots. Clin Chem 2008; 54: 657–664.
Gold L, Walker JJ, Wilcox SK, Williams S . Advances in human proteomics at high scale with the SOMAscan proteomics platform. N Biotechnol 2012; 29: 543–549.
Mehdiani A, Maier A, Pinto A, Barth M, Akhyari P, Lichtenberg A . An innovative method for exosome quantification and size measurement. J Vis Exp 2015; 17: 50974.
Lapuente-Brun E, Moreno-Loshuertos R, Acin-Perez R, Latorre-Pellicer A, Colas C, Balsa E et al. Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 2013; 340: 1567–1570.
Adams SH, Hoppel CL, Lok KH, Zhao L, Wong SW, Minkler PE et al. Plasma acylcarnitine profiles suggest incomplete long-chain fatty acid beta-oxidation and altered tricarboxylic acid cycle activity in type 2 diabetic African-American women. J Nutr 2009; 139: 1073–1081.
Chung J, Nguyen AK, Henstridge DC, Holmes AG, Chan MH, Mesa JL et al. HSP72 protects against obesity-induced insulin resistance. Proc Natl Acad Sci USA 2008; 105: 1739–1744.
Meckes Jr DG, Gunawardena HP, Dekroon RM, Heaton PR, Edwards RH, Ozgur S et al. Modulation of B-cell exosome proteins by gamma herpesvirus infection. Proc Natl Acad Sci USA 2013; 110: E2925–E2933.
Bosman GJ, Lasonder E, Luten M, Roerdinkholder-Stoelwinder B, Novotny VM, Bos H et al. The proteome of red cell membranes and vesicles during storage in blood bank conditions. Transfusion 2008; 48: 827–835.
Vega VL, Rodriguez-Silva M, Frey T, Gehrmann M, Diaz JC, Steinem C et al. Hsp70 translocates into the plasma membrane after stress and is released into the extracellular environment in a membrane-associated form that activates macrophages. J Immunol 2008; 180: 4299–4307.
de Jong OG, Verhaar MC, Chen Y, Vader P, Gremmels H, Posthuma G et al. Cellular stress conditions are reflected in the protein and RNA content of endothelial cell-derived exosomes. J Extracell Vesicles 2012; 1.
Dragovic RA, Gardiner C, Brooks AS, Tannetta DS, Ferguson DJ, Hole P et al. Sizing and phenotyping of cellular vesicles using nanoparticle tracking analysis. Nanomedicine 2011; 7: 780–788.
Sokolova V, Ludwig AK, Hornung S, Rotan O, Horn PA, Epple M et al. Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B Biointerfaces 2011; 87: 146–150.
Colombo M, Moita C, van Niel G, Kowal J, Vigneron J, Benaroch P et al. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J Cell Sci 2013; 126 (Pt 24): 5553–5565.
DeLany JP, Dube JJ, Standley RA, Distefano G, Goodpaster BH, Stefanovic-Racic M et al. Racial differences in peripheral insulin sensitivity and mitochondrial capacity in the absence of obesity. J Clin Endocrinol Metab 2014; 99: 4307–4314.
Kristensen JM, Skov V, Petersson SJ, Ortenblad N, Wojtaszewski JF, Beck-Nielsen H et al. A PGC-1alpha- and muscle fibre type-related decrease in markers of mitochondrial oxidative metabolism in skeletal muscle of humans with inherited insulin resistance. Diabetologia 2014; 57: 1006–1015.
Ritov VB, Menshikova EV, He J, Ferrell RE, Goodpaster BH, Kelley DE . Deficiency of subsarcolemmal mitochondria in obesity and type 2 diabetes. Diabetes 2005; 54: 8–14.
Gomez LA, Monette JS, Chavez JD, Maier CS, Hagen TM . Supercomplexes of the mitochondrial electron transport chain decline in the aging rat heart. Arch Biochem Biophys 2009; 490: 30–35.
Rosca MG, Vazquez EJ, Kerner J, Parland W, Chandler MP, Stanley W et al. Cardiac mitochondria in heart failure: decrease in respirasomes and oxidative phosphorylation. Cardiovasc Res 2008; 80: 30–39.
Antoun G, McMurray F, Thrush AB, Patten DA, Peixoto AC, Slack RS et al. Impaired mitochondrial oxidative phosphorylation and supercomplex assembly in rectus abdominis muscle of diabetic obese individuals. Diabetologia 2015; 58: 2861–2866.
Maranzana E, Barbero G, Falasca AI, Lenaz G, Genova ML . Mitochondrial respiratory supercomplex association limits production of reactive oxygen species from complex I. Antioxid Redox Signal 2013; 19: 1469–1480.
Koves TR, Ussher JR, Noland RC, Slentz D, Mosedale M, Ilkayeva O et al. Mitochondrial overload and incomplete fatty acid oxidation contribute to skeletal muscle insulin resistance. Cell Metab 2008; 7: 45–56.
Thyfault JP, Cree MG, Zheng D, Zwetsloot JJ, Tapscott EB, Koves TR et al. Contraction of insulin-resistant muscle normalizes insulin action in association with increased mitochondrial activity and fatty acid catabolism. Am J Physiol Cell Physiol 2007; 292: C729–C739.
Baker PR 2nd, Boyle KE, Koves TR, Ilkayeva OR, Muoio DM, Houmard JA et al. Metabolomic analysis reveals altered skeletal muscle amino acid and fatty acid handling in obese humans. Obesity 2015; 23: 981–988.
Aguer C, McCoin CS, Knotts TA, Thrush AB, Ono-Moore K, McPherson R et al. Acylcarnitines: potential implications for skeletal muscle insulin resistance. FASEB J 2015; 29: 336–345.
Salvi M, Morrice NA, Brunati AM, Toninello A . Identification of the flavoprotein of succinate dehydrogenase and aconitase as in vitro mitochondrial substrates of Fgr tyrosine kinase. FEBS Lett 2007; 581: 5579–5585.
Hebert-Chatelain E . Src kinases are important regulators of mitochondrial functions. Int J Biochem Cell Biol 2013; 45: 90–98.
Salvi M, Brunati AM, Bordin L, La Rocca N, Clari G, Toninello A . Characterization and location of Src-dependent tyrosine phosphorylation in rat brain mitochondria. Biochim Biophys Acta 2002; 1589: 181–195.
Zang QS, Martinez B, Yao X, Maass DL, Ma L, Wolf SE et al. Sepsis-induced cardiac mitochondrial dysfunction involves altered mitochondrial-localization of tyrosine kinase Src and tyrosine phosphatase SHP2. PLoS One 2012; 7: e43424.
Giorgio V, Soriano ME, Basso E, Bisetto E, Lippe G, Forte MA et al. Cyclophilin D in mitochondrial pathophysiology. Biochim Biophys Acta 2010; 1797: 1113–1118.
Kwong JQ, Molkentin JD . Physiological and pathological roles of the mitochondrial permeability transition pore in the heart. Cell Metab 2015; 21: 206–214.
Illig T, Gieger C, Zhai G, Romisch-Margl W, Wang-Sattler R, Prehn C et al. A genome-wide perspective of genetic variation in human metabolism. Nat Genet 2010; 42: 137–141.
Hopkinson DA, Mestriner MA, Cortner J, Harris H, Esterase D . A new human polymorphism. Ann Hum Genet 1973; 37: 119–137.
Mailloux RJ, McBride SL, Harper ME . Unearthing the secrets of mitochondrial ROS and glutathione in bioenergetics. Trends Biochem Sci 2013; 38: 592–602.
Mailloux RJ, Xuan JY, McBride S, Maharsy W, Thorn S, Holterman CE et al. Glutaredoxin-2 is required to control oxidative phosphorylation in cardiac muscle by mediating deglutathionylation reactions. J Biol Chem 2014; 289: 14812–14828.
Go YM, Jones DP . Redox theory of aging: implications for health and disease. Clin Sci 2017; 131: 1669–1688.
Benjamin IJ, McMillan DR . Stress (heat shock) proteins: molecular chaperones in cardiovascular biology and disease. Circ Res 1998; 83: 117–132.
Bruce CR, Carey AL, Hawley JA, Febbraio MA . Intramuscular heat shock protein 72 and heme oxygenase-1 mRNA are reduced in patients with type 2 diabetes: evidence that insulin resistance is associated with a disturbed antioxidant defense mechanism. Diabetes 2003; 52: 2338–2345.
Henstridge DC, Bruce CR, Drew BG, Tory K, Kolonics A, Estevez E et al. Activating HSP72 in rodent skeletal muscle increases mitochondrial number and oxidative capacity and decreases insulin resistance. Diabetes 2014; 63: 1881–1894.
Qian SB, McDonough H, Boellmann F, Cyr DM, Patterson C . CHIP-mediated stress recovery by sequential ubiquitination of substrates and Hsp70. Nature 2006; 440: 551–555.
Henstridge DC, Whitham M, Febbraio MA . Chaperoning to the metabolic party: The emerging therapeutic role of heat-shock proteins in obesity and type 2 diabetes. Mol Metab 2014; 3: 781–793.
Elia M . The inter-organ flux of substrates in fed and fasted man, as indicated by arterio-venous balance studies. Nutr Res Rev 1991; 4: 3–31.
Acknowledgements
We are deeply grateful to the study participants in this research. We thank J Quizi, M Pasqua, Dr F McMurray, H Doelle and A Beninato for assistance with laboratory and clinical protocols. ABT was supported by a Canadian Institutes of Health Research (CIHR) postdoctoral fellowship, GA by a Vanier Canada Graduate Scholarship and BB by a Natural Sciences and Engineering Research Council scholarship. This study was funded by CIHR (MOP57810, MOP136936).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on International Journal of Obesity website
Rights and permissions
This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it.The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Thrush, A., Antoun, G., Nikpay, M. et al. Diet-resistant obesity is characterized by a distinct plasma proteomic signature and impaired muscle fiber metabolism. Int J Obes 42, 353–362 (2018). https://doi.org/10.1038/ijo.2017.286
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2017.286
This article is cited by
-
Plasma proteomic profiles predict individual future health risk
Nature Communications (2023)
-
Position statement on nutrition therapy for overweight and obesity: nutrition department of the Brazilian association for the study of obesity and metabolic syndrome (ABESO—2022)
Diabetology & Metabolic Syndrome (2023)
-
Dopamine receptor D1- and D2-agonists do not spark brown adipose tissue thermogenesis in mice
Scientific Reports (2020)
-
Proteomic profiles before and during weight loss: Results from randomized trial of dietary intervention
Scientific Reports (2020)
-
Plasma protein patterns as comprehensive indicators of health
Nature Medicine (2019)