Abstract
Huanglongbing (HLB) is the most destructive bacterial disease of citrus worldwide. While most citrus varieties are susceptible to HLB, Poncirus trifoliata, a close relative of Citrus, and some of its hybrids with Citrus are tolerant to HLB. No specific HLB tolerance genes have been identified in P. trifoliata but recent studies have shown that constitutive disease resistance (CDR) genes were expressed at much higher levels in HLB-tolerant Poncirus hybrids and the expression of CDR genes was modulated by Candidatus Liberibacter asiaticus (CLas), the pathogen of HLB. The current study was undertaken to mine and characterize the CDR gene family in Citrus and Poncirus and to understand its association with HLB tolerance in Poncirus. We identified 17 CDR genes in two citrus genomes, deduced their structures, and investigated their phylogenetic relationships. We revealed that the expansion of the CDR family in Citrus seems to be due to segmental and tandem duplication events. Through genome resequencing and transcriptome sequencing, we identified eight CDR genes in the Poncirus genome (PtCDR1-PtCDR8). The number of SNPs was the highest in PtCDR2 and the lowest in PtCDR7. Most of the deletion and insertion events were observed in the UTR regions of Citrus and Poncirus CDR genes. PtCDR2 and PtCDR8 were in abundance in the leaf transcriptomes of two HLB-tolerant Poncirus genotypes and were also upregulated in HLB-tolerant, Poncirus hybrids as revealed by real-time PCR analysis. These two CDR genes seem to be good candidate genes for future studies of their role in citrus-CLas interactions.
Similar content being viewed by others
Introduction
Citrus (Citrus L.) is a widely grown and nutritious fruit, with oranges, grapefruit, lemons, limes, and tangerines among the well-known citrus varieties worldwide.1 China, Brazil, United States (U.S.), India, Mexico, and Spain are the world's leading citrus fruit-producing countries, representing close to two-thirds of the global citrus production.2 In the U.S., a total of 9.02 million metric tons of citrus production was reported in the 2014–2015 season, with Florida as the leading citrus-producing state (56%), followed by California (41%), Texas, and Arizona.3
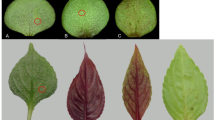
Huanglongbing (HLB) has become the most serious recent disease threat to the U.S. citrus industry.4,5 HLB was first reported in Florida in 2005 and has caused a cumulative loss of more than $2.9 billion in grower revenues between 2006–2007 and 2013–2014 (Florida Citrus Commission; http://www.floridacitrus.org/). HLB resulted in an average annual loss of more than $975 million in the Florida citrus industry output. HLB, also known as citrus greening, is presumably caused by the phloem-limited, gram-negative bacterium Candidatus Liberibacter asiaticus (CLas).6 Three Liberibacter species are known to cause HLB; among them, CLas is the most widespread species. While CLas is heat-tolerant, the African and American forms of Liberibacter are heat-sensitive. Ca. L. africanus is presently primarily distributed in Africa,7 and Ca. L. americanus has been found only in Brazil.8 The bacterium CLas is vectored by the Asian citrus psyllid (ACP; Diaphorina citri).9 ACPs also damage citrus directly by feeding on new citrus growth, resulting in twisted and curled leaves and shoots.10 Typical symptom of HLB in citrus is the asymmetrical pattern of blotchy yellowing or mottling on leaves. As the disease progresses, citrus fruit becomes smaller, and juice quality declines. Diseased mature fruit may remain partially green, which is why HLB is also called citrus greening. The fruit can become lopsided with dark aborted seeds.11,12 At present, there are no effective control methods for HLB. CLas attacks all important commercial varieties of citrus, including oranges, grapefruit, lemons, and tangerines. Sweet oranges, grapefruit, and mandarins are highly susceptible to HLB.4,13,14
Several studies have indicated that Poncirus trifoliata (L.) Raf., a close relative of Citrus, and some of its hybrids are tolerant to HLB.15–18 Poncirus trifoliata is sexually compatible with Citrus and has been widely used as a breeding parent in developing new citrus rootstock cultivars.19,20 It carries valuable genes for improving the resistance of citrus scion cultivars to biotic (for example, citrus tristeza virus) and abiotic stresses.21,22 In recent studies, several previously released Poncirus and Citrus hybrids, such as ‘US-897’ (C. reticulata L. Blanco Cleopatra × P. trifoliata Flying Dragon) and ‘US-942’ (C. reticulata Sunki×P. trifoliata Flying Dragon), have proven to be HLB-tolerant, even though these hybrids were not initially selected for HLB tolerance.15–17 It is expected that P. trifoliata will become a major source of valuable genes for use in sexual hybridization, somatic hybridization, and cisgenic approaches to improving Citrus toward resistance to HLB.20
Creating HLB-resistant/tolerant citrus cultivars is considered the most economic, environment-friendly approach for long-term management of HLB.23 However, it is very challenging to achieve this goal solely via conventional breeding due to the complex reproductive biology and long periods of juvenility.24 As has been demonstrated in major agronomic crops, integrating genomics, transcriptomics, proteomics, and metabolomics can facilitate the identification of differentially expressed genes as candidates for genetic transformation and for use in marker-assisted early selection of disease-resistant progeny in conventional breeding, and can provide much deeper insight about the complex interactions between the host plant citrus and the pathogen CLas. Mining large datasets from whole genome sequencing and transcriptome sequencing projects is an invaluable tool that can assist in the identification of plant resistance and defense-related genes. Several reports had been published on the identification of HLB-responsive genes in citrus using transcriptomic, proteomic, and metabolomic approaches.25–30 Recently, a meta-analysis and a gene-network co-expression analysis were conducted with published reports, suggesting constitutive disease resistance 1 genes (CDR1) as potential candidate genes for HLB resistance/tolerance in Poncirus.15,31,32 CDR1 was first identified and cloned in Arabidopsis. Its product has been implicated in disease resistance signaling.33 Overexpression of a rice (Oryza sativa L.) CDR1 gene, OsCDR1, led to constitutive activation of defense response and enhanced resistance in rice and Arabidopsis against bacterial and fungal pathogens.34
CDR genes belong to the plant aspartic proteinase (APs) gene family. Plant APs are widely distributed in the plant kingdom,35 and the majority of plant APs identified so far are translated as single-chain preproenzymes, which are subsequently converted to mature enzymes with single or two chains.36 Plant APs have a ‘typical’ precursor of ~100 amino acids in the protein, also known as the plant-specific insert, which is removed upon activation into the mature form of the enzymes.37 Plant APs are classified into three groups: typical APs, nucellin-like APs, and atypical APs.38 CDRs are apoplastic enzymes which belong to the ‘atypical’ plant APs. Atypical APs do not contain plant-specific inserts and display intermediate features between the typical and nucellin-like sequences.38 Thus far, only a small number of atypical and nucellin-like APs have been studied, such as nucellin and PCS1 involved in cell death regulation,39 CND41 involved in nitrogen remobilization,40 and CDR1 involved in disease resistance.33,34 Recently PvNod41, an AP from common bean (Phaseolus vulgaris) and closely related to CDR1, has been identified as a major player during plant defense and nodule development.41
The potential roles of CDR1 genes in HLB resistance/tolerance were first recognized in a microarray-based genome-wide gene expression study.15 In a subsequent study comparing CDR1 expression in CLas or citrus tristeza virus infection,17 one CDR1 ortholog was constitutively expressed at much higher levels in the HLB-tolerant Poncirus hybrids ‘US-897’ and ‘US-942’ than in HLB-susceptible Cleopatra mandarin (C. reticulata), while a second form of CDR1 was induced in Cleopatra after CLas infection to a level similar to that found in non-inoculated ‘US-897’ and ‘US-942’.
The present study is focused on genome-wide identification and analysis of CDR genes in Citrus and Poncirus genome and transcriptome sequences. The genomic organization, phylogenetic relationship, protein domain architecture and exon/intron junctions of the identified citrus CDR genes were comprehensively analyzed. Given the critical roles of plant CDR genes in defense, CDR gene variants in HLB-susceptible and HLB-tolerant Citrus and Poncirus genotypes are of great interest. Publicly available genome sequences are only available for HLB-susceptible sweet orange (C. sinensis) and Clementine mandarin (C. ×clementina), therefore we resequenced the genome of two P. trifoliata accessions (DPI 50-7 and Flying Dragon (FD)) and two Poncirus hybrid cultivars ‘US-897’ and ‘US-812’, sequenced the leaf transcriptome of the two P. trifoliata accessions DPI 50-7 and Flying Dragon, and identified the genetic variation in Citrus and Poncirus CDR genes. We analyzed the expression of the entire CDR gene family in HLB-susceptible and HLB-tolerant genotypes after CLas infection and obtained information about the response of each copy of CDR genes to HLB.
Materials and methods
Identification of CDR genes in Citrus genomes
We created an in-house MySQL database of the two citrus genomes: Citrus ×clementina v 1.0 and Citrus sinensis at https://phytozome.jgi.doe.gov/. CDR genes were identified using keyword searches such as ‘constitutive disease resistance’ and ‘aspartic protease family protein’, and the identified CDR gene sequences were retrieved. BLASTP (http://blast.ncbi.nlm.nih.gov/Blast.cgi) was conducted to identify CDR gene family members using the amino-acid sequence of previously characterized Arabidopsis CDR1 (NP_198319).33 We also downloaded the Poncirus EST unigene database (https://www.citrusgenomedb.org/species/trifoliata/unigene1.0) and searched for CDR gene-related ESTs using BLASTP. CDR genes with BLASTP search score values⩾100 and e-value⩽e -10 were used for further analysis. The aspartic protease domain (ASP) (PF00026) was downloaded from the Pfam database (http://pfam.sanger.ac.uk/). To exclude overlapping genes, all putative CDR genes were aligned using ClustalW.42 The ASP domain was then manually checked for each identified CDR gene.
CDR sequence alignment and phylogenetic analysis
The structural divergence among the identified CDR genes was examined by investigating the conserved motif in the encoded CDR proteins. The complete amino-acid sequences of CDR genes were subjected to MEME analysis online (http://meme.nbcr.net/meme). The subcellular localization of CDR proteins was predicted by subCELlular LOcalization predictor (CELLO v.2.5; http://cello.Life.nctu.edu.tw/). CDR genes were aligned using ClustalW.42 Phylogenetic trees were constructed with the PhyML software plugin in Geneious software (http://www.geneious.com/) using the neighbor-joining (NJ) method, and the bootstrap test was replicated 1000 times.
Domain and gene structure analysis
The Pfam domain and signal peptide of CDR genes were predicted using the conserved domain database (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) with default settings. The results were confirmed using CDART search (http://www.ncbi.nlm.nih.gov/Structure/lexington/lexington.cgi). Diagrams of protein structures were constructed with the DOG 1.0 software (http://dog.biocuckoo.org/). The cDNA and DNA sequences of CDR genes were retrieved from the citrus genome databases (see above) and diagrams of exon/intron structures of CDR genes were illustrated using the online Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/). Physical locations (scaffold) of these CDR genes in the citrus genomes were identified.
Plant material, DNA and RNA isolation and HiSeq sequencing
Two HLB-tolerant P. trifoliata accessions DPI 50-7 and Flying Dragon were selected for whole genome resequencing and leaf transcriptome sequencing. The accession Flying Dragon was the source of HLB tolerance in Poncirus hybrids including ‘US-897’ and ‘US-942’.15–17 The accession DPI 50-7 (previously labeled as DPI 9-6) is similar to Flying Dragon in HLB tolerance, but it is morphologically and isozymically distinct from Flying Dragon.21 Two HLB-tolerant Poncirus hybrids ‘US-897’ and ‘US-812’ (C. reticulata Sunki×P. trifoliata Benecke) were also selected for whole genome resequencing. Leaf samples were collected from two different sources. The first source of plants was grown under disease-free conditions in the greenhouse at the Chiefland Budwood Foundation, Bureau of Citrus Budwood Registration, Division of Plant Industry, Florida Department of Agriculture and Consumer Services, Chiefland, Florida. The second source of DPI 50-7 and Flying Dragon plants was grown in the orchard at the University of Florida’s Citrus Research and Education Center in Lake Alfred, Florida, and had been exposed to CLas through natural psyllid transmission. Leaves from both plant sources were collected in April 2015. Leaf tissues were immediately frozen in liquid nitrogen and stored at −80 °C. Genomic DNA was extracted from the leaves using the CTAB method43 and treated with RNase I (Qiagen, Hilden, Germany). The isolated DNA was shipped to Novogene Corporation (Beijing, China) where the DNA was fragmented and sequenced on an Illumina HiSeq 2500 platform (Illumina, Inc., San Diego, CA, USA) following Illumina protocols, to produce a paired-end library of sequence reads for each sample. Total RNA was isolated from leaves using the RNeasy plant isolation kit (Qiagen, Germany). RNA samples were shipped to Novogene Corporation where mRNA were enriched and converted into cDNAs, which were sequenced on an Illumina HiSeq 2500 using the Illumina ‘TruSeq RNA-Seq Sample Prep kit.’
Identification of CDR genes in Poncirus genomes and variant calling in resequenced genomic and transcriptomic sequencing data
Illumina shotgun DNA sequence reads were mapped to the annotated citrus genome assembly C. ×clementina v1.0 using Burrows-Wheeler Aligner (BWA) v0.7.12 software (for resequenced genomes) (bio-bwa.sourceforge.net; La Jolla, CA, USA) and TopHat v2.1.0 (for transcriptomes), respectively. Prior to read mapping, sequence reads were trimmed to remove sequence tags and filtered for low-quality reads based on the distribution of Phred-like scores in each sequencing cycle using trim galore (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Sequence reads of the four resequenced genomes were separately mapped to the annotated reference citrus genome assembly using BWA-mem software with default parameters.44 TopHat v2.1.0 (ref. 45) was run on RNA-Seq reads with default parameters; a citrus gene model annotation file in the GFF3 format was used to enable Bowtie2 v2.2.8 to first align transcript sequences to the transcriptome and then to map only unmapped reads to the genome. Single-nucleotide polymorphisms (SNP) in the assembled reads (DNA-Seq and RNA-Seq) were identified using SAMtools46 and Geneious software, with a minimum Phred quality score of 20. DNA resequencing data are available in the NCBI Sequence Read Archive repository (Acc. #SRP096286).
Identification of CDR-related ESTs from the Poncirus EST database
Poncirus ESTs were downloaded from the citrus genome database (https://www.citrusgenomedb.org/species/trifoliata/unigene1.0). The Arabidopsis CDR1 gene sequence was used in BLASTP searches against the EST database to identify CDR genes-related ESTs with an E value<1e−10. Poncirus ESTs (contigs) matching with CDR genes were manually checked for the presence of the ASP domain. RNA-Seq reads of DPI 50-7 and Flying Dragon were mapped to the identified CDR-related contigs using the TopHat (v2.1.0) and Bowtie2 (2.2.8) programs with default parameters.47
Estimation of the expression level of CDR genes in RNA-Seq data
The expression level and transcript abundance of CDR genes in the Poncirus transcriptomes were determined with Cufflinks v2.0.2. Changes in the abundance of CDR transcripts between two sources of Poncirus samples were estimated using the Cuffdiff program. FPKM (Fragments per Kilobase of exon per Million fragments mapped) were calculated for each CDR gene, but only transcripts showing an FPKM>1 were reported (Supplementary Table 1).
RNA isolation and real-time PCR
For real-time PCR analysis of CLas infected and non-infected plants, three plants each of greenhouse-grown two year-old seedlings of six genotypes were used: ‘US-812’, ‘US-897’, and ‘US-942’ (HLB-tolerant), and Valencia sweet orange (C. sinensis), Duncan grapefruit (C. paradisi), and Ruby Red grapefruit (C. paradisi) (HLB-susceptible). Plants were inoculated in June 2013, by grafting with CLas-infected budwood and were maintained in the U.S. Horticultural Research Laboratory greenhouses in Fort Pierce, Florida. The inoculum source used for CLas infection was tested to be negative for citrus tristeza virus (CTV) infection. Infection with CLas was verified via PCR analysis as described.29 Total RNA was isolated from leaves using the RNeasy plant kit (Qiagen). Total RNA was treated with DNase to remove potential residual DNA and converted to cDNA using the Superscript III kit (Invitrogen, Carlsbad, CA, USA). We designed 16 sets of gene-specific primers (Table 1) and also used the previously reported CDR primers15 for PCR amplification which we named UaCDR1 and UaCDR2 in the present study. Real-time PCR was carried out using the AriaMx system (Agilent Technologies, Santa Clara, CA, USA) and the SYBR green chemistry. Relative gene expression levels were calculated using the 2−ΔΔCT method.48 All real-time PCR experiments were conducted with three biological and two technical replicates. Each reaction was carried out in triplicate with a reaction volume of 20 μL containing 0.5 μL (10 μm) of each gene-specific primers (Table 1), 1.0 μl of cDNA (20 ng/μl), 10 μl of SYBR green (1X buffer), and 8 μl autoclaved deionized water. The PCR cycling parameters were 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s.
Results
Identification of 17 CDR genes in Citrus genomes
A total of 17 CDR genes were identified in two Citrus genomes: nine from the C. ×clementina genome and eight from the C. sinensis genome. The nomenclature system for CDR genes in the present study provisionally uses the names CcCDR1 to CcCDR9 for C. ×clementina CDR genes and CsCDR1 to CsCDR8 for C. sinensis CDR genes, to distinguish each of the CDR genes based on the homology between the citrus databases. The predicted CDR proteins varied in length from 185 (CcCDR2) to 480 (CcCDR1 and CsCDR1) amino acids. The predicted relative molecular mass varied from 20.4 kDa (CcCDR2) to 51.7 kDa (CcCDR1 and CsCDR1), and the pIs ranged from 4.67 (CcCDR8) to 9.71 (CcCDR1), with eight members having a pI<7 and nine CDR members having a pI>7 (Supplementary Table 2). Pair-wise analysis of predicted CDR protein sequences indicated that the overall amino-acid identity was 53.4%.
Phylogenetic and physical relationships among Citrus CDR genes
The 17 CDR genes were grouped into five major clades, clade I, II, III, IV and V, with well-supported bootstrap values (Figure 1a). Clade I and II had two members, one from C. ×clementina, and one from C. sinensis. Four members were clustered into clade III, three genes from C. ×clementina and one from C. sinensis. Three members were present in clade IV, with two genes from C. ×clementina and one from C. sinensis. Six members fell into clade V, two genes from C. ×clementina and four from C. sinensis. Fifteen of the predicted CDR proteins would be localized in the outer membrane of citrus cells while the predicted CcCDR8 and CcCDR9 proteins would be present extracellularly (Supplementary Table 3).
Phylogenetic analysis, distribution of conserved domains, and gene structure of Citrus CDR genes. (a) Neighbor-joining tree was created using PhyML software with 1000 bootstrap after aligning conserved ASP domain of CDR genes with ClustalW. A total of nine CDR genes from C. ×clementina (CcCDR1- CcCDR9) and eight CDR genes from C. sinensis genome (CsCDR1- CsCDR8) were used to construct the tree. Names in the bracket represent respective gene IDs from Citrus genome databases. Five different clades were color labeled. Number above and below branches indicate bootstrap values. (b) Conserved domains in the deduced protein sequences from CcCDR (C.×clementina) and CsCDR (C. sinensis) genes. The relative positions of each domain within each protein are marked with different colors. (c) The structures of CcCDR genes are presented from C.×clementina (light blue) from C. sinensis (dark blue). Gene lengths are displayed proportionally. The blue regions represent exons. Interrupted black lines are introns and red regions are UTRs.
Three CDR genes (CcCDR2, CcCDR3, and CcCDR4) are located within 50 kb on Scaffold 3 in the C.×clementina genome assembly. The predicted protein structure of CcCDR3 and CcCDR4 is almost identical. CcCDR5 and CcCDR6 are present at the same genomic scaffold location and have similar predicted protein structures, indicating a likely tandem duplication event. CsCDR3, CsCDR5 and CsCDR6 in the C. sinensis genome represent different alternate splicing isoforms of the CsCDR4.
Domains and structures of Citrus CDR genes
CDR proteins are characterized by the following domains: a signal peptide, a propeptide, and an ASP domain with two active sites.49 All 17 predicted citrus CDR proteins contain at least one ASP domain (Figure 1b). Except for CcCDR2, CcCDR8, CcCDR9, CsCDR2 and CsCDR8, 12 CDR proteins each contain a signal peptide of 24 amino acids. In addition, the 12 CDR proteins each carry one low complexity domain. All 17 CDR proteins contain a TAXI_N and TAXI_C domain along with the ASP domain.
The divergence of exon/intron structure often plays a key role in the evolution of gene families.50 To understand the structural components of Citrus CDR genes, their exon and intron organization were inferred by comparing the cDNA sequences with the corresponding genomic DNA sequences (Figure 1c). Among the nine CcCDR genes, one gene (CcCDR8) contains one intron, and five genes (CcCDR1, CcCDR3, CcCDR4, CcCDR6 and CcCDR7) have either 5’ or 3’ or both the untranslated regions (UTR) present in their transcripts. Among eight CsCDR genes, one gene (CsCDR4) has three transcription isoforms, three genes (CsCDR3, CsCDR5 and CsCDR6) each contains one or two introns and both 5′ and 3′ UTRs. The protein structures of these four CsCDR genes (CsCDR3-6) are highly similar. The other four CsCDR genes (CsCDR1-2 and CsCDR7-8) have no UTRs in their respective transcripts. CsCDR2 contained the largest number of introns (four introns) among all the CDR genes in this study.
CDR genes and SNPs in resequenced Poncirus genomes
We obtained 21 gigabases (Gb) and 10.9 Gb of clean sequence data for Poncirus accessions DPI 50-7 and Flying Dragon, respectively, which represent 55× and 25× depth of coverage of their genomes (Table 2). A total of 5411 sequence reads of DPI 50-7 and 2665 sequence reads of Flying Dragon matched to 15 Citrus CDR genes. We identified eight copies of CDR genes (named as PtCDR1 to PtCDR8) in the resequenced Poncirus samples. SNPs in the PtCDR genes were called with a read-depth level of 3 to 50 and a Phred quality score>20. The type of SNP (transition, transversion, substitution, deletion, and insertion) and location of the SNPs (5′-UTR, exon, intron and 3′-UTR) were characterized for the eight PtCDR genes (Supplementary Table 4). Most of deletions and insertions were present in the UTR regions. The ratio of transition/transversion was>1 for four PtCDR genes (PtCDR1, PtCDR2, PtCDR6 and PtCDR7), suggesting the presence of larger numbers of non-synonymous SNPs (nsSNPs) in these genes. The predicted proteins of all PtCDR genes contain the ASP domain. Poncirus CDR genes were clustered with their Citrus CDR orthologs (Figure 2).
Phylogenetic tree of CDR genes of Citrus and Poncirus. The phylogenetic tree was created using eight Poncirus CDR genes (PtCDR1-PtCDR8), Arabidopsis CDR1 gene and 17 Citrus CDR genes. The ASP domain was first aligned with ClustalW and a neighbor-joining tree was created using PHYML software with 1000 bootstrap value.
There are various numbers of amino-acid (aa) changes in PtCDR genes: PtCDR1 (4 aa), PtCDR2 (27 aa), PtCDR3 (21 aa), PtCDR4 (21 aa), PtCDR5 (19 aa), PtCDR6 (5 aa), PtCDR7 (7 aa), and PtCDR8 (16 aa). We observed insertion/deletion of amino acids in four PtCDR genes: PtCDR2 (insertion of 4 aa), PtCDR4 (deletion of 1 aa), PtCDR6 (insertion of 2 aa), and PtCDR8 (insertion of 7 aa). Overall, substitution and insertion/deletion of amino acids were more abundant in PtCDR2 and PtCDR8 compared to other PtCDR genes.
We obtained ~11 and 13 Gb of clean DNA sequences for the ‘US-897’ and ‘US-812’ genome, respectively, equivalent to an average of 26× depth of coverage per hybrid genotype. A total of 3142 sequence reads from ‘US-897’ and 4335 reads from ‘US-812’ matched to the 17 Citrus CDR genes, respectively. Most of the sequence variants found in ‘US-897’ and ‘US-812’ were heterozygous in nature. The numbers of homozygous SNPs present in these Poncirus hybrids for a given CDR gene were about half of that in DPI 50-7 and Flying Dragon, as would be expected due to the hybrid origin of ‘US-812’ and ‘US-897’.
Consensus DNA sequences were generated from the sequence reads for all PtCDR genes (Supplementary Table 5). ‘US-812’ carries two different alleles (one from Citrus and one from P. trifoliata) at PtCDR1, PtCDR2, PtCDR3, PtCDR4, PtCDR6 and PtCDR8 loci and only one allele from P. trifoliata at PtCDR5 and PtCDR7 loci. On the other hand, ‘US-897’ carries two different alleles at all PtCDR gene loci, except for PtCDR3, at which ‘US-897’ carries only the P. trifoliata allele. The absence of a Citrus allele at the PtCDR5 and PtCDR7 loci in ‘US-812’ and at the PtCDR3 locus in ‘US-897’ may suggest that these loci are hemizygous in ‘US-812’ and ‘US-897’.
CDR genes in Poncirus transcriptomes
We obtained ~20 million sequence reads for each of the four leaf transcriptomes (two Poncirus accessions (DPI 50-7 and Flying Dragon) grown under two different conditions). The FPKM values for each CDR gene were calculated (Supplementary Table 1). Two Poncirus CDR genes, PtCDR1 and PtCDR4, were in lower abundance in CLas-exposed, field-grown DPI 50-7 than in non-inoculated, greenhouse-grown DPI 50-7. PtCDR2 (for DPI 50-7 and Flying Dragon) and PtCDR8 (for DPI-50-7) were in higher abundance in CLas-exposed, field-grown plants than in non-inoculated, greenhouse-grown plants.
CDR-related ESTs from the Poncirus EST database
Out of the 61 874 Poncirus ESTs from the citrus genome database, 10 ESTs (contigs) match with the Arabidopsis CDR genes and contain the ASP domain. Mapping RNA-Seq reads of DPI 50-7 and Flying Dragon with the 10 identified CDR-related contigs allowed for retrieval of 7032 reads from the RNA-Seq data that have 98% identity with the 10 CDR-related contigs. The 10 Poncirus ESTs correspond to two Poncirus CDR genes, PtCDR2 and PtCDR8, indicating that these two genes are expressed in Poncirus leaves.
Real-time PCR amplification of CDR genes in HLB-susceptible and tolerant genotypes
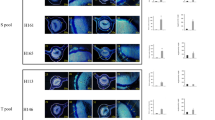
Results for 14 pairs of CDR primers are represented in Figure 3. Expression of most of the CDR genes was higher in CLas infected ‘US-812’ and ‘US-897’ when compared with their respective non-infected samples (Figure 3, Supplementary Table 6). Moreover, CDR transcripts amplified by primer pairs CDR14 (CsCDR3), CDR15 (CsCDR6), CDR16 (CsCDR5/PtCDR2), UaCDR1 and UaCDR2 had higher expression levels in the HLB-tolerant genotypes. PtCDR2 was also found to be upregulated in RNA-Seq data. All upregulated CDR genes belong to clade V of the phylogenetic tree and share similar gene structures (Figure 1a).
Relative expression of CDR genes in HLB-susceptible and HLB-tolerant genotypes. The results of real-time PCR of each CDR gene with three HLB-tolerant (US-812, US-897 and US-942) and three HLB-susceptible (V, DG and RR) genotypes are presented as a heat map. ‘Green’ blocks represent down-regulation while ‘red’ blocks represent up-regulation in CLas-infected plant when compared with its respective non-inoculated plant. Citrus and Poncirus genotypes are represented horizontally and primers used for Real-time PCR analysis are shown vertically.
Discussion
CDR genes in Citrus
Recently, several studies were carried out to identify potential candidate genes in Citrus and Poncirus associated with HLB tolerance, and CDR genes were among those identified.15 In this study, 17 CDR genes were identified in two Citrus genomes, and they fell into five clades. Fifteen of the 17 CDR genes contained no or only one intron. Citrus CDR genes in the same clades share similar motifs. On the basis of the high similarity at both protein and gene structure levels and their locations in the citrus genome, it appears that CcCDR5 and CcCDR6 have arisen from a recent tandem duplication and may have a redundant function in citrus. These two genes have the same gene structure except for the presence of a 57-bp 3′-UTR in CcCDR6. CsCDR3-CsCDR6 were present within the same phylogenetic clade (Figure 1a), and they may represent paralogs. Based on the results presented above, it is clear that the expansion of the Citrus CDR family was the result of segmental and tandem duplication, which could be the result of chromosomal rearrangement and fusions. Similar results were found with the aspartic gene family in rice.51
CDR genes in Poncirus
Genome resequencing and analysis led to the identification of eight CDR genes in Poncirus. Compared to Citrus CDR genes, Poncirus CDR genes carry a multitude of SNPs. Most of the SNPs were present in exonic regions of PtCDR genes, and deletion and insertion-type mutations were prominent in the regulatory and non-coding regions (5′ and 3′-UTR regions). Most of the identified SNPs were of the transition type, and the ratio between transition and transversion was more than one for four PtCDR genes. The degeneracy of the genetic code and the selective pressure for gene conservation are likely accountable for the dominance of transitions over transversions. In general, transition mutations are generated at higher frequency than transversions in synonymous substitutions, and there is a stronger selection against replacement substitutions, leading to higher occurrences of transitions. SNPs in protein-coding sequences have potential effects on gene function, especially the nsSNPs that could lead to amino-acid residue changes and altered functional or structural properties of the protein. Results from the present study indicate that the predicted protein structure and function may be altered in some of the Poncirus CDR genes, particularly in PtCDR2 and PtCDR8, as they contain the largest numbers of amino-acid changes among all Poncirus CDR genes. Li et al.52 analyzed the contributions of genic and non-genic sequence polymorphisms to natural variation of quantitative traits in maize and revealed that 79% of the explained variation could be attributed to SNPs located in genes or within 5 Kb upstream of genes. The SNPs in the genic and promotor regions of Poncirus CDR genes may be useful for genetic dissection of complex traits such as HLB tolerance.
Transcripts of four PtCDR genes were detected in Poncirus leaf transcriptomes. While PtCDR1 and PtCDR4 were in low abundance, PtCDR2 and PtCDR8 showed high abundance in plants grown in the field and exposed to natural HLB disease pressure. Both PtCDR2 and PtCDR8 were grouped in clade V of the phylogenetic tree and have similar gene structures. A total of 70 SNPs were revealed in these four expressed PtCDR genes when compared with their Citrus CDR orthologs. Among the SNPs, 20% were nsSNPs. Most of these nsSNPs are present in regulatory regions. These nsSNPs should be closely examined in future studies of the roles of CDR genes in HLB tolerance.
In a previous study,17 the expression of one CDR probe set (Cit.23704.1.S1_at) was 300-1000 fold higher in ‘US-897’ and ‘US-942’ than in Cleopatra (Citrus reshni hort. Ex Tanaka), regardless of CLas infection. In that study, another CDR probe set (Cit.28117.1.S1_s_at) was found to be expressed similarly in the three cultivars without CLas infection, and increased moderately in all three cultivars through the 30 weeks following infection. An earlier study15 using microarray analysis also found higher expression of these CDR probe sets in ‘US-897’ as compared with Cleopatra, independent of CLas infection. The present study shows that probe sets Cit.23704.1.S1_at and Cit.28117.1.S1_s_at bind to CcCDR3 or PtCDR8 genes with 90% homology. In contrast to the present study, the primers (UaCDR1 and UaCDR2) used to validate these probe sets bind to multiple CDR genes instead of individual CDR genes. We observed that the CDR genes in clade V were highly expressed in HLB-tolerant genotypes compared with HLB-susceptible genotypes. PtCDR2 and PtCDR8 identified with relative high abundance in Poncirus transcriptome sequencing data as well in real-time PCR analysis were clustered with clade V, making these genes very interesting candidates for further functional characterization.
It is worth noting that many CDR genes (including CcCDR1, 7 and 8) showed low expressions when compared with other CDR genes after CLas infection. A possible explanation for higher induction for Clade V CDR genes could be that these genes have characteristic UTR regions in their gene sequences which are not common in genes present in other clades, which in turn, could result in diverse regulatory mechanisms of clade V genes. The expression patterns of CDR genes support our findings that the distribution of specific motifs or specific patterns for a motif in proteins is associated with a specific clade in the phylogram. These motifs may be involved in regulation of HLB-related gene expression.
In summary, we identified 17 CDR genes in two citrus genomes and revealed key features of their gene structure. Separation of these CDR genes into five clades was mutually supported by their phylogeny and exon/intron structure and the distribution of conserved domains. Whole genome resequencing led to the identification of eight CDR genes in Poncirus. PtCDR2 and PtCDR8 were found in high abundance in Poncirus leaf transcriptomes. The expression of PtCDR2 and PtCDR8 genes responded to CLas infection differently in HLB-tolerant and susceptible genotypes. The information gained in this study provides new insights regarding the potential roles of CDR genes in mediating citrus responses to CLas infection and HLB development. Gene sequences and sequence variants from this analysis should be very valuable for functional validation of CDR genes and potentially for engineering HLB resistance/tolerance in new citrus cultivars.
References
Liu Y, Heying E, Tanumihardjo SA . History, global distribution, and nutritional importance of citrus fruits. Compr Rev Food Sci Food Saf 2012; 11: 530–545.
FAO 2009. Press release, 19 June 2009. Available at http://www.fao.org/news/story/en/item/20568/icode/.
United States Department of Agriculture-National Agricultural Statistical Service (USDA-NASS). Citrus Fruits 2015 Summary. USDA: Washington DC. 2015.
Halbert SE, Manjunath KL . Asian citrus psyllids (Sternorrhyncha: Psyllidae) and greening disease of citrus: a literature review and assessment of risk in Florida. Fla Entomol 2004; 87: 330–353.
Bové JM . Huanglongbing: a destructive, newly-emerging, century-old disease of citrus. J Plant Pathol 2006; 88: 7–37.
Gottwald TR, da Graça JV, Bassanezi RB . Citrus Huanglongbing: the pathogen and its impact. Plant Health Prog 2007; 6: 1–18.
Jagoueix S, Bove JM, Garnier M . The phloem-limited bacterium of greening disease of citrus is a member of the α subdivision of the Proteobacteria. Int J Syst Evol Microbiol 1994; 44: 379–386.
Texeira DD, Ayres J, Kitajima EW et al. First report of a Huanglongbing-like disease of citrus in São Paulo State, Brazil and association of a new Liberibacter species, “Candidatus Liberibacter americanus”, with the disease. Plant Dis 2005; 89: 107–107.
Hall DG, Richardson ML, Ammar ED, Halbert SE . Asian citrus psyllid, Diaphorina citri, vector of citrus Huanglongbing disease. Entomol Exp Appl 2013; 146: 207–223.
Lopes SA, Bertolini E, Frare GF et al. Graft transmission efficiencies and multiplication of ‘Candidatus Liberibacter americanus' and 'Ca. Liberibacter asiaticus' in citrus plants. Phytopathology 2009; 99: 301–306.
Da Graça JV, Korsten L . Citrus Huanglongbing: Review, present status and future strategiesIn:. Diseases of Fruits and Vegetables Volume I. Springer: Netherlands. 2004, 229–245.
Gottwald TR . Current epidemiological understanding of citrus Huanglongbing. Annu Rev Phytopathol 2010; 48: 119–139.
Folimonova SY, Robertson CJ, Garnsey SM, Gowda S, Dawson WO . Examination of the responses of different genotypes of citrus to Huanglongbing (citrus greening) under different conditions. Phytopathology 2009; 99: 1346–1354.
Fan J, Chen C, Yu Q et al. Comparative transcriptional and anatomical analyses of tolerant rough lemon and susceptible sweet orange in response to ‘Candidatus Liberibacter asiaticus’ infection. Mol Plant-Microbe Interact 2012; 25: 1396–1407.
Albrecht U, Bowman KD . Transcriptional response of susceptible and tolerant citrus to infection with Candidatus Liberibacter asiaticus. Plant Sci 2012a; 185: 118–130.
Albrecht U, Bowman KD . Tolerance of trifoliate citrus rootstock hybrids to Candidatus Liberibacter asiaticus. Sci Horticult 2012b; 147: 71–80.
Bowman KD, Albrecht U . Comparison of gene expression changes in susceptible, tolerant and resistant hosts in response to infection with Citrus tristeza virus and Huanglongbing. J Citrus Pathol 2015; 2: 1–6.
Ramadugu C, Keremane ML, Halbert SE et al. Long-term field evaluation reveals Huanglongbing resistance in Citrus relatives. Plant Disease 1996; 100: 1858–1869.
Bowman KD, Faulkner L, Kesinger M . New citrus rootstocks released by USDA 2001-2010. HortScience 2016; 51: 1208–1214.
Grosser JW, Gmitter FG, Castle WS . Breeding citrus rootstocks to mitigate Huanglongbing (HLB, or citrus greening disease). Acta Hortic 2016; 1127: 83–88.
Gmitter FG, Xiao SY, Huang S et al. A localized linkage map of the citrus tristeza virus resistance gene region. Theor Appl Genet 1996; 92: 688–695.
Weber CA, Moore GA, Deng Z, Gmitter FG . Mapping freeze tolerance quantitative trait loci in a Citrus grandis×Poncirus trifoliata F1 pseudo-testcross using molecular markers. J Amer Soc Hort Sci 2003; 128: 508–514.
National Research Council. Strategic Planning for the Florida Citrus Industry: Addressing Citrus Greening Disease. The National Academies Press: Washington, DC. 2010.
Gmitter FG Jr, Soneji JR, Rao MN . Citrus breeding. In: Jain SM, Priyadarshan PM (ed.). Breeding Plantation Tree Crops: Temperate Species. Springer Science + Business Media, LLC: Berlin, Heidelberg, Germany. 2009, 105–134.
Albrecht U, Bowman KD . Gene expression in Citrus sinensis (L.) Osbeck following infection with the bacterial pathogen Candidatus Liberibacter asiaticus causing Huanglongbing in Florida. Plant Sci 2008; 175: 291–306.
Kim JS, Sagaram US, Burns JK, Li JL, Wang N . Response of sweet orange (Citrus sinensis) to 'Candidatus Liberibacter asiaticus' infection: microscopy and microarray analyses. Phytopathology 2009; 99: 50–57.
Fan J, Chen C, Brlansky RH, Gmitter FG Jr, Li ZG . Changes in carbohydrate metabolism in Citrus sinensis infected with ‘Candidatus Liberibacter asiaticus’. Plant Pathol 2010; 59: 1037–1043.
Fan J, Chen C, Yu Q, Brlansky RH, Li ZG, Gmitter FG . Comparative iTRAQ proteome and transcriptome analyses of sweet orange infected by ‘Candidatus Liberibacter asiaticus’. Physiol Plant 2011; 143: 235–245.
Albrecht U, Fiehn O, Bowman KD . Metabolic variations in different citrus rootstock cultivars associated with different responses to Huanglongbing. Plant Physiol Biochem 2016; 107: 33–44.
Aritua V, Achor D, Gmitter FG, Albrigo G, Wang N . Transcriptional and microscopic analyses of citrus stem and root responses to Candidatus Liberibacter asiaticus infection. PloS ONE 2013; 8: e73742.
Du D, Rawat N, Deng Z, Gmitter FG Jr . Construction of citrus gene coexpression networks from microarray data using random matrix theory. Horticult Res 2015; 2: 15026.
Rawat N, Kiran SP, Du D, Gmitter FG, Deng Z . Comprehensive meta-analysis, co-expression, and miRNA nested network analysis identifies gene candidates in citrus against Huanglongbing disease. BMC Plant Biol 2015; 15: 1.
Xia Y, Suzuki H, Borevitz J et al. An extracellular aspartic protease functions in Arabidopsis disease resistance signaling. EMBO J 2004; 23: 980–988.
Prasad BD, Creissen G, Lamb C, Chattoo BB . Overexpression of rice (Oryza sativa L.) OsCDR1 leads to constitutive activation of defense responses in rice and Arabidopsis. Mol. Plant-Microbe Interact 2009; 22: 1635–1644.
Rawlings ND, Barrett AJ . MEROPS: the peptidase database. Nuc Acids Res 1999; 27: 325–331.
Simões I, Faro C . Structure and function of plant aspartic proteinases. Eur J Biochem 2004; 271: 2067–2075.
Mutlu A, Gal S . Plant aspartic proteinases: enzymes on the way to a function. Physiol Plant 1999; 105: 569–576.
Faro C, Gal S . Aspartic proteinase content of the Arabidopsis genome. Curr Protein Pept Sci 2005; 6: 493–500.
Ge X, Dietrich C, Matsuno M et al. An Arabidopsis aspartic protease functions as an anti‐cell‐death component in reproduction and embryogenesis. EMBO Rep 2005; 6: 282–288.
Kato Y, Murakami S, Yamamoto Y et al. The DNA-binding protease, CND41, and the degradation of ribulose-1, 5-bisphosphate carboxylase/oxygenase in senescent leaves of tobacco. Planta 2004; 220: 97–104.
Olivares JE, Díaz-Camino C, Estrada-Navarrete G et al. Nodulin 41, a novel late nodulin of common bean with peptidase activity. BMC Plant Biol 2011; 11: 134.
Thompson JD, Higgins DG, Gibson TJ . CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 1994; 22: 4673–4680.
Cheng YJ, Guo WW, Yi HL, Pang XM, Deng X . An efficient protocol for genomic DNA extraction from Citrus species. Plant Mol Biol Rep 2003; 21: 177–178.
Li H, Durbin R . Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Trapnell C, Pachter L, Salzberg SL . TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 2009; 25: 1105–1111.
Li H . A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011; 27: 2987–2993.
Langmead B, Trapnell C, Pop M, Salzberg SL . Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 2009; 10: R25.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 2001; 25: 402–408.
Barrett AJ, Woessner JF, Rawlings ND . Handbook of Proteolytic Enzymes Vol. 1. Elsevier: Amsterdam, Netherlands, 2012.
Babenko VN, Rogozin IB, Mekhedov SL, Koonin EV . Prevalence of intron gain over intron loss in the evolution of paralogous gene families. Nuc Acids Res 2004; 32: 3724–3733.
Chen J, Ouyang Y, Wang L, Xie W, Zhang Q . Aspartic proteases gene family in rice: gene structure and expression, predicted protein features and phylogenetic relation. Gene 2009; 442: 108–118.
Li X, Zhu C, Yeh C-T et al. Genic and nongenic contributions to natural variation of quantitative traits in maize. Genome Res 2012; 22: 2436–2444.
Acknowledgements
We thank Ravi Pratap Singh (MySQL, Teradata database administrator) for writing custom design scripts helping in RNA-Seq and DNA-Seq data analysis pipelines. We also thank Benjamin Rosson (Operations and Management Consultant-Manager, Bureau of Citrus Budwood Registration, Division of Plant Industry, Florida Department of Agricultural and Consumer Services) and Michael Kesinger (Chief, Bureau of Citrus Budwood Registration, Division of Plant Industry, Florida Department of Agriculture and Consumer Services) for providing Poncirus (DPI-5-7 and Flying Dragon) leaf samples. ZD acknowledges financial support of this study from the Citrus Research and Development Foundation, Inc. (CDRF) (Project #108766 and #105077) and from the USDA-NIFA Citrus Disease Research and Extension (CDRE) Program (Grant No. 2015-70016-23027).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information for this article can be found on the Horticulture Research website
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Rawat, N., Kumar, B., Albrecht, U. et al. Genome resequencing and transcriptome profiling reveal structural diversity and expression patterns of constitutive disease resistance genes in Huanglongbing-tolerant Poncirus trifoliata and its hybrids. Hortic Res 4, 17064 (2017). https://doi.org/10.1038/hortres.2017.64
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2017.64
This article is cited by
-
Transcriptome-wide Identification of CDR Family in Citrus Latifolia and its Expression During HLB Disease
Tropical Plant Biology (2023)
-
Whole genome re-sequencing of sweet cherry (Prunus avium L.) yields insights into genomic diversity of a fruit species
Horticulture Research (2020)
-
Wide-ranging transcriptomic analysis of Poncirus trifoliata, Citrus sunki, Citrus sinensis and contrasting hybrids reveals HLB tolerance mechanisms
Scientific Reports (2020)
-
Comparative transcriptome analysis reveals synergistic and disparate defense pathways in the leaves and roots of trifoliate orange (Poncirus trifoliata) autotetraploids with enhanced salt tolerance
Horticulture Research (2020)