Abstract
The basic helix-loop-helix (bHLH) transcription factors (TFs) play important roles in regulating multiple biological processes in plants. However, there are few reports about the function of bHLHs in flower senescence. In this study, a bHLH TF, PhFBH4, was found to be dramatically upregulated during flower senescence. Transcription of PhFBH4 is induced by plant hormones and abiotic stress treatments. Silencing of PhFBH4 using virus-induced gene silencing or an antisense approach extended flower longevity, while transgenic petunia flowers with an overexpression construct showed a reduction in flower lifespan. Abundance of transcripts of senescence-related genes (SAG12, SAG29) was significantly changed in petunia PhFBH4 transgenic flowers. Furthermore, silencing or overexpression of PhFBH4 reduced or increased, respectively, transcript abundances of important ethylene biosynthesis-related genes, ACS1 and ACO1, thereby influencing ethylene production. An electrophoretic mobility shift assay showed that the PhFBH4 protein physically interacted with the G-box cis-element in the promoter of ACS1, suggesting that ACS1 was a direct target of the PhFBH4 protein. In addition, ectopic expression of this gene altered plant development including plant height, internode length, and size of leaves and flowers, accompanied by alteration of transcript abundance of the gibberellin biosynthesis-related gene GA2OX3. Our results indicate that PhFBH4 plays an important role in regulating plant growth and development through modulating the ethylene biosynthesis pathway.
Similar content being viewed by others
Introduction
Flower senescence is an important coordinated process regulated by internal and environmental changes.1 Microarray studies of gene expression have been used to generate genome-wide transcriptome profiles of senescing petals in Arabidopsis.2 The data revealed that hundreds of upregulated and downregulated genes, including various transcription factors (TFs), are involved in flower senescence progress. TFs play critical roles in plant growth and development. The most represented families amongst TFs specifically upregulated during petal senescence in Arabidopsis were AP2-EREBP, homeobox (HB), and AUX-IAA.2 The upregulation of the AP2-EREB TFs establishes the role of ethylene in Arabidopsis. In petunia flowers, expression profiles of the ethylene-responsive element-binding factor (ERF) family genes were studied in detail.3 Some of ERFs appear to be associated with fruit ripening and with corolla senescence.4–7 Among the HB TFs upregulated in Arabidopsis petals was KNAT1, a member of the Class I KNOX family known to modulate cytokinin levels.8 The expression of genes encoding AUX-IAA proteins in Arabidopsis suggests the role of auxin in petal senescence. Nevertheless, exact roles of these regulatory elements in the process of flower senescence are still largely unknown.
Petunia is an ideal model system for studies of flower senescence due to its short life cycle, vast genetic resources, and large number of amenities for biochemical and molecular analysis.9 We identified a cluster of genes upregulated during development and senescence of petunia flowers, including several transcription factors.10,11 In addition, we have successfully employed tobacco rattle virus (TRV)-based virus-induced gene silencing (VIGS) to study the function of senescence-related genes in petunia corollas.12,13 By microarray analysis, we previously determined that a homeodomain-leucine-zipper TF, PhHD-ZIP was upregulated during flower senescence.10 Silencing PhHD-Zip using VIGS resulted in extended flower longevity. Transcript abundances of ethylene biosynthesis-related genes and ethylene production were dramatically reduced in the PhHD-Zip-silenced flowers. On the other hand, overexpression of PhHD-Zip in petunia caused early flower senescence. Furthermore, PhHD-Zip transcript levels in petunia flower were increased by hormones (ethylene, ABA) and abiotic stresses (dehydration, NaCl, and cold). The results suggest that PhHD-Zip plays an important role in regulating petunia flower senescence.11 From the same microarray study,10 a basic helix-loop-helix (bHLH) TF was also found to be highly expressed in flower petals and upregulated during the flower senescence process, suggesting that it may also play a role in the regulation of flower senescence.
The bHLH TFs constitute a large family of regulatory proteins found in plants and animals. One hundred and sixty two genes in Arabidopsis and 167 genes in rice have been predicted to encode bHLHs.14–16 These proteins have a characteristic, highly conserved bHLH domain and comprise two functionally distinct regions comprising approximately 60 amino acids.17 The basic region, located at the N-terminal end of the domain, consists of 13–17 primarily basic amino acids and is involved in DNA binding. The HLH region, at the C-terminal end, is rich in hydrophobic residues and contributes to the formation of homodimers or heterodimers.18,19 Outside the conserved bHLH domain, these proteins exhibit considerable sequence divergence.20 So far, functional analysis shows that bHLH proteins play important roles in fruit dehiscence, epidermal cell development, flavonoid biosynthesis, phytochrome signaling, plant hormone signaling, and biotic/abiotic stress responses.18,21–26 For example, FLOWERING bHLH 1 (FBH1), FBH2, FBH3, and FBH4 bind to the E-box cis-elements in the CONSTANS (CO) promoter. Overexpression of all FBH genes increased CO expression levels and resulted in early flowering.27 However, there has previously been no report of a function for bHLH TFs in flower senescence.
Here, we report the functional characterization of a bHLH TF, PhFBH4. Petunia plants in which bHLH expression was downregulated by VIGS or antisense silencing showed extended flower longevity while overexpression of the PhFBH4 in petunia resulted in earlier flower senescence. These data suggest an important role for PhFBH4 in the control of flower senescence.
Materials and methods
Plant material
Petunia hybrida cv. ‘Primetime Blue’ was used for VIGS experiments. Four-week-old seedlings were transferred to pots and used for VIGS inoculation. After inoculation, the seedlings were placed in a growth chamber with a day (16 h)/night (8 h) cycle and temperature regime of 25/20 °C. Petunia hybrida cv. ‘Mitchell diploid’ was used for stable transformation. Wild-type (WT) and transgenic plants were grown under standard greenhouse conditions.
Hormone and abiotic stress treatments
Hormone and abiotic stress treatments were carried out as described in detail by Chang et al.11 Briefly, the whole flowers were harvested from the plant prior to anther dehiscence. For each treatment, at least three individual flowers were used. The harvested flowers were immediately placed in 2-ml microtubes with or without water, 0.1 mM ABA, 50 µM GA3, or 100 mM NaCl at room temperature. For low and high temperature treatments, harvested flowers were placed in the tubes with water and treated in a 4 °C cold room or in a 29 °C room. For ethylene treatment, flowers were placed in tubes with water, then sealed in a large glass container and treated with 3 µl/l ethylene. Corollas were collected at 0, 3, 6, and 12 h after treatment. For 1-methylcyclopropene (1-MCP) treatment, flowers were first treated with 50 nl/l 1-MCP for 4 h, and then exposed to 3 µl/l ethylene. Corollas were collected at 0, 3, 6, and 12 h after the ethylene treatment. Harvested tissues were immediately frozen in liquid nitrogen, and then kept at –80 °C until RNA extraction.
VIGS vector construction and petunia inoculation
The chalcone synthase (CHS)/TRV2 construct was generated previously.28 To generate the PhFBH4/CHS/TRV2 construct, a 326 bp DNA fragment was amplified from petunia cDNA with gene-specific primers (Supplementary Table S1) and then cloned into the CHS/TRV2 vector as described previously by Chen et al.12 and Chang et al.11 Agrobacterium tumefaciens strain GV3101 was transformed with the VIGS constructs. The presence of the gene fragments in the bacteria was confirmed by polymerase chain reaction (PCR) with gene-specific primers.
Four-week-old plants (when the first 4 true leaves had emerged) were used for VIGS inoculation following the protocol that has been previously described by Jiang et al.13
Plasmid construction and stable transformation of petunias
To generate PhFBH4 silencing transgenic plants, a 281 bp fragment was cloned into the BamHΙ and SpeΙ site of the pGSA1403 vector in the antisense orientation. To generate PhFBH4 overexpression transgenic plants, a 1056 bp DNA sequence containing the ORF region of PhFBH4 gene was cloned into the BamHΙ and SpeΙ site of the pGSA1403 vector in the sense orientation. The constructs were transformed into A. tumefaciens strain LBA4404 using electroporation. PCR amplification was performed to confirm the destination vectors had integrated with the PhFBH4 gene.
Transformation and regeneration of petunia ‘Mitchell Diploid’ was performed using the leaf disk method and cultivation process described previously by Wang et al.10 At least 12 independent transgenic lines were generated. Homozygous lines from T2 or later generations were used for later experiments.
Flower longevity
Flower longevity was recorded as the time from anthesis until the corolla was completely wilted.11 At least 10 flowers from different plants were used for longevity studies. Data were statistically analyzed using JMP10.0 software package (SAS Institute, Cary, NC, USA).
RNA extraction, semi-quantitative, and quantitative RT-PCR
Petals of wild–type (WT) and transgenic flowers under the different treatments were collected. Total RNA was extracted from petunia corollas using Trizol Reagent according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA, USA). RNA was treated with RNase-free DNase (Promega, Madison, WI, USA) to remove DNA contamination according to the manufacturer’s instructions. Two micrograms of total RNA was used to synthesize first-strand cDNA using the SuperScript III kit (Invitrogen). Semi-quantitative RT-PCR was performed in 25 μl reactions containing 0.3 μM of each primer and 1 μl of template cDNA. Amplification by qPCR was performed in 25 μl reactions containing 1 μl cDNA, 0.3 μM of each primer, and 12.5 μl of SYBR Green PCR Master Mix (Toyobo, New York, NY, USA). 26S ribosomal RNA served as an internal control.10,12
Measurement of ethylene production
Ethylene production was measured as described previously by Chang et al.11 Briefly, at 5 days after anthesis (D5), each individual flower was placed in a tube with water and incubated for 3 h at 25 °C. A 1 ml sample of gas was collected using a gas-tight hypodermic syringe and injected into a gas chromatographer (GC-8A; Shimadzu, Kyoto, Japan) for ethylene detection and measurement. Five biological replicates were used for each measurement.
Electrophoretic mobility shift assay
The electrophoretic mobility shift assay (EMSA) was performed according to Luo et al.29 Recombinant pGEX-PhFBH4 proteins were produced in Escherichia coli strain BL21. The E. coli cells were lysed by sonication and purified with glutathione–Sepharose 4B beads (GE Healthcare, Sunnyvale, CA, USA). The proACO1 probe 5′-TTAAATCACACATGAATAATATCAAATGTTTTGGTC-3′, the proACS1 probe 5′-TTTCTTTCTCACGTGTAGCTTCTA-3′, the mutated proACS1 probe 5‘-TTTCTTTCTGCTACTTAGCTTCTA-3’ and their complementary probes were labeled with biotin. For each binding reaction, 1 μg aliquot recombinant protein and 2 nM biotin-labeled probe were used. The LightShift chemiluminescent EMSA kit (Pierce, Rockford, IL, USA) was used for detection.29
Results
Isolation and expression pattern of PhFBH4
One transcript, annotated as bHLH TF, was identified among genes upregulated during flower senescence using a custom Nimblegen microarray (Supplementary Figure S1).10 To analyze the function of this putative transcription factor, full-length cDNA was cloned by rapid amplification of cDNA ends. Sequence analysis showed that this TF encodes 345 amino acids with a bHLH domain. Phylogenetic analysis suggested that this bHLH protein belongs to the bHLH IX family17 and has a high homology with AtFBH4, SlFBH4, and StFBH4 (Supplementary Figure S2). Therefore, the gene was named PhFBH4.
To confirm the microarray data, flowers at different stages were collected (Figure 1A) and qRT-PCR was used to detect the expression level of PhFBH4. The PhFBH4 transcript level constantly increased in petals from the just fully opened flower (D0) to early wilted flower (D6) and then remained high until wilting completed (D7; Figure 1B). Furthermore, PhFBH4 transcripts were detected in the leaf, stem, root, and all flower organs (Supplementary Figure S3). One interesting feature of PhFBH4 is that expression level in old leaves is much higher than that in young leaves.
PhFBH4 transcripts in different flower stages. (A) Different stages of petunia flower after anthesis. D0: the day of anthesis, D2, D4, D6, D7: 2, 4, 6, and 7 days after anthesis, respectively. (B) Expression level of PhFBH4 in different flower stages by qRT-PCR. Error bars show SD of the means of three biological replicates.
PhFBH4 expression is regulated by ethylene, ABA and abiotic stresses
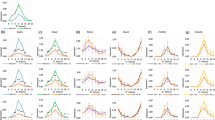
As hormones and stresses play important roles in flower senescence, we determined the expression pattern of PhFBH4 in petals after different hormone and abiotic stress treatments. Flowers treated with ethylene, ABA, drought, high temperature, and salt displayed higher expression of PhFBH4 than untreated controls (Figure 2). Cold treatment inhibited PhFBH4 expression by 50% after 3 h. Application of GA3 did not affect PhFBH4 expression (Figure 2).
Expression pattern of PhFBH4 under hormone and abiotic treatment by qRT-PCR. (A) Expression of PhFBH4 under hormone treatment. Petunia flowers harvested at anthesis were placed in tubes with water (mock), 0.1 mM ABA, 50 μM GA3, treated with ethylene (3 µl/l), or with 1-MCP for 4 h before ethylene treatment. (B) Expression of PhFBH4 under abiotic stress treatment. Petunia flowers were placed in tubes with water at 29 °C and 4 °C, without water (drought), or with 100 mM NaCl at room temperature (salt). Error bars show SD of the means of three biological replicates.
Virus-induced PhFBH4 silencing extended flower longevity
The TRV-based VIGS system has proven to be an efficient and fast method to silence target genes in petunia.12 Therefore, to quickly study the function of PhFBH4, a 308 bp fragment of PhFBH4 was cloned into a silencing construct bearing a fragment of the petunia CHS gene as a visual reporter. Five weeks after inoculation, white sectors were observed on the normally purple corollas indicating silencing of CHS (Figure 3A). Semi-quantitative RT-PCR was used to detect the gene expression in different plants. Compared with WT and CHS/TRV controls, the PhFBH4 gene transcript was clearly downregulated in the white flowers of the PhFBH4/CHS/TRV plants (Figure 3C). Silencing PhFBH4 extended flower longevity by two more days in comparison to the controls (WT flowers and white flowers from CHS/TRV) (Figure 3B). Transcripts of the senescence marker genes, SAG12 and SAG29, were barely detected in white flowers of PhFBH4/CHS/TRV plants 7 days after anthesis, indicating senescence progress was delayed by silencing PhFBH4 (Figure 3C).
Silencing PhFBH4 using the VIGS system extended flower longevity. (A) Phenotype of plants using different constructs at 5 weeks after inoculation. (B) Flower longevity of attached flowers in different plants. CHS/TRV (W), white flowers in CHS/TRV plants; PhFBH4/CHS/TRV (W), white flowers in PhFBH4/CHS/TRV plants. Error bars indicate SD (n ≥ 10). Different letters denote significant differences at p > 0.05 analyzed by Tukey's test (C) Abundance of related gene expression in flowers on D7 in different plants.
PhFBH4 regulates flower senescence by modulating ethylene biosynthesis
To further investigate the function of PhFBH4, we generated PhFBH4 overexpression and antisense silencing transgenic plants in petunia. Expression level of PhFBH4 was determined by qRT-PCR (Figure 4A). To confirm whether PhFBH4 is involved in flower senescence, the flower longevity of WT and transgenic plants was recorded (Figure 4B). The longevity of intact WT flowers was approximately 7 days. Overexpression of PhFBH4 accelerated flower senescence and shortened flower longevity to 5.5 days, while silencing of PhFBH4 extended flower longevity to about 9 days (Figure 4C). Since petunia plants of PhFBH4-OX-2 and PhFBH4-AS-1 exhibited the strongest expression and flower longevity difference, they were chosen for further analysis. Transcript abundance of the senescence marker genes, SAG12 and SAG29, were correlated with the flower longevity of WT and transgenic plants (Figure 4D).
Ectopic expression of PhFBH4 affected flower longevity. (A) Expression of PhFBH4 in WT, PhFBH4 overexpression and antisense silencing transgenic plants by qRT-PCR. PhFBH4-OX-2, PhFBH4-OX-7, different lines of PhFBH4 overexpression. PhFBH4-AS-1, PhFBH4-AS-3, different lines of PhFBH4 antisense silencing. Error bars show SE of the means of three biological replicates. (B) Different flower stages in WT petunia and PhFBH4 transgenic petunia. D0: the day of anthesis; D5, D7: 5 and 7 days after anthesis, respectively. (C) Flower longevity in WT and PhFBH4-OX and AS transgenic plants. Error bars indicate SD (n ≥ 10). Different letters denote significant differences at p ≤ 0.05 analyzed by Tukey's test. (D) Senescence marker genes, SAG12 and SAG29, expression in the flower of wide type petunia and transgenic petunia on D5.
As ethylene is important in flower senescence, we then investigated the abundance of ACO and ACS gene transcripts and ethylene production in WT and transgenic plants. ACO1 and ACS1 transcript levels were significantly upregulated in PhFBH4-OX plants compared with those of WT. On the other hand, ACO1 and ACS1 expression levels were decreased in PhFBH4-AS plants (Figure 5). Ethylene production further confirmed the transcriptional differences. Ethylene production on day 5 was significantly reduced in the PhFBH4 silenced flowers and was much higher in PhFBH4 overexpression flowers than that in WT flowers (Figure 6). These results suggested that PhFBH4 regulates flower senescence by mediating the ethylene biosynthetic genes ACO1 and ACS1.
Expression of ethylene biosynthesis-related genes in D5 petunia flowers. Abundance of transcripts of genes associated with ethylene biosynthesis was determined at D5 in WT and PhFBH4 transgenic plants. Error bars show SD of the means of three biological replicates. Different letters denote significant differences at p ≤ 0.05 analyzed by Tukey's test.
To test the direct binding between PhFBH4 protein and promoters of ACO1/ACS1, an EMSA was carried out. A search for potential PhFBH4-binding motifs revealed the presence of the E-box motif CANNTG and the G-box motif CACGTG in the 1.5 kb-upstream promoter regions of the petunia ACS1 and ACO1 genes. The biotin-labeled probes were designed to bind to the E/G-box element in the promoter of ACO1 and ACS1 (Figure 7). The EMSA result showed that the PhFBH4 protein was capable of binding to the biotin-labeled probe of the G-box in the promoter of ACS1 (Figure 7, lane 2). Binding was gradually abolished by the addition of an unlabeled oligonucleotide competitor in 10-fold (lane 3), 100-fold (lane 4), and 1000-fold (lane 5) molar excess. No binding was observed when mutant oligonucleotide probes were used (lane 6), whereas specific binding was maintained when the same amount of the excess mutant competitor was added (lane 7). However, there was no binding between the PhFBH4 protein and the E-boxes in the promoter of ACO1 (Figure 7, lane 8). These results suggest that ACS1 is a direct target of PhFBH4.
Electrophoretic mobility shift assay of the PhFBH4 protein. The biotin-labeled oligonucleotide of proACS1 was mixed with GST-tagged protein control (lane 1) or PhFBH4 proteins prepared from cells transfected with pGST::PhFBH4 plasmid (lanes 2–7). PhFBH4 protein physically binds a cis-element (G-box) in the PhACS1 promoter (lane 2). Binding was gradually abolished by the addition of an unlabeled oligonucleotide competitor in 10-fold (lane 3), 100-fold (lane 4), and 1000-fold (lane 5) molar excess, whereas specific binding was maintained when the same amount of the excess mutant oligonucleotide competitor was added (lanes 6 and 7). PhFBH4 protein failed to bind the E-boxes in the promoter of ACO1 (lane 8).
Ectopic expression of PhFBH4 affects plant growth
At 50 days after transplanting, phenotypical differences were clearly observed between WT and transgenic plants (Figure 8). Leaves and flowers of PhFBH4-AS plants were significantly larger than WT control while leaves and flowers of PhFBH4-OX plants were smaller (Figure 8). Changes in PhFBH4 expression also affected plant height. Stem and internode lengths of the antisense transgenic lines were significantly greater than those of the controls, and were significantly smaller in the over-expressing transgenic lines (Table 1). There were no differences in the number of internodes or flowering time between WT and transgenic plants (Table 1).
The plant hormone gibberellin (GA) plays an important role in many aspects of plant development, particularly in plant height and stem elongation.30 To test whether the phenotypic differences between WT and PhFBH4 transgenic plants were associated with the GA pathway, transcript abundances of GA biosynthesis-related genes were determined. qRT-PCR results showed that the transcript abundances of the GA biosynthetic genes, GA20ox1 and GA20ox2, were not changed in WT and PhFBH4 transgenic plants. However, one of the GA metabolic genes GA2ox3 displayed a significant difference in abundance. The transcriptional level of GA2ox3 in PhFBH4-OX plants was increased 2.74-fold and was reduced 48% in PhFBH4-AS plants, compared to that of WT controls (Figure 9). The significant difference in GA2ox3 transcript levels could result in a change of bioactive GA, thereby influencing the growth in PhFBH4 transgenic plants.
Expression analysis of GA-related genes in the flower of WT and transgenic petunia on D4 by qRT-PCR. Abundance of transcripts of genes associated with GA was determined at D4 in WT and PhFBH4 transgenic plants. Error bars show SD of the means of three biological replicates. Different letters denote significant differences at p ≤ 0.05 analyzed by Tukey's test.
Discussion
PhFBH4 is involved in flower senescence by modulating ethylene biosynthesis
In this study, we identified a bHLH TF, named PhFBH4 because of its sequence similarity to Arabidopsis FBH4. We found that its transcript abundance increased dramatically during flower senescence, and increased in response to a range of abiotic stressors and to treatments with plant hormones, particularly ethylene. The gaseous phytohormone ethylene is known to play a critical role in flower senescence.31 In many flowers, the onset of floral senescence is initiated by a climacteric rise in ethylene production.32 Application of ethylene accelerates flower senescence, while ethylene inhibitors such as 1-MCP can significantly delay the senescence process.33–35 The biosynthetic pathway of ethylene has been well studied.36 Two important enzymes, 1-aminocyclopropane-1-carboxylate synthase (ACS), which catalyzes the conversion of S-adenosylmethionine (AdoMet) to ACC, and ACC oxidase (ACO), which converts ACC into ethylene, are encoded by multiple gene families.37 In carnation, petunia, and tomato, the increase in the ethylene synthesis is accompanied by increased ACS and ACO gene expression and elevated enzyme activities.38–41 Downregulation of the ACO gene in carnation and petunia causes low ethylene production and markedly delayed petal senescence.12,42 However, the transcriptional regulation of the ethylene biosynthesis pathway during the flower senescence has been little studied. In this study, manipulation of PhFBH4 expression using overexpression and silencing approaches altered flower longevity, accompanied by alterations in the transcript abundances of ethylene biosynthesis genes ACO1 and ACS1 and ethylene production. Furthermore, our results in the EMSA suggest that PhFBH4 directly binds to the promoter of ACS1 (Figure 7). Our data demonstrates that PhFBH4 is involved in the regulation of flower senescence progress through its interaction with the ethylene biosynthesis pathway.
In Arabidopsis thaliana, FLOWERING BHLH 1 (FBH1), FBH2, FBH3, and FBH4 were identified as four CO transcriptional activators. All FBH proteins are related to the bHLH-type TFs that preferentially bind to the E-box cis-elements in the CO promoter. Overexpression of all FBH genes caused early flowering regardless of photoperiod. Furthermore, FBH homologs in poplar and rice induced CO expression in Arabidopsis.27 However, the early flowering phenotype seen in Arabidopsis FBH-OX was not observed in PhFBH4-OX transgenic petunia plants. Interestingly, PhFBH4 only binds the G-box cis-elements in the ACS1 promoter. This may explain the difference in flowering timing between Arabidopsis and petunia.
PhFBH4 may be involved in crosstalk between ethylene and GA during plant development
In addition to its positive role in flower senescence, ethylene is generally considered a growth inhibitor.43 After ethylene treatment, rapid inhibition of elongation was reported in stems of Pisum sativum,44 leaves of Poa species45 and roots of Cucumis sativus.46 Ethylene overproduction reduces internode length through modification of ACS and ACO gene expression levels in Nicotiana tabacum.47
Gibberellins and ethylene are both involved in the control of plant developmental processes from seed germination and cell elongation in hypocotyls to formation of stomata and flower senescence.48,49 Crosstalk between GAs and ethylene, as well as with other hormones, has been demonstrated in Arabidopsis. Bioactive GA levels are low in the ctr1 mutant and after ACC treatment but increase in the ethylene-insensitive etr1-2 mutant.50–52 Furthermore, ethylene regulates the maintenance and exaggeration of the apical hook by modifying DELLA degradation.53 Active ethylene signaling results in decreased GA content, thus stabilizing DELLA proteins.51 Recent evidence suggests that reduction in the bioactive GA content enhances ethylene-mediated flower senescence in rose.54 These results suggest that the antagonism effect between ethylene and GA is mediated by regulating bioactive GA levels and the stability of DELLA proteins.55 In carnation, exogenous application of GA can delay the senescence of cut flowers by reducing ethylene production.56 In this study, overexpression of PhFBH4 increased the abundance of transcripts of ethylene biosynthesis genes (ACO1/ACS1; Figure 5) and also increased ethylene production (Figure 6). Moreover, the increased expression of the GA metabolic gene GA2ox3 in PhFBH4-OX transgenic plants would raise bioactive GAs content, while silencing PhFBH4 would reduce their levels (Figure 9). Our data support this hypothesis. In PhFBH4-OX transgenic plants, which produced more ethylene and would hypothetically have less bioactive GA than WT petunia, we observed reduced stem, leaf and flower size. In contrast, silencing of PhFBH4 resulted in longer internode length and larger leaves and flowers compared with WT. Furthermore, overexpression of PhFBH4 accelerated flower senescence and shortened flower longevity, while silencing of PhFBH4 extended flower longevity (Figure 4C). These results suggest that PhFBH4 mediates an antagonistic relationship of ethylene and GA in plant growth and flower senescence. It would be interesting to comprehensively analyze the relationships between plant growth and concentrations of some of bioactive GAs or expression levels of expanded GA-related genes such as GA3ox genes in the future.
References
Tripathi SK, Tuteja N . Integrated signaling in flower senescence: an overview. Plant Signal Behav 2007; 2: 437–445.
Wagstaff C, Yang TJW, Stead AD, Buchanan-Wollaston V, Roberts JA . A molecular and structural characterization of senescing Arabidopsis siliques and comparison of transcriptional profiles with senescing petals and leaves. Plant J 2009; 57: 690–705.
Liu J, Li J, Wang H et al. Identification and expression analysis of ERF transcription factor genes in petunia during flower senescence and in response to hormone treatments. J Exp Bot 2011; 62: 825–840.
El-Sharkawy I, Sherif S, Mila I, Bouzayen M, Jayasankar S . Molecular characterization of seven genes encoding ethylene-responsive transcriptional factors during plum fruit development and ripening. J Exp Bot 2009; 60: 907–922.
Liu M, Diretto G, Pirrello J et al. The chimeric repressor version of an ethylene response factor (ERF) family member, Sl-ERF.B3, shows contrasting effects on tomato fruit ripening. New Phytol 2014; 203: 206–218.
Yin XR, Allan AC, Chen KS, Ferguson IB . Kiwifruit EIL and ERF genes involved in regulating fruit ripening. Plant Physiol 2010; 153: 1280–1292.
Li X, Zhu X, Mao J et al. Isolation and characterization of ethylene response factor family genes during development, ethylene regulation and stress treatments in papaya fruit. Plant Physiol Biochem 2013; 70: 81–92.
Hay A, Tsiantis MA . KNOX family TALE. Curr Opin Plant Biol 2009; 12: 593–598.
Reid M, Chen J-C, Jiang C-Z . Virus-Induced Gene Silencing for Functional Characterization of Genes in Petunia. In: Gerats T, Strommer J, editors. Petunia. New York: Springer; 2009. pp381–394.
Wang H, Stier G, Lin J et al. Transcriptome changes associated with delayed flower senescence on Transgenic Petunia by inducing expression of etr1-1, a mutant ethylene receptor. PLoS One 2013; 8: e65800.
Chang X, Donnelly L, Sun D et al. A Petunia homeodomain-leucine zipper protein, PhHD-Zip, plays an important role in flower senescence. PLoS One 2014; 9: e88320.
Chen J-C, Jiang CZ, Gookin TE et al. Chalcone synthase as a reporter in virus-induced gene silencing studies of flower senescence. Plant Mol Biol 2004; 55: 521–530.
Jiang C-Z, Chen J-C, Reid M . Virus-induced gene silencing in ornamental plants. In: Kodama H, Komamine A, editors. RNAi and Plant Gene Function Analysis Vol. 744. Methods in Molecular Biology. Springer; 2011; pp81–96.
Bailey PC, Martin C, Toledo-Ortiz G et al. Update on the basic helix-loop-helix transcription factor gene family in Arabidopsis thaliana. Plant Cell 2003; 15: 2497–2502.
Li X, Duan X, Jiang H et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol 2006; 141: 1167–1184.
Toledo-Ortiz G, Huq E, Quail PH . The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 2003; 15: 1749–1770.
Heim MA, Jakoby M, Werber M et al. The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 2003; 20: 735–747.
Fernández-Calvo P, Chini A, Fernández-Barbero G et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 2011; 23: 701–715.
Ellenberger T, Fass D, Arnaud M, Harrison SC . Crystal structure of transcription factor E47: E-box recognition by a basic region helix-loop-helix dimer. Genes Dev 1994; 8: 970–980.
Morgenstern B, Atchley WR . Evolution of bHLH transcription factors: modular evolution by domain shuffling? Mol Biol Evol 1999; 16: 1654–1663.
Abe H, Urao T, Ito T et al. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 2003; 15: 63–78.
Fairchild CD, Schumaker MA, Quail PH . HFR1 encodes an atypical bHLH protein that acts in phytochrome A signal transduction. Genes Dev 2000; 14: 2377–2391.
Riechmann JL, Heard J, Martin G et al. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 2000; 290: 2105–2110.
Li F, Guo S, Zhao Y et al. Overexpression of a homopeptide repeat-containing bHLH protein gene (OrbHLH001) from Dongxiang wild rice confers freezing and salt tolerance in transgenic Arabidopsis. Plant Cell Rep 2010; 29: 977–986.
Liljegren SJ, Roeder AH, Kempin SA et al. Control of fruit patterning in Arabidopsis by INDEHISCENT. Cell 2004; 116: 843–853.
Payne CT, Zhang F, Lloyd AM . GL3 encodes a bHLH protein that regulates trichome development in Arabidopsis through interaction with GL1 and TTG1. Genetics 2000; 156: 1349–1362.
Ito S, Song YH, Josephson-Day AR et al. FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis. Proc Natl Acad Sci U S A 2012; 109: 3582–3587.
Chen JC, Jiang CZ, Gookin TE et al. Chalcone synthase as a reporter in virus-induced gene silencing studies of flower senescence. Plant Mol Biol 2004; 55: 521–530.
Luo J, Ma N, Pei H et al. A DELLA gene, RhGAI1, is a direct target of EIN3 and mediates ethylene-regulated rose petal cell expansion via repressing the expression of RhCesA2. J Exp Bot 2013; 64: 5075–5084.
Martin DN, Proebsting WM, Hedden P . The SLENDER gene of pea encodes a gibberellin 2-oxidase. Plant Physiol 1999; 121: 775–781.
Reid MS, Jiang CZ . Postharvest biology and technology of cut flowers and potted plants. Hortic Rev 2012; 40: 3–56.
Woltering EJ, Somhorst D, Van Der Veer P . The role of ethylene in interorgan signaling during flower senescence. Plant Physiol 1995; 109: 1219–1225.
Lovell PJ, Lovell PH, Nichols R . The control of flower senescence in petunia (Petunia hybrids). Ann Bot 1987; 60: 49–59.
Porat R, Borochov A, Halevy AH . Enhancement of petunia and dendrobium flower senescence by jasmonic acid methyl ester is via the promotion of ethylene production. Plant Growth Regul 1993; 13: 297–301.
Serek M, Sisler EC, Reid MS . Effects of 1-MCP on the vase life and ethylene response of cut flowers. Plant Growth Regul 1995; 16: 93–97.
Yang SF, Hoffman NE . Ethylene biosynthesis and its regulation in higher plants. Annu Rev Plant Physiol 1984; 35: 155–189.
Zarembinski T, Theologis A . Ethylene biosynthesis and action: a case of conservation. Plant Mol Biol 1994; 26: 1579–1597.
Tang X, Gomes AMTR, Bhatia A, Woodson WR . Pistil-specific and ethylene-regulated expression of 1-aminocyclopropane-1-carboxylate oxidase genes in petunia flowers. Plant Cell 1994; 6: 1227–1239.
Tang X, Woodson WR . Temporal and spatial expression of 1-aminocyclopropane-1-carboxylate oxidase mRNA following pollination of immature and mature petunia flowers. Plant Physiol 1996; 112: 503–511.
Woodson WR, Park KY, Drory A, Larsen PB, Wang H . Expression of ethylene biosynthetic pathway transcripts in senescing carnation flowers. Plant Physiol 1992; 99: 526–532.
Llop-Tous I, Barry CS, Grierson D . Regulation of ethylene biosynthesis in response to pollination in tomato flowers. Plant Physiol 2000; 123: 971–978.
Savin KW, Baudinette SC, Graham MW . Antisense ACC oxidase RNA delays carnation petal senescence. HortSci 1995; 30: 970–972.
Pierik R, Tholen D, Poorter H, Visser EJW, Voesenek LACJ . The Janus face of ethylene: growth inhibition and stimulation. Trends Plant Sci 2006; 11: 176–183.
Fuchs Y, Lieberman M . Effects of kinetin, IAA, and gibberellin on ethylene production, and their interactions in growth of seedlings. Plant Physiol 1968; 43: 2029–2036.
Fiorani F, Bögemann GM, Visser EJW, Lambers H, Voesenek LACJ . Ethylene emission and responsiveness to applied ethylene vary among Poa species that inherently differ in leaf elongation rates. Plant Physiol 2002; 129: 1382–1390.
Pierik R, Verkerke W, Voesenek R, Blom K, Visser E . Thick root syndrome in cucumber (Cucumis sativus L.): a description of the phenomenon and an investigation of the role of ethylene. Ann Bot 1999; 84: 755–762.
Knoester M, Linthorst HJM, Bol JF, van Loon LC . Modulation of stress-inducible ethylene biosynthesis by sense and antisense gene expression in tobacco. Plant Sci 1997; 126: 173–183.
Fleet CM, Sun TP . A DELLAcate balance: the role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 2005; 8: 77–85.
Richards DE, King KE, Ait-ali T, Harberd NP . HOW GIBBERELLIN REGULATES PLANT GROWTH AND DEVELOPMENT: a molecular genetic analysis of gibberellin signaling. Annu Rev Plant Physiol Plant Mol Biol 2001; 52: 67–88.
Chiwocha SDS, Cutler AJ, Abrams SR et al. The etr1-2 mutation in Arabidopsis thaliana affects the abscisic acid, auxin, cytokinin and gibberellin metabolic pathways during maintenance of seed dormancy, moist-chilling and germination. Plant J 2005; 42: 35–48.
Achard P, Baghour M, Chapple A et al. The plant stress hormone ethylene controls floral transition via DELLA-dependent regulation of floral meristem-identity genes. Proc Natl Acad Sci U S A 2007; 104: 6484–6489.
Vandenbussche F, Vancompernolle B, Rieu I et al. Ethylene-induced Arabidopsis hypocotyl elongation is dependent on but not mediated by gibberellins. J Exp Bot 2007; 58: 4269–4281.
Achard P, Vriezen WH, Van Der Straeten D, Harberd NP . Ethylene regulates Arabidopsis development via the modulation of DELLA protein growth repressor function. Plant Cell 2003; 15: 2816–2825.
Lü P, Zhang C, Liu J et al. RhHB1 mediates the antagonism of gibberellins to ABA and ethylene during rose (Rosa hybrida) petal senescence. Plant J 2014; 78: 578–590.
De Grauwe L, Chaerle L, Dugardeyn J et al. Reduced gibberellin response affects ethylene biosynthesis and responsiveness in the Arabidopsis gai eto2-1 double mutant. New Phytol 2008; 177: 128–141.
Saks Y, Staden J . Evidence for the involvement of gibberellins in developmental phenomena associated with carnation flower senescence. Plant Growth Regul 1993; 12: 105–110.
Acknowledgements
This work was funded in part by the United States Department of Agriculture (USDA) Floriculture Initiative (5306-21000-019-00D). We thank Linda Donnelly for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplemental Information for this article can be found on the Horticulture Research website (http://www.nature.com/hortres).
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yin, J., Chang, X., Kasuga, T. et al. A basic helix-loop-helix transcription factor, PhFBH4, regulates flower senescence by modulating ethylene biosynthesis pathway in petunia. Hortic Res 2, 15059 (2015). https://doi.org/10.1038/hortres.2015.59
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2015.59
This article is cited by
-
Expression of ethylene biosynthetic genes during flower senescence and in response to ethephon and silver nitrate treatments in Osmanthus fragrans
Genes & Genomics (2024)
-
A combined transcriptomic and proteomic analysis of chrysanthemum provides new insights into petal senescence
Planta (2022)
-
Disruption of transcription factor RhMYB123 causes the transformation of stamen to malformed petal in rose (Rosa hybrida)
Plant Cell Reports (2022)
-
Phosphoproteome analysis reveals the involvement of protein dephosphorylation in ethylene-induced corolla senescence in petunia
BMC Plant Biology (2021)
-
Genome and transcriptome-based characterization of high energy carbon-ion beam irradiation induced delayed flower senescence mutant in Lotus japonicus
BMC Plant Biology (2021)