Abstract
The genetic relatedness between individuals because of their recent common ancestry is now routinely estimated from marker genotype data in molecular ecology, evolutionary biology and conservation studies. The estimators developed for this purpose assume that marker allele freque218 in a population are known without errors. Unfortunately, however, these frequencies, upon which both the definition and the estimation of relatedness are based, are rarely known in reality. Frequently, the only data available in a relatedness analysis are a sample of multilocus genotypes from which both allele frequencies and relatedness must be deduced. Furthermore, because of various constraints, sample sizes of individuals can be quite small (say <50 individuals) in practice. This study shows, for the first time, that the widely used relatedness estimators become severely biased when they use allele frequencies calculated from an extremely small sample (say <10 individuals). The extent of bias depends on the sample size, the (unknown) population allele frequencies, the actual relatedness and the estimators. It also shows that relatedness estimators become even more biased when they use allele frequencies calculated from a sample by omitting a focal pair of individuals whose relatedness is being estimated. This study modifies two estimators to suit small samples and shows, both analytically and by analysing simulated and empirical data, that the two modified estimators are much less biased, more precise and more accurate than the original estimators. These performance advantages of the modified estimators are shown to increase with a decreasing sample size of individuals and with an increasing value of actual relatedness.
Similar content being viewed by others
Introduction
Inbreeding and relatedness are pivotal concepts in population genetics theory (Wright, 1921, 1922), and have important applications in many research areas in quantitative genetics, conservation genetics, forensics, evolution and ecology (Weir et al., 2006). Two individuals are genetically related because they have recently shared genealogical history, or have common ancestors in the recent past. The number of common ancestors and their distances (that is, the number of generations) to a pair of individuals determine the (expected) extent of relatedness between the individuals. Related individuals have more similar genotypes at each locus because their alleles have a higher probability of identity by descent (IBD) than unrelated individuals. As a result, they also tend to have a higher similarity in the phenotype of a quantitative trait (Falconer and Mackay, 1996; Lynch and Walsh, 1998).
Relatedness is traditionally calculated from pedigree data, as exemplified by the analysis of Wright (1922) of a Shorthorn cattle pedigree. Unfortunately, pedigree is rarely available and complete from natural populations. With the rapid development of genetic markers, quite a few methods have been proposed (see, for example, Lynch, 1988; Queller and Goodnight, 1989; Li et al., 1993; Loiselle et al., 1995; Ritland, 1996; Lynch and Ritland, 1999; Wang, 2002, 2007; Milligan, 2003; Thomas, 2010), implemented in computer programs (see, for example, Hardy and Vekemans, 2002; Wang, 2011a) and applied to estimating the genetic relatedness between a pair of individuals from their marker genotypes. Compared with pedigree data, marker data are easier to collect and do not have to be accumulated over a prolonged number of generations. A single sample of individuals taken from a population and genotyped at a number of marker loci provides all the information necessary for assessing the relative relatedness between the sampled individuals (Weir et al., 2006). Furthermore, markers can yield realized rather than expected relatedness as calculated from pedigrees, and can produce much better relatedness estimates than pedigrees when they are numerous (Kardos et al., 2015; Wang, 2016). This marker-based approach has enabled many genetics studies of natural populations of various plant and animal species (DeWoody, 2005; Garant and Kruuk, 2005), and has made many analyses (such as estimating heritability) traditionally based on pedigrees much more powerful (see, for example, Manolio et al., 2009).
Marker-based relatedness estimators are developed on the assumption that marker allele frequencies in a suitably defined reference population (see, for example, Ritland, 1996; Lynch and Ritland, 1999; Wang, 2014) are known without errors. This allele frequency information is supposed to be independent of the sample multilocus genotype data, and the reference population is implicitly assumed to be large and at random mating such that all homologous genes within or between reference individuals are not IBD. A shift of reference populations (that is, reference allele frequencies) will change the biological meaning and the estimated values of relatedness among sampled individuals (Anderson and Weir, 2007; Wang, 2011b, 2014). Strictly under this assumption, a number of relatedness estimators are shown to be unbiased (see, for example, Lynch and Ritland, 1999; Van de Casteele et al., 2001; Wang, 2002, 2011b) and consistent, providing increasingly accurate estimates with an increase in marker information.
In reality, however, the strictly defined reference allele frequencies are never available. Although allele frequency data independent of sample genotype data can be available in the rare cases of some well-studied populations (model organisms), these populations are invariable structured genetically. Most frequently, the only data one has are samples of multilocus genotypes, from which one has to deduce allele frequencies by assuming all individuals are unrelated and non-inbred. Using these estimated allele frequencies to infer relatedness creates several problems (Wang, 2014).
First, the average relatedness among sampled individuals becomes close to zero, and a substantial proportion of the pairwise relatedness estimates are negative. This is true regardless of the actual genetic structure of the sample. These results are expected because marker-based relatedness is more appropriately interpreted as a correlation coefficient, as originally conceived by (Wright 1921, 1922), rather than the probability of IBD, as used in developing various estimators (Wang, 2014). A negative relatedness estimate means the individuals are less related than the average. Such relatedness estimates are expected to be lower, more or less proportionally, than those when ancestral reference allele frequencies would have been used. However, this underestimation of relatedness does not cause problems in the majority of relatedness analyses (for example, regression and correlation analyses) in which it is the relative rather than absolute values of relatedness that matters (Wang, 2014).
Second, the relatedness picture of a sample of individuals is distorted when the sample, small or large, contains a substantial proportion of close relatives. With allele frequencies estimated by naively assuming all individuals in the sample are non-inbred and unrelated, relatedness tends to be more underestimated for closely related individuals than for loosely related or unrelated individuals (Wang, 2014). The distorted relatedness estimates could derail or cause bias of all downstream analyses, no matter whether they depend on relative (for example, in a correlation analysis) or absolute (for example, in inferring genealogical relationship) relatedness.
Third, estimating allele frequencies and relatedness from the same sample of individuals introduces circularity, and results in an underestimation of relatedness in the order of 1/N, where N is the number of sampled individuals (Queller and Goodnight, 1989; Loiselle et al., 1995; Ritland, 1996; Lynch and Ritland, 1999). The underestimation is true no matter whether the sample is genetically structured or not. It is suggested that this bias can be simply removed by adding a correction factor of order 1/N (Loiselle et al., 1995), or by calculating and using allele frequencies obtained by omitting the focal pair of individuals (Queller and Goodnight, 1989; Lynch and Ritland, 1999). In the latter approach, however, it is recognized that pathological behaviour will occur when an allele appears only in the focal individuals (Lynch and Ritland, 1999). In such a case, allele frequency will be estimated to be zero by this exclusion procedure, causing some relatedness estimators to become undefined. However, no study has been conducted to check whether these two bias correction procedures are effective or not.
In the pre-genomics era, small sample sizes and thus the resultant biases of a relatedness estimator were not a serious problem, compared with other problems, except when N is extremely small (say, N<10). When a typical set of only 10–20 microsatellites is used in calculating relatedness, the sampling variance is expected to overwhelm the small bias in the order of 1/N (Lynch and Ritland, 1999) in determining estimation accuracy. With the rapid development and applications of next-generation sequencing, however, hundreds of thousands of single-nucleotide polymorphisms (SNPs) can be genotyped at ease for an individual. With this vast volume of data, marker-based relatedness estimates can be highly precise, and much more accurate than pedigree-based estimates (Kardos et al., 2015; Wang, 2016). Furthermore, a dramatic increase in the number of markers usually accompanies a dramatic decrease in the number of sampled and genotyped individuals because of practical constraints such as cost. Moreover, the extremely sparse nature of SNP data given by next-generation sequencing makes the small sample size problem even more acute: the number of usable genotypes at any locus is typically much smaller than the number of sampled individuals. In such a situation of many more sampled markers (say, millions) than sampled genotypes at a locus (say, 50 or less), which is typical in the genomic era, the bias due to a small sample size of individuals becomes prominent. The number of sampled individuals can also be very small in other practical situations, such as ancient samples (for example, museum samples, excavated fossil bones), mixed samples of unknown sources (for example, confiscated animal products, victims of a disaster) and samples from highly endangered species.
In this study, I analyse the bias of several popular moment estimators of relatedness when a small sample of multilocus genotypes is used for calculating both allele frequencies and relatedness. I also investigate whether omitting a focal pair of individuals in estimating allele frequencies can remove or reduce the bias or not. Finally, I propose a method to modify some of the estimators such that they provide unbiased and accurate relatedness estimates even when sample size is extremely small. Simulated and empirical data sets were analysed to study the behaviour of the original and modified estimators.
Bias due to small sample size
In this section, I quantify the bias caused by calculating both allele frequencies and relatedness from a small sample of individual genotypes. For generality, relatedness estimators are described for a locus with any number of alleles and for multiple loci. For simplicity, however, the bias of the estimators is investigated by considering a single locus with two codominant alleles, A and B, with frequencies p and q (=1−p) in a large random mating population at Hardy–Weinberg equilibrium. The bias is confirmed by simulations, described in the next section, for multiallelic and multiple loci.
In a sample of N individuals drawn at random from the population, the counts i, j and k for the three genotypes AA, BB and AB, respectively, at a diallelic locus follow the multinomial distribution

For a given sample configuration characterized by genotype counts i, j and k, with i+j+k≡N, allele frequencies can be estimated by assuming unrelated and non-inbred individuals such that

The average relatedness of the N individuals calculated using  and
and  is expected to be negative, roughly in the order of 1/N, because of the circularity in allele frequency and relatedness estimation (Wang, 2014). In the following, I quantify analytically this bias of several estimators, when allele frequencies are estimated by including and omitting a focal pair of individuals whose relatedness is being estimated.
is expected to be negative, roughly in the order of 1/N, because of the circularity in allele frequency and relatedness estimation (Wang, 2014). In the following, I quantify analytically this bias of several estimators, when allele frequencies are estimated by including and omitting a focal pair of individuals whose relatedness is being estimated.
Queller and Goodnight estimator
Originally developed by Queller and Goodnight (1989), the estimator (denoted by QG or  ) has a number of variants in use. Here I use the symmetrical one obtained by averaging the estimates using each of the two individuals as reference (see Lynch and Ritland, 1999). If individuals X and Y have genotypes {a, b} and {c, d}, respectively, at a locus with n alleles, the estimator is
) has a number of variants in use. Here I use the symmetrical one obtained by averaging the estimates using each of the two individuals as reference (see Lynch and Ritland, 1999). If individuals X and Y have genotypes {a, b} and {c, d}, respectively, at a locus with n alleles, the estimator is

where allele indexes a, b, c, d=1, 2, …, n, pi is the frequency of allele i (=a, b, c, d), and the Kronecker delta variable δij=1 if i=j and δij=0 otherwise. The first and second terms on the right side of (3) give the estimates when X and Y are used as reference, respectively. For a number of L loci, the sum of the 2L numerator terms and the sum of the 2L denominator terms are calculated separately before the division is conducted to give the final multilocus estimate (Queller and Goodnight, 1989).
For a diallelic locus (that is, n=2) with alleles A and B, estimator (3) is undefined when the reference is a heterozygote, because the denominator is zero. For example, when X is the reference and it has a heterozygous genotype {a, b}={A, B}, then the denominator 2(1+δab−pa−pb)=0 because δab=0 and pa+pb≡1. In such a case, the undefined part of the estimator is set as zero. Alternatively, the undefined part is abandoned and only the defined part is used as the estimator. When both parts are undefined, the estimator is regarded as undefined and abandoned. This alternative treatment could increase bias (see below) and is thus not used in this study.
A diallelic locus has three possible genotypes and six possible (ordered) genotype pairs. These are {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB}, and their corresponding relatedness estimates calculated by (3) are 1, 1, 0,  , respectively. Using the genotype distribution (1), the expected average relatedness in a sample of N individuals is
, respectively. Using the genotype distribution (1), the expected average relatedness in a sample of N individuals is

where f[i,j,k|p] is calculated by (1), k≡N−i−j,  is the frequency that two individuals show the same genotype whose count is g in the sample,
is the frequency that two individuals show the same genotype whose count is g in the sample,  is the frequency that two individuals show different genotypes whose counts are g and h (g, h=i, j, k), respectively, in the sample. It can be shown that Equation (4) reduces to rr̄QG ≡ 0, irrespective of the sample size N (>1) and population allele frequencies p and q. This means QG estimator is unbiased when population allele frequencies are known and are used in calculating the relatedness of the N sampled individuals.
is the frequency that two individuals show different genotypes whose counts are g and h (g, h=i, j, k), respectively, in the sample. It can be shown that Equation (4) reduces to rr̄QG ≡ 0, irrespective of the sample size N (>1) and population allele frequencies p and q. This means QG estimator is unbiased when population allele frequencies are known and are used in calculating the relatedness of the N sampled individuals.
When allele frequencies are unknown and are estimated from the genotypes of the sample of individuals being considered for relatedness, the average relatedness estimate is still calculated by (4), with p and q in the brackets (that is, in the estimators) being replaced by  and
and  , respectively, calculated by (2). It turns out that
, respectively, calculated by (2). It turns out that  does not reduce to zero in general. Its value depends on both population allele frequencies p and q and sample size N. When N=3 and 4, for example,
does not reduce to zero in general. Its value depends on both population allele frequencies p and q and sample size N. When N=3 and 4, for example,  and
and  respectively, where h=pq. The effects of N and p on
respectively, where h=pq. The effects of N and p on  are shown in Figure 1. As can be seen,
are shown in Figure 1. As can be seen,  can be both positively and negatively biased, depending on values of N and p. With an increasing sample size N, rr̄QG always asymptotes to 0 regardless of p, as expected.
can be both positively and negatively biased, depending on values of N and p. With an increasing sample size N, rr̄QG always asymptotes to 0 regardless of p, as expected.
Means of different estimators as a function of sample size N. For each estimator, population allele frequencies are p=0.1 or p=0.4, and sample allele frequencies and relatedness are estimated by either excluding (Exc) or including (Inc) the focal pair of individuals. The expected values of estimator LL or W are not affected by population allele frequency p, so each line is for both p=0.1 and p=0.4. Note estimators W and LL are equivalent for a diallelic locus as shown by the graph in the lower right corner.
When allele frequencies are estimated from the sample by omitting the focal pair of individuals, the average relatedness estimate is expected to be

where allele frequency estimates are

for m=1 and 2. Note that  can be zero or one when the focal pair of individuals are excluded, leaving the estimator undefined. When i=N−1 and j=1, for example, excluding the individual with genotype {BB} will lead to
can be zero or one when the focal pair of individuals are excluded, leaving the estimator undefined. When i=N−1 and j=1, for example, excluding the individual with genotype {BB} will lead to  and
and  . In such a case, the estimator for genotype pair {AA, BB} becomes undefined and is set as zero.
. In such a case, the estimator for genotype pair {AA, BB} becomes undefined and is set as zero.
Again (5) does not reduce to zero, and  depends on both N and p, as shown by Figure 1. Calculating allele frequencies by excluding focal individuals actually increases bias substantially, in contrast to the usual perception (Queller and Goodnight, 1989; Lynch and Ritland, 1999). When P=0.1 and N=20, for example,
depends on both N and p, as shown by Figure 1. Calculating allele frequencies by excluding focal individuals actually increases bias substantially, in contrast to the usual perception (Queller and Goodnight, 1989; Lynch and Ritland, 1999). When P=0.1 and N=20, for example,  values are −0.028 and −0.222 when the focal pair of individuals are included and excluded in calculating allele frequencies, respectively.
values are −0.028 and −0.222 when the focal pair of individuals are included and excluded in calculating allele frequencies, respectively.
In Figure 1,  calculated by (3) is set to zero when it is undefined (because the denominator is zero). This is the most favourable treatment because (in theory) the mean estimate across all pairs of individuals in a sample should be close to zero, and because (in practice) most individuals in a natural population are expected to be unrelated. The alternative treatment (that is, abandoning undefined
calculated by (3) is set to zero when it is undefined (because the denominator is zero). This is the most favourable treatment because (in theory) the mean estimate across all pairs of individuals in a sample should be close to zero, and because (in practice) most individuals in a natural population are expected to be unrelated. The alternative treatment (that is, abandoning undefined  estimates) most often increases the bias. For the case of using all N=4 sampled individuals in calculating allele frequencies,
estimates) most often increases the bias. For the case of using all N=4 sampled individuals in calculating allele frequencies,  for the proposed treatment (that is, setting undefined estimate to zero) and
for the proposed treatment (that is, setting undefined estimate to zero) and  for the alternative treatment, when p=0.05, 0.1, 0.2 and 0.4, respectively. The alternative treatment often leads to a highly negative
for the alternative treatment, when p=0.05, 0.1, 0.2 and 0.4, respectively. The alternative treatment often leads to a highly negative  , especially when p is close to zero or one.
, especially when p is close to zero or one.
Ritland estimator
It was proposed by Li and Horvitz (1953) and was made popular by Ritland (1996) (denoted by R or  hereafter). For individuals X and Y with genotypes {a, b} and {c, d} respectively at a locus with n alleles, it is calculated by
hereafter). For individuals X and Y with genotypes {a, b} and {c, d} respectively at a locus with n alleles, it is calculated by

where the Kronecker delta δji=1 (for i, j=a, b, c, d) if j=i, and δji=0 if otherwise. In contrast to  ,
,  is always defined for a segregating locus (n>1). The variance of (7) is proportional to 1/(n−1), derived by assuming zero relatedness (Ritland, 1996). The multilocus estimator is obtained by weighting the locus-specific estimates by the inverses of their sampling variances. When locus l has nl alleles and relatedness estimate
is always defined for a segregating locus (n>1). The variance of (7) is proportional to 1/(n−1), derived by assuming zero relatedness (Ritland, 1996). The multilocus estimator is obtained by weighting the locus-specific estimates by the inverses of their sampling variances. When locus l has nl alleles and relatedness estimate  calculated by (7), the multilocus estimator is
calculated by (7), the multilocus estimator is  .
.
Note that Ritland (1996) estimated coancestry (θ) that is the probability that two homologous genes, one taken at random from one individual and another taken at random from another individual, are IBD. For non-inbred individuals, r=2θ and (7), when divided by 2, reduces to Ritland (1996) estimator of θ.
For the 6 possible genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} at a diallelic locus,  , respectively, calculated by (7). The expected average relatedness estimate in a sample of N individuals is
, respectively, calculated by (7). The expected average relatedness estimate in a sample of N individuals is

It can be shown that, with any known values of p and q, (8) reduces to  , irrespective of the sample size N (>1). This means R estimator is unbiased when allele frequencies are known.
, irrespective of the sample size N (>1). This means R estimator is unbiased when allele frequencies are known.
It can be shown that  does not reduce to zero when allele frequencies estimated from the sample by Equation (2) are used in calculating
does not reduce to zero when allele frequencies estimated from the sample by Equation (2) are used in calculating  in (8). When N=3 and 4, for example, (8) reduces to
in (8). When N=3 and 4, for example, (8) reduces to  ) and
) and  respectively, where h=pq. The effects of N and p on
respectively, where h=pq. The effects of N and p on  are shown in Figure 1. As can be seen,
are shown in Figure 1. As can be seen,  , unlike
, unlike  , is always negative. The magnitude of bias depends on values of N and p. With an increasing sample size N,
, is always negative. The magnitude of bias depends on values of N and p. With an increasing sample size N,  asymptotes to 0, regardless of population allele frequency p, as expected.
asymptotes to 0, regardless of population allele frequency p, as expected.
When allele frequencies are estimated by omitting the focal pair of individuals,  becomes
becomes

where  and
and  are calculated by (6) for m=2–4. When all copies of an allele (A or B) in the sample appear in the focal individuals, the frequency of the allele is estimated to be zero by omitting the focal individuals. In such cases,
are calculated by (6) for m=2–4. When all copies of an allele (A or B) in the sample appear in the focal individuals, the frequency of the allele is estimated to be zero by omitting the focal individuals. In such cases,  becomes undefined and is set to zero. Like (8),
becomes undefined and is set to zero. Like (8),  in (9) does not reduce to zero in general. It varies with both sample size N and allele frequency p. Some numerical examples showing the effects of N and p on
in (9) does not reduce to zero in general. It varies with both sample size N and allele frequency p. Some numerical examples showing the effects of N and p on  are shown in Figure 1. As can be seen, excluding focal individuals in calculating allele frequencies leads to an overestimation of relatedness in general. At the same sample size, more bias is induced by excluding than including focal individuals for allele frequency estimation.
are shown in Figure 1. As can be seen, excluding focal individuals in calculating allele frequencies leads to an overestimation of relatedness in general. At the same sample size, more bias is induced by excluding than including focal individuals for allele frequency estimation.
Lynch and Ritland estimator
For individuals X and Y with genotypes {a, b} and {c, d} respectively, the estimator (Lynch and Ritland, 1999, denoted by LR or  hereafter) is
hereafter) is

for a single locus. For multiple loci, estimates (10) are weighted by the inverses of their sampling variances derived by assuming zero relatedness (Lynch and Ritland, 1999).
For a diallelic locus with equal allele frequencies (p=q=1/2), the denominator 2(1+δab)(pa+pb)−8papb=0 when the reference individual X is a heterozygote and thus δab=0. In such a case, this part of the estimate is undefined and is set to zero. This is also true with individual Y when it is used as the reference.
For the 6 possible genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} at a diallelic locus,  =1, 1, 1,
=1, 1, 1,  ,
,  ,
,  , respectively, calculated by (10). It can be shown that, similar to
, respectively, calculated by (10). It can be shown that, similar to  in (8), the average
in (8), the average  in a sample of N individuals is always zero when population allele frequency p is known and is used in the calculation. It means
in a sample of N individuals is always zero when population allele frequency p is known and is used in the calculation. It means  is unbiased with known allele frequencies.
is unbiased with known allele frequencies.
When allele frequencies are calculated from the sample, the average relatedness among the N sampled individuals is expected to be

where  and
and  are calculated by (2). It can be shown
are calculated by (2). It can be shown  in general. The value of
in general. The value of  varies with both sample size N and population allele frequency p. When N=3 and 4, for example,
varies with both sample size N and population allele frequency p. When N=3 and 4, for example,  and
and  , respectively, where h=pq. The effects of N and p on
, respectively, where h=pq. The effects of N and p on  are shown in Figure 1. As can be seen,
are shown in Figure 1. As can be seen,  can be both negatively and positively biased, depending on values of N and p. With an increasing sample size N,
can be both negatively and positively biased, depending on values of N and p. With an increasing sample size N,  asymptotes to 0 regardless of population allele frequency p, as expected.
asymptotes to 0 regardless of population allele frequency p, as expected.
When allele frequencies are estimated by omitting the focal pair of individuals,  becomes
becomes

where  and
and  are calculated by (6) for m=2–4. When
are calculated by (6) for m=2–4. When  =0 or
=0 or  =0,
=0,  is undefined and is set to zero. Some numerical examples in Figure 1 show that more bias results from excluding than including focal individuals in calculating allele frequencies.
is undefined and is set to zero. Some numerical examples in Figure 1 show that more bias results from excluding than including focal individuals in calculating allele frequencies.
Lynch and Li estimator
Proposed by Lynch (1988) and improved by Li et al. (1993), this estimator (denoted as LL or  hereafter) calculates the relatedness between individuals X and Y using their similarity index SXY, defined as the average fraction of alleles at a locus in one individual for which there is another allele in the other individual that is identical in state. For individuals X and Y with genotypes {a, b} and {c, d}, respectively, at a locus, the similarity index is
hereafter) calculates the relatedness between individuals X and Y using their similarity index SXY, defined as the average fraction of alleles at a locus in one individual for which there is another allele in the other individual that is identical in state. For individuals X and Y with genotypes {a, b} and {c, d}, respectively, at a locus, the similarity index is

where the first and second terms in the brackets give the similarity indexes when individual X and Y are used as the reference, respectively.
The relatedness between individuals X and Y is

where SXY is calculated by (13), and S0 is the expected similarity index for unrelated individuals (Li et al., 1993). For a locus with n alleles of frequencies pi in a population, S0 is calculated by (Li et al., 1993)

where

for m=2, 3. For multiple loci, the sum of the numerators (one for each locus) and the sum of the denominators are calculated before making the division to give the final estimate (Li et al., 1993).
For the 6 possible genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} at a diallelic locus,  , respectively, as calculated by (13), and
, respectively, as calculated by (13), and  =1, 1, 1,
=1, 1, 1,  ,
,  ,
,  , respectively, as calculated by (14). It can be shown, similar to
, respectively, as calculated by (14). It can be shown, similar to  in (8), that the average
in (8), that the average  in a sample of N individuals is always zero, regardless of the value of N, when population allele frequency p is known and is used in the calculation. It means
in a sample of N individuals is always zero, regardless of the value of N, when population allele frequency p is known and is used in the calculation. It means  is unbiased with known allele frequencies.
is unbiased with known allele frequencies.
When allele frequencies estimated from the sample are used in calculating S0, the expected value of S0 is

where  is
is  calculated by (15) and (16) from a sample containing i, j and k (≡N−i−j) genotypes of AA, BB and AB,
calculated by (15) and (16) from a sample containing i, j and k (≡N−i−j) genotypes of AA, BB and AB,  and
and  are calculated by (2). After some algebra,
are calculated by (2). After some algebra,  is simplified to
is simplified to  , where h=pq. Similarly, the average observed similarity, SXY, among the N sampled individuals is expected to be
, where h=pq. Similarly, the average observed similarity, SXY, among the N sampled individuals is expected to be

which simplifies to  . Inserting
. Inserting  and
and  derived above into (14) yields the expected value of the average relatedness among the N sampled individuals,
derived above into (14) yields the expected value of the average relatedness among the N sampled individuals,

Similar to  ,
,  and
and  ,
,  is biased when allele frequencies are estimated from the same sample of individuals whose relatedness is being estimated. The smaller the sample size N is, the larger the bias will be. Different from
is biased when allele frequencies are estimated from the same sample of individuals whose relatedness is being estimated. The smaller the sample size N is, the larger the bias will be. Different from  ,
,  and
and  , however, the bias of
, however, the bias of  depends on N only, is always negative and is independent of the underlying population allele frequencies (p, q). For comparison, some numerical values of
depends on N only, is always negative and is independent of the underlying population allele frequencies (p, q). For comparison, some numerical values of  are shown in Figure 1.
are shown in Figure 1.
It can be derived similarly that, when allele frequencies are calculated from the sample by omitting the two focal genotypes,  ,
,  and
and

Equation (18) shows that  underestimates relatedness, the extent of the underestimation depends on N only and is unaffected by the actual population allele frequencies (p, q). For a given sample size N, more bias is induced by excluding than including the two focal genotypes in allele frequency estimation (Figure 1). When N=10, for example, the bias is −0.0526 and −0.0667 when the two focal individuals are included and excluded in calculating allele frequencies, respectively.
underestimates relatedness, the extent of the underestimation depends on N only and is unaffected by the actual population allele frequencies (p, q). For a given sample size N, more bias is induced by excluding than including the two focal genotypes in allele frequency estimation (Figure 1). When N=10, for example, the bias is −0.0526 and −0.0667 when the two focal individuals are included and excluded in calculating allele frequencies, respectively.
Wang estimator
This estimator (Wang, 2002, denoted by W or  hereafter) uses the same similarity index, (13), defined in estimator LL but can estimate both two- (Φ) and four-gene (Δ) relatedness, and thus the total relatedness r. It is much more complicated than
hereafter) uses the same similarity index, (13), defined in estimator LL but can estimate both two- (Φ) and four-gene (Δ) relatedness, and thus the total relatedness r. It is much more complicated than  . In the case of a diallelic locus, the estimator is
. In the case of a diallelic locus, the estimator is



for individuals X and Y with genotypes {a, b} and {c, d} respectively, where  and
and  when SXY=1 and SXY≠1, respectively, and
when SXY=1 and SXY≠1, respectively, and  and
and  when SXY=3/4 and SXY≠3/4, respectively. It can be shown that
when SXY=3/4 and SXY≠3/4, respectively. It can be shown that  and
and  and
and  are identical for a diallelic locus. However,
are identical for a diallelic locus. However,  and
and  are different for a locus with more than two alleles (Wang, 2002).
are different for a locus with more than two alleles (Wang, 2002).
Estimators (19), (20) and (21) or their multiallelic forms (Wang, 2002) are calculated for a single locus. Following previous work (Ritland, 1996; Lynch and Ritland, 1999, Wang (2002) derived the variances of these estimators by assuming zero relatedness. Weighting single-locus estimates by the inverses of their variances yields multilocus estimators (Wang, 2002).
For a single diallelic locus,  and
and  have identical properties as shown above for the latter. For multiple loci, they are slightly different because different weighting schemes were applied to loci with different allele frequencies (Wang, 2002). It can be shown that, for a sample of N unrelated individuals,
have identical properties as shown above for the latter. For multiple loci, they are slightly different because different weighting schemes were applied to loci with different allele frequencies (Wang, 2002). It can be shown that, for a sample of N unrelated individuals,  when population allele frequencies are known and used in the estimation. However, when allele frequencies are estimated from the sample with the focal individuals either included or excluded,
when population allele frequencies are known and used in the estimation. However, when allele frequencies are estimated from the sample with the focal individuals either included or excluded,  is positively and
is positively and  is negatively biased in general (Supplementary Figure S1). Much of the opposite biases cancel each other that
is negatively biased in general (Supplementary Figure S1). Much of the opposite biases cancel each other that  is much less biased (Figure 1).
is much less biased (Figure 1).
Loiselle estimator
Loiselle et al. (1995) proposed an estimator to calculate the average coancestry among a group of individuals from their marker genotypes. The estimator can also be used for two individuals, as shown by Heuertz et al. (2003), and multiplying the estimator by 2 gives the relatedness for non-inbred individuals. An important characteristic of the estimator, denoted as LS or  hereafter, is that it uses a correction for small sample sizes. For two individuals X and Y in a sample of N individuals genotyped at L loci, the relatedness estimator is
hereafter, is that it uses a correction for small sample sizes. For two individuals X and Y in a sample of N individuals genotyped at L loci, the relatedness estimator is

where Xli (Yli) is the frequency (=1, 0.5, 0) of allele i (=1, 2, …, nl) at locus l (=1, 2, …, L) in individual X (Y), pli is the frequency of allele i at locus l estimated from the sample of N individuals and nl is the number of alleles at locus l. The first term of the estimator gives the relatedness when allele frequencies are either known (that is, not estimated from the sample) or estimated from a large sample (that is, N large). For a single diallelic locus (that is, L=1 and n1=2), it is essentially identical to the estimator of Yang et al. (2011). The second term of the estimator corrects for the bias caused by estimating pli from a small sample of N individuals.
For the 6 possible genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} at a diallelic locus,  ,
,  ,
,  ,
,  ,
,  ,
,  , respectively, when allele frequencies p and q are known (that is, dropping the correction 2/(2N−1) from (22)). It can be shown that the average
, respectively, when allele frequencies p and q are known (that is, dropping the correction 2/(2N−1) from (22)). It can be shown that the average  in a sample of N individuals is always zero when known population allele frequency p is used in the calculation. It means
in a sample of N individuals is always zero when known population allele frequency p is used in the calculation. It means  is unbiased with known allele frequencies.
is unbiased with known allele frequencies.
When allele frequencies are calculated from the sample of N individuals,  for genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} are
for genotype pairs {AA, AA}, {BB, BB}, {AB, AB}, {AA, BB}, {AA, AB} and {BB, AB} are  ,
,  ,
,  ,−C,
,−C,  ,
,  , respectively, where C=4−2/(2N−1). The average relatedness among the N individuals is expected to be
, respectively, where C=4−2/(2N−1). The average relatedness among the N individuals is expected to be

where  and
and  are calculated by (2). It can be shown
are calculated by (2). It can be shown  , despite the correction for sample size N. The value of
, despite the correction for sample size N. The value of  depends on both sample size N and population allele frequency p. When N=3 and 4, for example,
depends on both sample size N and population allele frequency p. When N=3 and 4, for example,  and
and  , respectively, where h=pq. The effects of N and p on
, respectively, where h=pq. The effects of N and p on  are shown in Figure 1. As can be seen,
are shown in Figure 1. As can be seen,  is most often negatively biased. The extent of underestimation depends on values of N and p. The bias of LS estimator is usually smaller than the other estimators for the same values of N and p, thanks to the correction for sample size. With an increasing sample size N,
is most often negatively biased. The extent of underestimation depends on values of N and p. The bias of LS estimator is usually smaller than the other estimators for the same values of N and p, thanks to the correction for sample size. With an increasing sample size N,  asymptotes to 0 regardless of population allele frequency p, as expected.
asymptotes to 0 regardless of population allele frequency p, as expected.
When allele frequencies are estimated by omitting the focal pair of individuals,  becomes
becomes

where  and
and  are calculated by (6) for m=2–4 and C=4−2/(2N−5). The denominator of C becomes 2N−5 because 2 focal individuals are omitted in calculating allele frequencies. Note that
are calculated by (6) for m=2–4 and C=4−2/(2N−5). The denominator of C becomes 2N−5 because 2 focal individuals are omitted in calculating allele frequencies. Note that  and
and  can be zero when the focal pair of individuals are excluded, leaving the estimator undefined. In such cases, the estimator is set to zero.
can be zero when the focal pair of individuals are excluded, leaving the estimator undefined. In such cases, the estimator is set to zero.
Like (23),  in (24) does not reduce to zero but varies with both sample size N and allele frequency p. Some numerical examples showing the effects of N and p on
in (24) does not reduce to zero but varies with both sample size N and allele frequency p. Some numerical examples showing the effects of N and p on  are shown in Figure 1. As can be seen, excluding focal individuals in calculating allele frequencies leads to an overestimation of relatedness in general. At the same sample size, more bias is induced by excluding than including focal individuals for allele frequency estimation. The large adverse effect of omitting focal individuals in calculating allele frequencies is still substantial even when sample size is N=50.
are shown in Figure 1. As can be seen, excluding focal individuals in calculating allele frequencies leads to an overestimation of relatedness in general. At the same sample size, more bias is induced by excluding than including focal individuals for allele frequency estimation. The large adverse effect of omitting focal individuals in calculating allele frequencies is still substantial even when sample size is N=50.
Unbiased estimators
Estimators  and
and  can be modified to become unbiased when population allele frequencies are estimated from the same small sample of individuals whose relatedness is being estimated. Consider a locus with n alleles, and suppose the number of copies of allele i (i=1, 2, …, n) is Ni in a sample of N individuals. The sample allele count configuration is N={N1, N2,…, Nn}, with
can be modified to become unbiased when population allele frequencies are estimated from the same small sample of individuals whose relatedness is being estimated. Consider a locus with n alleles, and suppose the number of copies of allele i (i=1, 2, …, n) is Ni in a sample of N individuals. The sample allele count configuration is N={N1, N2,…, Nn}, with  . The sum of estimated allele frequencies to the mth power, tm[N], can be calculated from the sample N as
. The sum of estimated allele frequencies to the mth power, tm[N], can be calculated from the sample N as

for m=2, 3. Equation (25) corresponds to Equation (16) for the case of known population allele frequencies. It reduces asymptotically to Equation (16) with an increasing sample size N, as expected. It is derived by considering sampling without replacement. Let us consider the estimation of  as an example. The probability that the first gene drawn at random from the sample is of allele type i is
as an example. The probability that the first gene drawn at random from the sample is of allele type i is  . Given the first allele i, the probability that the second gene drawn at random from the remaining sample is also of allele type i is
. Given the first allele i, the probability that the second gene drawn at random from the remaining sample is also of allele type i is  . Therefore, the probability of sampling two alleles of type i from the sample without replacement is
. Therefore, the probability of sampling two alleles of type i from the sample without replacement is  . Summing
. Summing  over i for i=1–n gives
over i for i=1–n gives  of Equation (25) for m=2. Similarly,
of Equation (25) for m=2. Similarly,  is derived by considering the probability of sampling three genes of the same allele type without replacement from sample N.
is derived by considering the probability of sampling three genes of the same allele type without replacement from sample N.
Using  calculated by Equation (25) instead of tm calculated by Equation (16) leads to an unbiased LL estimator. For a diallelic locus, the expected value of S0 for a sample of N individuals is
calculated by Equation (25) instead of tm calculated by Equation (16) leads to an unbiased LL estimator. For a diallelic locus, the expected value of S0 for a sample of N individuals is

where i+j+k≡N and  is calculated by (25). After some algebra,
is calculated by (25). After some algebra,  is simplified to
is simplified to  , where h=pq. The average observed similarity, SXY, among the N sampled individuals is obtained in deriving (17), which is
, where h=pq. The average observed similarity, SXY, among the N sampled individuals is obtained in deriving (17), which is  Inserting
Inserting  and
and  into estimator
into estimator  (14) leads to
(14) leads to  , irrespective of sample size N, and population allele frequencies p and q.
, irrespective of sample size N, and population allele frequencies p and q.
Similarly, using  by Equation (25) instead of t2 calculated by Equation (16) leads to unbiased
by Equation (25) instead of t2 calculated by Equation (16) leads to unbiased  . However,
. However,  and
and  are still biased in opposite directions. Their effects on
are still biased in opposite directions. Their effects on  cancel out exactly such that
cancel out exactly such that  is always unbiased. From hereafter, the modified LL and W estimators calculated by using (25) are denoted as LLM or
is always unbiased. From hereafter, the modified LL and W estimators calculated by using (25) are denoted as LLM or  and WM or
and WM or  , respectively.
, respectively.
Simulations
Simulations were conducted to check the above analytical results, and to investigate other cases such as multiallelic locus, multiple loci and a mixed sample containing both unrelated and closely related individuals. A sample of N individuals was drawn from a large outbred population at Hardy–Weinberg equilibrium and linkage equilibrium. Two types of samples were considered. For an unrelated sample, all pairs of sampled individuals were unrelated, as assumed in the analytical study above. For a mixed sample, one pair of individuals were related as full sibs (FS), half sibs (HS) or parent offspring (PO) and the rest of the pairs were unrelated (UR). Each sampled individual was genotyped at a number of L loci, and each locus had a fixed number of n codominant alleles with a uniform, equal or triangular frequency distribution in the population.
All of the sampled genotypes were used in calculating allele frequencies (that is, no omitting of the focal pair of individuals) and relatedness estimators. For each parameter combination, R=105 replicate data sets were simulated and analysed. The quality of a relatedness estimator was measured by its bias and accuracy RMSE (root mean squared errors),


for each relationship (FS, HS, PO, UR), where M is the number of pairs of individuals in a sample having the relationship, r is the true value and  is the estimated value of relatedness. The simulated true value of r is 0.5, 0.25, 0.5 and 0 for FS, HS, PO and UR, respectively.
is the estimated value of relatedness. The simulated true value of r is 0.5, 0.25, 0.5 and 0 for FS, HS, PO and UR, respectively.
The simulations (Figure 2 and Supplementary Figure S2) confirm the analytical results (above) that all estimators give biased r estimates for different types of relationships (FS, HS, PO, UR) when the same genotype data of a small sample of individuals are used to calculate allele frequencies and relatedness. With a uniform allele frequency distribution for each of L=10 000 SNPs, relatedness is always underestimated except for the case of LS estimator and UR, regardless of sample sizes in the range (4, 64), estimators and types of relationships. The underestimation increases rapidly with a decreasing sample size. When N=4, for example, the means of LL (or W), QG, R, LR and LS estimators are 0.41, 0.20, 0.05, 0.07 and 0.36 for FS dyads; 0.13, −0.09, −0.13, −0.17 and 0.18 for HS dyads; and −0.18, −0.43, −0.34, −0.41 and 0.06 for UR dyads. Results for PO dyads are similar to those for FS dyads.
Means and RMSEs of different estimators as a function of sample size (N). The genotype data of N individuals at a number of 10 000 SNPs with uniform allele frequency distribution were simulated and used to estimate allele frequencies and the relatedness of full-sib (FS, r=0.5), parent–offspring (PO, r=0.5), half-sib (HS, r=0.25) and unrelated (UR, r=0) pairs of individuals.
Relatively, LL (or W) and LS have smaller biases than QG, R and LR for different relationships (FS, HS, PO, UR). The bias correction, 2/(2N−1), for small sample size N does not ensure LS is unbiased. However, the correction works well and reduces the bias of the estimator substantially for all types of relationships (FS, HS, PO, UR). Without the correction, LS would have been highly biased for small samples, just like QG, R and LR. As shown in Figure 2 (and Supplementary Figure S2), the extent of bias varies with the true relatedness. The relatedness of closely related individuals (for example, FS and PO) tends to be much more underestimated than that of unrelated individuals (UR). As a result, no single correction in terms of N exists that can make an estimator unbiased for all types of relationships. The correction of LS results in underestimated and overestimated relatedness for closely related (for example, FS) and unrelated (UR) individuals in a mixed sample. Overall, the correction 2/(2N−1) makes the LS estimator less biased and more accurate than most of the unmodified estimators when the sample is small (Figure 2).
All estimators become less biased with an increase in sample size N. However, the rate of decline in bias with N is slow. Even at a reasonably large sample size of N=64 individuals, QG, R and LR estimators still underestimate relatedness slightly, giving an average r estimate of ∼0.48, 0.23 and −0.02 for FS (or PO), HS and UR dyads, respectively. The bias patterns of different estimators for multiallelic loci (Supplementary Figure S2) are generally similar to those for diallelic loci (Figure 2). The accuracy patterns, however, are different because, owing to the huge difference in the number of loci (L), accuracy (measured by RMSE) is mainly determined by bias and sampling variance for the diallelic (L=10 000 in Figure 2) and multiallelic (L=20 in Supplementary Figure S2) cases. With an increase in L, bias should be increasingly more important than variance in determining accuracy, regardless of the number and frequency distribution of alleles per locus, and the accuracy pattern for multiallelic loci shown in Supplementary Figure S2 for L=20 should approach that for diallelic loci shown in Figure 2 for L=10 000.
The modified estimators, LLM and WM, are almost unbiased, irrespective of N and types of relationships (Figure 2). Because of the much reduced bias and some reduction in sampling variance (see Supplementary Figure S2), LLM and WM are much more accurate than the original estimators when sample size N is small, except for the case of low relatedness (HS, UR) and few loci (Supplementary Figure S2). The RMSEs of LLM and WM can be smaller than those of other estimators by several orders when N is small and true relatedness is high (Figure 2 and Supplementary Figure S2).
Analysis of an empirical data set
To investigate the genetic structure of Atlantic salmon in the entire North American range of the species, Moore et al. (2014) sampled 1080 individuals from 50 populations and genotyped each individual at 3192 SNP loci. Individuals sampled from within a population were not studied for relatedness. If they were, relatedness estimates would be substantially biased because the sample size for each population is only ⩽25 individuals.
To demonstrate the bias of the original estimators and the sample size-independent properties of the modified estimators, a sample of 25 individuals taken from a single population was analysed. First, the 25 individuals were used to calculate allele frequencies at each locus, and these estimated frequencies were used to obtain pairwise relatedness estimates. Second, the 25 individuals were partitioned into 5 non-overlapping subsamples, each containing 5 individuals. Each subsample was then analysed for allele frequencies that were then used in calculating relatedness. If an estimator is robust to small sample size, then relatedness estimates for a given dyad obtained from the original sample (25 individuals) and from the subsamples (each having 5 individuals) should be similar.
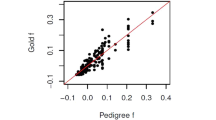
Figure 3 (see also Supplementary Figure S3) plots these estimates for different estimators. The modified estimators, WM and LLM, and the estimator with bias correction, LS, give very similar, although not identical, estimates calculated from the subsamples and the original sample. Most of the points (each showing the relatedness estimates of a dyad calculated from the original sample and a subsample) are centred on the diagonal line (Figure 3), and there is no obvious trend that estimates from the subsamples are uniformly smaller or larger than those from the original sample. In contrast, all pairwise estimates obtained from subsamples are much smaller than estimates from the original sample for each of the five unmodified estimators without bias correction. For example, the relatedness of a highly related dyad was estimated by WM, LLM, LS, LR, R and QG to be 0.44, 0.44, 0.23, 0.20, 0.13 and 0.36, respectively, in the original sample, and to be 0.38, 0.39, 0.24, −0.02, −0.01 and 0.11, respectively, in the subsamples. A decrease in sample size reduces the original estimators without bias correction by ∼0.2, reduces the modified estimators only by 0.05 and increases LS estimator by ∼0.01. Despite that LS gives consistent estimates that are little affected by sample size, it could underestimate the relatedness of close relatives (as shown in simulations in Figure 2 and Supplementary Figure S2). All estimates from LS tend to shrink toward 0, with the highest and lowest related dyads whose MW estimates are 0.44 and −0.25 having LS estimates of 0.24 and −0.05, respectively.
Scatter graph of relatedness estimates obtained from subsamples (N=5, y axis) and the original sample (N=25, x axis) of a salmon data set. An original sample of 25 individuals was taken from a single population, with each individual genotyped at 3192 SNP loci. Five non-overlapping subsamples, each having five individuals, were obtained from the original sample. Each point plots the relatedness estimates for each of 50 dyads obtained from an estimator using the original sample (x axis) and a subsample (y axis). The thin diagonal line shows the ideal case when relatedness estimates made from the original sample and subsamples are equal across the 50 dyads. Estimators W and LL are not shown in the figure because they are similar to estimators LR, R and QG. Instead, the modified estimators WM and LLM are shown in the figure. The slope and intercept of the six estimators are 0.775 and 0.017 for WM, 0.848 and 0.001 for LLM, 1.046 and −0.207 for LR, 1.202 and −0.172 for R, 0.944 and −0.213 for QG and 1.034 and 0.001 for LS.
Discussion
Estimating pairwise relatedness from genetic marker data is now a routine analysis in molecular ecology, evolutionary biology and conservation studies. The estimators developed for this purpose invariably assume that population allele frequencies of markers are known without errors, and the behaviours of these estimators were usually investigated under this assumption (see, for example, Lynch and Ritland, 1999; Wang, 2002; Milligan, 2003). Unfortunately, however, population allele frequencies are rarely known in reality. Frequently, the only data one has in a relatedness analysis are a sample of multilocus genotypes. In such a case, we have to calculate both allele frequencies and relatedness from the same sample. Furthermore, because of various constraints, sample sizes of individuals (or numbers of genotypes at a locus, to be precise) can be quite small.
Current relatedness estimators were developed in the pre-genomic era mainly for application to microsatellite data. Although their bias due to small sample size is well recognized (see, for example, Queller and Goodnight, 1989; Ritland, 1996; Lynch and Ritland, 1999), it is deemed unimportant except when sample size N is small (say, N<100; Ritland, 1996). Furthermore, with just L=10–30 microsatellites typically used in a relatedness analysis, the accuracy is dominated by sampling variance rather than bias even when N is small. In the genomic era, however, the N»L situation is reversed; a typical large-yet-sparse data set given by next-generation sequencing could have millions of SNP loci, with each having a small number of genotypes because of a small number of sampled individuals and a high rate of missing data. This study showed, for the first time, that the popular relatedness estimators can become highly biased and their accuracy is dominated by bias rather than sampling error when they are applied to such SNP data sets (that is, L»N). The direction (that is, over- or under-estimation) and extent of bias depends on sample sizes, the underlying (unknown) population allele frequencies, the estimators and the true relatedness. With regard to sample size N, the bias is roughly on the order of 1/N. For example, the relatedness of first-degree relatives (PO, FS) is expected to be 0.5. However, it is estimated on average to be ∼0.27 by R estimator (Figure 2) when N=10. As a possible consequence, first-degree relatives may be mistaken as second-degree relatives if one is unaware of the bias. The same is observed in the analysis of the salmon SNP data set (Figure 3 and Supplementary Figure S3).
This study also showed that omitting the focal individuals in calculating allele frequencies, as suggested in the literature (see, for example, Queller and Goodnight, 1989; Lynch and Ritland, 1999), cannot remove the bias of popular relatedness estimators. On the contrary, this ad hoc treatment in estimating allele frequencies not only causes a high frequency of undefined estimators but also induces more biased estimates (Figure 1). This is perhaps not too surprising. At a small sample size, allele frequencies are estimated without bias by allele counting method, although estimates of higher-order terms of the frequencies can be biased (Nei and Chesser, 1983; Weir, 1996). When a focal pair of individuals is omitted, however, both allele frequencies and their higher order terms are biased, leading to worse estimates of relatedness than those when all sampled individuals are used in calculating allele frequencies.
Among the estimators investigated in this study, the one described in Loiselle et al. (1995), LS, is the only one that uses a correction for small sample sizes. Both analytical and simulation results show that, compared with other estimators, LS has substantially reduced biases for all types of relationships and for different sample sizes. As a result, it is more accurate than most of the unmodified estimators (Figure 2 and Supplementary Figure S2). However, the correction is insufficient to make the estimator unbiased (Figure 1). In a mixed sample containing both related and unrelated individuals, LS tends to underestimate and overestimate relatedness for related and unrelated individuals, respectively. This is not surprising because the extent of bias of a relatedness estimator varies with true relatedness, and it is impossible to apply a single correction for small sample size, such as  , to obtain unbiased relatedness estimates for all possible relationships. In contrast, the modified estimators,
, to obtain unbiased relatedness estimates for all possible relationships. In contrast, the modified estimators,  and
and  , are almost unbiased for all relationships, sample sizes and allele frequency distributions.
, are almost unbiased for all relationships, sample sizes and allele frequency distributions.
Estimating both allele frequencies and relatedness from the same sample has three problems (see Introduction; Wang, 2014). This study has addressed the third problem, underestimation of relatedness due to small sample sizes. The first problem (that is, negative relatedness estimates, mean of relatedness estimates across dyads in a sample being close to zero) is no longer pertinent when relatedness is defined, understood and used in terms of a correlation coefficient rather than a probability of IBD (Wright, 1965; Wang, 2014). The second problem comes from the genetic structure of a sample, no matter whether it is small or large. When a sample containing both related and unrelated individuals is used in calculating allele frequencies by (naively) assuming unrelated individuals, relatedness will be underestimated because of the biased allele frequency estimates. Indeed, my simulation in Figure 2 shows that the modified estimators, WM and LLM, underestimate r for all relationships (FS, PO, UR, …) when sample size is extremely small such that sample genetic structures become substantial. However, the bias is rather small. The smallest sample in Figure 2 has N=4 individuals or 6 dyads. For the case of FS, the sample contains one FS dyad and five UR dyads, the FS dyad frequency being 1/6=0.167. Despite the high FS frequency, however, the biases of LLM and WM are rather small. For example, the mean LLM estimates are 0.48 and −0.04 for FS and UR, respectively. I also simulated even smaller samples, with each containing two full siblings and one unrelated individual and thus a proportion of 1/3=0.33 full-sib dyads. The mean LLM estimates are 0.46 and −0.06 for FS and UR, respectively, the biases being still reasonably small. Compared with the huge bias caused by small sample size as shown in this study, the bias caused by the genetic structure of a sample is negligible.
This study modified the LL and W estimators and showed, using analytical (Figure 1), simulated (Figure 2 and Supplementary Figure S2) and empirical data (Figure 3 and Supplementary Figure S3), that relatedness can be reliably estimated by the modified estimators with little bias even when sample size is extremely small (say, 3 individuals). Because of the great reduction in bias and some decrease in sampling variance (Supplementary Figure S2), the modified estimators are always much more accurate (that is, smaller RMSE) than the original estimators, except when few loci are used (such that RMSE is dominated by sampling variance rather than bias) and true relatedness is low (for example, UR). The smaller the sample size is, the greater the accuracy improvements the modified estimates make. When sample sizes are large or when population allele frequencies are known, the relative performances of different estimators depend on the true relationship being estimated. Whereas LL and W usually give the best estimates for highly related dyads (for example, FS and PO), LR and R usually give the most accurate estimates for unrelated or loosely related dyads (for example, UR) (Wang, 2002; Thomas, 2010). When sample sizes are small and many loci are used, however, the modified estimators, WM and LLM, always perform better than all of the original estimators, regardless of the actual relatedness being estimated. The modified estimators are now implemented in the software package Coancestry (https://www.zsl.org/science/software/coancestry).
This study assumes an outbred population in which close inbreeding, due to close relative mating such as sib mating and selfing, is absent or rare. All estimators investigated in this study are valid under this assumption, whereas estimators LS and R do not require the assumption and apply to both outbred and inbred populations (Wang, 2007). Similarly, genotyping artefacts causing excessive individual homozygosity, such as allelic dropouts and null alleles in microsatellite data and SNPs genotypes called from low-coverage next-generation sequencing data, could affect the accuracy of estimators WM, LLM, LL, W, LR and QG, but not of estimators LS and R (Wang, 2007). However, all estimators are robust to pervasive inbreeding (that is, due to genetic drift from the finite size or structure of a population), and to low levels of excessive homozygosity due to either close inbreeding or genotyping artefacts (say, <10% increase in homozygosity over that expected under Hardy–Weinberg equilibrium), as demonstrated for some of the estimators before (Wang, 2007). When close inbreeding is deemed important in a population, then estimators LS and R could be preferred over other estimators.
Data archiving
All simulated data and the simulation code are available upon request to the author.
References
Anderson AD, Weir BS . (2007). A maximum-likelihood method for the estimation of pairwise relatedness in structured populations. Genetics 176: 421–440.
DeWoody JA . (2005). Molecular approaches to the study of parentage, relatedness, and fitness: practical applications for wild animals. J Wildl Manage 69: 1400–1418.
Falconer DS, Mackay TFC . (1996) Introduction to Quantitative Genetics. 4th edn. Longman: Harlow, UK.
Garant D, Kruuk LE . (2005). How to use molecular marker data to measure evolutionary parameters in wild populations. Mol Ecol 14: 1843–1859.
Hardy OJ, Vekemans X . (2002). SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Res 2: 618–620.
Heuertz M, Vekemans X, Hausman JF, Palada M, Hardy OJ . (2003). Estimating seed vs. pollen dispersal from spatial genetic structure in the common ash. Mol Ecol 12: 2483–2495.
Kardos M, Luikart G, Allendorf FW . (2015). Measuring individual inbreeding in the age of genomics: marker-based measures are better than pedigrees. Heredity 115: 63–72.
Li CC, Horvitz DG . (1953). Some methods of estimating the inbreeding coefficient. Am J Hum Genet 5: 107–117.
Li CC, Weeks DE, Chakravarti A . (1993). Similarity of DNA fingerprints due to chance and relatedness. Hum Hered 43: 45–52.
Loiselle BA, Sork VL, Nason J, Graham C . (1995). Spatial genetic structure of a tropical understory shrub, Psychotria officinalis Rubiaceae. Am J Bot 82: 1420–1425.
Lynch M . (1988). Estimation of relatedness by DNA fingerprinting. Mol Biol Evol 5: 584–599.
Lynch M, Ritland K . (1999). Estimation of pairwise relatedness with molecular markers. Genetics 152: 1753–1766.
Lynch M, Walsh JB . (1998) Genetics and Analysis of Quantitative Traits. Sinauer Associates: Sunderland, MA.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ et al. (2009). Finding the missing heritability of complex diseases. Nature 461: 747–753.
Milligan BG . (2003). Maximum-likelihood estimation of relatedness. Genetics 163: 1153–1167.
Moore JS, Bourret V, Dionne M et al. (2014). Conservation genomics of anadromous Atlantic salmon across its North American range: outlier loci identify the same patterns of population structure as neutral loci. Mol Ecol 23: 5680–5697.
Nei M, Chesser RK . (1983). Estimation of fixation indices and gene diversities. Ann Hum Genet 47: 253–259.
Queller DC, Goodnight KF . (1989). Estimating relatedness using molecular markers. Evolution 43: 258–275.
Ritland K . (1996). Estimators for pairwise relatedness and inbreeding coefficients. Genet Res 67: 175–186.
Thomas SC . (2010). A simplified estimator of two and four gene relationship coefficients. Mol Ecol Res 10: 986–994.
Van de Casteele T, Galbusera P, Matthysen E . (2001). A comparison of microsatellite-based pairwise relatedness estimators. Mol Ecol 10: 1539–1549.
Wang J . (2002). An estimator for pairwise relatedness using molecular markers. Genetics 160: 1203–1215.
Wang J . (2007). Triadic IBD coefficients and applications to estimating pairwise relatedness. Genet Res 89: 135–153.
Wang J . (2011a). COANCESTRY: a program for simulating, estimating and analysing relatedness and inbreeding coefficients. Mol Ecol Res 11: 141–145.
Wang J . (2011b). Unbiased relatedness estimation in structured populations. Genetics 187: 887–901.
Wang J . (2014). Marker‐based estimates of relatedness and inbreeding coefficients: an assessment of current methods. J Evol Biol 27: 518–530.
Wang J . (2016). Pedigrees or markers: Which are better in estimating relatedness and inbreeding coefficient? Theor Popul Biol 107: 4–13.
Weir BS . (1996) Genetic Data Analysis II: Methods for Discrete Population Genetic Data. Sinauer Associates: Sunderland, MA, USA.
Weir BS, Anderson AD, Hepler AB . (2006). Genetic relatedness analysis: modern data and new challenges. Nat Rev Genet 7: 771–780.
Wright S . (1921). Systems of mating. Genetics 6: 111–178.
Wright S . (1922). Coefficients of inbreeding and relationship. Am Nat 61: 330–338.
Wright S . (1965). The interpretation of population structure by F‐statistics with special regard to systems of mating. Evolution 19: 395–420.
Yang J, Lee SH, Goddard ME, Visscher PM . (2011). GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88: 76–82.
Acknowledgements
I am grateful to the editor, Hardy OJ, and three anonymous referees for insightful and constructive comments and suggestions on earlier versions of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Wang, J. Estimating pairwise relatedness in a small sample of individuals. Heredity 119, 302–313 (2017). https://doi.org/10.1038/hdy.2017.52
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.52
This article is cited by
-
Vocal convergence and social proximity shape the calls of the most basal Passeriformes, New Zealand Wrens
Communications Biology (2024)
-
Remnant kenngoor (Phascogale calura) retain genetic connectivity and genetic diversity in a highly fragmented landscape
Conservation Genetics (2024)
-
Maximizing Eucalyptus pilularis progeny selection using a parentage matrix obtained with microsatellite markers
Euphytica (2024)
-
Genetic diversity, genetic differentiation and demographic history of Cryptomeria (Cupressaceae), a tertiary relict plant in East Asia based on RAD sequencing
European Journal of Forest Research (2024)
-
SNP heterozygosity, relatedness and inbreeding of whole genomes from the isolated population of the Faroe Islands
BMC Genomics (2023)