Abstract
The Scandinavian wolf population descends from only five individuals, is isolated, highly inbred and exhibits inbreeding depression. To meet international conservation goals, suggestions include managing subdivided wolf populations over Fennoscandia as a metapopulation; a genetically effective population size of Ne⩾500, in line with the widely accepted long-term genetic viability target, might be attainable with gene flow among subpopulations of Scandinavia, Finland and Russian parts of Fennoscandia. Analytical means for modeling Ne of subdivided populations under such non-idealized situations have been missing, but we recently developed new mathematical methods for exploring inbreeding dynamics and effective population size of complex metapopulations. We apply this theory to the Fennoscandian wolves using empirical estimates of demographic parameters. We suggest that the long-term conservation genetic target for metapopulations should imply that inbreeding rates in the total system and in the separate subpopulations should not exceed Δf=0.001. This implies a meta-Ne of NeMeta⩾500 and a realized effective size of each subpopulation of NeRx⩾500. With current local effective population sizes and one migrant per generation, as recommended by management guidelines, the meta-Ne that can be reached is ~250. Unidirectional gene flow from Finland to Scandinavia reduces meta-Ne to ~130. Our results indicate that both local subpopulation effective sizes and migration among subpopulations must increase substantially from current levels to meet the conservation target. Alternatively, immigration from a large (Ne⩾500) population in northwestern Russia could support the Fennoscandian metapopulation, but immigration must be substantial (5–10 effective immigrants per generation) and migration among Fennoscandian subpopulations must nevertheless increase.
Similar content being viewed by others
Introduction
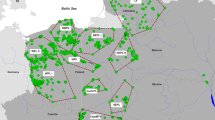
The gray wolf (Canis lupus) was hunted to extinction in Scandinavia (Norway and Sweden; Figure 1) during the mid-1900s, and the present population was established in the early 1980s. This population descends from few individuals and has been highly isolated with census size remaining low for decades (Laikre et al., 2013), which has resulted in extreme inbreeding and inbreeding depression (Liberg et al., 2005; Räikkönen et al., 2013). The conservation case of the Scandinavian wolves is well known scientifically and politically (Liberg et al., 2005; Darpö, 2011; Laikre et al., 2013; Chapron, 2014). In this paper we address the topical issue of if and how conservation genetic goals with respect to genetically effective population size can be reached via increased connectivity with wolf populations further east.
The area of Fennoscandia (medium gray) comprises Norway and Sweden (Scandinavia), Finland and the Russian regions of Kola and Karelia. Detailed wolf distribution data are available for Finland and Scandinavia, and the dark gray areas indicate current wolf occurrence. Such information is missing for Karelia and Kola and it is thus unclear where in those areas wolves occur (marked with stripes). Numbers indicate estimated census sizes (cf. Supplementary Appendices 1 and 2).
The Scandinavian wolf population originates from a founder pair from Finland/Russia that had their first litter in Sweden in 1983. The population has been closely monitored; tracking data combined with molecular genetic analyses of biological material have provided an almost complete pedigree of the population (Vilà et al., 2003; Liberg et al., 2005). After the establishment in 1983 and up to 2012, a total of 18 additional wolves are known to have immigrated into Scandinavia from Finland/Russia, but only three of these have managed to reproduce; one male that first reproduced in 1990, and two males that both first reproduced in 2008. Thus, there is a total of five founders of the present population (Åkesson et al., 2014; Bruford, 2015; Mills and Feltner, 2015).
Population levels remained low during the first decades with substantial poaching, and authorities granting permissions to kill individuals considered to cause problems. Poaching is also viewed as the main reason for the restricted gene flow from populations in Finland and further east (Liberg et al., 2012). This has resulted in very high inbreeding levels, and the current c. 400 Scandinavian wolves are on average as inbred as offspring from full sib matings (f≈0.25; Jansson et al., 2015). Indications of inbreeding depression manifested as reduced litter size and a high frequency of spinal disorders has been reported (Liberg et al., 2005; Räikkönen et al., 2013).
Swedish authorities have issued several licensed wolf hunts during the past few years (SEPA Decisions 2009), which have been strongly criticized by the European Commission (Potočnik, 2011; EC, 2015) because within the European Union (EU), where Sweden is a member, the wolf is listed as a priority species under the conservation legislation of the EU Habitats Directive (Council Directive 92/43/EEC). Similarly, according to national policy, Sweden is committed to maintain naturally occurring species at population sizes large enough to permit harboring genetic variation for long-term persistence (Swedish Environmental Objectives, Government Proposition 2009/10:155). The genetic situation of the Scandinavian wolves has been discussed for decades both nationally and internationally (Laikre, 1999; Liljelund, 2011; Laikre et al., 2013; Bruford, 2015; Chapron, 2015), particularly due to the EU legal discussions (Darpö, 2011; EEB, 2015; EC, 2015).
The Swedish Parliament recently decided that a ‘Favourable Reference Population’ (FRP) for the Swedish wolf population is in the interval 170–270 animals (Swedish Parliament Decision 2013/14:99). FRP is a concept under the EU Habitats Directive referring to the population size ‘considered the minimum necessary to ensure long-term viability of the species’ (Evans and Arvela, 2011). The Swedish Parliament commissioned the Swedish Environmental Protection Agency (SEPA) to make the final decision regarding where in this interval FRP is reached, and the SEPA decided 270 (SEPA PM 2013-12-16, NV-09661-12).
Both scientists and policy makers have suggested that if long-term viability is to be achieved for the Scandinavian wolf population it must be managed as a non-isolated part of a subdivided population interconnected via gene flow covering geographic areas over multiple countries (Laikre and Ryman, 2009; Hansen et al., 2011; Liljelund, 2011; Laikre et al., 2013; Bruford, 2015). We refer to such a series of interconnected subpopulations as a ‘metapopulation’ in line with the broad definition by Hastings and Harrison (1994; ‘any set of conspecific populations linked by dispersal’).
A geographic area proposed as reasonable for harboring a genetically viable wolf metapopulation refers to Fennoscandia (Hansen et al., 2011), which includes Norway, Sweden, Finland and the Russian areas of Karelia and Kola (Figure 1). Requirements for a Fennoscandian wolf metapopulation in terms of subpopulation sizes and degree of connectivity to meet long-term conservation goals regarding effective population size (Ne) have not been dealt with, however. The aim of the present paper is to address this issue. The current EU management recommendation on connectivity relies on the one-migrant-per-generation rule (OMPG; Franklin, 1980; Mills and Allendorf, 1996; Wang, 2004). This rule suggests that one genetically effective immigrant per generation and subpopulation is needed to prevent excessive inbreeding (Linnell et al., 2008).
The genetically effective population size (Ne) is the most widely used parameter for quantifying and monitoring rates of inbreeding and loss of genetic variation through drift. The concept was developed for single, isolated populations; Ne is defined as the size of an ideal population without the evolutionary forces of mutation, selection or migration that exhibits the same expected rate of genetic drift or inbreeding per generation as the actual, real life population of interest (Wright, 1931). In conservation genetics the ‘50/500 rule’ has become widely established suggesting that for a single isolated population Ne>50 is needed for short-term conservation and Ne>500 for long-term conservation (Franklin, 1980; Allendorf and Ryman, 2002; Jamieson and Allendorf, 2012).
Considerable research has been devoted to modeling Ne of subdivided populations, but most of these efforts have used simplifying assumptions such as subpopulations of equal size, standard models of migration and drift–migration equilibrium (Wright, 1938; Maruyama and Kimura, 1980; Whitlock and Barton, 1997; Nunney, 1999; Wang and Caballero, 1999; Tufto and Hindar, 2003; Waples, 2010). Results from that work include demonstrating that under the conditions of Wright’s island model global Ne can increase above the sum of the local Nes, while deviations from these idealized conditions with respect to asymmetrical migration patterns and/or asymmetries in population size can reduce the total effective size below the sum of the Nes of the separate subpopulations (Whitlock and Barton, 1997; Nunney, 1999; Tufto and Hindar, 2003).
We have recently developed a general analytical framework for exploring inbreeding dynamics and effective population sizes in more complex metapopulations than has previously been theoretically or computationally feasible (Hössjer et al., 2014, 2015). Our analytical approach allows modeling systems with any number of subpopulations of arbitrary census and effective size. Migration patterns are also optional, as are initial degrees of inbreeding and relatedness (coefficient of kinship; Wright, 1931) within and among populations.
Here we use this new analytical framework to first suggest how to apply the Ne⩾500 rule to a subdivided population. We suggest new criteria for long-term conservation goals that take both subpopulations and the metapopulation as a whole into account. In a second step, we model the case of the Fennoscandian wolf to address the following questions: (i) what metapopulation effective size can currently be reached over Fennoscandia based on empirical estimates of parameters of interest? (ii) What is needed in terms of population sizes and migration rates for the Fennoscandian wolf to attain the suggested long-term conservation genetic goal for a subdivided population?
Materials and methods
We used the mathematical framework of Hössjer et al. (2014, 2015) that allows modeling the expected change of inbreeding and effective size of arbitrary metapopulations. Key expressions applied here include equation 25 of Hössjer et al. (2014) and equation 48 of Hössjer et al. (2015). Parameter notations used in the present paper are given in Table 1. We considered a diploid organism without selfing where mating occurs after migration. Migration is stochastic, and rates are expressed as the actual number of immigrants per generation (m) from one subpopulation to another (cf. Figure 2). Thus, one migrant per generation, for example, refers to an average of one immigrant per generation based on binomial sampling. Because we focused on issues relating to rates of inbreeding only, the effective population sizes (rather than census sizes) are of concern. Thus, immigrants are regarded as ‘ideal’ individuals with the same characteristics as the ‘ideal’ individuals of the recipient population (Hössjer et al., 2014, 2015), and we ignored the forces of selection and mutation. We also assumed constant population sizes.
Schematic illustration of the Fennoscandian wolf metapopulation used for meta-Ne modeling. Local subpopulation effective sizes are based on empirical estimates (Figure 1; Supplementary Appendices 1 and 2; Supplementary Table 1). Arrows indicate gene flow and m represents the number of migrants per generation; two-headed arrows indicate two-way exchange. In some models one-way migration from Finland to Scandinavia is used (dotted arrow) as suggested by empirical observations. Starting levels of average subpopulation inbreeding are indicated by f. In some models initial f=0 is used for all subpopulations; in other scenarios initial inbreeding levels of the Scandinavian and Finnish subpopulations are based on empirical estimates of fSc=0.25 and fFi=0.1. In the latter cases a kinship coefficient of fSc–Fi=0.05 is also assumed. All other kinship coefficients are assumed to be 0 (zero) in all models. We assume an isolated Fennoscandian metapopulation (a) or one with one-way gene flow from neighboring northwestern Russian provenances (b).
We dealt exclusively with inbreeding effective size (NeI; Wright, 1931, 1938) which quantifies the rate at which inbreeding is increasing. There are also several other types of effective sizes (Crandall et al., 1999; Hössjer et al., 2015) but they are beyond the scope of the present focus. We analyzed how expected values of average inbreeding (f) change over time in the separate subpopulations as well as for the system as a whole under various scenarios of subpopulation sizes and migration rates and patterns. Average inbreeding of the metapopulation (fMeta) corresponds to the probability that the two alleles at a locus are identical by descent in an individual drawn at random from the metapopulation as a whole (here weighting subpopulations according to effective size; Hössjer et al., 2015).
While the short-term conservation genetic rule of thumb Ne⩾50 clearly refers to the inbreeding effective size (Franklin 1980), the Ne⩾500 rule is not directly stated in terms of a particular type of Ne, and Franklin (1980) exclusively considered isolated populations of constant size where inbreeding and variance (NeV) effective sizes always are the same (that is, NeI=NeV, where NeV quantifies the rate of genetic drift, that is, allele frequency change). The Ne⩾500 rule relies on the assumption that the loss of additive genetic variation for quantitative traits due to restricted population size is balanced by mutation when Ne⩾500, and that additive genetic variation is lost at the same rate as heterozygosity (Franklin, 1980).
We quantified expected inbreeding rates in terms of effective population size. Effective size in generation t is defined as Ne=1/(2Δf), where Δf=(ft−ft−1)/(1−ft−1) and ft is the inbreeding coefficient in generation t. In an isolated population of constant size, where Δf is constant and exclusively determined by drift (in the absence of immigration), this definition of Ne corresponds to the traditional one. In contrast, in a population receiving immigrants Δf is affected by both immigration and drift, and the corresponding quantity 1/(2Δf) is referred to here as realized effective size (NeR; Table 1).
The realized effective size of the total metapopulation is denoted NeMeta. This quantity can be computed using various schemes for weighting of the separate Δfs of the different subpopulations (Hössjer et al., 2014, 2015), and here we weighted by local effective size (Nex) such that a large subpopulation contributes more to NeMeta than a small one. Note that Nex is the effective size of subpopulation x if it were completely isolated.
We also computed the eigenvalue effective size NeE (Ewens, 1982; Hössjer et al., 2014), that is the effective size of the metapopulation as a whole when migration–drift equilibrium has been attained. In a metapopulation where each subpopulation both receives immigrants from, and sends emigrants to, the rest of the system (through one or more subpopulations) the rate of inbreeding will eventually be the same (1/(2NeE)) in all subpopulations as well as for the system as a whole (regardless of how the Δfs of the different subpopulations are weighted; Hössjer et al., 2014, equation 61; Hössjer et al., 2015, equation 49).
The Hössjer et al. (2014, 2015) approach allows for specifying degree of relatedness between subpopulation x and y (fxy) at the starting point (generation t=0). Subpopulation census size (Nx), and within subpopulation average inbreeding (fx) and kinship (fxx) affect the upper bound for fxy (Hössjer et al., 2015; equation 2).
Parameterization
We reviewed the literature to obtain empirical estimates from Fennoscandian wolf populations of parameters needed for modeling metapopulation-Ne (Supplementary Appendix 1; Supplementary Table 1). We translated the empirical data into a schematic metapopulation (Figure 2a), with local effective population sizes (Nex) for Scandinavia (Sc), Finland (Fi), Karelia (Ka) and Kola (Kol) of NeSc=100, NeFi=50, NeKa=50 and NeKol=25, respectively. All but the NeKol have an empirical basis; for Kola we used the census estimate of 100 and assumed an Ne/N ratio of 0.25 as estimated for the Scandinavian population (Supplementary Table 1).
We analyzed scenarios both with and without initial inbreeding in the system (Figure 2). With initial inbreeding we used starting point inbreeding averages of fSc=0.25, fFi=0.10, fKa=0 and fKol=0. We used fKa=0 rather than 0.07 estimated from FIS (Supplementary Table 1) because that estimate did not deviate significantly from zero (Jansson et al., 2012). With respect to kinship among populations no direct estimates exist as far as we have been able to find; thus, we used approximations of fxy=0 between all populations except between Scandinavia and Finland where we used fSc–Fi=0.05 for those scenarios assuming initial inbreeding (Figure 2) because the Scandinavian founders have immigrated from Finland (Supplementary Appendix 1).
With respect to migration we assumed m=1 as a basis. Immigration from Finland to Scandinavia (mFi–Sc) has been monitored intensely and indicates mFi-Sc≈1 (Supplementary Appendix 1; Supplementary Table 1). With respect to migration from Scandinavia to Finland (mSc–Fi) only three cases have been documented and two of these migrants were killed before being able to reproduce; the location of the third one—with radio collar—is unknown (Supplementary Appendix 1; Dr Jouni Aspi, University of Oulu, Finland pers. com.) suggesting mSc–Fi=0. Empirical data support immigration between Finland and Karelia of mFi–Ka=mKa–Fi≈1–3 but with a recent decrease (Supplementary Table 1; Aspi et al., 2009; Jansson et al., 2012). No data appear to exist on migration to and from Kola. In addition to empirical support for mxy≈1, the OMPG referred to in EU management guidelines for large carnivores (Linnell et al., 2008) stresses the need for evaluating the effect on meta-Ne under this migration rate.
Results
General aspects of metapopulation-Ne in conservation
First, some general aspects of effective population size in the context of metapopulations should be highlighted for clarity. Ne is only defined for populations where (i) there exists genetic variation and (ii) inbreeding increases or remains constant (when Ne=∞). For populations with no genetic variation or with a decreasing level of inbreeding Ne is undefined. In a metapopulation, inbreeding level of a separate subpopulation can decrease as an effect of immigration, and in such situations the realized effective size of the receiving subpopulation (NeR) becomes undefined. For instance, if subpopulation x with fx>0 starts receiving immigrants from an unrelated donor population y (fxy=0) and with an immigration rate large enough for fx to decrease, then the realized Ne of subpopulation x (NeRx) becomes undefined. In many situations fx will eventually start to increase again, and at that ‘turning point’, when inbreeding goes from decreasing to increasing, NeRx will become definable again; initially it may be very large because the rate of increase of inbreeding is typically low at the turning point.
Similarly, for a metapopulation where subpopulations are completely isolated from each other (that is, m=0) we have NeE=∞ because the subpopulations will eventually become fixed for different alleles; identity by descent of alleles of different subpopulations will not increase but remain the same generation after generation in the system as a whole. Thus, metapopulation-Ne is irrelevant for systems without migration between subpopulations, and as m approaches 0 and the system is close to migration–drift equilibrium, meta-Ne will approach its undefined state.
Conservation genetic goals for metapopulation-Ne
To our knowledge, conservation genetic goals reflecting long-term genetic viability have not been dealt with specifically for metapopulations. Rather, an implicit suggestion has been that for a subdivided population the same rule of thumb should apply as for a single, isolated one, that is, NeMeta⩾500 should reflect long-term viability for the metapopulation as a whole (Hansen et al., 2011; Laikre et al., 2013). We suggest, however, that NeMeta⩾500 cannot be the only focus for long-term genetic viability of a metapopulation. Rather, the inbreeding rates within the separate subpopulations must also be considered. We base this suggestion on the dynamics of a purely hypothetical subdivided population analyzed as an example here. We consider a subdivided population of constant size that consists of eight subpopulations, two of effective size Ne=50 (subpopulations 1 and 2) and six of Ne=25 (subpopulations 3–8; no figure). In this example, migration follows an island model such that in every generation each subpopulation receives m immigrants drawn at random from an infinitely large migrant pool to which the other subpopulations have contributed equally.
In the absence of subdivision, that is, under panmixia, the effective size of this population is Ne=250, corresponding to the sum of the separate subpopulation effective sizes. Subdivision increases the total effective size (NeMeta), but this increase is rather modest at high migration rates, and NeMeta as well as the realized effective sizes (NeR) of the separate subpopulations will approach the eigenvalue effective size (NeE) within a few generations. For m=5, for example, we have NeE=260, and all the realized effective sizes are within 90% of this value in <15 generations (not shown). At lower migration rates, however, NeE grows progressively larger and the approach to equilibrium is much slower (Figure 3a). At m=1 we have NeE=299 and it takes about 100 generations for the system to approach equilibrium; the corresponding value for m=0.2 (one migrant every fifth generation) is NeE=504 and NeMeta is clearly below this value even after 200 generations (Figure 3a).
Realized (NeR) and eigenvalue (NeE) effective size (a) and inbreeding (b) over 200 generations in a hypothetical subdivided population of constant size that consists of eight subpopulations, two of effective size Ne=50 (subpopulations 1 and 2) and six of Ne=25 (subpopulations 3–8). Note that this is a hypothetical example that does not relate to the Fennonscandian wolves. Migration follows an island model such that in every generation each subpopulation receives on average m immigrants drawn at random from an infinitely large migrant pool to which the other subpopulations have contributed equally. Initial inbreeding and kinship is zero (0) within and between all subpopulations. The two curves in b marked Ne=50 and 500 show the accumulation of inbreeding in isolated populations of that size (short- and long-term targets). Black curves, m=1; gray curves, m=0.2. See text for details.
As indicated in Figure 3, however, NeE and NeMeta are poor indicators of the rate of inbreeding in the separate subpopulations before equilibrium has been attained. At m=0.2 the eigenvalue effective size is NeE=504 for this system, thus reflecting an overall rate of inbreeding of Δf<0.001 per generation, so eventually the long-term genetic target will be reached for the metapopulation. Before this has happened, however, the realized effective size of subpopulations has been low for extended periods of time, resulting in an initial accumulation of inbreeding that for the smaller subpopulations (3–8) and for the system as a whole even exceeds that of the short-term conservation goal (Figure 3b). Thus, although meta-Ne eventually will reach the long-term conservation genetic goal, the inbreeding rates of separate subpopulations will initially (over 50–75 generations) exceed acceptable levels for even short-term conservation goals.
We suggest that the conservation genetic target for metapopulations to reflect long-term genetic viability should imply that the rate of inbreeding in the system as a whole, as well as in the separate subpopulations, should not exceed Δf=0.001 (as for an Ne of 500), that is,
-
1
metapopulation effective size is NeMeta⩾500 and
-
2
realized effective size of each subpopulation equals or exceeds 500 (NeRx⩾500).
Further we suggest that goals 1–2 should be reached within, say, 5–20 generations, that is, within the time frames typically considered in the context of short-term conservation (for example, Allendorf and Ryman, 2002). We now apply this proposed conservation genetic target for long-term viability to the Fennoscandian wolf metapopulation.
Meta-Ne of a closed Fennoscandian population assuming no initial inbreeding
First we modeled a ‘closed’ Fennoscandian population using current estimates of local effective sizes, no inbreeding or kinship, and migration patterns implying that all subpopulations both receive and contribute migrants (that is, two-way migration; Figure 2a; Supplementary Appendix 1; Supplementary Table 1). The results indicate that it is not possible to reach a metapopulation NeMeta⩾500 for this system without a very restricted gene flow that results in excessive rates of inbreeding within the separate subpopulations as well as for the system as a whole (Figure 4). With the very restricted migration rate of 1 migrant every 20th generation (m=0.05), the equilibrium metapopulation Ne is NeE=871 (Figure 4a). However, the system is not even close to this equilibrium within the first 200 generations (corresponding to c. 1000 years assuming a generation length of 5 years as estimated for the Scandinavian population; Forslund, 2009).
Expected realized effective size (NeR; left) and inbreeding (f; right) over 200 generations in a population system of four interconnected wolf populations meant to mimic that of Fennoscandia (cf. Figure 2a). All migration is two way as indicated in Figure 2a, and migration rates are m=0.05 (a, b), 1 (c, d) or 3 (e, f) individuals per generation. Initial inbreeding and kinship is zero (0) between and within all populations. In the right panel (b, d, f) the two curves Ne=50 and 500 show the accumulation of inbreeding in isolated populations of that size (short- and long-term targets).
Further, for m=0.05 metapopulation-Ne (NeMeta) increases from 57 to 161 over the initial period of 200 generations and stays below 115 for the first 100 generations, as do all the subpopulations. Thus, inbreeding rates are much above the long-term target rate over this period, and over the first c. 50 generations inbreeding accumulates almost as fast as for the short-term conservation goal of Ne=50 in the Karelian and Finnish subpopulations, as well as in the metapopulation as a whole (Figure 4b). In Kola, the effect of immigration does not reduce the accumulation of inbreeding below the short-term goal until after more than 100 generations, and the rate of inbreeding in the Scandinavian population is, of course, also much higher than the target of Δf=0.001 for the initial 200 generation period (Figure 4b).
With the higher migration rates of m=1 and 3, the rate of inbreeding drops below the short-term target in all subpopulations, as well as for the system as a whole within the first few generations (Figures 4d and f). However, the long-term target of metapopulation Ne⩾500 is never reached, as NeE=250 and 233 for m=1 and 3, respectively (Figures 4c and e). Attaining equilibrium is much quicker than at low migration rates, however; the system as a whole is close to equilibrium within 50–150 generations, and at that point realized effective size of all the subpopulations are close to the eigenvalue effective size (NeRx≈NeMeta≈NeE).
Clearly, meta-Ne increases above the sum of the local effective sizes at low migration rates and decreases with increasing migration. When migration rates are high enough to make the system as a whole approach panmixia, meta-Ne will approach the sum of local population sizes. In the present case the system is close to NeMeta=NeE=225 (100+50+50+25) at m>3. Thus, low migration results in a high NeE at the cost of high rates of inbreeding in the subpopulations for many generations, and thereby of the system as a whole.
In a next step, we doubled the subpopulation effective sizes (that is, NeSc=200, NeFi=100, NeKa=100 and NeKol=50) and found that NeE becomes close to or above 500 for all three migration rates (not shown). Again, NeE is large (≈1742) with the lowest migration rate of m=0.05, but it takes a very long time to approach equilibrium; NeMeta is far below 500 and only increases from 113 to 210 over the first 200 generations. Also, with the higher migration rates of m=1 and 3 the equilibrium values of NeE=499 (m=1) and NeE=466 (m=3) are not reached until after c. 100 generations or more.
In another model we increased subpopulations of Scandinavia and Finland proportionally to a total size of 400 (NeSc=267 and NeFi=133), while assuming that Kola and Karelia remain at NeKa=50 and NeKol=25, and using m=3. The rationale for increasing only Scandinavia and Finland is that both Sweden and Finland are members of the EU and have agreed to the EU target of maintaining long-term viable populations of naturally occurring species, whereas Russia is not a member of EU. The sum of subpopulation effective sizes is now close to 500 (475) and NeE becomes 489, which is reached by NeMeta at t≈100.
Clearly, the conservation genetic target that we suggest for metapopulations is not possible to reach for a closed Fennoscandian wolf population unless subpopulation effective sizes (in isolation) increase so that their sum approaches 500. Further, substantial gene flow among subpopulations is necessary (m>3) to reach the target within reasonable time and to avoid excessive inbreeding during the approach to equilibrium.
Effects of one-way vs two-way migration between Scandinavia and Finland
Empirical observations suggest that gene flow may have been unidirectional from Finland to Scandinavia since the current Scandinavian population was founded in the early 1980s (Supplementary Appendix 1). Thus, we investigated the potential effects of one-way vs two-way migration between Finland and Scandinavia on the Fennoscandian metapopulation, still assuming no initial inbreeding in the system. We used current estimates of subpopulation effective sizes (Figure 2a) and m=1 implying that Finland, Kola and Karelia receive one migrant each from the other two, whereas the migration between Finland and Scandinavia is either two-way or one-way from Finland to Scandinavia (Figure 2a).
Eigenvalue effective size (NeE) is almost doubled under two-way migration as compared with one-way migration from Finland to Scandinavia (NeE=250 and 132, respectively; Figure 5). On the way to equilibrium, however, the Scandinavian population benefits from one-way migration over the first c. 100 generations. The realized Ne for Scandinavia increases more rapidly with one-way migration, and at generation t=75, for example, we have NeRSc=244 and 214 for one-way and two-way migration, respectively. This benefit for the Scandinavian population occurs at a cost for the other subpopulations and for the metapopulation as a whole, however; at t=75 we have NeMeta=169 vs 221 for one-way vs two-way migration (Figure 5). Similarly, realized Ne for Finland, Karelia and Kola are all at NeR=132 under one-way migration, as compared with NeRFi=241, NeRKa=222 and NeRKola=214 with two-way (not shown). In fact, neither of Finland, Karelia or Kola reaches a realized Ne above 132 with one-way migration from Finland to Sweden, while they are above this value already in generation 22 under two-way migration (not shown).
Effect of one-way vs two-way migration between Finland and Scandinavia on realized (NeR) and eigenvalue (NeE) effective size over 200 generations in the metapopulation system of four interconnected wolf populations meant to mimic that of Fennoscandia (cf. Figure 2a). Migration rate is m=1 implying that Finland, Kola and Karelia receive one migrant each from the other two, whereas the migration between Finland and Scandinavia is either two way or one way from Finland to Scandinavia. Initial inbreeding and kinship is zero (0) within and between all subpopulations. To avoid cluttering curves are shown only for Finland and Scandinavia (see text for details). Black curves, one-way migration; gray curves, two-way migration.
At first glance these results may appear surprising, but they can be explained as follows. Under one-way migration (from Finland to Scandinavia) the Scandinavian population behaves as a ‘genetic sink’ only receiving migrants from a metapopulation comprising Finland, Kola and Karelia. The eigenvalue effective size of this smaller metapopulation (without Scandinavia) is NeEFi,Kol,Ka=132, and with the present population sizes and migration rates the ultimate fate of the Scandinavian subpopulation is completely determined by the Finland–Kola–Karelia population system. Thus, under one-way migration each of the four populations will eventually approach a realized effective size of NeR=132. In contrast, under two-way migration all the four subpopulations are connected through two-way exchange of migrants with at least one of the other subpopulations. This results in a larger metapopulation with an eigenvalue effective size of NeE=250, and the realized effective size of each of the subpopulations will eventually approach this number (NeR=250). Further, the reason why NeRSc is larger under one-way migration than under two-way is that Scandinavia then receives immigrants that are unrelated during a longer time than with two-way migration when Scandinavia exports migrants to Finland and can receive descendants of those migrants back (cf. Hössjer, 2015).
Meta-Ne of a closed Fennoscandian population assuming initial inbreeding
Next, we explored meta-Ne for the Fennoscandian wolf when there is initial inbreeding in Scandinavia (fSc=0.25) and Finland (fFi=0.1) as observed in these populations (Supplementary Table 1), including a kinship between Scandinavia and Finland of fSc–Fi=0.05. Other starting parameters were as before (Figure 2a). We examined both one-way and two-way migration between Finland and Scandinavia, using m=1 in all cases of migration.
Now, Ne becomes undefined for the Scandinavian and Finnish subpopulations during an initial period when inbreeding decreases from the starting values of fSc=0.25 and fFi=0.1 (Figure 6a). With one-way migration the decrease of inbreeding continues over 39 generations for the Scandinavian population when fSc levels off at around 0.24 and then begins to increase again (not shown). In the Finnish subpopulation the decrease only occurs for two generations; fFi=0.092 in t=2 and then starts to increase slowly again.
Effect of one-way vs two-way migration between Finland and Scandinavia on realized (NeR) and eigenvalue effective size (NeE) when assuming initial inbreeding in Scandinavia (fSc=0.25) and Finland (fFi=0.1), and kinship between Scandinavia and Finland of fSc–Fi=0.05 (cf. Figure 2a). Migration rate is m=1 implying that Finland, Kola and Karelia receive one migrant each from the other two, whereas the migration between Finland and Scandinavia is either two way or one way from Finland to Scandinavia. To avoid cluttering, curves for Scandinavia and Finland are shown in a and curves for Karelia and Kola are shown in b. Curves for the total metapopulation are included in both a and b. Gray curves, two-way migration; black curves, one-way migration.
With two-way migration the decrease of inbreeding continues until t=7 for Scandinavia when fSc reaches its lowest value at fSc≈0.24, and until t=4 for Finland when fFi≈0.09. At the turning points where the decrease of inbreeding levels off and inbreeding starts to increase again, the increase is initially very small and NeRSc and NeRFi are therefore very large. For instance, at generation t=8 with two-way migration NeRSc≈18 000, and NeRFi=493 (as compared with NeRSc≈116, and NeRFi=113 in the case of no initial inbreeding; cf. Figure 6a vs Figure 4c). The final equilibrium values are, however, the same as in the case of no initial inbreeding, that is, NeE=132 and 250 for one-way and two-way migration, respectively (Figure 6). Thus, the initial reduction of inbreeding in Scandinavia and Finland results in realized effective population sizes of those subpopulations that to begin with are larger than when starting with no inbreeding, but with m=1 this early effect on NeR has leveled out around t=150–200 both in the case of one-way and two-way migration (Figure 6a).
As observed for the case with no initial inbreeding (Figure 5), one-way migration from Finland to Scandinavia also results in a ‘favorable’ situation for Scandinavia (with respect to the rate of inbreeding). The reason is that the immigrants are less related to the Scandinavian wolves than in the situation with two-way migration when Scandinavia exports genes, which can later migrate back again. For example, at t=75 and initial inbreeding we have NeRSc≈570 and 355 for one-way and two-way migration, respectively (Figure 6a).
When starting with inbreeding in the Finnish and Scandinavian populations the average inbreeding coefficient of the metapopulation as a whole increases from fMeta≈0.13 to 0.49 and 0.43 in generation t=200 for one- and two-way migration between Finland and Scandinavia, respectively. Clearly, after 200 generations the effect of initial inbreeding is minor relative to the case of no initial inbreeding (cf. Figure 4d).
With two-way migration and initial inbreeding the level in Finland decreases from the starting value of fFi=0.1 to 0.088 in generations t=5–8 and then increases again, passes fFi=0.1 in t=16 and reaches fFi=0.16 in t=50. In Kola and Karelia fKol and fKa pass 0.1 in generation t=16 and 21, respectively. At t=50, fKol≈0.20 and fKa≈0.19.
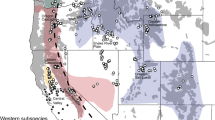
Meta-Ne with gene flow from Russia east of Fennoscandia
Clearly, a Fennoscandian wolf metapopulation cannot reach the conservation genetic target we propose unless the sum of subpopulation effective sizes approaches 500 and migration between subpopulations is extensive. Further, the results above indicate that inbreeding levels will remain high if the Fennoscandian population is a closed population. We modeled the effects of inflow of genes from the Russian population east of the Russian provinces of Kola and Karelia (Figure 1). Census estimates indicate around 3000 wolves in this area (Supplementary Appendix 2), and assuming Ne/N=0.25 this corresponds to an effective size of NeRu=750 (Figure 2b).
As expected, currently estimated inbreeding levels in Fennoscandia can be reduced with inflow from Russia. However, for a positive effect of immigration from Russia the migration within the Fennoscandian metapopulation must be extensive. For example, with a bidirectional gene flow within Fennoscandia of m≈1, a unidirectional inflow from Russia of 10 wolves per generation (Figure 2b) will never result in a reduction of average f of the metapopulation (fMeta) below the initial value of 0.13. In contrast, with m=5 within Fennoscandia, fMeta drops to 0.1 within the first few generations, and it is reduced to this level also in Scandinavia around generation 30, with fSc below 0.15 in generation 12 (not shown).
Metapopulation effective size of a Fennoscandian wolf population with currently estimated population sizes can increase to NeMeta>500 with immigration from Russia. Again, gene flow within Fennoscandia is critical for how long this will take. For example, even if 10 immigrants from Russia enter Fennoscandia via Karelia each generation NeMeta will not reach 500 until generation t=90 if migration within Fennoscandia is m=1. Assuming a generation interval of 5 years (Forslund, 2009) this translates into 450 years. With m=2, 3 or 5 within Fennoscandia the time to NeMeta⩾500 is 52, 37 and 26 generations, respectively (that is, in 130–260 years). With only one immigrant from Russia every generation NeMeta=500 is reached at generation t=99 (that is, in ~500 years) with a genetic exchange within Scandinavia of m=5. Clearly, if applying a shorter generation length of, for example, 3.4 years as estimated for the Finnish population (Supplementary Appendix I) these time frames in years are shorter.
Discussion
We used recently developed theory to address issues of long-term conservation genetic targets of metapopulations, with particular focus on the Fennoscandian wolf population. We suggest that the well-established rule of thumb for long-term genetic conservation of Ne⩾500, corresponding to an increase of inbreeding of Δf⩽0.001, should apply to the metapopulation as a whole as well as to each of the subpopulations. Thus, long-term genetic viability of metapopulations includes meeting the criteria of (i) metapopulation effective size NeMeta⩾500 and (ii) realized effective size of each subpopulation is NeRx⩾500. We also suggest as a conservation target that these two goals should be reached within the time frames of short-term conservation, that is, typically 5–20 generations. We argue that less rigorous criteria than those of long-term conservation would be acceptable during short-term conservation periods, but long-term criteria should be met when short-term conservation periods have passed.
When applying these criteria to the empirical case of the Fennoscandian wolf our findings can be summarized as follows:
-
1
The current Fennoscandian wolf population cannot reach the above long-term conservation target. Rather, assuming generally recommended rates of gene flow (m≈1; the OMPG rule) and current local effective population sizes, the meta-Ne that can be reached is 132 or 250 (see next point).
-
2
Two-way migration between all subpopulations is essential for the genetic health of the metapopulation. Unidirectional gene flow from Finland to Scandinavia, rather than bidirectional, greatly reduces metapopulation effective size. With one-way migration between those subpopulations NeE=132, while NeE=250 with two-way migration (assuming one migrant per generation).
-
3
To reach the above conservation criteria subpopulation effective sizes must increase such that their sum approaches 500. Further, the bidirectional migration rates among subpopulations must increase substantially above one migrant per generation. Thus, the OMPG rule is not sufficient in this particular case.
-
4
If subpopulation effective sizes are not allowed to increase, the genetic viability of the Fennoscandian wolf population must be dependent on gene flow from the northwestern Russian population. Several conditions must be met for such gene flow to have the wanted effect, however. First, migration from northwestern Russia must be substantial, c. 5–10 migrants per generation. Second, gene flow among the Fennoscandian subpopulations must be high (c. 3–5 migrants per generation). Further, the Russian population supporting migrants to the Fennoscandian metapopulation must be large and comprise an effective size of at least Ne=500 (data not shown).
Several general aspects of the concept of effective population size and its application to metapopulations have become apparent in this study. First, asymmetrical gene flow can have a great impact on metapopulation effective size and realized Ne of separate subpopulations. In our case study of Fennoscandian wolves this is exemplified by the markedly reduced NeE resulting from one-way migration from Finland to Scandinavia. Second, the parameter Ne can be difficult to interpret for metapopulations where inbreeding decreases in some subpopulations, because it then becomes undefined and can take extremely large values at the turning points when inbreeding starts to increase regardless of actual inbreeding levels (cf. Figure 6), thus poorly reflecting the actual conservation genetic situation. Third, the OMPG rule is not generally applicable in the context of metapopulation genetic conservation. Rather, necessary migration rates to avoid excessive inbreeding depend on the number of subpopulations, their effective sizes, as well as the pattern of gene flow among them.
We have suggested two criteria for long-term genetic viability of metapopulations: (i) NeMeta⩾500; and (ii) NeRx⩾500. In fact, it is sufficient to apply the second criterion because when this criterion is met the first one will also be satisfied for the diploid model of inbreeding effective size (cf. Hössjer et al., 2015). We chose to keep both criteria for clarity.
International conservation policies agreed by Finland, Norway and Sweden stipulate that viable populations of wolves should be maintained in the area. Such agreements are particularly strong for Finland and Sweden as members of the EU, but also for Norway, which has signed and ratified the Bern Convention (as have Finland, Sweden and other EU members) that lists the wolf in its Appendix II of strictly protected fauna (www.coe.int/bernconvention). In contrast, Russia is not part of either EU or the Bern Convention.
Our results indicate that criteria for long-term genetic viability are not met for the current Fennoscandian wolf population. In order to meet such criteria, local population sizes as well as migration rates between subpopulations need to increase. Alternatively, genetic support from Russia east of the province of Karelia is needed. Even with such gene flow, however, considerably improved connectivity among the Fennoscandian populations is necessary.
Knowledge is scarce on population genetic characteristics, dynamics and connectivity of wolf populations in the Russian provinces of Kola and Karelia of Fennoscandia, as well as of those in neighboring Russian provinces (Supplementary Appendix 1; Supplementary Table 1), and research to generate such information is urgently needed. Similarly, management cooperation is not well developed (Dr Jouni Aspi, University of Oulu, Finland pers.com.). Thus, it may be dangerous for Scandinavia and Finland to rely on gene flow from northwestern Russia in order to meet conservation goals for wolves in Fennoscandia. Further, due to the lack of common conservation policy, management collaboration and information it can be argued that Scandinavia and Finland alone should take responsibility for maintaining a long-term viable wolf population (Nilsson, 2015). This requires that the sum of local subpopulation-Nes is ≈500, with a high degree of connectivity. For instance, if NeSc=300 and NeFi=200 the time for NeMeta and NeRx to reach 500 varies in the range 15–50 generations for two-way migration in the interval 5<m<20. Assuming an Ne/N ratio of 0.25 (Forslund, 2009; Bruford, 2015) this would translate into 1200 wolves in Sweden and 800 in Finland, with an exchange of 20–80 individuals per generation.
The present study represents an application of a new mathematical framework to a practical scenario; in fact, it was the case of the Fennoscandian wolves that initiated the mathematical developments presented by Hössjer et al. (2014, 2015). Clearly, however, this framework can be applied to many other situations where exploration of metapopulation effective size is of interest.
Also, there are several general aspects of metapopulation effective size that we have not been able to address here. For instance, we have noted that it is unclear from the original presentation by Franklin (1980) which Ne his Ne⩾500 rule applies to. Some authors have suggested that it is the variance effective size (Harmon and Braude, 2010), but since loss of additive genetic variation is associated with loss of heterozygosity (Franklin 1980) it is actually the inbreeding effective size that is applicable here. The haploid inbreeding effective size might be most relevant because it quantifies loss of gene diversity (cf. Hössjer et al., 2014). We elaborate on this in forthcoming work (Hössjer et al., unpublished). NeMeta is larger for the haploid inbreeding model than for the diploid one because rate of inbreeding is quantified based on two alleles drawn at random from the whole population rather than from individuals. However, the difference for NeRx between the haploid and diploid models is negligible.
Further, in the present model we have ignored the forces of mutation and selection. Adding mutation to the model is not expected to change our basic conclusions because we consider relatively short time frames from an evolutionary perspective—even our long-term perspectives in conservation is typically to be regarded as short-term from an evolutionary standpoint. Selection is difficult to include because a wide range of potential selective regimes are possible. With directional selective pressures, however, rates of inbreeding are expected to increase at a higher rate than in a situation without selection. Similarly, growing immigration into inbred populations such as Scandinavia and Finland may initially result in a genetic rescue effect where less inbred individuals contribute more progeny than others, thus increasing the variance of reproductive success. Such a higher variance is expected to increase the rate of inbreeding and thus reduce realized effective population sizes during the initial phases of the approach to equilibrium. This kind of genetic rescue effect with extensive spread of genes from an immigrating individual followed by increased inbreeding and associated inbreeding depression was documented empirically in the isolated and highly inbred Isle Royale wolf population (Hedrick et al., 2014).
More theoretical work is needed with respect to several aspects of metapopulation effective size such as exploring the effects of selection and the behavior of other types of effective sizes (such as haploid vs diploid Nei, variance, linkage disequilibrium, coalescence effective sizes and so on) in metapopulations. We are in the process of continuing research along the lines of both to explore metapopulation effective size dynamics, including applying our current framework to other practical conservation cases of concern, and to further address the mathematical theory of metapopulation effective sizes.
Concluding recommendations on Fennoscandian wolves
On the basis of the results from this study we find the following management recommendations warranted for the Fennoscandian wolves.
-
1
Two-way migration between Finland and Scandinavia needs to be secured.
-
2
Monitoring of migration rates among all subpopulations of Fennoscandia is required.
-
3
The degree of connectivity needed among subpopulations is in the order of c. 3–5 genetically effective migrants per generation.
-
4
Local effective population sizes need to increase such that the sum of the subpopulation Nes that comprise the metapopulation approaches 500.
-
5
Owing to the lack of a common conservation policy between EU member states (Sweden and Finland) and Russia, we propose that the conservation target of meta-Ne⩾500 should be reached for wolf populations of Scandinavia and Finland.
Data archiving
There were no data to deposit.
References
Allendorf FW, Ryman N . (2002). The role of genetics in population viability analysis. In: Beissinger SR, McCullough DR (eds) Population Viability Analysis. University of Chicago Press: Chicago, IL, USA. pp 50–85.
Aspi J, Roininen E, Kiiskilä J, Ruokonen M, Kojola I, Bljudnik L et al. (2009). Genetic structure of the northwestern Russian wolf populations and gene flow between Russia and Finland. Conserv Genet 10: 815–826.
Bruford MW . (2015). Additional population viability analysis of the Scandinavian wolf population. Swedish Environmental Protection Agency Report 6639, July 2015. Available at https://www.naturvardsverket.se/Documents/publikationer6400/978-91-620-6639-0.pdf?pid=14226 (accessed 5 October 2015).
Chapron G . (2014). Challenge the abuse of science in setting policy. Science 516: 289.
Chapron G . (2015). Clarification of the concept of Favourable Conservation Status. Report to the Swedish Environmental Protection Agency NV-02945-5.
Crandall KA, Posada D, Vasco D . (1999). Effective population sizes: missing measures and missing concepts. Anim Conserv 2: 317–319.
Darpö J . (2011). Brussels advocates Swedish grey wolves. SIEPS—Swedish Institute for European Policy Studies. European Policy Analysis 2011:8epa, 20 pp.
EC. (2015). Commission requests Sweden to bring wolf hunting into line with EU standards. European Commission Infringement Decision of June 18, 2015. Available at europa.eu/rapid/press-release_MEMO-15-5162_en.htm (accessed June 2015).
EEB. (2015). URGENT—stop the Swedish licensed hunt on endangered wolves. Letter of appeal to EU Commissioner Karmenu Vella from the European Environmental Bureau and the World Wide Fund for Nature (WWF). Available at www.eeb.org/EEB/?LinkServID=0B8B5A08-5056-B741-DB32358B9EA57469 (accessed 5 May 2015).
Evans D, Arvela M (2011). Assessment and reporting under Article 17 of the Habitats Directive. Explanatory notes and guidelines for the period 2007–2012, final draft prepared for the Habitats Committee in April 2011. European Topic Center on Biological Diversity, Copenhagen, Denmark.
Ewens WJ . (1982). On the concept of effective population size. Theor Popul Biol 21: 373–378.
Forslund P . (2009). Modeling growth and inbreeding in the Scandinavian wolf population. Appendix 1:3 of Suggestions for actions to strengthen the genetic status of the wolf population, Report by the Swedish Environmental Protection Agency to the Swedish Government Dnr 429-8585-08 Nv (In Swedish).
Franklin IR . (1980). Evolutionary change in small populations. In: Soulé M, Wilcox B (eds), Conservation Biology: An Evolutionary-Ecological Perspective. Sinauer Associates: Sunderland, MA, USA. pp 135–149.
Hansen MM, Andersen LW, Aspi J, Fredrickson R . (2011). Evaluation of the conservation genetic basis of management of grey wolves in Sweden. Report from the international evaluation panel of the Swedish Large Carnivore Inquiry, Swedish Government Investigation SOU 2011:37. Statens Offentliga Utredningar (the Swedish Government’s Official Investigations). Available from the Swedish Government at http://www.regeringen.se (accessed September 2015).
Harmon LJ, Braude S . (2010) Conservation of small populations: Effective population size, inbreeding, and the 50/500 rule. In: Braude S, Low SB (eds), An Introduction to Methods and Models in Ecology and Conservation Biology. Princeton University Press: Princeton, NJ, USA. pp 125–138.
Hastings A, Harrison S . (1994). Metapopulation dynamics and genetics. Annu Rev Ecol Syst 25: 167–188.
Hedrick PW, Peterson RO, Vucetich LM, Adams JF, Vucetich JA . (2014). Genetic rescue in Isle Royale wolves: genetic analysis and the collapse of the population. Conserv Genet 15: 1111–1121.
Hössjer O . (2015). On the eigenvalue effective size of structured populations. J Math Biol 71: 595–646.
Hössjer O, Olsson F, Laikre L, Ryman N . (2014). A new general analytical approach for modeling patterns of genetic differentiation and effective size of subdivided populations over time. Math Biosci 258: 113–133.
Hössjer O, Olsson F, Laikre L, Ryman N . (2015). Metapopulation inbreeding dynamics, effective size and subpopulation differentiation—a general analytical approach for diploid organisms. Theor Popul Biol 102: 40–59.
Jamieson IG, Allendorf FW . (2012). How does the 50/500 rule apply to MVPs? Trends Ecol Evol 27: 578–584.
Jansson E, Ruokonen M, Kojola I, Aspi J . (2012). Rise and fall of a wolf population: Genetic diversity and structure during recovery, rapid expansion, and drastic decline. Mol Ecol 21: 5178–5193.
Jansson M, Amundin M, Laikre L . (2015). Genetic contribution from a zoo population can increase genetic variation in the highly inbred wild Swedish wolf population. Conserv Genet 16: 1501–1505.
Laikre L . (1999). Conservation genetics of Nordic carnivores: lessons from zoos. Hereditas 130: 203–216.
Laikre L, Ryman N . (2009). Review of existing knowledge and conservation genetic status of today’s Swedish wolf population. Appendix 1:4 of Suggestions for actions to strengthen the genetic status of the wolf population. Report by the Swedish Environmental Protection Agency to the Swedish Government Dnr 429-8585-08 Nv (In Swedish).
Laikre L, Jansson M, Allendorf FW, Jakobsson S, Ryman N . (2013). Hunting effects on favourable conservation status of highly inbred Swedish wolves. Conserv Biol 27: 248–253.
Liberg O, André H, Pedersen C-H, Sand H, Sejberg D, Wabakken P et al. (2005). Severe inbreeding depression in a wild wolf Canis lupus population. Biol Lett 1: 17–20.
Liberg O, Chapron G, Wabakken P, Pedersen HC, Thompson Hobbs N et al. (2012). Shoot, shovel and shut up: cryptic poaching slows restoration of a large carnivore in Europe. Proc R Soc B 279: 910–915.
Liljelund L-E . (2011). Rovdjurens bevarandestatus. Delbetänkande av Rovdjursutredningen. (Large carnivore conservation status) interim report of the Large Carnivore Inquiry, Swedish Government Investigation SOU 2011:37 (in Swedish). Statens Offentliga Utredningar (the Swedish Government's Official Investigations). Available from the Swedish Government at http://www.regeringen.se (accessed September 2015).
Linnell J, Salvatori V, Boitani L . (2008). Guidelines for population level management plans for large carnivores in Europe. Contract no. 070501/2005/424162/MAR/B2. Final version European Commission: Brussels, Belgium.
Maruyama T, Kimura M . (1980). Genetic variability and effective population size when local extinction and recolonisation of subpopulations are frequent. Proc Natl Acad Sci USA 77: 6710–6714.
Mills LS, Allendorf FW . (1996). The one-migrant-per-generation rule in conservation and management. Conserv Biol 10: 1509–1518.
Mills LS, Feltner J . (2015). An updated synthesis on appropriate science-based criteria for “favorable reference population” of the Swedish wolf (Canis lupus) population. Report to the Swedish Environmental Protection Agency NV-02945-15.
Nilsson T . (2015). Peer-review statement in Bruford MW: additional population viability analysis of the Scandinavian wolf population. Swedish Environmental Protection Agency Report 6639, July 2015, pp 67–69.
Nunney L . (1999). The effective size of a hierarchically structured population. Evolution 53: 1–10.
Potočnik J . (2011). Statement by Janez Potočnik, European Commissioner for Environment, on the issue of conservation of the wolf in Sweden. Available at http://ec.europa.eu/archives/commission_2010-2014/potocnik/media/pdf/Swedish%20wolves.pdf accessed 5 October 2015.
Räikkönen J, Vucetich JA, Vucetich LM, Peterson RO, Nelson MP . (2013). What the inbred Scandinavian wolf population tells us about the nature of conservation. PLoS ONE 8: e67218.
SEPA Decisions 2009–2013. Decisions on license hunting for wolves 2010, 2011, 2013, 2014. Swedish Environmental Protection Agency, Dnr 411-7484-09 Nv, NV-03454-10, NV-01007-13, NV-08512-13 (In Swedish).
Tufto J, Hindar K . (2003). Effective size in management and conservation of subdivided populations. J Theor Biol 222: 273–281.
Vilà C, Sundqvist A-K, Flagstad Ø, Seddon J, Brörnerfeldt S, Kojola I et al. (2003). Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant. Proc R Soc London B 270: 91–97.
Wang J . (2004). Application of the one-migrant-per-generation rule in conservation and management. Conserv Biol 18: 332–343.
Wang J, Caballero A . (1999). Developments in predicting the effective size of subdivided populations. Heredity 82: 212–226.
Waples RS . (2010). Spatial-temporal stratifications in natural populations and how they affect understanding and estimation of effective population size. Mol Ecol Resour 10: 785–796.
Whitlock MC, Barton NH . (1997). The effective size of a subdivided population. Genetics 146: 427–441.
Wright S . (1931). Evolution in Mendelian populations. Genetics 16: 97–159.
Wright S . (1938). Size of population and breeding structure in relation to evolution. Science 87: 430–431.
Åkesson M, Hedmark E, Liberg O, Svensson L . (2014). Summary of the pedigree of the Scandinavian wolf population up until 2013. Report from the Grimsö Wildlife Research Station, Department of Ecology, Swedish University of Agricultural Sciences (In Swedish).
Acknowledgements
We thank Frank E. Zachos, Frank Hailer and three anonymous referees for valuable comments on earlier versions of this manuscript. This work was supported by the Swedish Environmental Protection Agency (contract no. 11/116; LL), the Swedish Research Council Formas (215-2012-1550; LL), the Swedish Research Council (621-2011-3715; NR and 621-2013-4633; OH). We are grateful to Eugenia Bragina, Jouni Aspi and Peep Männil for valuable information, Lovisa Wennerström for help with illustrations and Anastasia Andersson for Russian translations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Laikre, L., Olsson, F., Jansson, E. et al. Metapopulation effective size and conservation genetic goals for the Fennoscandian wolf (Canis lupus) population. Heredity 117, 279–289 (2016). https://doi.org/10.1038/hdy.2016.44
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.44
This article is cited by
-
Assessment of the Global Variance Effective Size of Subdivided Populations, and Its Relation to Other Effective Sizes
Acta Biotheoretica (2023)
-
Genetic signature of immigrants and their effect on genetic diversity in the recently established Scandinavian wolf population
Conservation Genetics (2022)
-
Biological and Sociopolitical Sources of Uncertainty in Population Viability Analysis for Endangered Species Recovery Planning
Scientific Reports (2019)
-
Nutraceutical values of fish demand their ecological genetic studies: a short review
The Journal of Basic and Applied Zoology (2018)
-
Genomic consequences of intensive inbreeding in an isolated wolf population
Nature Ecology & Evolution (2017)