Abstract
Changes in chromosome number have a critical role in the evolution and formation of plant species. Triploids, which carry three complete sets of chromosomes, in particular produce offspring with different chromosome numbers, including diploid and tetraploid progeny, as well as a swarm of aneuploid progeny, which carry incomplete chromosome sets. In this study, we investigated the mechanisms shaping these swarms at the population level through a detailed characterization of the progeny of triploid Arabidopsis thaliana. We report that triploid meiosis predominately produced aneuploid gametes, most of which were viable. We performed reciprocal crosses between triploid and either diploid or tetraploid plants and karyotyped all surviving individuals. This allowed us to dissect the parent-of-origin (cross-direction) effects and also the effect of the dosage of the crossing partner on the inheritance of each chromosome type. Overall, our data indicate that the chromosomal composition of the swarms produced by the triploid A. thaliana were strongly influenced by selection acting against specific gamete combinations, but not necessarily associated with aneuploidy. Finally, each of the five chromosome types responded differently to this selection, suggesting the presence of dosage-sensitive factor(s) critical for viability and encoded on different chromosomes.
Similar content being viewed by others
Introduction
Variation in chromosome number has played an important role in shaping the genomes of extant species. For example, whole-genome duplication, resulting in polyploid genomes, is very common in flowering plants (Wolfe and Shields, 1997; Soltis et al., 2003). Changes in chromosome number can also be caused by errors in chromosome partitioning during meiosis and result in aneuploid individuals, carrying incomplete chromosome sets. In mammals, viable aneuploidies are rare and associated with severe developmental defects (Epstein, 1986), as exemplified by trisomy 21 (Down' syndrome) in humans. In plants, aneuploidy is more common and may have a role in both speciation and genome evolution (Ramsey and Schemske, 2002).
Aneuploidy is very frequent in plant polyploid populations (Khush, 1973; Ramsey and Schemske, 2002; Henry et al., 2006). This frequency is even higher in areas where diploid and tetraploid populations coexist and can intercross to produce triploid individuals (Ramsey and Schemske, 1998; Henry et al., 2005). Triploids are characterized by the presence of three complete sets of chromosomes. They are meiotically unstable and produce diploid, triploid or tetraploid progeny, as well as a swarm of aneuploid individuals of intermediate genome content (GC) (Punyasingh, 1947; Khush, 1973; Singh, 2003; Henry et al., 2005). By virtue of producing populations of aneuploid individuals of variable chromosome composition (karyotypes), triploid plants can thus be used to investigate the mechanisms underlying changes in chromosome number and composition (karyotype) at the population level, as well as cellular phenomena, such as meiotic mechanisms and dosage-sensitive interactions originating from aneuploidy in gametes and in zygotes.
In previous reports, we have shown that triploids of Arabidopsis thaliana are fertile, producing 65–75% of viable seeds (Henry et al., 2005). The progeny of selfed triploids of A. thaliana included diploids, triploids and a few tetraploid individuals, as well as a high percentage of aneuploid individuals of intermediate GC, which we refer to as the aneuploid swarm. Furthermore, we showed that the progenies of genotypically different triploids varied in the distribution of GC that was present in the aneuploid swarm, suggesting a genetic basis to the mechanisms shaping them. To facilitate further study, we have developed a PCR-based high-throughput method allowing rapid karyotyping of aneuploid individuals and showed that it could be used to characterize the chromosomal composition of aneuploid populations derived from triploid or tetraploid individuals (Henry et al., 2006). Using this method, we have shown that the progeny of the tetraploid A. thaliana comprised 25% of aneuploid individuals. This highlighted the need for a better understanding of the production and selection of aneuploid individuals and for an increased awareness of aneuploidy within polyploid populations. Furthermore, this finding suggested that, in research studies using polyploid individuals or populations, it would be advisable to consider the manner in which the presence of aneuploid individuals could affect the outcome.
In this report, we further investigated the mechanisms regulating karyotype selection in the aneuploid swarms produced by the triploid A. thaliana. Given that the swarms produced by selfed triploid plants are complicated by the possible origin of aneuploidy from both gametes, we chose to characterize simpler swarms. To this end, we generated pseudo-backcross (pBC) populations between triploid and euploid individuals and characterized the progeny of each cross. Specifically, we investigated the influence of parent-of-origin effects on karyotype distribution by comparing crosses in which the triploid was used as the seed parent with those in which it was used as the pollen parent. Similarly, the influence of genomic dosage of the gamete contributed by the crossing partner was examined by comparing crosses of triploids with those of diploid or tetraploid individuals. The strength of our analysis lies in the ability to compare results obtained from these pBCs to distinguish the role of different mechanisms and developmental stages (Figure 1). For example, the effect of any mechanism acting before fertilization, such as gametic selection, cannot be influenced by the ploidy of the crossing partner, that is, it should be visible regardless of whether the backcross parent is a diploid or a tetraploid. Similarly, selection due to altered dosage of factors that are expressed in the embryo post fertilization and that are not parentally imprinted, may not be influenced by the direction of the cross but can be influenced by the ploidy of the other gamete.
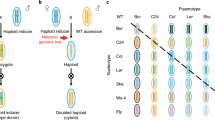
Crossing strategy for the analysis of parent-of-origin effects and effects of the dosage of crossing partner on the progeny of triploid individuals. Crosses between triploids and euploids (pseudo-backcrosses) were used. The parent-of-origin effect was tested by comparing the output of reciprocal crosses. The effect of the dosage of the gamete produced by the crossing partner was tested by comparing the output of similar crosses involving backcross parents of different ploidy (Col-0 or 4x-Col).
Materials and methods
Plant growth conditions, lines and crosses
All plants were grown on soil (Sunshine Professional Peat-Lite mix 4; SunGro Horticulture, Vancouver, BC, Canada) in a growth room lit by fluorescent lamps (Model TL80; Phillips, Sunnyvale, CA, USA) at 22±3 °C with a 16:8 h light:dark photoperiod or in a greenhouse at similar temperature and light regimes, with supplemental light provided by sodium lamp illumination as required.
The tetraploid Col-0 line (4x-Col) was produced by colchicine treatment as described previously (Henry et al., 2005). Wa-1 is the naturally occurring tetraploid ecotype Warschau-1 (Accession number CS6885, Arabidopsis Biological Resources Center (ABRC), Columbus, OH, USA). C and W refer to basic genomes or alleles of Col-0 and Wa-1, respectively. Triploid individuals were produced by crossing diploid and tetraploid individuals. The pBCs were produced by crossing the CWW triploids to either diploid Col-0 or tetraploid 4x-Col, in both directions. The four types of pBCs and the number of progeny analyzed for each type of cross were as follows: Col-0 × CWW (2 × 3; N=80), CWW × Col-0 (3 × 2; N=102), 4x-Col × CWW (4 × 3; N=33) and CWW × 4x-Col (3 × 4; N=47). Crosses are represented in the conventional way throughout this paper, with the seed parent indicated first.
Phenotypic analysis of seed viability
Siliques were harvested into individual tubes and all the seeds obtained from each fruit were counted using a dissecting microscope. Seeds were characterized as ‘plump’ if they contained a visible embryo structure that was at least 20% the size of wild-type seed or as ‘shriveled’ if they did not. Mean values for each group of plants were compared pairwise using Student's t-test and P-values <0.05 were considered significant.
Chromosome counts in developing microspores
Microspores obtained from developing flower buds were harvested and used for chromosome counts by FISH (fluorescent in situ hybridization) as described in Henry et al. (2006). Briefly, FITC (fluorescein isothiocyanate)-labeled A. thaliana centromeric DNA probes (Comai et al., 2003) were hybridized to slides prepared as follows: The slides were first incubated in 100 μg ml−1 ribonuclease A (cat. no. R-4642, Sigma-Aldrich, Milwaukee, WI, USA) in 2 × SSC for 30 min at 37 °C. Next, they were washed thrice for 5 min each in 2 × SSC and for 1 min in 10 mM of HCl. The slides were then incubated with 2 μg ml−1 of pepsin (cat. no. P6887, Sigma-Aldrich) in 10 mM of HCl for 10 min at 37 °C, rinsed in distilled water and washed thrice for 5 min each in 2 × SSC. Finally, the slides were dipped in 4% formaldehyde for 10 min at room temperature and washed thrice for 5 min each in 2 × SSC before being dehydrated through a series of ethanol dilutions (70, 90 and 100%). After these initial steps, the slides were washed in 2 × SSC and then in 70% formamide in 2 × SSC and subsequently treated as described previously (Comai et al., 2003), starting with denaturation at 65 °C in 70% formamide for 15 min. For hybridization, 10 μl of hybridization mix was deposited on each slide. After hybridization, the slides were stained with 0.25 μg ml−1 of DAPI (4′,6-diamidino-2-phenylindole). For an example of the type of picture obtained using this protocol, see Figure 2 in Henry et al. (2006).
Several individual triploid plants were used for microspore observations: CWW (N=11) and WWC (N=10), for a total of 3003 microspores (detailed counts for each triploid plant can be found in Supplementary Table 1). For each microspore, the number of chromosomes was recorded. The expected distribution of gametes in each chromosome number class was calculated by assuming that for each chromosome type, two of the three chromosomal copies migrated toward one pole, whereas the remaining copy migrated toward the other pole. Migration of one or two copies toward a specific pole was assumed to be independent for each of the five chromosome types. Distributions of chromosome numbers in microspores produced by the two triploids were compared with each other and with the expected distributions using χ2-tests. A P-value <0.05 was considered significant.
Flow cytometric measurement of nuclear DNA content
Each individual in the pBC progeny was analyzed for nuclear DNA content. Flow cytometry of isolated nuclei stained with propidium iodide was performed as described previously (Henry et al., 2005, 2006). Standard curves were calculated from multiple independent measurements on diploid, triploid and tetraploid individuals. Using these curves, GC values of the samples were expressed as multiples of the haploid GC of Col-0 (Henry et al., 2005, 2006). On this scale, a value of 2.0 corresponds to a diploid individual and a 4.0 value corresponds to a tetraploid individual. The first approximation of chromosome number was obtained from these measurements of nuclear genome DNA content (or GC) by dividing the GC value by the average GC of a single chromosome. For example, a GC value of 2.37 corresponds to 12 chromosomes (2.37/0.2=11.85). All the GC values obtained using flow cytometry are available in Supplementary Table 2.
Determination of chromosome composition (karyotyping)
Karyotyping was performed to more precisely determine the number of chromosomes present in each of the pBC progeny as well as to determine their chromosomal composition. A detailed description of the technique used to karyotype the populations analyzed in this study was published previously (Henry et al., 2006). Briefly, each individual in the progenies of the four types of pBCs were genotyped at 12 markers using quantitative fluorescent PCR. These 12 markers are located such that at least 1 marker is present on each chromosome arm. Details regarding the primer sequence and location of these markers can be found in Henry et al. (2006). Next, for each marker and for each individual, the fluorescence intensities of the PCR peaks associated with the C and W alleles were recorded and their ratio was calculated. These ratios were divided into distinct heterozygous categories. Each category was then associated with specific heterozygote combinations from which the number of chromosome copies carrying the marker could be inferred. For example, a Col/Wa ratio of 0.25 (1:3) at marker nga1145 located on chromosome 2, indicated one Col-0 allele and three Wa-1 alleles, for a total of four alleles and therefore four copies of chromosome 2. The number of copies of each chromosome type and the total number of chromosomes could thus be inferred in this manner for each of the individuals. Finally, the individuals were categorized according to zygote chromosome number (Table 2). The individuals with 10, 15 or 20 chromosomes were classified as euploids, whereas all other individuals were classified as aneuploids. All raw fluorescence values obtained from the microsatellite analysis as well as inferred gametic and zygotic genotypes are available in Supplementary Table 2.
Comparison of GC distributions
The numbers of gametes in the different ploidy classes (euploid vs aneuploid and haploid vs diploid) were calculated (Table 2). These ratios were then compared either with expected ratios (see below) or with ratios observed in another pBC using Fisher's exact tests to accommodate for N<5 in some categories. Two-tailed P-values are shown in Table 3. The expected numbers of gametes in each category were calculated on the basis of the observed percentages of microspores observed in the FISH experiments (Figure 3). Specifically, from the CWW triploid, the expected percentages of haploid, aneuploid and diploid gametes were 6.5, 88.8 and 4.7%, respectively. When comparing observed frequencies in two different pBCs, numbers obtained from the cross with the lowest N value were used as observed data, and numbers of the other crosses were adjusted and used as expected data.
Distribution of chromosome numbers in microspores produced by triploid individuals. Chromosomes were counted in microspores from CWW (1380 microspores from 11 individuals, gray bars) and WWC (1623 microspores from 10 individuals, white bars) triploids and the mean frequency of microspores in each chromosome number class was calculated. The s.e. are indicated. The black line connects the percentages (gray circles) of microspores expected by assuming a random assortment of three sets of chromosomes during meiosis and no selection.
Statistical analyses of chromosome inheritance
The contribution of the triploid parent to the karyotype of the individuals in the progenies was derived assuming that the euploid parent always contributed one (for diploid Col-0) or two (for 4x-Col) copies of each chromosome type.
For each type of cross, individuals were classified according to whether they received one or two copies of a given chromosome type from the triploid parent. The percentages of individuals who received one or two copies of a given chromosome type are summarized in Figure 4 and in Table 4. The effect of cross-direction and dosage of the euploid parent on chromosome inheritance was tested by comparing these percentages between different pBC populations. Proportions were compared using χ2-tests, and results were considered significantly different when the P-value was <0.01 to correct for five independent tests on five chromosome types (Figure 4).
Effect of cross-direction (parent-of-origin effect) and effect of the ploidy of the crossing partner on the chromosomal composition of the successful aneuploid gametes. For each chromosome type, gametes that contributed to the viable progeny were classified into two categories depending on the number of copies of that specific chromosome type received from the CWW triploid. The numbers of gametes in both categories are expressed as the percentage of the total number. Each cross is described five times, for each chromosome type. Seed parents are indicated first. The effect of the ploidy of the crossing partner (left columns) and the effect of the cross-direction (parent-of-origin effect, right columns) are assessed by pairwise comparison between crosses, as indicated by brackets. The numbers of gametes are compared using χ2 tests, and P-values are considered significant (depicted in bold) if <0.01 to control for testing on five independent chromosome types.
Results
Triploids of A. thaliana were produced by performing interploidy crosses between Col-0 and the natural tetraploid Wa-1 (WWW.W). These individuals were then crossed in both directions with either diploid Col-0 or 4x-Col. To characterize the mechanisms shaping triploid output, the following traits were followed: chromosome counts in microspores, seed viability after crossing and GC distribution and karyotype of the surviving progeny.
Triploid A. thaliana produces a majority of aneuploid microspores
To investigate the products of triploid meiosis, we documented the distribution of chromosome numbers in developing microspores using FISH to centromeric DNA probes and recorded the number of centromeres in microspores obtained from the CWW and WWC triploids. Both triploids produced mostly aneuploid gametes carrying between 6 and 9 chromosomes, as well as a few haploid and diploid gametes and a very low number of microspores with 4 or 11 chromosomes (Figure 3). The distributions of chromosome numbers were compared to an expected distribution that was obtained assuming a random assortment of chromosomes during the meiotic divisions and no selection. The CWW triploid produced a greater proportion of aneuploid microspores than did the WWC triploid (88.8 vs 84.1%, χ2 P-value <0.0001). The observed distributions were qualitatively similar to the expected symmetrical binomial distribution, providing no evidence of massive chromosome loss. The observed distributions exhibited an overrepresentation of euploids when compared with the expected distribution (χ2 P-values <0.0001 for both triploids that were tested), suggestive of early selection against aneuploidy during microspore development.
Most aneuploid karyotypes are viable
To document which of the gametes produced by the CWW triploid generated viable progeny, it was crossed with either diploid Col-0 or tetraploid 4x-Col in both directions (Figure 1). All individuals were karyotyped to infer the karyotypes of the gametes contributed by the triploid parent (see Materials and methods section for details).
We recorded the number of successful gametes corresponding to each of the 30 different possible aneuploid karyotypes (5 single disomics, 10 double disomics, 10 triple disomics and 5 quadruple disomics) as well as the two euploid karyotypes (haploid and diploid) (Table 1). The number of observed aneuploid karyotypes in pollen grains (16 of 30 observed in either the CC × CWW or the CCCC × CWW cross) was lower than in ovules (27 of 30 observed in either the CWW × CC or the CWW × 4x-Col cross), consistent with stronger selection against aneuploid pollen grains. Nevertheless, even in this moderately sized population, all but 2 of the 30 possible types of aneuploid gametes were represented at least once. This demonstrates functionality for most aneuploid gamete karyotypes in at least one of the two gametophytes.
In terms of aneuploidy in zygotes, 24 of the 30 possible ‘low’ GC aneuploids (GC between diploidy and triploidy) were observed at least once in the crosses to Col-0. Similarly, 20 of the 30 possible ‘high’ GC aneuploids (GC between triploidy and tetraploidy) were observed at least once in the crosses to 4x-Col (Table 1). Hence, our results show that 44 of the 60 possible aneuploid karyotypes are viable. Given the high number of possible aneuploid zygotic karyotypes, it remains to be seen whether specific karyotypes are fully lethal or merely not represented in our limited sample size.
Paternal excess limits seed viability in pBCs to triploids
To investigate the importance of selection acting on the aneuploid swarm during seed development, we measured seed viability in each of the pBC populations. Three of the four types of pBCs exhibited high seed viability (84.4–91.1%, Figure 5). The cross involving 4x-Col as a pollen parent exhibited significantly lower seed viability than did the other crosses (60.5% seed viability, pairwise t-test P ⩽0.0332). The CWW triploid was also crossed as seed parents with tetraploid Wa-1. Seed viability in this cross was even lower as only 26.8% viable seeds were recovered. Crosses using tetraploid pollen parents thus exhibited lower seed viability than did the three other pBCs (all t-test P ⩽0.0332), suggesting that paternal genomic excess, that is, crosses involving a pollen parent of higher ploidy than the seed parent, induced seed lethality.
Seed viability in the pseudo-backcrosses. The mean percentage of plump seed (as a measure of seed viability) was measured for each of the four pseudo-backcrosses. The mean (indicated by the height of each column) and s.e. (indicated by the error bars) are shown. Means were compared by Student's t-test. The different letters above two columns indicate significantly different means for these two measurements (P-value <0.05).
GC distribution in the pBCs
Chromosome number distributions were compared between crosses (Figure 2 and Tables 2 and 3). Chromosome number distributions in the pBCs were also compared with the distribution observed in microspores produced by CWW (Figure 3). Indeed, a slight selection against aneuploid microspores had already been observed in developing microspores. Although we do not know whether the ovule population exhibits a similar distribution, comparing with the pollen population ensured a more conservative analysis than using the theoretical distribution. Three of the four cross types produced fewer aneuploid progeny than expected on the basis of the number of aneuploid microspores that were observed (Fisher's exact test P⩽0.0012; Table 3), thus confirming the deleterious effect of aneuploidy. The percentage of aneuploid progeny varied depending on the cross type (Table 2). It was lowest in the 4x-Col × CWW cross (48%) and greatest in the CWW × 4x-Col cross (74%), in which it was not significantly different from the frequency observed in microspores. Comparison between crosses highlighted a parent-of-origin effect in the crosses to 4x-Col (Fisher's exact test P=0.02, Table 3) but not in the crosses to diploid Col-0.
Our data provided no evidence that increased overall genome dosage acted to buffer the effect of chromosomal imbalance. Indeed, although the percentage of aneuploid progeny in the CWW × 4x-Col cross (74.5%) was slightly higher than in the CWW × Col-0 cross (61.8%), this trend was reversed in the reciprocal crosses (48.5% for the 4x-Col × CWW cross compared with 61.3% for the Col-0 × CWW cross). Hence, the crosses to 4x-Col did not result in more aneuploid progeny than crosses to 2x-Col, but produced a different distribution of individuals of aneuploid GC.
Indeed, karyotype distribution within the progeny of crosses to diploid Col-0 exhibited a strong bias toward aneuploids of low chromosome numbers (average number of chromosomes in the gametes of 6.4 and 6.8 for the 2 × 3 and 3 × 2 crosses, respectively, Table 2). However, crosses to 4x-Col generated many triploid and tetraploid progeny and varying proportions of aneuploids of intermediate chromosome numbers (average number of chromosomes in the gametes of 7.9 and 7.3 for the 4 × 3 and 3 × 4 crosses, respectively, Table 2). Thus, the ploidy of the crossing partner strongly influenced the survival of the fertilization products derived from aneuploid gametes.
Selection on euploid gametes
Observations of microspores showed that the number of haploid and diploid gametes produced by the CWW triploid is similar although slightly more haploid microspores were observed (6.5 haploid vs 4.7% diploid, Figure 3). Considering that these gametes are not aneuploid, a similar ratio of diploid and triploid (in the case of the crosses to diploid Col-0) or triploid and tetraploid (in the case of the crosses to 4x-Col) individuals should be expected in the progeny of pBCs. Yet, our results demonstrate otherwise (Table 2). Specifically, crosses to diploid Col-0 produced a high percentage of diploids and a low percentage of triploids (35 vs 3.8% for the 2 × 3 cross and 35.3 vs 2.9% for the 3 × 2 cross, respectively, Fisher's exact test P=0.0078 and P=0.0011, respectively, Table 3). On the other hand, crosses to 4x-Col produced ratios closer to the expected 1:1 ratio of triploid and tetraploid individuals (Fisher's exact test P=1 for both crosses, Table 3). Accordingly, no parent-of-origin effect could be detected for this effect, but the effect of the dosage of the other gamete was significant whether the CWW triploid was used as a seed or a pollen parent (Fisher's exact test P=0.022 and P=0.003, respectively, Table 3). There was thus a strong selection against imbalance in gamete ploidy in the crosses between the triploid and diploid Col-0, even in situations that were devoid of aneuploidy.
Chromosome-specific effects are visible in the aneuploid progeny
We investigated the inheritance of each chromosome type by comparing the number of successful aneuploid gametes receiving one copy (monosomic) with the number of gametes that received two copies (disomic) from the triploid parent. Therefore, the surviving aneuploid individuals were classified according to the number of copies of each chromosome type that they had received (Figure 4 and Table 4). Within a pBC population, the percentage of disomic gametes varied considerably for different chromosome types. Similarly, for a specific chromosome type, the ratio of monosomic to disomic gametes varied depending on the type of cross (Figure 4). These results suggest the existence of chromosome-specific factors influencing the rate of chromosomal transmission.
The parent-of-origin effect and the effect of the ploidy of the crossing partner on transmission rates were assessed by comparing the proportions of monosomic and disomic gametes using pairwise χ2-tests, as presented in Figure 4. Each of the five chromosome types exhibited a specific pattern of transmission, as described below.
The most dramatic differences in the apparent transmission rate were observed for chromosome 1. The transmission of chromosome 1 was strongly affected by parent-of-origin effect irrespective of dosage. Transmission was also affected by the ploidy of the pollen grains in crosses to triploid seed parent. The effect of chromosome 1 dosage in males could not be assessed robustly because of the low number of individuals that were disomic for chromosome 1 in these crosses. In other words, pollen grains monosomic for chromosome 1 were strongly favored, irrespective of the ploidy of the other gamete, whereas ovules disomic for chromosome 1 were favored when crossed with diploid pollen.
The inheritance of chromosome 2 was unaffected by the dosage of the other gamete. The direction of the cross did affect the inheritance of chromosome 2 in crosses to 4x-Col. In this case, pollen grains that carried two copies of chromosome 2 were favored in crosses to tetraploid individuals.
Chromosome 3 was the only chromosome type whose inheritance was significantly affected by the ploidy of the other gamete when transmitted both through the male and the female gametophytes. This effect favored disomic gametes for chromosome 3 when crossed with 4x-Col and monosomic gametes when crossed with diploid Col-0. No parent-of-origin effects could be detected.
The inheritance of chromosome 4 uncovered both a parent-of-origin effect and an effect of the ploidy of the other gamete. Specifically, pollen grains that carried two copies of chromosome 4 were favored in crosses to tetraploid individuals.
Finally, chromosome 5 was the only one for which no significant differences could be detected on the basis of either parent-of-origin effect or genomic dosage of the crossing partner.
Discussion
The progeny of a triploid is expected to be shaped by various mechanisms acting sequentially (Figure 6). In this study, we have attempted to better characterize the role of each of these mechanisms that shape the progeny of triploid A. thaliana. We have performed crosses between triploids and either diploid Col-0 or 4x-Col (Figure 1), such that we can identify parent-of-origin effects as well as the effect of the ploidy of the other (euploid) gamete on karyotype selection in the progeny of the triploid.
Triploid A. thaliana produces mostly aneuploid microspores
By definition, triploids are meiotically unstable because three sets of chromosomes must be resolved to two poles. For each triplet type, the simplest model involves movement of one chromosome to one pole and of the other two chromosomes to the other pole at anaphase I. However, many synaptic arrangements of the triplet and crossing-over locations are possible and can result in chromosome loss (McClintock, 1929; Satina and Blakeslee, 1937b; Singh, 2003) or in the migration of all three copies to a single pole. In this study, we have determined the distribution of chromosome numbers in microspores produced by triploid A. thaliana (Figure 3). We detected a mild selection for euploid gametes but no significant chromosome loss. In addition, we never detected the migration of all three copies of a given chromosome type to a single pole in the 262 fully karyotype plants under study.
In triploids of many species, differences in male and female meiosis produce male and female gametes with different sets of karyotypes (McClintock, 1929; Satina and Blakeslee, 1937a, 1938; Singh, 2003). Owing to the difficulty in obtaining and observing high numbers of ovules, we did not perform a similar analysis on megaspores. Therefore, it is possible that the distribution of chromosome numbers in the ovules is different from that observed in microspores.
Aneuploidy in the gametes or the zygote is generally viable
After gamete production, the chromosomal imbalance associated with aneuploidy can lead to modified ratios of dosage-sensitive factors required for viability, resulting in selection against specific karyotypes (Birchler et al., 2001).
Aneuploidy in the gamete was expected to be particularly deleterious because of the more drastic change in relative dosage in a haploid background compared with a diploid or polyploid background for the zygote or endosperm. In addition, cell divisions associated with gamete maturation into gametophytes provide an opportunity for dosage imbalance to disrupt development and cellular function. Yet, of the 30 possible aneuploid gametic karyotypes, 27 were recovered in the ovule and 16 were recovered in the pollen grains (Table 1). The fact that selection against aneuploid pollen grains appears to be stronger than the selection against aneuploid ovules does not necessarily indicate that aneuploidy is less viable in pollen grains. More probably, dosage imbalance reduces the fitness of aneuploid pollen grains, making them less likely to compete successfully for ovule fertilization. This is especially true in situations in which pollen grains are in excess, such as in hand-pollinated Arabidopsis (Cole et al., 2005).
Of the 60 possible aneuploid zygotic karyotypes associated with a GC intermediate between diploidy and tetraploidy, 44 were recovered in the progeny of the pBCs. Aneuploidy was expected to be less severe in high-dosage backgrounds than in low-dosage backgrounds (Khush, 1973; Ramsey and Schemske, 1998; Vizir and Mulligan, 1999; Birchler et al., 2001). Yet, the overall percentage of aneuploid individuals produced in the crosses to 4x-Col (63.7%) and in the crosses to 2x-Col (61.5%) was similar, suggesting that the effects of genome doubling on buffering the effects of aneuploidy are minimal.
Finally, although it is possible that some of the missing gametic or zygotic karyotypes are indeed deleterious, it is also possible that those karyotypes were simply not recovered due to low sample size. Indeed, in a recent study, 35 individuals produced from selfed triploids of Col-0 were karyotyped using comparative genome hybridization (Huettel et al., 2008). Among the karyotypes observed in this population, there were three karyotypes which were not observed in our populations: one individual was disomic for all chromosome types except for chromosome 4 (14 chromosomes), two individuals were triploids for all chromosomes and tetrasomic for chromosome 5 (16 chromosomes) and one individual was trisomic for chromosomes 1, 3 and 5 but was tetrasomic for chromosomes 2 and 5 (17 chromosomes) (see Supplementary Table S1 in (Huettel et al., 2008)). Whether these karyotypes were recovered in the other study merely by chance, because they were produced by a selfed triploid or because they were genotypically Col-0, remains undetermined. Yet, these results combined with those of our study suggest that most aneuploid karyotypes are viable, both in gametes and in zygotes.
Mechanisms of selection operating at or after fertilization
During or soon after fertilization, dosage effects that are dependent on the chromosomal composition of both mating gametophytes can affect seed development, either by affecting the zygote itself or the endosperm (Figure 6). These gametophytic effects can be due to maternally stored products, products contributed by the gametophytes (both in equal or unequal proportions) and products of imprinted genes (Dilkes and Comai, 2004). In addition, seed development can also be affected by biparental determinants, expressed in the zygote or the endosperm (Figure 6). When the action of these factors is dosage sensitive, their altered contributions by gametes of different genomic constitutions, aneuploid or not, can result in seed failure. For example, the altered dosage of specific gene products is believed to be responsible for the failure of interploidy crosses (Birchler, 1993; Scott et al., 1998; Carputo et al., 2003; Dilkes and Comai, 2004) and crosses involving aneuploid individuals (Khush, 1973; Ramsey and Schemske, 1998; Vizir and Mulligan, 1999; Birchler et al., 2001). Strikingly, species exhibiting high triploid fertility and a highly diverse set of aneuploid progeny are also successful in interploidy crosses (Levan, 1942; Ramsey and Schemske, 1998), suggesting that the mechanisms regulating the success of interploidy crosses and those shaping triploid output might be closely related.
Selection on aneuploid survival
All crosses produced less aneuploid progeny than expected, with the exception of the 3 × 4 cross, which produced the highest percentage of aneuploid progeny (74.5%), which was indistinguishable from the observed frequency of aneuploids in microspores (88%, Table 3). Considering that the distribution of aneuploid ovules produced by the CWW triploid in the 3 × 2 cross are similar to those produced in the 3 × 4 cross, this suggests that selection against aneuploidy occurred post fertilization. Moreover, the 3 × 4 cross exhibited the highest percentage of visible seed death, much higher than the 3 × 2 cross (Figure 5). Taken together, these results suggest that selection against aneuploidy in the 3 × 2 cross occurred very early after fertilization, such that seed development did not proceed long enough to produce visible dead seeds. Similarly, the 2 × 3 and 4 × 3 crosses produced very different populations of aneuploid individuals. The population of pollen grains produced by the CWW triploid must have thus been very diverse in terms of karyotypes. Yet, only certain pollen karyotypes, different depending on the cross, went on to produce viable progeny. Again, these data along with the low seed death recorded for those two crosses argue for the very early failure of seed development after fertilization. Microscopic observations of immature siliques will be necessary to confirm this hypothesis.
The distribution of GC of the aneuploids recovered from the different crosses was affected by the dosage of the crossing partner. Specifically, crosses to haploid gametes strongly favored the survival of aneuploids of low GC (average chromosome number of 6.6), whereas crosses to diploid gametes did not (average chromosome number of 7.5). Selection of the ‘low’ GC aneuploids in crosses to diploid Col-0 could be explained by a need to minimize the difference in GC between the two gametes. However, the lack of similar selection in the CWW × 4x-Col cross cannot be explained by this hypothesis.
Selection on euploid gametes
Within the euploid progeny produced by any given cross, the number of progeny of high and low chromosome numbers is expected to be the same because fully haploid and fully diploid gametes have equal probabilities. For example, the 3 × 2 cross is expected to produce as many diploids as triploid progeny. As the production of diploid or triploid individuals does not involve aneuploidy, variation from this expectation would indicate selection against fertilization of gametes of different ploidy even in the absence of aneuploidy in the gametes or in the fertilization products. This effect was very strong in the pBCs (Table 3, comparison of haploid ratio with diploid gametes). Specifically, triploids crossed with diploid Col-0 produced much fewer triploid individuals than expected. On the other hand, in crosses to 4x-Col, the expected ratio (close to 1:1) of triploid and tetraploid individuals was produced. Again, as the populations of gametes produced by the triploid are the same irrespective of whether it will be crossed with diploid Col-0 or 4x-Col, these data suggest selection against specific ploidy mismatches between the gametes after fertilization.
The low percentage of seed viability in the CWW × 4x-Col cross is reminiscent of that observed when crossing individuals of different ploidy, and which are believed to result from altered paternal to maternal ratios in the zygote or in the endosperm (Birchler, 1993; Scott et al., 1998; Carputo et al., 2003). Such phenomena have been previously observed for A. thaliana in which crosses between diploid and tetraploid individuals result in low seed viability, especially in cases of paternal excess (Scott et al., 1998; Dilkes et al., 2008). A similar phenomenon is observed in this study, in which the CWW × 4x-Col and CWW × Wa-1 crosses both exhibited very low seed viability, as compared with the other crosses.
Nevertheless, paternal excess does not suffice to explain the observed ratios of euploid individuals. For example, similar proportions of triploids and tetraploid individuals in the 3 × 4 crosses are unexpected. Indeed, considering how deleterious paternal excess is in a 2 × 4 scenario, one would have expected a much higher number of tetraploid individuals than triploid individuals in the progeny of the 3 × 4 cross. Interestingly, this trend is consistent with that observed in aneuploid chromosome numbers, as outlined in the previous section, where no selection against low chromosome number aneuploid gametes was visible in the crosses to 4x-Col. Moreover, if the 3 × 4 cross can produce triploid individuals by fertilization of a haploid ovule by a diploid pollen grain, why are there so few triploids produced in the 2 × 3 cross? Perhaps the finding that gene expression in the maternal sporophyte is a critical determinant of paternal-excess survival (Dilkes et al., 2008) can explain these results. The imbalance between the triploid maternal sporophyte and the triploid embryo and tetraploid endosperm in a 3 × 4 cross is substantially less than the diploid sporophyte to the same embryo and endosperm ploidies in the 2 × 3 cross. Indeed, the latter example is identical in seed makeup to the interploidy paternal excess crosses between diploid and tetraploid A. thaliana, which results in substantial seed lethality (Dilkes et al., 2008). Similarly, 4 × 2 interploidy crosses (in Col-0) normally exhibit high seed viability and readily produce triploid individuals. Yet, very few triploid individuals are produced from the 3 × 2 cross, possibly because of the lower ploidy of the maternal sporophyte.
Chromosome-specific patterns of inheritance in aneuploids
In many species, the rate of transmission of extra chromosomes in trisomics is highly dependent on the chromosome type (Khush, 1973; Singh, 2003). In addition, transmission of extra chromosomal copies through males is often severely reduced compared with transmission through females (Khush, 1973; Singh, 2003). Similarly, in maize, extra copies of chromosomal segments are not transmitted as efficiently through males as through females (Auger and Birchler, 2002). Our analysis of chromosome transmission in the aneuploid individuals of the pBCs suggests that these observations are also true in the progeny of triploids, in which all chromosome types are potentially imbalanced (Figure 4 and Table 4). Moreover, comparison of different crosses allows for the identification of potential dosage-sensitive loci on specific chromosomes.
As mentioned earlier, the pattern of inheritance of chromosome 1 varied mostly between crosses. The overrepresentation of progeny from ovules that were disomic for chromosome 1 in the 3 × 4 cross and from those monosomic for chromosome 1 in the 2 × 3 cross was consistent with selection against paternal excess of chromosome 1 after fertilization. Such a phenotype could be caused by a need for balance between a maternal factor encoded on chromosome 1 and paternal factors. This maternal factor encoded on chromosome 1 could either be contributed by the ovule or paternally imprinted, such as MEDEA situated at the top of chromosome 1 (Grossniklaus et al., 1998). To the contrary and somewhat surprisingly, maternal excess was overall preferred to balanced numbers of chromosome 1 in the 3 × 2 and 4 × 3 crosses (Table 4). Whether the two effects can be attributed to the same or different loci on chromosome 1 remains to be determined. Opposite but weaker trends were observed for the inheritance of chromosome 2 in which disomy was favored through the pollen and monosomy was favored in the ovule, irrespective of the ploidy of the other gamete (Table 4 and Figure 4). It is possible that the analysis of larger populations from reciprocal crosses in combination with marker transmission analysis could lead to the identification of maternally and paternally expressed dosage-sensitive loci required for seed viability.
The pattern of inheritance of chromosome 4 was also noteworthy. Specifically, the percentage of successful gametes that were disomic for chromosome 4 was significantly higher in the 4 × 3 cross than in all other crosses, suggesting a strong selection against maternal excess at high dosage. This effect could be linked, for example, to a gene located on chromosome 4 and expressed only in the male gametophyte. Alternatively, the gene could be maternally imprinted and require a balance with maternal factors encoded elsewhere in the genome. However, this effect was not observed at a low dosage (the 3 × 2 cross).
Finally, the inheritance of chromosome 3 was strongly affected by the dosage of the crossing partner, irrespective of the direction of the cross and such that gametes containing the same number of copies of chromosome 3 as the other gamete were strongly favored. This behavior would be consistent with the presence on chromosome 3 of a dosage-sensitive factor either contributed equally by both gametophytes or expressed in the fertilization products.
Analysis of bigger populations in a similar manner might uncover additional effects, such as the preferential inheritance of specific combinations of chromosomes for example, which could lead to the identification of interactions between dosage-sensitive loci located on different chromosomes.
In summary, our results show a strong tolerance of Arabidopsis gametes and zygotes not only to trisomy but also to all types of aneuploidy, including complex karyotypes in which more than one chromosome type is imbalanced. Although the selection against aneuploidy is visible, our results suggest that it is mostly due to a very early failure of seed development. Dosage of the crossing partner most influenced the type of surviving aneuploids and euploid progeny. Finally, chromosome-specific effects could be detected, suggesting that a genotypic analysis of populations similar to those presented in this study may result in the identification of dosage-sensitive and possibly imprinted factors, required for successful fertilization events.
References
Auger D, Birchler J (2002). Maize tertiary trisomic stocks derived from B_A translocations. J Heredity 93: 42–47.
Birchler JA (1993). Dosage analysis of maize endosperm development. Annu Rev Genet 27: 181–204.
Birchler JA, Bhadra U, Bhadra MP, Auger DL (2001). Dosage-dependent gene regulation in multicellular eukaryotes: implications for dosage compensation, aneuploid syndromes, and quantitative traits. Dev Biol 234: 275–288.
Carputo D, Frusciante L, Peloquin SJ (2003). The role of 2n gametes and endosperm balance number in the origin and evolution of polyploids in the tuber-bearing Solanums. Genetics 163: 287–294.
Cole RA, Synek L, Zarsky V, Fowler JE (2005). SEC8, a subunit of the putative Arabidopsis exocyst complex, facilitates pollen germination and competitive pollen tube growth. Plant Physiol 138: 2005–2018.
Comai L, Tyagi AP, Lysak MA (2003). FISH analysis of meiosis in Arabidopsis allopolyploids. Chromosome Res 11: 217–226.
Dilkes BP, Comai L (2004). A differential dosage hypothesis for parental effects in seed development. Plant Cell 16: 3174–3180.
Dilkes BP, Spielman M, Weizbauer R, Watson B, Burkart-Waco D, Scott RJ et al. (2008). The maternally expressed WRKY transcription factor TTG2 controls lethality in interploidy crosses of Arabidopsis. PLoS Biol 6: 2707–2720.
Epstein C (1986). The Consequences of Chromosomal Imbalance, Vol 18. Cambridge University Press: Cambridge, 486pp.
Grossniklaus U, Vielle-Calzada JP, Hoeppner MA, Gagliano WB (1998). Maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis. Science 280: 446–450.
Henry IM, Dilkes BP, Comai L (2006). Molecular karyotyping and aneuploidy detection in Arabidopsis thaliana using quantitative fluorescent polymerase chain reaction. Plant J 48: 307–319.
Henry IM, Dilkes BP, Young K, Watson B, Wu H, Comai L (2005). Aneuploidy and genetic variation in the Arabidopsis thaliana triploid response. Genetics 170: 1979–1988.
Huettel B, Kreil DP, Matzke M, Matzke AJ (2008). Effects of aneuploidy on genome structure, expression, and interphase organization in Arabidopsis thaliana. PLoS Genet 4: e1000226.
Khush G (1973). Cytogenetics of Aneuploids. Academic Press: New York, 301pp.
Levan A (1942). The effect of chromosomal variation in sugar beets. Hereditas 28: 345–399.
McClintock B (1929). A cytological and genetical study of triploid maize. Genetics 14: 180–222.
Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomajian C, Zheng H et al. (2005). The pattern of polymorphism in Arabidopsis thaliana. PLoS Biol 3: e196.
Punyasingh K (1947). Chromosome numbers in crosses of diploid, triploid and tetraploid maize. Genetics 32: 541–554.
Ramsey J, Schemske DW (1998). Pathways, mechanisms, and rates of polyploid formation in flowering plants. Ann Rev Ecol Syst 29: 467–501.
Ramsey J, Schemske DW (2002). Neopolyploidy in flowering plants. Annual Review of Ecology and Systematics 33: 589–639.
Satina S, Blakeslee A (1937a). Chromosome behavior in triploids of Datura stramonium. I. The male gametophyte. Am J Bot 24: 518–526.
Satina S, Blakeslee A (1937b). Chromosome behavior in triploids of Datura stramonium. II. The female gametophyte. Am J Bot 24: 621–627.
Satina S, Blakeslee A (1938). Chromosome behavior in triploid Datura. III. The seed. Am J Bot 25: 595–602.
Scott RJ, Spielman M, Bailey J, Dickinson HG (1998). Parent-of-origin effects on seed development in Arabidopsis thaliana. Development 125: 3329–3341.
Singh R (2003). Plant Cytogenetics. CRC Press: Boca Raton.
Soltis D, Soltis P, Tate J (2003). Advances in the study of polyploidy since plant speciation. New Phytol 161: 173–191.
Vizir I, Mulligan B (1999). Genetics of gamma-irradiation-induced mutations in Arabidopsis thaliana: large chromosomal deletions can be rescued through the fertilization of diploid eggs. J Heredity 90: 412–417.
Wolfe KH, Shields DC (1997). Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387: 708–713.
Acknowledgements
We thank the many anonymous reviewers who contributed considerable time and insight into the analysis of the data. We thank the Comparative Genome Center (Department of Biology, University of Washington) and the Murdock Foundation for use of the ABI3100 Genetic Analyzer and of software for data analysis. We thank the Cell Analysis Facility (Department of Immunology, University of Washington supported by NIH S10 RR021167) and Biology Greenhouse (Biology Department, University of Washington) for material support. We thank the Magnus Nordborg Laboratory for the sequence database of A. thaliana ecotypes (Nordborg et al., 2005). This work was supported by the National Science Foundation ‘Functional genomics of plant polyploids’ Project (NSF DBI 0077774 and DBI 0501712 to LC), the USDA National Research Initiative (2003-35300-13248 to BPD) and the Washington Research Foundation (to IMH).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Rights and permissions
About this article
Cite this article
Henry, I., Dilkes, B., Tyagi, A. et al. Dosage and parent-of-origin effects shaping aneuploid swarms in A. thaliana. Heredity 103, 458–468 (2009). https://doi.org/10.1038/hdy.2009.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.81
Keywords
This article is cited by
-
Potato Introgressive Hybridisation Breeding for Bacterial Wilt Resistance Using Solanum commersonii Dun. as Donor: Genetic and Agronomic Characterisation of a Backcross 3 Progeny
Potato Research (2022)
-
Genome evolution and phylogenetic relationships in Opuntia tehuacana (Cactaceae, Opuntioideae)
Brazilian Journal of Botany (2022)
-
Do pentaploid hybrids mediate gene flow between tetraploid Senecio disjunctus and hexaploid S. carniolicus s. str. (S. carniolicus aggregate, Asteraceae)?
Alpine Botany (2021)
-
Transgenerational effects of inter-ploidy cross direction on reproduction and F2 seed development of Arabidopsis thaliana F1 hybrid triploids
Plant Reproduction (2019)
-
Pairing analysis and in situ Hybridisation reveal autopolyploid-like behaviour in Solanum commersonii × S. tuberosum (potato) interspecific hybrids
Euphytica (2017)