Abstract
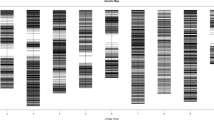
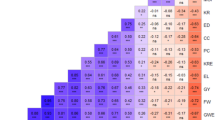
Amplified fragment length polymorphisms (AFLPs) produced with EcoRI and PstI both in combination with MseI restriction enzymes have been studied in the parents of four barley mapping populations. Averages of 15.9 and 18.7 polymorphic products per assay were produced for the EcoRI/MseI and PstI/MseI combinations, respectively. There was some evidence of interaction between cross combinations and restriction enzyme combinations, with PstI/MseI generating relatively more polymorphic products than EcoRI/MseI in the Blenheim × E224/3 cross combination, the least polymorphic of the four. Three hundred and ninety-eight AFLP products, using both restriction enzyme combinations, were generated in a doubled haploid population of 68 lines produced from the Blenheim × E224/3 cross. These were added to existing marker data for the cross to study the effects of incorporation of AFLPs produced by different restriction enzyme combinations upon genetic maps. Addition of the AFLP data resulted in greater genome coverage, both through linking previously separate groups and extensions to other groups. This increase in coverage appeared to result from AFLPs sampling some different regions of the genome compared to RAPDs and RFLPs, as the map distances spanned by the RAPD and RFLP linkage groups were similar with and without incorporation of AFLPs. There was also evidence that the EcoRI and PstI restriction enzymes sampled different regions of the genome. The revised maps were used in scanning for QTLs controlling a subset of 12 economically important traits measured in the cross. Overall, the QTLs accounted for an average of 53 per cent of the phenotypic variation for the characters. Positive and negative alleles were present in each parent for each character, apart from hot water extract corrected to 1.5 per cent nitrogen (HWEc). Several regions of the genome appeared to be involved in the control of several characters, notably chromosome 2, the denso locus on chromosome 3, the short arm of chromosome 5 and chromosome 7. Although there was considerable similarity to previous results of QTL mapping for the subset of characters, the greater genome coverage afforded by the inclusion of the AFLPs revealed some new QTL locations.
Similar content being viewed by others
Article PDF
References
Amasino, R M, John, M C, Klaas, M, and Crowell, D M. 1990. Role of DNA methylation in the regulation of gene expression in plants. In: Clawson G. A., Willis, D. B., Weisbach, A. and Jones, P. (eds) Nucleic Acid Methylation, pp. 187–198. Wiley-Liss, New York.
Becker, J, and Heun, M. 1995. Barley microsatellites: allele variation and mapping. Plant Mol Biol, 27, 835–845.
Becker, J, Vos, P, Kuiper, M, Salamini, F, and Heun, M. 1995. Combined mapping of AFLP and RFLP markers in barley. Mol Gen Genet, 249, 65–73.
Botstein, D, White, R L, Skolnick, M H, and Davis, R W. 1980. Construction of a genetic map in man using restriction fragment length polymorphisms. Am J Hum Genet, 32, 314–331.
Devaux, P, Kiuan, A, and Kleinhofs, A. 1993. Anther culture and Hordeum bulbosum derived barley doubled haploids: mutations and methylation. Mol Gen Genet, 241, 674–679.
Folkertsma, R T, Van Der Voort Rouppe, J N A M, De Groot, K E, Van Zandvoort, P M, Schots, A, and Gommers, F J. et al. 1996. Gene pool similarities of potato cyst nematode populations assessed by AFLP analysis. Mol Pl-Micr Interact, 9, 47–54.
Graner, A, Jahoor, A, Schondelmaier, J, Siedler, H, Pillen, K, and Fischbeck, G. et al. 1991. Construction of an RFLP map in barley. Theor Appl Genet, 83, 250–256.
Gruenbaum, Y, Naveh-Many, T, Cedar, H, and Razin, A. 1981. Sequence specificity of methylation in higher plant DNA. Nature, 292, 860–862.
Helentjaris, T, and Burr, B. 1989. Development and Application of Molecular Markers to Problems in Plant Genetics Current Communications in Molecular Biology. Cold Spring Harbor, Laboratory Press, Cold Spring Harbor. NY.
Heun, M, Kennedy, A E, Anderson, J A, Lapitan, N L V, Sorells, M E, and Tanksley, S D. 1991. Construction of a restriction fragment length polymorphism map for barley (Hordeum vulgare). Genome, 34, 437–447.
Lin, J J, and Kuo, J. 1995. AFLPTM: A novel PCR-based assay for plant and bacterial DNA fingerprinting. Focus, 17, 51–56.
Meksem, K, Leister, D, Pepeman, J, Zabeau, M, Salamini, F, and Gebhardt, C. 1995. A high resolution map on potato chromosome V based on RFLP and AFLP markers. Mol Gen Genet, 249, 74–81.
Mullis, K, Faloona, S, Schrf, S, Saiki, R, Horn, G, and Erlich, H. 1986. Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harb Symp Quant Biol, 51, 263–273.
Powell, W, Morgante, M, Vogel, J, Tingey, S, and Rafalski, J A. 1994. Technology for plant genome analysis and breeding. In: Javornik, B., Bohanec, B. and Kreft, I. (eds) Proc Int Colloq Impact of Plant Biotechnology on Agriculture, pp. 177–180. Planprint, Ljubljana.
Powell, W, Bonar, N, Baird, E, Russell, J, and Waugh, R. 1995. Molecular marker techniques for barley genome analysis and breeding. Ann Rep Scot Crop Res Inst 1994, 57–58.
Powell, W, Morgante, M, Andre, C, Hanafey, M, Vogel, J, Tingey, S, and Rafalski, A. 1996. The utility of RFLP. RAPD. AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed, 2, 225–238.
Rafalski, A J, Morgante, M, Vogel, J, Powell, W, and Tingey, S V. 1996. Generating new DNA markers in plants. In: Birren, B. and Lai, E. (eds) Analysis of Non-mammalian Genomes, pp. 75–129, Academic Press, London.
Saghai-Maroof, M A, Soliman, K M, Jorgensen, R A, and Allard, R W. 1984. Ribosomal DNA spacer length polymorphism in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA, 81, 8014–8018.
Stam, P, and Van Oouen, J W. 1995. JoiniTiap (tm) version 2.0: Software for the calculation of genetic linkage maps. CPRO-DLO Wageningen.
Tautz, D. 1989. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucl Acids Res, 17, 6463–6471.
Thomas, W T B, Powell, W, and Swanston, J S. 1991. The effects of major genes on quantitatively varying characters in barley. 4. The GPert and denso loci and quality characters. Heredity, 66, 381–389.
Thomas, C M, Vos, P, Zabeau, M, Jones, D A, Norcott, K A, Chadwick, B P, and Jones, J D G. 1995a. Identification of amplified restriction fragment polymorphism (AFLP) markers tightly linked to the tomato Cf-9 gene for resistance to Cladosporium fulvum. Plant J, 8, 785–794.
Thomas, W T B, Powell, W, Waugh, R, Chalmers, K J, and Barua, U M, Jack, P. et al. 1995b. Detection of quantitative trait loci for agronomic, yield, grain and disease characters in spring barley (H. vulgare L.). TheorAppl Genet, 91, 1037–1047.
Thomas, W T B, Powell, W, Swanston, J S, Ellis, R P, Chalmers, K J, and Barua, U M. et al. 1996. Quantitative trait loci for germination and malting quality characters in a spring barley cross. Crop Sci, 36, 265–273.
Tinker, N A, and Mather, D E. 1995. Methods for QTL analysis in progeny replicated in multiple environments. J Quant Trait Loci, http://probe.nalusda.gov:8000/otherdocs/jqtl/jqtl1995-01/jqtl15.html
Tinker, N A, Mather, D E, Rossnagel, B G, Kasha, K J, Kleinhofs, A, and Hayes, P. et al. 1996. Regions of the genome than affect agronomic performance in two row barley. Crop Sci, 36, 1053–1062.
Vos, P, Hogers, R, Bleeker, M, Reuans, M, Van Der Lee, T, and Hornes, M. et al. 1995. AFLP: a new concept for DNA fingerprinting. Nucl Acids Res, 23, 4407–4414.
Weber, J, and May, P E. 1989. Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am J Hum Genet, 44, 388–396.
Welsh, J, and McClelland, J. 1990. Fingerprinting genomes using PCR with arbitrary primers. Nucl Acid Res, 18, 6531–6535.
Williams, J G K, Kubelik, A R, Livak, K J, Rafalski, A J, and Tingey, S V. 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl Acids Res, 18, 6531–6535.
Zabeau, M, and Vos, P. 1993. Selective restriction fragment amplification: a general method for DNA fingerprinting. European Patent Application 92402629.7 Publication Number EP 0534858 A1.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Powell, W., Thomas, W., Baird, E. et al. Analysis of quantitative traits in barley by the use of Amplified Fragment Length Polymorphisms. Heredity 79, 48–59 (1997). https://doi.org/10.1038/hdy.1997.122
Received:
Issue Date:
DOI: https://doi.org/10.1038/hdy.1997.122
Keywords
This article is cited by
-
Genetic characterisation, expression and association of quality traits and grain texture in barley (Hordeum vulgare L.)
Euphytica (2016)
-
Molecular characterization and functional analysis of barley semi-dwarf mutant Riso no. 9265
BMC Genomics (2015)
-
Mapping a major QTL for malt extract of barley from a cross between TX9425 × Naso Nijo
Theoretical and Applied Genetics (2015)
-
A genetic linkage map based on AFLP and SSR markers and mapping of QTL for dry-matter content in sweetpotato
Molecular Breeding (2013)
-
Genome-wide association mapping of agronomic traits in relevant barley germplasm in Uruguay
Molecular Breeding (2013)