Abstract
Purpose: Lynch syndrome is a genetic disease that predisposes to colorectal tumors, caused by mutation in mismatch repair genes. The use of genetic tests to identify mutation carriers does not always give perfectly clear results, as happens when an unclassified variant is found. This study aimed to define the pathogenic role of 35 variants present in MSH2, MLH1, MSH6, and PMS2 genes identified in our 15-year case study.
Methods: We collected clinical and molecular data of all carriers, and then we analyzed the variants pathogenic role with web tools and molecular analyses. Using a Bayesian approach, we derived a posterior probability of pathogenicity and classified each variant according to a standardized five-class system.
Results: The MSH2 p.Pro349Arg, p.Met688Arg, the MLH1 p.Gly67Arg, p.Thr82Ala, p.Lys618Ala, the MSH6 p.Ala1236Pro, and the PMS2 p.Arg20Gln were classified as pathogenic, and the MSH2 p.Cys697Arg and the PMS2 p.Ser46Ile were classified as likely pathogenic. Seven variants were likely nonpathogenic, 3 were nonpathogenic, and 16 remained uncertain.
Conclusion: Quantitative assessment of several parameters and their integration in a multifactorial likelihood model is the method of choice for classifying the variants. As such classifications can be associated with surveillance and testing recommendations, the results and the method developed in our study can be useful for helping laboratory geneticists in evaluation of genetic tests and clinicians in the management of carriers.
Similar content being viewed by others
Main
Lynch syndrome, also known as Hereditary Non-Polyposis Colorectal Cancer, is an autosomal dominant syndrome caused by mutations in one of the mismatch repair (MMR) genes, namely MLH1, MSH2, MSH6, and PMS2.1 These genes are involved in the repair of DNA polymerase errors, especially those involving one or few base pairs. The loss of MMR function causes the accumulation of mutations, particularly in tandem repeat sequences leading to microsatellite instability (MSI), a typical marker of Hereditary Non-Polyposis Colorectal Cancer tumors.
Genetic tests are offered to probands that fulfill Amsterdam clinical criteria and/or that develop MSI tumors at young ages1; the finding of a clearly pathogenic mutation in a family allows a better surveillance of carriers. However, mutation screening with advanced DNA sequence technologies leads to the detection of an increasing number of missense, silent, and intronic variants. These changes often do not produce truncated proteins upon translation, like most disease-causing pathogenic mutations, and are so-called unclassified variants (UVs). Their relation to disease development is often not clear, but it is very important for the risk determination of the carrier to offer an adequate follow-up. This is the reason why there has been an intense debate for many years, and especially in the past 2 years, on how to assess mutation pathogenicity in cancer susceptibility genes.2,3
This work is in line with most recent publications and aims to provide some helpful data for the purpose of UVs classification.4–12 It simply represents an initial attempt to evaluate UVs of MMR genes based on a relatively elementary quantitative assessment of several parameters, integrated in a multifactorial likelihood model.13
MATERIALS AND METHODS
Patients and controls
From 1994 to 2008, we performed genetic tests of MMR genes in 306 unrelated probands that fulfilled Amsterdam and/or Bethesda criteria (Table 1) who were recruited mainly at the C.R.O. National Cancer Institute in Aviano (recorded as A-AV), and at collaborating Centers in Padova (A-PD), Varese (A-VA), Modena (A-MD), and Montecchio (A-VR). Immunohistochemistry (IHC) and MSI were not used as selection criteria for enrollment. We selected all patients with one or more UV and, when available, tested the relatives of the proband. For every person involved in the UV analysis, we collected, under informed consent, blood samples from which we obtained DNA. Where possible, we also obtained RNA from lymphoblastoid cell lines and DNA from tumor tissues. As controls, we collected DNA from 90 unrelated subjects with negative colonoscopy (clean colon) at ages ranging from 23 to 86 years (mean 63.42 ± 13.36).
Mutation, MSI, and IHC
The DNA samples of probands were sequenced for the entire open reading frame and flanking intronic sequences of MSH2, MLH1, MSH6, and PMS2 genes. Tumor DNAs were tested only for the mutation detected in blood DNA and evaluated for loss of heterozygosity (LOH) by DNA sequencing. MSI and IHC analyses were carried out and evaluated on tumor tissues by conventional methods.14,15
cDNA splicing and primer extension analyses
cDNA obtained from mRNA extracted from lymphoblastoid cell lines was polymerase chain reaction (PCR)-amplified and analyzed by agarose gel electrophoresis to evaluate a potential involvement of the UV nucleotide in the correct splicing signaling. Primers used can be requested from authors.
A primer extension method was used to analyze cDNA allelic expression. Single nucleotide primer extension was carried out in a final volume of 10 μl containing 0.25 pmol of purified PCR product, 0.2 μM of reverse primer located downstream of the mutation site, and 5 μl of SNaPshot Multiplex Ready Reaction Mix (Applied Biosystems, Foster City, CA). The reaction was performed as recommended by the manufacturer in a thermal cycler (10 cycles), run on the ABI PRISM 3100 Genetic Analyzer (Applied Biosystems) and evaluated with GeneScan software. The electropherogram was used to calculate the ratio between mutated and wild-type peak areas in both gDNA and cDNA.
Denaturing high-performance liquid chromatography analysis
The same PCR reactions set up for mutational screening were prepared for healthy controls, with a reference sample carrying the UV for every amplification. If a variant was present, denaturation followed by slow renaturation leads to the formation of heteroduplexes that are detected by a denaturing high-performance liquid chromatography analysis instrument (Transgenomics Inc., Omaha, NE). To confirm UV presence, every sample with a different chromatogram was sequenced.
In silico analysis
For the in silico evaluation of the selected UVs, the bioinformatics tools reported in Table 2 were used. We used the default threshold and collected all the results supplied by the tools exactly as they emerged. The A-GVGD value was obtained through the Alamut software (Interactive Biosoftware, Rouen, France) that uses International Agency for Research on Cancer (IARC)-style curated alignments. The A-GVGD Web site alignments were not used because the MSH6 and PMS2 protein alignments have not yet been released by IARC. Data on in vitro functional studies were collected from the available literature.16–23
Statistical evaluation
UVs were classified using likelihood of pathogenicity for both quantitative and qualitative data. For individual variants, we used a Bayesian approach to incorporate direct and indirect evidence of pathogenicity in a single model,4,13 starting from a prior probability of 0.524 and assuming statistical independence of the sources of information. Posterior probability of pathogenicity was obtained from the likelihood ratio and categorized in five classes.6 To test our method, a well-known common polymorphism (MLH1 p.Ile219Val) was introduced as the negative control. For the positive control, there are no well-known pathogenic missense mutations, and hence, we used a truncating one (MSH2 p.Arg406Ter).
Prior odds ratios (OR) of pathogenicity for each variant were calculated from published work applying a continuity correction.25 For in silico analyses, OR were obtained from published results based on different tools: A-GVGD,12 PolyPhen, Pmut, SIFT,26 and splicing tools.27 As in silico predictions are not independent, we calculated one average OR for the three exon splicing enhancer (ESE)-related software tools, one average OR for the two splicing site-related software, and one for the protein-related tools.
For MSI and IHC, we estimated OR from data published by Engel et al.,28 who evaluated these molecular features as predictive of mutations in 1119 unrelated patients. Because of the correlation of absence of protein expression and presence of MSI, a single OR for these two combined data was estimated. When the UV co-occurs with a clearly pathologic mutation, it is impossible to assess whether MSI phenotype is due to the UV, to the pathologic mutation, or both, and we accordingly assigned an OR of 1. This neutral score was not applied when a typical compound heterozygous phenotype was evident.
OR for pathogenicity based on familiarity was derived from a set of patients previously screened for pathogenic mutations: the event was the presence of a clear-cut pathogenic mutation and groups were based on the fulfillment of Amsterdam criteria.
For OR related to co-occurrence, we used the formula proposed by Easton et al.29 Co-occurrence was considered exclusively as the presence of a mutation in trans of the same gene, with the exception of cases with a recognizable biallelic phenotype. For families where the genetic test could be extended to at least one relative, OR of cosegregation of disease and variant were evaluated according to Thompson et al.30 For this analysis, we assumed the age-specific risks and penetrance estimated by Marroni et al.31 for the Italian population.
In instances in which calculating OR was not possible, we arbitrarily assigned an OR of 2 (see Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A127); the value is conservative and was chosen so that data for which a quantitative assessment is difficult would not heavily bias quantifiable evidence. This estimate was tentatively applied with the aim of also incorporating “qualitative data,” as LOH, allelic imbalance, absence in healthy controls, and in vitro functional effect, into the multifactorial model.
RESULTS
We analyzed 35 UVs that are listed in Table 3: 12 MSH2, 15 MLH1, 4 MSH6, and 4 PMS2. Twenty UVs were missense, 7 were silent, and 8 were intronic mutations. To test our system, we also included in the analysis the p.Arg406Ter nonsense mutation of the MSH2 gene and the common p.Ile219Val polymorphism of the MLH1 gene (Table 3). Bibliographic references for some UVs can be found online in the MMR variants database (http://www.med.mun.ca/MMRvariants/).
Globally, we found UVs in a total of 40 probands, corresponding to about 13% of the screened patients. In most cases, every patient carried only one UV, whereas five patients carried two or three UVs each.
Copresence of a clear-cut pathogenic mutation was ascertained for nine UVs (Table 3). Data concerning disease-UV segregation, presence of Amsterdam criteria in the family, MSI, and IHC are also reported in Table 3.
The denaturing high-performance liquid chromatography analysis of at least 50 individuals with clean colon revealed the presence of only 4 UVs, all within the MSH2 gene. Three of them (p.Gly322Asp, p.Leu556Leu, and p.Lys579Lys) were present in only one person and the last one (c.1077-10T>C) was present in four individuals (Table 3). In addition, our previous studies of the PMS2 gene revealed the presence of p.Arg20Gln in 2 of 70 control chromosomes, whereas the p.Ser46Ile variant was not detected in any control.32,33 The LOH analysis was only possible for 21 samples, and we obtained results from 16 tumors. The MSH2 p.Arg406Ter truncating mutation and 8 UVs (MSH2: p.Ala328Ala, p.Met688Arg, and p.Cys697Arg; MLH1: p.Gly67Arg, p.Arg265His, and p.Lys618Ala; MSH6: p.Ala1236Pro; and PMS2: p.Arg20Gln) showed loss of the wild-type allele, MSH2 p.Lys113Lys revealed loss of the variant allele, and the remaining five UVs, as well as the p.Ile219Val polymorphism, did not show LOH (Table 3).
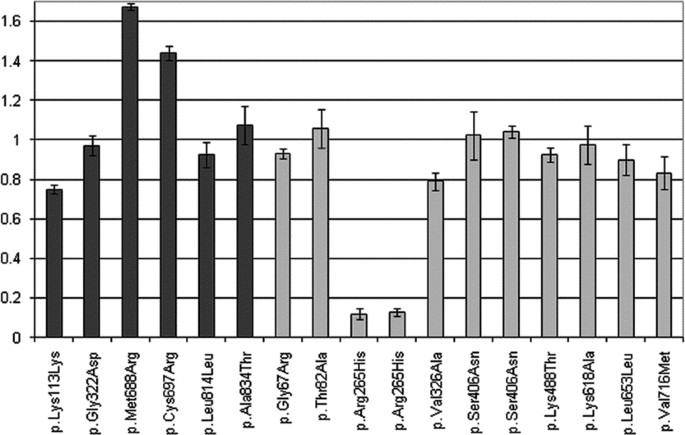
None of the UVs analyzed showed cDNA alteration of the physiological splicing (data not shown). The allelic expression analysis revealed a high imbalance level in the MLH1 p.Arg265His sample, but it was probably due to the in cis presence of the MLH1 c.1011delC frameshift mutation that is predicted to cause mRNA decay. The other UVs did not show a comparably high imbalance level, but MSH2 p.Lys113Lys, p.Met688Arg, p.Cys697Arg, and MLH1 p.Val326Ala showed imbalance ratios higher than 1.2 or lower than 0.8. Data are shown in Figure 1 and reported in Table 3.
Every web-based prediction software used for the analysis rendered a value that is indicative of the predicted effect of the nucleotidic/aminoacidic substitution on the splicing (Table 4) or on the protein (Table 5).
We categorized variants according to the IARC classification based on the posterior probability of causality (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A127). Seven UVs were classified as definitely pathogenic and two as likely pathogenic, whereas three and seven UVs resulted not pathogenic or likely not, respectively. The additional 16 UVs remained of uncertain significance (Table 6). The polymorphism used to test our method was included in the class with the lowest probability, while the truncating mutation, even if lacking the in silico protein prediction, was attributed to the highest class.
DISCUSSION
We evaluated all the UVs collected in our 15 years of genetic testing for Lynch syndrome. This integrated analysis aimed to develop a model for assigning each UV to a pathogenic or a nonpathogenic category and to elucidate their role in Lynch syndrome development. All evaluations were made under the arbitrary assumption that the analyzed variants could have the same impact on molecular and clinical phenotype as the proven pathogenic MMR gene mutations resulting in protein truncation or instability.
Among the UVs, the MSH2 p.Pro349Arg, p.Met688Arg, the MLH1 p.Gly67Arg, p.Thr82Ala, p.Lys618Ala, the MSH6 p.Ala1236Pro, and the PMS2 p.Arg20Gln were the most likely to be pathogenic. All these UVs and two others in the likely pathogenic class (MSH2 p.Cys697Arg and PMS2 p.Ser46Ile) are missense variants, whereas deep intronic and silent variants never reached the highest probability of pathogenicity. In some cases, this lack is associated with a real absence of mRNA alteration, but in other situations, it could be partially due to data absence, because in silico protein prediction is not possible for nonmissense variants.
Ten UVs resulted nonpathogenic or likely nonpathogenic, whereas 16 remained uncertain. However, these three categories could be biased because of paucity of data or conflicting results. Therefore, we cannot exclude that some of the uncertain and nonpathogenic variants could represent low penetrance alleles, unable to fully manifest their (low) pathogenic role in our limited-size families.
Importantly, the positive control p.Arg406Ter presented the highest probability of pathogenicity, whereas the negative control, p.Ile219Val, had one of the lowest. Moreover, three of the class 1 or 2 UVs (MSH2 p.Gly322Asp, MLH1 p.Ser406Asn and p.Val716Met) were classified as benign in a structured assessment study on MMR ambiguous mutations by Barnetson et al.34MSH2 p.Gly322Asp and MLH1 c.1039-8T>A were also considered by Arnold et al.,24 but not studied further, because they were found at polymorphic frequency in unaffected controls. This points to the potential value of our classification scheme.
All the class 4 and class 5 UVs present MSI and/or IHC absence of the UV-carrying protein while almost all showed LOH of the wild-type allele. All are present in at least one family fulfilling the Amsterdam criteria and only one was present in healthy controls. Some of them fall in fundamental functional domains, where the aminoacidic sequence is highly conserved and the perfect folding of the domain is very important. This is the case for MSH2 p.Pro349Arg and p.Cys697Arg, the latter located in the highly conserved adenosine triphosphate domain where a similar variant, p.Cys697Phe, is reported to completely inactivate MSH2.35 It is also the case for MLH1 p.Gly67Arg that is located in the adenosine triphosphate binding region and showed a reduced MMR activity of the mutated protein in functional in vitro assays.16–18 p.Cys697Arg was overexpressed in cDNA. The increased expression of the allele with p.Cys697Arg could be due to an increased transcription or mRNA stability associated with a reduced protein functionality. Some other UVs fall in domains involved in repair function, like MSH2 p.Met688Arg that is located in a site that, when changed, leads to a moderate reduction of repair capacity, as reported for p.Met688Ile.36MLH1 p.Thr82Ala has been reported to reduce MMR activity in vitro19 and seems to cosegregate in this family with p.Lys618Ala, an UV that modifies protein stability leading to increased degradation, without perturbing PMS2 binding and functionality.20 However, in the literature, the outcome of the functional tests for the latter variant are contradictory and it is possible that both UVs have some pathogenic role and they could act in synergy. According to an alternative classification,34 p.Lys618Ala should be a benign variant and consequently only p.Thr82Ala could be truly responsible for the observed phenotype. For MSH6 and PMS2, only p.Ala1236Pro and p.Arg20Gln, respectively, resulted as class 5 pathogenic, but this conclusion should be considered with caution. There are no functional information on these UVs, MSH6 and PMS2 variants are less studied, and the literature data are truly scant. Both p.Ala1236Pro and p.Arg20Gln are present in families fulfilling the Amsterdam criteria and in probands with highly unstable tumors that are features associated with the highest probability of carrying a pathogenic mutation. This could be the reason that p.Arg20Gln was class 5 even if it is present in healthy controls and, in a family, occurs with a pathogenic mutation. Inclusion of p.Ala1236Pro class 5 seems to be further justified by a high probability of causality derived also from multiple OR values, sometimes low, but always concordant and in favor of pathogenicity. p.Ser46Ile was placed in class 4 by our evaluation. We have already described it to be present in a peculiar Turcot Syndrome family,33 where the carrier showed the typical clinical signs of a compound heterozygote. This feature, together with clinical and molecular data, strongly suggests the pathogenic role of the UV. In addition, it has been described as present in a patient with early onset sporadic colorectal cancer.37
There are several critical aspects in the evaluation of UVs that have been well-debated several times. Every positive datum is useful in recognizing pathogenic variants, whereas the negative data increase the UV's probability of being classified as nonpathogenic, even if a negative datum is not always an index of nonpathogenicity. MSI is a useful marker for MMR deficiency; however, MSI absence is not only related to nonpathogenic UVs but also to mutations that induce low or null degrees of instability, like some MSH6 ones.38 Conversely, MSI presence may not be related to the UV, as a coexisting pathogenic mutation may not have been identified or MLH1 promoter hypermethylation is present. IHC is also useful, but a lack of expression is not always related to functional alteration of the protein: a missense variant does not truncate the protein but it could induce a conformational change that alters the binding of the antibody, although not necessarily inactivating the protein. Conversely, a MMR protein with a single amino acid change could be functionally inactivated but still recognized by the antibody. Future studies would warrant a likelihood model accounting for these caveats.
UV-disease segregation is considered one of the most important parameters to be evaluated for the assessment of UVs pathogenicity.4 However, this is not always easy to analyze as it may be difficult to obtain DNA samples from several members of the same family: if the genetic test reveals only an UV in the proband, lack of data for pathogenicity does not allow a complete and correct risk evaluation and genetic counseling in the family. In fact, we found many difficulties in recruiting relatives of the UV carriers, mainly due to the unclear pathogenic role of the UV itself and the consequent uncertainty of the genetic test result. The small number of relatives is the cause of poor informative value of segregation data in some families. Moreover, it is also possible that an UV segregates with the disease because it is in linkage with a pathogenic mutation: this occurred in two UVs (MSH2 p.Leu814Leu and MLH1 pArg265His). Alternatively, the variant could cosegregate with disease by chance, as is probable for those UVs that were shared by two members of the same sibship (MSH2 p.Lys113Lys, MLH1 p.Thr82Ala and p.Lys618Ala, and MSH6 p.Glu983Gln and p.Pro1082Pro).
Most UVs are rare and, consequently, their frequency in healthy controls should be determined in prohibitively large samples, not available for this study. However, if a variant showed a frequency of 1% or more in our limited-size control sample, we could infer that it most likely does not have a pathogenic role.4
The experimental analysis of splicing alteration was complicated by the difficulty in obtaining viable lymphocytes, necessary for the establishment of lymphoblastoid cell lines. Moreover, the only extra bands seen in our RNA assays seemed to be physiologic alternative splicings,39 but we cannot exclude that the reverse transcriptase-PCR we used for the analysis was unable to highlight the existing alterations.
A number of databases and software to evaluate UVs have been developed. None of them, individually, is able to predict the real molecular and functional effect in all cases. However, a combination of analyses seems to be more reliable than a single-software test, even if an analysis performed using different web tools is complicated by prediction discordance, particularly for ESE/exon splicing silencer creation or elimination, due to different algorithms used to evaluate nucleotide variations. A study conducted on ESEfinder and then confirmed on RescueESE8,40,41 demonstrated that the prediction does not always correlate with an in vitro effect. Protein-prediction software tools are more reliable and there is greater agreement in their output data.
Our choice to consider all these clinical, molecular, and bioinformatics features is an attempt to remedy every single method defect and lack of an acknowledged evaluation. To this aim, an assessment of all these features in very large cohorts is important for establishing the true power of each one. It is especially fundamental to have access to large and appropriate reference sample sets for derivation of the OR necessary to predict the probability of pathogenicity of mutations that do not clearly affect mRNA or protein.
To be as objective as possible, we calculated the probability of pathogenicity of every UV with the method proposed by Goldgar et al.,4,13 calculating OR for every feature that was possible. The probabilities calculated were then classified in five classes, following the guidelines of the Special Issue of Human Mutation.3–12 An increasing number of groups are trying to classify UVs using these recommendations, but most of them focus on the BRCA1 and BRCA2 genes, for which a multifactorial likelihood classification has already been developed and refined. Instead, for MMR genes, there are not well-established models or well-characterized features42 so that, at the time of writing, a very few groups have attempted to classify MMR UVs with the Bayesian likelihood method. The study by Arnold et al.24 investigated several variants, three of which are in common with our data (MLH1 c.307-29C>A, c.1039-8 T>A and MSH2 p.Gly322Asp): only c.307-29C>A was analyzed by a similar comprehensive approach and assigned to class 3, as it is in our study.
As each pathogenicity class is associated with surveillance and testing recommendations, these integrated analyses could help laboratory geneticists better understand the pathogenic role of UVs and obtain useful data for genetic counselors. However, our work has to be considered among the pioneers in this framework and most of our evaluations need to be further refined. At the moment, this evaluation has to be considered only for research purposes and not for clinical use.
REFERENCES
Lynch HT, Lynch PM, Lanspa SJ, Snyder CL, Lynch JF, Boland CR . Review of the Lynch syndrome: history, molecular genetics, screening, differential diagnosis, and medicolegal ramifications. Clin Genet 2009; 76: 1–18.
Genuardi M, Carrara S, Anti M, Ponz de Leòn M, Viel A . Assessment of pathogenicity criteria for constitutional missense mutations of the hereditary nonpolyposis colorectal cancer genes MLH1 and MSH2. Eur J Hum Genet 1999; 7: 778–782.
Tavtigian SV, Greenblatt MS, Goldgar DE, Boffetta P . Assessing pathogenicity: overview of results from the IARC Unclassified Genetic Variants Working Group. Hum Mutat 2008; 29: 1261–1264.
Goldgar DE, Easton DF, Byrnes GB, Spurdle AB, Iversen ES, Greenblatt MS . Genetic evidence and integration of various data sources for classifying uncertain variants into a single model. Hum Mutat 2008; 29: 1265–1272.
Greenblatt MS, Brody LC, Foulkes WD, et al. Locus-specific databases and recommendations to strengthen their contribution to the classification of variants in cancer susceptibility genes. Hum Mutat 2008; 29: 1273–1281.
Plon SE, Eccles DM, Easton D, et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 2008; 29: 1282–1291.
Hofstra RM, Spurdle AB, Eccles D, et al. Tumor characteristics as an analytic tool for classifying genetic variants of uncertain clinical significance. Hum Mutat 2008; 29: 1292–1303.
Spurdle AB, Couch FJ, Hogervorst FB, Radice P, Sinilnikova OM . Prediction and assessment of splicing alterations: implications for clinical testing. Hum Mutat 2008; 29: 1304–1313.
Couch FJ, Rasmussen LJ, Hofstra R, Monteiro AN, Greenblatt MS, de Wind N . Assessment of functional effects of unclassified genetic variants. Hum Mutat 2008; 29: 1314–1326.
Tavtigian SV, Greenblatt MS, Lesueur F, Byrnes GB . In silico analysis of missense substitutions using sequence-alignment based methods. Hum Mutat 2008; 29: 1327–1336.
Ou J, Niessen RC, Vonk J, Westers H, Hofstra RM, Sijmons RH . A database to support the interpretation of human mismatch repair gene variants. Hum Mutat 2008; 29: 1337–1341.
Tavtigian SV, Byrnes GB, Goldgar DE, Thomas A . Classification of rare missense substitutions, using risk surfaces, with genetic- and molecular-epidemiology applications. Hum Mutat 2008; 29: 1342–1354.
Goldgar DE, Easton DF, Deffenbaugh AM, et al. Integrated evaluation of DNA sequence variants of unknown clinical significance: application to BRCA1 and BRCA2. Am J Hum Genet 2004; 75: 535–544.
Baudhuin LM, Burgart LJ, Leontovich O, Thibodeau SN . Use of microsatellite instability and immunohistochemistry testing for the identification of individuals at risk for Lynch syndrome. Fam Cancer 2005; 4: 255–265.
Laghi L, Bianchi P, Malesci A . Differences and evolution of the methods for the assessment of microsatellite instability. Oncogene 2008; 27: 6313–6321.
Shimodaira H, Filosi N, Shibata H, et al. Functional analysis of human MLH1 mutations in Saccharomyces cerevisiae. Nat Genet 1998; 19: 384–389.
Kondo E, Suzuki H, Horii A, Fukushige S . A yeast two-hybrid assay provides a simple way to evaluate the vast majority of hMLH1 germ-line mutations. Cancer Res 2003; 63: 3302–3308.
Raevaara TE, Korhonen MK, Lohi H, et al. Functional significance and clinical phenotype of nontruncating mismatch repair variants of MLH1. Gastroenterology 2005; 129: 537–549.
Takahashi M, Shimodaira H, Andreutti-Zaugg C, Iggo R, Kolodner RD, Ishioka C . Functional analysis of human MLH1 variants using yeast and in vitro mismatch repair assays. Cancer Res 2007; 67: 4595–4604.
Perera S, Bapat B . The MLH1 variants p.Arg265Cys and p.Lys618Ala affect protein stability while p.Leu749Gln affects heterodimer formation. Hum Mutat 2008; 29: 332.
Drost M, Zonneveld JB, van Dijk L, et al. A cell-free assay for the functional analysis of variants of the mismatch repair protein MLH1. Hum Mutat 2010; 31: 247–253.
Ollila S, Dermadi Bebek D, Jiricny J, Nyström M . Mechanisms of pathogenicity in human MSH2 missense mutants. Hum Mutat 2008; 29: 1355–1363.
Trojan J, Zeuzem S, Randolph A, et al. Functional analysis of hMLH1 variants and HNPCC-related mutations using a human expression system. Gastroenterology 2002; 122: 211–219.
Arnold S, Buchanan DD, Barker M, et al. Classifying MLH1 and MSH2 variants using bioinformatic prediction, splicing assays, segregation, and tumor characteristics. Hum Mutat 2009; 30: 757–770.
Sweeting MJ, Sutton AJ, Lambert PC . What to add to nothing? Use and avoidance of continuity corrections in meta-analysis of sparse data. Stat Med 2004; 23: 1351–1375.
Karchin R, Agarwal M, Sali A, Couch F, Beattie MS . Classifying variants of undetermined significance in BRCA2 with protein likelihood ratios. Cancer Inform 2008; 6: 203–216.
Lastella P, Surdo NC, Resta N, Guanti G, Stella A . In silico and in vivo splicing analysis of MLH1 and MSH2 missense mutations shows exon- and tissue-specific effects. BMC Genomics 2006; 7: 243.
Engel C, Forberg J, Holinski-Feder E, et al. Novel strategy for optimal sequential application of clinical criteria, immunohistochemistry and microsatellite analysis in the diagnosis of hereditary nonpolyposis colorectal cancer. Int J Cancer 2006; 118: 115–122.
Easton DF, Deffenbaugh AM, Pruss D, et al. A systematic genetic assessment of 1,433 sequence variants of unknown clinical significance in the BRCA1 and BRCA2 breast cancer-predisposition genes. Am J Hum Genet 2007; 81: 873–883.
Thompson D, Easton DF, Goldgar DE . A full-likelihood method for the evaluation of causality of sequence variants from family data. Am J Hum Genet 2003; 73: 652–655.
Marroni F, Pastrello C, Benatti P, et al. A genetic model for determining MSH2 and MLH1 carrier probabilities based on family history and tumor microsatellite instability. Clin Genet 2006; 69: 254–262.
Viel A, Novella E, Genuardi M, et al. Lack of PMS2 gene-truncating mutations in patients with hereditary colorectal cancer. Int J Oncol 1998; 13: 565–569.
Agostini M, Tibiletti MG, Lucci-Cordisco E, et al. Two PMS2 mutations in a Turcot syndrome family with small bowel cancers. Am J Gastroenterol 2005; 100: 1886–1891.
Barnetson RA, Cartwright N, van Vliet A, et al. Classification of ambiguous mutations in DNA mismatch repair genes identified in a population-based study of colorectal cancer. Hum Mutat 2008; 29: 367–374.
Ollila S, Sarantaus L, Kariola R, et al. Pathogenicity of MSH2 missense mutations is typically associated with impaired repair capability of the mutated protein. Gastroenterology 2006; 131: 1408–1417.
Gammie AE, Erdeniz N, Beaver J, Devlin B, Nanji A, Rose MD . Functional characterization of pathogenic human MSH2 missense mutations in Saccharomyces cerevisiae. Genetics 2007; 177: 707–721.
Nakagawa H, Lockman JC, Frankel WL, et al. Mismatch repair gene PMS2: disease causing germline mutations are frequent in patients whose tumors stain negative for PMS2 protein, but paralogous genes obscure mutation detection and interpretation. Cancer Res 2004; 64: 4721–4727.
Lagerstedt Robinson K, Liu T, Vandrovcova J, et al. Lynch syndrome (hereditary nonpolyposis colorectal cancer) diagnostics. J Natl Cancer Inst 2007; 99: 291–299.
Genuardi M, Viel A, Bonora D, et al. Characterization of MLH1 and MSH2 alternative splicing and its relevance to molecular testing of colorectal cancer susceptibility. Hum Genet 1998; 102: 15–20.
Lastella P, Resta N, Miccolis I, Quagliarella A, Guanti G, Stella A . Site directed mutagenesis of hMLH1 exonic splicing enhancers does not correlate with splicing disruption. J Med Genet 2004; 41: e72.
Auclair J, Busine MP, Navarro C, et al. Systematic mRNA analysis for the effect of MLH1 and MSH2 missense and silent mutations on aberrant splicing. Hum Mutat 2006; 27: 145–154.
Spurdle AB . Clinical relevance of rare germline sequence variants in cancer genes: evolution and application of classification models. Curr Opin Genet Dev 2010; 20: 315–323.
Acknowledgements
This work was supported by Alleanza Contro il Cancro (INTEF project) and AIRC. We gratefully acknowledge our colleagues D. Barana, A. M. Chiaravalli, M. Pedroni, and I. Mammi for their help in collecting samples and laboratory analyses.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The authors declare no conflict of interest.
Supplemental digital content is available for this article. Direct URL citations appear in the printed text and are provided in the HTML and PDF versions of this article on the journal's Web site (www.geneticsinmedicine.org).
Rights and permissions
About this article
Cite this article
Pastrello, C., Pin, E., Marroni, F. et al. Integrated analysis of unclassified variants in mismatch repair genes. Genet Med 13, 115–124 (2011). https://doi.org/10.1097/GIM.0b013e3182011489
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3182011489
Keywords
This article is cited by
-
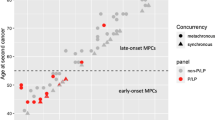
An unusual case of familial adenomatous polyposis with very early symptom occurrence
Familial Cancer (2014)
-
CoDP: predicting the impact of unclassified genetic variants in MSH6 by the combination of different properties of the protein
Journal of Biomedical Science (2013)
-
Incidence, clinical features and possible etiology of early onset (≤40 years) colorectal neoplasms
Internal and Emergency Medicine (2013)
-
Frequency of mutations in mismatch repair genes in a population-based study of women with ovarian cancer
British Journal of Cancer (2012)
-
Clinical and molecular detection of inherited colorectal cancers in northeast Italy
Tumor Biology (2012)