Abstract
Purpose
To describe our experience with a large cohort (411 patients from 288 families) of various forms of skeletal dysplasia who were molecularly characterized.
Methods
Detailed phenotyping and next-generation sequencing (panel and exome).
Results
Our analysis revealed 224 pathogenic/likely pathogenic variants (54 (24%) of which are novel) in 123 genes with established or tentative links to skeletal dysplasia. In addition, we propose 5 genes as candidate disease genes with suggestive biological links (WNT3A, SUCO, RIN1, DIP2C, and PAN2). Phenotypically, we note that our cohort spans 36 established phenotypic categories by the International Skeletal Dysplasia Nosology, as well as 18 novel skeletal dysplasia phenotypes that could not be classified under these categories, e.g., the novel C3orf17-related skeletal dysplasia. We also describe novel phenotypic aspects of well-known disease genes, e.g., PGAP3-related Toriello–Carey syndrome–like phenotype. We note a strong founder effect for many genes in our cohort, which allowed us to calculate a minimum disease burden for the autosomal recessive forms of skeletal dysplasia in our population (7.16E-04), which is much higher than the global average.
Conclusion
By expanding the phenotypic, allelic, and locus heterogeneity of skeletal dysplasia in humans, we hope our study will improve the diagnostic rate of patients with these conditions.
Similar content being viewed by others
Introduction
Heritable generalized disorders of bone and cartilage development, collectively known as skeletal dysplasias, are relatively common birth defects with an incidence of 1.3–3.2 per 10,000.1,2,3,4 They range in severity from those that are embryonically lethal to those with minimum morbidity. The International Skeletal Dysplasia Society has attempted to classify the large number of skeletal dysplasia phenotypes into groups based on clinical and radiological patterns, although the more recent classification placed a strong emphasis on the molecular ontology.5,6
Until recently, the diagnosis of skeletal dysplasia relied almost exclusively on careful phenotyping, often aided by consultation with experienced radiologists. Seeking the opinion of international experts was not uncommon given the overwhelming number of overlapping phenotypes and the degree of variation thereof. However, the advent of genomic tests has the potential to alleviate this bottleneck because these tests scan a large number of target disease genes irrespective of the suspected clinical diagnosis.7 In a recent study, we showed that the use of a gene panel that encompasses most of the known skeletal dysplasia genes was indeed a very helpful diagnostic aid in nearly 38% of cases.8
Through the widespread use of this panel, as well as reflex exome sequencing in those who test negative, in patients referred to our center with suspected skeletal dysplasia phenotypes, we accumulated a large number of molecularly characterized cases. As expected, the diagnosis in many of these cases had to be revised as a result, and atypical phenotypes were not uncommon. Furthermore, we encountered a number of interesting variants in genes not previously established as skeletal dysplasia genes, sometimes with resulting novel phenotypes. Here, we share detailed phenotypic and genotypic features of this large cohort of molecularly characterized individuals with skeletal dysplasia. Our results expand the phenotypic expression of known diseases, establish new phenotypes, increase the number of pathogenic/likely pathogenic alleles of various skeletal dysplasias, and suggest or establish novel disease loci, with the ultimate goal of improving the diagnostic rate among patients with these conditions.
Materials and methods
Human subjects
Affected patients and available family members were recruited using a King Faisal Specialist Hospital and Research Centre institutional review board–approved protocol (RAC 2121053, 2080006, 2090035 and 2070023) with informed consent. Patients were usually referred by their primary care physicians to clinical geneticists with interest in skeletal dysplasias and were evaluated clinically and radiologically. Only those with suspected skeletal dysplasia based on clinical and/or radiographic evidence were included. We grouped and classified cases according to the most recently published international classification of skeletal dysplasia.5 Demographics (sex, age, race, consanguinity, family history) are provided in Supplementary Table S1 online. Venous blood samples were collected in EDTA tubes and, in a few patients, in PAXgene for DNA and RNA extraction, respectively.
Panel testing
The dysmorphology/skeletal dysplasia panel was our first-tier test as described previously.8 Briefly, this is a multigene panel (296 genes) that comprises genes known to cause various forms of gross facial and skeletal dysmorphism and is run on the Ion Proton next-generation sequencing platform (Thermo Fisher, Waltham, MA, USA).
Autozygome analysis and exome sequencing
Cases that tested negative on the panel were subjected to combined autozygome/exome analysis as described previously.9,10 Briefly, genome-wide genotyping of the patients and their available family members using the Axiom platform was pursued to determine the candidate autozygome using long stretches (>2 Mb) of homozygosity as surrogates of autozygosity, as determined by AutoSNPa (http://dna.leeds.ac.uk/autosnpa).
Whole-exome sequencing was performed using TruSeq Exome Enrichment kit (Illumina) to build Illumina sequencing library with enrichment for the desired target using the Illumina Exome Enrichment protocol. The libraries were sequenced using IlluminaHiSeq 2000 Sequencer and single-nucleotide polymorphisms and indels were identified by SAMtools. Sequenced reads were assembled against UCSC hg19.
Variant interpretation
Variants from gene panels and whole-exome sequencing were filtered based on (i) candidate autozygome where applicable; (ii) allele frequency (only those with <0.001 were considered based on publicly available databases Exome Variant Server, 1000 Genomes, and gnomAD, as well as our in-house ethnically matched exomes (2,363); and (iii) predicted pathogenicity according to the nature of the variant (splicing and coding) as well as three in silico tools (PolyPhen, SIFT, and CADD) for missense variants. Candidate variants were classified as per the American College of Medical Genetics and Genomics guidelines.11 For the sporadic cases or those born to nonconsanguineous parents, all the rare (minor allele frequency <0.001) heterozygous and homozygous variants were investigated for candidacy as well. Variants were classified as novel if absent in the Human Gene Mutation Database, ClinVar, and, to the best of our knowledge, in the literature.
Results
Molecular characterization of a large skeletal dysplasia cohort
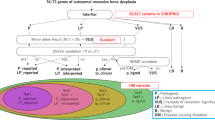
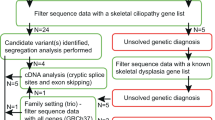
We describe in this communication 411 skeletal dysplasia patients (288 families) in whom a potentially causal variant was identified (Supplementary Table S1 online). These variants were found in 123 established disease-causing genes and 5 that we propose as novel candidate genes (Supplementary Figure S1 online). The number of unique pathogenic/likely pathogenic alleles in established disease-related genes was 224, including 54 that were novel (24%). The breakdown of all variants identified was 77.4% (222 families) recessive (98% of which were homozygous), 22.3% (64 families) dominant (72% were de novo), and only 0.3% X-linked (Figure 1). Although most of the encountered recessive alleles were private (64%), the contribution of founder alleles was disproportionately high, accounting for approximately half (51%) of familial cases with recessive forms of skeletal dysplasia. These founder variants ranged in their minor allele frequency between 0.000212 and 0.0032 among Saudis.12
While genes with established links to skeletal dysplasia accounted for the majority of cases (91%) (Supplementary Figure S2, Table S1 online), our analysis revealed potential confirmation of previously reported candidates. Specifically, C3orf17 (NEPRO) was previously proposed by us as a novel candidate gene in an autosomal recessive Larsen-like phenotype.13 In addition to the previously published family, we were able to identify a second family (15DG2238 and 15DG2239) with an identical phenotype of facial dysmorphism (midfacial hypoplasia, depressed nasal bridge, frontal bossing, wide anterior fontanel), chest narrowing with pectus excavatum, and acromelia (see Figure 2 and Supplementary Table S2 online) in whom we identified the same homozygous variant (haplotype analysis confirmed the founder nature of the variant). In addition, we propose the following as novel candidate genes: WNT3A (osteopenia), SUCO (osteogenesis imperfecta), DIP2C (a novel dysplasia), RIN1 (cranioectodermal dysplasia), and PAN2 (craniosynostosis, severe early-onset scoliosis, imperforate anus, and double urinary collecting system) based on single mutational events (clinical details of the patients, pedigrees, and supporting evidence for the candidate genes are shown in Figure 3, Supplementary Tables S1, S3, and Supplementary Figures S3, S44 online).
Clinical features of NEPRO (C3ORF17)-related phenotype. Images from 15DG2238 (family I), shown for the first time (a–f), and the previously reported 15DG0765 from family II (g–k), showing a similar skeletal profile including narrow chest, micromelia, reduced vertebral body height, short iliac wings, narrow acetabular angle, shortening and broadening of the phalanges, generalized osteopenia and metaphyseal cupping. (l) Sequence chromatogram of the mutation (c.280C>T). (m) Cartoon representation of NEPRO protein with the position of the founder mutation (p.Arg94Cys) along with pedigrees. (n) Genome-wide linkage analysis (four affected members from families I and II) showing a single peak with pLOD score of ~3.5 that spans NEPRO (C3ORF17) locus.
(a–d) A patient with a mutation in DIP2C showing (a) short humerus; (b) bilateral shortening of femora; (c) clinodactyly with shortening of metacarpal bones, a hypoplastic middle phalanx of the middle finger, and absent 1st left phalanx; and (d) bilateral metatarsus adduction, bilateral shortening of 1st metatarsals, which is more pronounced in the left with the first toe appearing small and rotated. (e and f) A patient with a mutation in PAN2 showing plagiocephaly, malar hypoplasia, short upslanted palpebral fissures, micrognathia, long philtrum, epicanthus, anteverted nares, short and depressed nasal bridge, and scoliosis. (g–j) A patient with a mutation in RIN1 showing plagiocephaly, frontal bossing, long philtrum, and epicanthus. 3D computed tomography skull of the same patient showing craniosynostosis. (k–q) A patient with a mutation in WNT3A showing osteopenia, bilateral coxa valga deformity, mild left radial and ulnar bowing, broadening of metaphases, and bilateral shortening of the great toes and thumbs. (r–x) A patient with a mutation in SUCO showing diffuse osteopenia, multiple fractures with limb deformities, and short long bones.
Phenotypic expansion of skeletal dysplasia
Our large cohort spans 36 of the 42 categories in the International Skeletal Dysplasia Nosology classification (Supplementary Figures S4–43 online show many representative images). It also spans phenotypes that could not fit a specific category (Supplementary Table S1, Figure S2 online). We also note a surprising array of phenotypes that are sufficiently different from those previously ascribed to known disease genes to qualify as allelic disorders. The most striking example is the phenotype associated with a founder mutation in PGAP3. We ascertained five families (eight patients) who are homozygous for one founder variant (NM_033419.3:c.850C>T: p.(His284Tyr)) and share a phenotype of Toriello–Carey syndrome (Pierre Robin sequence, cerebellar hypoplasia, short neck, and developmental delay) (Figure 4 and Supplementary Table S4 online). The significant craniofacial and cerebellar involvement of these children prompted the clinical diagnosis of Toriello–Carey syndrome but the finding of a PGAP3 mutation prompted an investigation of their alkaline phosphatase level, which revealed the expected hyperphosphatasia finding (Supplementary Table S4 online).
Clinical images ofPGAP3-related Toriello–Carey syndrome phenotype patients, all with the same founder mutation. (a–l) Facial dysmorphia, bulbous nose, epicanthal folds, upslanted palpebral fissures, megalocornea, microcephaly, U-shaped cleft palate, a small tongue, overlapping toes, abnormal dentition, hypoplastic corpus callosum, hypoplastic cerebellum with absent vermis, and a communication between the third ventricle and the posterior fossa.
We also took advantage of the presence of multiplex families in our cohort as well as the presence of founder mutations to examine the consistency of phenotypes caused by the same recessive variants and found that they are indeed consistent (Supplementary Table S1 online). The founder mutation in B3GALT6 was a noteworthy exception. Mutations in this gene have been linked to two phenotypes: Ehlers–Danlos syndrome progeroid type 2 and spondyloepimetaphyseal dysplasia with joint laxity type 1. We note that one family presented with one phenotype while three families, who all shared the same founder variant, presented with the other (Figure 5 and Supplementary Table S5 online).
(a–k) Diffuse osteopenia, hip dysplasia, C-shaped levoconvexity scoliosis of the thoracolumbar spine, exaggerated lumbar lordosis, bilateral posterior radial head dislocation, lacunar skull appearance, bilateral acetabular dysplasia, bilateral femoral head subluxation, bilateral hallux valgus, metatarsus adductus, and fused bilateral proximal tibial physis. (l–n) Progeroid facial appearance, depressed nasal bridge with upturned nostrils, smooth lips, arching eyebrows, deep-set eyes, long eyelashes, arthrogryposis, and arachnodactyly.
Finally, it has always been assumed that recessive forms of skeletal dysplasia are common in Saudi Arabia due to consanguinity.14 Because this is the most comprehensive catalogue of skeletal dysplasias in the country, we attempted to quantify the disease burden using our previously published methods that rely on the unbiased carrier frequency of recessive alleles in the population.15 The resulting estimate of 1:1,397, already much higher than the global average, is probably a gross underestimate because private mutations (which account for the majority of recessive alleles as explained above) and dominant mutations were not included.
Discussion
Although a few skeletal dysplasia cohorts have been published to date, the present cohort is the largest molecularly characterized cohort from a single population ethnic group.16,17 This allowed us to observe a number of important patterns. First, consistent with our experience with other genetically heterogeneous disorders, we see a preponderance of autosomal recessive forms of phenotypes that can be caused by both dominant and recessive mutations.18,19,20,21 For instance, autosomal recessive forms of osteogenesis imperfecta accounted for 64% of the osteogenesis imperfecta cases, a markedly different distribution compared with outbred populations where de novo dominant mutations in collagen genes account for 85%.22 Second, reverse phenotyping is very common in skeletal dysplasia, where nearly half (48%) had been misclassified clinically when the correct molecular lesion was identified. This argues for the value of a “genomics-first” approach in these disorders as proposed for similarly heterogeneous disorders, e.g., developmental delay.21 Third, despite the impact of founder mutations, private mutations remain more common, which supports the view that targeted mutation analysis is not the ideal testing strategy.14,15,23
The novel candidate genes that we propose deserve a special mention. WNT3A encodes the first Wnt protein to be purified, and signals through the canonical signaling pathway.24,25 Its deficiency in mouse has been linked to reduced bone mass.26In vivo and in vitro studies have demonstrated a role of this Wnt protein in inducing mesenchymal stem cells (which have a dual adipogenic and osteogenic potential) toward osteogenesis, and it has been proposed as a potential target to reduce age-related increased adipogenic potential at the expense of osteogenic potential, a derangement that results in osteoporosis.27 Other proteins in the same pathway, such as LRP5 and WNT1, are known to play a role in osteoporosis and osteogenesis imperfecta.28 Similarly, available data on the function of SUCO (SUN domain containing ossification factor) make it another compelling candidate gene in osteogenesis imperfecta. Specifically, Suco Gt(KST050)Byg/Suco Gt(KST050)Byg mice exhibit low bone mass with multiple fractures and have been proposed as a good model for osteogenesis imperfecta in humans.29 The mechanistic connection between PAN2, RIN1, and DIP2C and the observed syndromes is less clear. Candidate gene PAN2 encodes a ubiquitously expressed nuclease that possesses a 3′ to 5′ RNase activity against poly(A) messenger RNA.30,31 This activity is carried out by the RNase motif in the C-terminus, which is abolished by the homozygous truncating mutation we detected in the patient. Although no further details are provided, the embryonic lethality of the knockout Pan2 mouse listed by Mouse Genome Informatics strongly supports an important developmental role by this gene. Syndromes with multiple congenital anomalies have been linked to mutations in genes encoding components of RNA degradation, although we are not aware of a human disease caused by mutations in a ribonuclease that specifically targets poly(A) messenger RNA.32,33 For the proposed RIN1-craiosynostosis link, it is worth highlighting that RIN1 regulates RAS signaling, which is a critical regulator of bone formation and has been implicated in syndromic craniosynostosis in humans as evidenced by patients with de novo mutations in this pathway.34 Perhaps the least obvious among the candidates is DIP2C, although its proposed role in transcriptional and methylation regulation leaves open the possibility that the observed phenotype is caused by the indirect perturbation of downstream genes as we have previously suggested for GZF1-related Larsen syndrome.35,36,37 As with all other novel candidate genes we report here, future cases will be needed to confirm the gene–disease link and to shed light onto potential mechanisms.
We should highlight that our utilization of autozygosity mapping was key to the identification of a number of challenging alleles, many of which evaded detection by clinical exome sequencing.38 These include the previously reported deep intronic variant in C21orf2 13 and exonic deletion in TMEM38B,20 as well as the novel CLCN7 (NM_001287.5:c.739-18G>A) and GALNS (NM_000512.4:c.567_1002dup:p.(Val335Ilefs*3)) variants. In addition, the founder mutation B3GALT6 was initially dismissed due to an apparently conflicting in silico prediction but its strong segregation with the phenotype in several families led us to reconsider its classification. This is consistent with our previously published experience with “negative” clinical exomes that can be successfully reinterpreted in light of autozygome analysis.38 It is likely that transcriptomics can also aid in the reinterpretation of those cases in the future especially when autozygosity mapping is not applicable.
In conclusion, by sharing the phenotypic and genotypic data of a large molecularly characterized skeletal dysplasia cohort, we hope to improve the diagnostic rate of these patients in the target population and beyond.
References
Dolk H, Loane M & Garne EThe prevalence of congenital anomalies in Europe. In: Posada de la Paz M, Groft SC (eds). Rare Diseases Epidemiology. Springer: Dordrecht, The Netherlands, 2010: 349–64.
Rimoin DL, Cohn D, Krakow D, Wilcox W, Lachman RS & Alanay Y. The skeletal dysplasias. Ann NY Acad Sci 2007;1117:302–9.
Spranger JW, Brill PW & Poznanski AK. Bone Dysplasias: An Atlas of Genetic Disorders of Skeletal Development. Oxford University Press: New York, 2002.
Barbosa-Buck CO, Orioli IM, da Graça Dutra M, Lopez-Camelo J, Castilla EE & Cavalcanti DP. Clinical epidemiology of skeletal dysplasias in South America. Am J Med Genet A 2012;158:1038–45.
Bonafe L, Cormier-Daire V, Hall C et al. Nosology and classification of genetic skeletal disorders: 2015 revision. Am J Med Genet A 2015;167:2869–92.
Hall CM. International nosology and classification of constitutional disorders of bone (2001). Am J Med Genet A 2002;113:65–77.
Lazarus S, Zankl A & Duncan E. Next-generation sequencing: a frameshift in skeletal dysplasia gene discovery. Osteoporos Int 2014;25:407–22.
Group SM. Comprehensive gene panels provide advantages over clinical exome sequencing for Mendelian diseases. Genome Biol 2015;16:1–14.
Alkuraya FS. The application of next-generation sequencing in the autozygosity mapping of human recessive diseases. Hum Genet 2013b;132:1197–211.
Alkuraya FS. Discovery of mutations for Mendelian disorders. Hum Genet 2016;135:615–23.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–23.
Abouelhoda M, Faquih T, El-Kalioby M & Alkuraya FS. Revisiting the morbid genome of Mendelian disorders. Genome Biol 2016a;17:235.
Shaheen R, Patel N, Shamseldin H et al. Accelerating matchmaking of novel dysmorphology syndromes through clinical and genomic characterization of a large cohort. Genet Med 2015;18:686–95.
Alkuraya FS. Genetics and genomic medicine in Saudi Arabia. Mol Genet Genomic Med 2014;2:369–78.
Abouelhoda M, Sobahy T, El-Kalioby M et al. Clinical genomics can facilitate countrywide estimation of autosomal recessive disease burden. Genet Med 2016b;18:1244–9.
Wang SR, Carmichael H, Andrew SF et al. Large-scale pooled next-generation sequencing of 1077 genes to identify genetic causes of short stature. J Clin Endocrinol Metab 2013;98:E1428–E37.
Bae J-S, Kim NK, Lee C et al. Comprehensive genetic exploration of skeletal dysplasia using targeted exome sequencing. Genet Med 2016;18:563.
Patel N, Aldahmesh MA, Alkuraya H et al. Expanding the clinical, allelic, and locus heterogeneity of retinal dystrophies. Genet Med 2016;18:554.
Patel N, Anand D, Monies D et al. Novel phenotypes and loci identified through clinical genomics approaches to pediatric cataract. Hum Genet 2017a;136:205–25.
Shaheen R, Alazami AM, Alshammari MJ et al. Study of autosomal recessive osteogenesis imperfecta in Arabia reveals a novel locus defined by TMEM38B mutation. J Med Genet 2012;49:630–5.
Anazi S, Maddirevula S, Faqeih E et al. Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield. Mol Psychiatry 2017;22:615–24.
Marini JC, Forlino A, Bachinger HP et al. Osteogenesis imperfecta. Nat Rev Dis Primers 2017;3:17052.
Alkuraya F. Impact of new genomic tools on the practice of clinical genetics in consanguineous populations: the Saudi experience. Clin Genet 2013a;84:203–8.
Staal FJ, Luis TC & Tiemessen MM. WNT signalling in the immune system: WNT is spreading its wings. Nat Rev Immunol 2008;8:581.
Willert K, Brown JD, Danenberg E & Duncan AW. Wnt proteins are lipid-modified and can act as stem cell growth factors. Nature 2003;423:448.
Wang Y, Li Y-P, Paulson C et al. Wnt and the Wnt signaling pathway in bone development and disease. Front Biosci (Landmark Ed) 2014;19:379.
Leucht P, Jiang J, Cheng D et al. Wnt3a reestablishes osteogenic capacity to bone grafts from aged animals. J Bone Joint Surg Am 2013;95:1278.
Faqeih E, Shaheen R & Alkuraya FS. WNT1 mutation with recessive osteogenesis imperfecta and profound neurological phenotype. J Med Genet 2013;50:491–2.
Sohaskey ML, Jiang Y, Zhao JJ, Mohr A, Roemer F & Harland RM. Osteopotentia regulates osteoblast maturation, bone formation, and skeletal integrity in mice. J Cell Biol 2010;189:511–25.
Belyaeva OV & Kedishvili NY. Human pancreas protein 2 (PAN2) has a retinal reductase activity and is ubiquitously expressed in human tissues. FEBS Lett 2002;531:489–93.
Uchida N, Hoshino S-i & Katada T. Identification of a human cytoplasmic poly (A) nuclease complex stimulated by poly (A)-binding protein. J Biol Chem 2004;279:1383–91.
Shaheen R, Anazi S, Ben-Omran T et al. Mutations in SMG9, encoding an essential component of nonsense-mediated decay machinery, cause a multiple congenital anomaly syndrome in humans and mice. Am J Hum Genet 2016;98:643–52.
Yan YB. Deadenylation: enzymes, regulation, and functional implications. Wiley Interdiscip Rev RNA 2014;5:421–43.
Timberlake AT, Furey CG, Choi J et al. De novo mutations in inhibitors of Wnt, BMP, and Ras/ERK signaling pathways in non-syndromic midline craniosynostosis. Proc Natl Acad Sci USA 2017;114:201709255.
Bian C, Chen Q & Yu X. The zinc finger proteins ZNF644 and WIZ regulate the G9a/GLP complex for gene repression. Elife 2015;4:e05606.
Lindroth AM, Larsson C, He L, Ali MA, Pandzic T & Sjöblom T. Loss of DIP2C in RKO cells stimulates changes in DNA methylation and epithelial-mesenchymal transition. BMC Cancer 2017;17:487.
Patel N, Shamseldin HE, Sakati N et al. GZF1 mutations expand the genetic heterogeneity of Larsen syndrome. Am J Hum Genet 2017b;100:831–6.
Shamseldin HE, Maddirevula S, Faqeih E et al. Increasing the sensitivity of clinical exome sequencing through improved filtration strategy. Genet Med 2016;19:593–8.
Acknowledgments
This work was supported by the King Salman Center for Disability Research (F.S.A.), King Abdulaziz City for Science and Technology (13-BIO1113-20, F.S.A.), and the Saudi Human Genome Program (F.S.A.). We also thank the study families for their enthusiastic participation and the Sequencing and Genotyping Core Facilities at King Faisal Specialist Hospital and Research Centre for their technical help.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Maddirevula, S., Alsahli, S., Alhabeeb, L. et al. Expanding the phenome and variome of skeletal dysplasia. Genet Med 20, 1609–1616 (2018). https://doi.org/10.1038/gim.2018.50
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.50
Keywords
This article is cited by
-
Dyssegmental dysplasia Rolland–Desbuquois type is caused by pathogenic variants in HSPG2 - a founder haplotype shared in five patients
Journal of Human Genetics (2024)
-
Biallelic PAN2 variants in individuals with a syndromic neurodevelopmental disorder and multiple congenital anomalies
European Journal of Human Genetics (2022)
-
An automated 13.5 hour system for scalable diagnosis and acute management guidance for genetic diseases
Nature Communications (2022)
-
Phenome-based approach identifies RIC1-linked Mendelian syndrome through zebrafish models, biobank associations and clinical studies
Nature Medicine (2020)