Abstract
Disclaimer: ACMG Clinical Laboratory Practice Resources are developed primarily as an educational tool for clinical laboratory geneticists to help them provide quality clinical laboratory genetic services. Adherence to these practice resources is voluntary and does not necessarily assure a successful medical outcome. This Clinical Laboratory Practice Resource should not be considered inclusive of all proper procedures and tests or exclusive of other procedures and tests that are reasonably directed to obtaining the same results. In determining the propriety of any specific procedure or test, the clinical laboratory geneticist should apply his or her own professional judgment to the specific circumstances presented by the individual patient or specimen. Clinical laboratory geneticists are encouraged to document in the patient’s record the rationale for the use of a particular procedure or test, whether or not it is in conformance with this Clinical Laboratory Practice Resource. They also are advised to take notice of the date any particular guideline was adopted, and to consider other relevant medical and scientific information that becomes available after that date. It also would be prudent to consider whether intellectual property interests may restrict the performance of certain tests and other procedures.
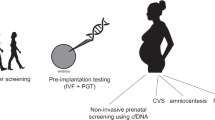
Noninvasive prenatal screening (NIPS) using cell-free DNA has been rapidly adopted into prenatal care. Since NIPS is a screening test, diagnostic testing is recommended to confirm all cases of screen-positive NIPS results. For cytogenetics laboratories performing confirmatory testing on prenatal diagnostic samples, a standardized testing algorithm is needed to ensure that the appropriate testing takes place. This algorithm includes diagnostic testing by either chorionic villi sampling or amniocentesis samples and encompasses chromosome analysis, fluorescence in situ hybridization, and chromosomal microarray.
Similar content being viewed by others
Background
This document was generated to support clinical cytogenetics laboratories in the testing and management of positive noninvasive prenatal screening (NIPS) results and is designed to be a rubric that can guide laboratory practice. The American College of Medical Genetics and Genomics (ACMG) revised its position statement on the use of NIPS for fetal aneuploidy in July 2016.1 This is meant to be a companion to that revised statement.
NIPS, also referred to as cell-free DNA (cfDNA) or noninvasive prenatal testing, has been available as a clinical screening option for pregnant women since 2011.2 Initially, NIPS was available primarily for the detection of trisomy 21,2, 3 but it rapidly evolved to include the detection of trisomies 13 and 18, sex chromosome identification, and sex chromosome aneuploidies.4, 5 NIPS has better performance as a screening test for trisomy 21 than for trisomies 13 or 18, or for sex chromosome aneuploidies.6 Recently, select microdeletion syndromes and smaller copy-number changes, as well as other autosomal aneuploidies, have been added by some laboratories as additional screening options.7, 8 Various factors affect the accuracy of NIPS results, including confined placental mosaicism (CPM), maternal genomic contribution and technical or statistical issues.9 Follow-up diagnostic testing is uniformly recommended for all patients with positive NIPS results.1, 10, 11 This document establishes a standardized testing algorithm that is essential for the cytogenetics laboratory to ensure that the appropriate diagnostic testing has occurred and that the results are reliable, accurate, and reflective of the fetal karyotype.
Diagnostic testing
Follow-up prenatal diagnostic testing is recommended for all patients with positive NIPS results. This can be accomplished by either chorionic villus sampling (CVS) or amniocentesis. In general, diagnostic testing should be appropriate for the suspected anomaly (i.e., chromosomal microarray (CMA) for smaller copy-number changes). Some laboratories may opt to perform fluorescence in situ hybridization (FISH) for the aneuploidy or copy-number change in question and then reflex to either chromosome analysis or CMA, dependent on the FISH results. While FISH is possible for either type of copy-number change, it may not be as accurate, depending on the exact size of the anomaly or structural rearrangements.
Chromosome analysis on either CVS or amniocentesis demonstrating nonmosaic trisomy or sex chromosome aneuploidy consistent with the NIPS result is considered confirmation of a positive NIPS and therefore of an affected fetus. A full study (as defined by the ACMG laboratory guidelines12) on CVS or amniocentesis demonstrating a normal karyotype would not typically warrant additional metaphase cell counts or other analyses. However, a mosaic result on CVS should not be considered confirmatory. There are known physiological limitations of CVS that include the possibility of CPM and rare case reports of complete discordancy between the CVS karyotype and the fetal karyotype.13, 14, 15, 16 While NIPS can be performed in the late first trimester of pregnancy, and CVS is a possibility for confirmatory studies (and often desired by the patient due to timing), CVS may simply reflect the same DNA/cells that were detected by NIPS, as both are derived from the placenta.17 Certain aneuploidies, including trisomy 13 and monosomy X, are more likely to be found in the mosaic form on CVS, which may influence genetic counseling about the preferred diagnostic test for confirmatory studies.18 When CVS shows mosaicism for the suspected trisomy, it is impossible to determine if this is CPM or true fetal mosaicism (TFM). Therefore, a mosaic CVS result cannot be treated as confirmation of an affected fetus and a follow-up amniocentesis is warranted, as is recommended in all cases of mosaicism observed on CVS12, 15, 16 (Table 1).
Similarly, CMA testing on either CVS or amniotic fluid may be used as confirmatory diagnostic testing in cases with positive NIPS results, or as reflex testing in cases with initial normal results from chromosome analysis. Smaller copy-number changes are ideally confirmed by this method. Again, if the NIPS results and CMA results are concordant, no further testing is recommended. However, given that structural information is not available from CMA analysis, a reflex to chromosome analysis may be considered to evaluate the structural arrangement to inform recurrence risks, especially for those cases with trisomies 21 and 13.
On occasion, prenatal diagnostic testing may not be performed due to loss of the pregnancy before testing is possible. In such instances, testing of the products of conception and/or the fetus by either chromosome analysis or CMA should be considered on a case-by-case basis.
Other forms of abnormal result exist, such as “no calls” and the unanticipated findings rendered by special maternal medical circumstances (e.g., obesity, oocyte donations and prior transfusions). These are discussed at length in the revised ACMG position statement1 but are beyond the scope of this laboratory algorithm.
Positive predictive value
While most NIPS laboratories report a greater than 99% specificity and sensitivity for trisomy 21, the positive predictive value (PPV) is essential for patient care. The 2016 ACMG position statement1 recommends that all laboratories reporting NIPS results also include the PPV, as well as the negative predictive value, detection rate, clinical specificity and fetal fraction. PPV answers an important question: “If NIPS is positive, what is the chance that the fetus is affected?” The PPV is affected by the analytic specificity and sensitivity of the test, as well as the prevalence of the disorder in the population.6, 11, 19 When the prevalence is age dependent, as it is for trisomies 13, 18 and 21, maternal age is a determining factor in the reliability of the test. Due to the higher incidence of aneuploidy associated with advanced maternal age, a 35-year-old woman with a positive result by NIPS has a higher PPV than a 25-year-old woman with a positive result. In addition, less common disorders, such as trisomy 13 and trisomy 18, would be expected to have lower PPVs than the more common trisomy 21. Very rare disorders (e.g., microdeletion syndromes and smaller copy-number changes), which are typically not associated with increased maternal age, would be expected to have even lower PPVs.7 The PPV can also be determined by comparing the NIPS results with the diagnostic testing results, with the caveat that these studies involve relatively low numbers of cases with wide confidence intervals.20, 21, 22, 23
Sources of discordant results
The source of cfDNA in the maternal circulation is primarily of maternal origin, with a much lower proportion (typically around 10%) being derived from degraded trophoblastic cells of the placenta.24 The primary reason for discordant NIPS and diagnostic cytogenetic testing is that the DNA tested is not solely representative of the fetus. This could be due to CPM or to a resorbed or unrecognized twin pregnancy. Furthermore, it has been reported that discordant results can be due to variations in the maternal DNA contribution, including low-level sex chromosome and autosomal chromosome mosaicism, maternal malignancies, and maternal copy-number variants.9, 17, 25, 26, 27 It is well known that some women may have low-level age-related losses and gains of the X chromosome.28, 29 There are a few reports of concurrent maternal malignancies when multiple or rare aneuploidies (e.g., autosomal monosomies) are detected by NIPS.30, 31 Other reasons for discordance might be technical or statistical.9 Since analytic algorithms differ between testing platforms and providers, there could be inconsistency in the reporting of aneuploidy results from the same pregnancy reported from different laboratories due to the utilization of different cutoffs, z-scores and/or comparison to different normalization controls. By necessity, reporting algorithms include screen-positive cases that are true negatives, to ensure that nearly all true positives would be identified by the screening test.
Cpm and tfm
When mosaicism is detected by CVS, cytogenetics laboratories attempt to distinguish between CPM and TFM. In general, regardless of the chromosome involved, this requires follow-up amniocentesis and often an extended chromosome analysis of this specimen with adherence to standard guidelines for distinguishing between pseudomosaicism and TFM.12 This extended analysis could include screening additional cells (or colonies) from independent cultures. Screening additional metaphase cells, however, has its limitations, and a very low level of fetal mosaicism can essentially never be ruled out. Theoretically, analyzing 15 amniotic fluid colonies from at least two independent coverslips will rule out a 19% level of mosaicism at the ninety-fifth confidence interval, while screening an additional 15 colonies will rule out a 10% level of mosaicism.32 Alternatively, interphase FISH for the mosaic aberration found at CVS might be useful, although it should be noted that laboratories need to validate and establish cutoff values for positivity for each probe utilized. Any value below these cutoff values or thresholds would be considered negative.33 CMAs may also be ordered as part of the follow-up testing, although detection of low-level mosaicism may be more challenging than by chromosome analysis and/or interphase FISH analysis34 (Table 1).
Uniparental disomy of known imprinted chromosomes
CPM can occur as a result of either postzygotic nondisjunction or aneusomy rescue. Given the latter, it is important to determine if the normal cell line represents uniparental disomy if an imprinted chromosome is involved.16 In these cases, discordance between the positive NIPS result and the diagnostic test result should be followed up with testing appropriate for detecting uniparental disomy of the particular chromosome of interest.
Nips results with multiple aneuploidies or rare aneuploidies
Although reportedly rare, any NIPS result that is positive for more than one aneuploidy or one that shows rare aneuploidies, such as an autosomal monosomy, should include consideration of the possibility of a maternal malignancy. A wide variety of maternal malignancies have been described in the literature in association with unusual NIPS results30, 31 and there are currently no guidelines for clinical evaluation following these rare results. Further evaluation and referral to an oncologist may be warranted.
Smaller copy-number changes
Some NIPS laboratories offer screening for rare microdeletion syndromes and smaller copy-number changes. Again, diagnostic testing is necessary in these cases, particularly as most will be falsely positive due to lower PPVs, and some may represent variants of uncertain significance. In most cases that are positive by NIPS for smaller copy-number changes, the breakpoints and the base pair coordinate positions and sizes are not provided or reported by the testing laboratory.35, 36 As a result, specific microdeletion FISH is not the appropriate diagnostic test, due to the possibility of incorrect or incomplete FISH probe coverage. In the vast majority of cases, a whole-genome CMA analysis should be used to determine the true fetal result. As well, it should be noted that maternal contribution may also play a role in discordant results, either due to low-level maternal mosaicism or maternal copy-number changes27 (Table 1).
NIPS cases positive for imprinted genetic disorders (e.g., Angelman or Prader−Willi syndrome) may come with the acknowledgment that the laboratory cannot distinguish between a deletion and uniparental disomy of the region in question. In such cases, methylation analysis, including methylation-specific multiplex ligation-dependent probe amplification or similar methodology is the appropriate diagnostic test to confirm the fetal result. It should be noted that methylation may not be complete for all loci at the time of CVS, and amniocentesis or neonatal testing may be warranted. Laboratories performing methylation analyses should be consulted regarding the appropriate specimen type and requirements.
Newborns
In some cases, parents with a fetus suspected to have an anomaly by NIPS will decline diagnostic testing and choose the option of a neonatal assessment. If possible, at birth, a genetics consultation should be requested and a detailed physical examination performed. In NIPS cases positive for trisomies 13, 18 or 21, normal findings on a physical examination by a clinical geneticist may be sufficient to preclude further testing. Any suspicion of an abnormal phenotype related to the aneuploidy in question should prompt a cytogenetics evaluation. When warranted, an extended chromosome analysis to rule out low levels of mosaicism or FISH may be performed. If the neonate has an abnormal physical examination that is not suggestive of the trisomy in question, CMA is recommended. For sex chromosome aneuploidies, chromosome analysis or CMA is recommended, with the possibility of additional interphase FISH analysis if mosaicism is suspected. Any NIPS result indicating smaller copy-number changes should be confirmed by CMA. Extensive testing of placental tissue is not recommended, as this is not important in the clinical care of the infant. In most cases, peripheral blood chromosome analysis of the infant’s mother to screen for mosaic sex chromosome gains or losses also may not be relevant and is typically not needed for patient care or for reproductive management.28, 29 However, maternal chromosome analysis or CMA may be warranted depending on the maternal phenotype or medical history.
Finally, sex designation by NIPS may be discordant with physical examination. While sex designation by NIPS is relatively accurate, there are cases of XX or XY NIPS results with the opposite sexed infant.37, 38 Blood chromosome analysis is recommended (Table 2). Clinical findings suggestive of a disorder of sexual differentiation may warrant follow-up by CMA or an appropriate gene panel.
Importance of genetic counseling
Pretest counseling by the provider offering NIPS should include both the advantages and limitations of this screening test, as well as the alternatives.1 For patients with positive NIPS results, posttest access to genetic counseling by a trained genetics professional is essential for guiding management. To ensure an informed decision regarding testing and diagnostic follow-up, patients undergoing this screening should be provided with up-to-date, balanced and accurate information about the limitations of NIPS, the implications of both negative and positive NIPS results, the potential for false positives and false negatives, and the role of diagnostic testing. Patients should understand that diagnostic testing is both available and voluntary. Furthermore, the education of providers is of paramount importance.
Several points to consider following a positive noninvasive screening result are listed in Box 1.
References
Gregg AR, Skotko BG, Benkendorf JL et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med 2016;18:1056–1065.
Palomaki GE, Kloza EM, Lambert-Messerlian GM et al. DNA sequencing of maternal plasma to detect Down syndrome: an international clinical validation study. Genet Med 2011;13:913–920.
Chiu RWK, Lo YMD . Noninvasive prenatal diagnosis empowered by high-throughput sequencing. Prenat Diagn 2012;32:401–406.
Palomaki GE, Deciu C, Kloza EM et al. DNA sequencing of maternal plasma reliably identifies trisomy 18 and trisomy 13 as well as Down syndrome: an international collaborative study. Genet Med 2012;14:296–305.
Chen EZ, Chiu RW, Sun H et al. Noninvasive prenatal diagnosis of fetal trisomy 18 and trisomy 13 by maternal plasma DNA sequencing. PLoS ONE 2011;6:e21791.
Bianchi DW, Parker RL, Wentworth J et al. DNA sequencing versus standard prenatal aneuploidy screening. N Engl J Med 2014;370:799–808.
Wapner RJ, Babiarz JE, Levy B et al. Expanding the scope of noninvasive prenatal testing: detection of fetal microdeletion syndromes. Am J Obstet Gynecol 2015;212:332.e1–332.e9.
Helgeson J, Wardrop J, Boomer T et al. Clinical outcome of subchromosomal events detected by whole-genome noninvasive prenatal testing. Prenat Diagn 2015;35:999–1004.
Bianchi DW, Wilkins-Haug L . Integration of noninvasive DNA testing for aneuploidy into prenatal care: what has happened since the rubber met the road? Clin Chem 2014;60:78–87.
Society for Maternal-Fetal Medicine (SMFM) Publications Committee. #36: renatal aneuploidy screening using cell-free DNA. Am J Obstet Gynecol 2015;212:711–716.
The American College of Obstetricians and Gynecologists and the Society of Maternal Fetal Medicine. Committee Opinion Number 640: cell-free DNA screening for fetal aneuploidy. Obstet Gynecol 2015;126:e31–e37.
American College of Medical Genetics. Standards and Guidelines for Clinical Genetics Laboratories, Section E4, Prenatal Diagnosis, 2009 edn. http://www.acmg.net/StaticContent/SGs/Section_E_2011.pdf.
Crane JP, Cheung SW . An embryogenic model to explain cytogenetic inconsistencies observed in chorionic villus versus fetal tissue. Prenat Diagn 1988;8:119–129.
Ledbetter DH, Zachary JM, Simpson JL et al. Cytogenetic results from the US collaborative study on CVS. Prenat Diagn 1992;12:317–345.
Hahnemann JM, Vejerslev LO . European collaborative research on mosaicism in CVS (EUCROMIC) – fetal and extrafetal cell lineages in 192 gestations with CVS mosaicism involving single autosomal trisomy. Am J Med Genet 1997;70:179–187.
Gardner RJM, Sutherland GR, Shaffer LG . Chromosome Abnormalities and Genetic Counseling, 4th edn. Oxford University Press: New York, 2012: 444–452 351–374.
Ashoor G, Poon L, Ayngelaki A et al. Fetal fraction in maternal plasma cell-free DNA at 11-13 weeks’ gestation: effect of maternal and fetal factors. Fetal Diagn Ther 2012;31:237–243.
Grati FR, Bajaj K, Malvestiti F et al. The type of feto-placental aneuploidy detected by cfDNA testing may influence the choice of confirmatory diagnostic procedure. Prenat Diagn 2015;35:994–998.
Mersy E, Smits LJM, van Winden LAAP et al. Noninvasive detection of fetal trisomy 21: systematic review and report of quality and outcomes of diagnostic accuracy studies performed between 1997 and 2012. Hum Reprod Update 2013;19:318–329.
Dar P, Curnow KJ, Gross SJ et al. Clinical experience and follow-up with large scale single-nucleotide polymorphism-based noninvasive prenatal aneuploidy testing. Am J Obstet Gynecol 2014;211:527.e1–527.e17.
Cheung SW, Patel A, Leung TY . Accurate description of DNA-based noninvasive prenatal screening. N Engl J Med 2015;372:1675–1677.
Meck JM, Kramer Dugan E, Matyakhina L et al. Noninvasive prenatal screening for aneuploidy: positive predictive values based on cytogenetic findings. Am J Obstet Gynecol 2015;213:214.e1–214.e5.
Wang JC, Sahoo T, Schonberg S et al. Discordant noninvasive prenatal testing and cytogenetic results: a study of 109 consecutive cases. Genet Med 2015;17:234–236.
Bianchi DW . Circulating fetal DNA: its origin and diagnostic potential – a review. Placenta 2004;60:S93–S101.
Wang Y, Chen Y, Tian F et al. Maternal mosaicism is a significant contributor to discordant sex chromosomal aneuploidies associated with noninvasive prenatal testing. Clin Chem 2014;60:251–259.
Curnow KJ, Wilkins-Haug L, Ryan A et al. Detection of triploid, molar, and vanishing twin pregnancies by a single-nucleotide polymorphism-based noninvasive prenatal test. Am J Obstet Gynecol 2015;212:79.e1–79.e9.
Snyder MW, Simmons LE, Kitzman JO et al. Copy-number variation and false positive prenatal aneuploidy screening results. N Engl J Med 2015;372:1639–1645.
Horsman DE, Dill FJ, McGillivray BC, Kalousek DK . X chromosome aneuploidy in lymphocyte cultures from women with recurrent spontaneous abortions. Am J Med Genet 1987;28:981–987.
Nowinski GP, Van Dyke DL, Tilley BC et al. The frequency of aneuploidy in cultured lymphocytes is correlated with age and gender but not with reproductive history. Am J Hum Genet 1990;46:1101–1111.
Osborne CM, Hardisty E, Devers P et al. Discordant noninvasive prenatal testing results in a patient subsequently diagnosed with metastatic disease. Prenat Diagn 2013;22:609–611.
Bianchi DW, Chudova D, Sehnert AJ et al. Noninvasive prenatal testing and incidental detection of occult maternal malignancies. JAMA 2015;314:162–169.
Hook EB . Exclusion of chromosomal mosaicism: tables of 90%, 95%, and 99% confidence limits and comments on use. Am J Hum Genet 1977;29:94–97.
Mascarello JT, Hirsch B, Kearney HM et al. Section E9 of the American College of Medical Genetics technical standards and guidelines: fluorescence in situ hybridization. Genet Med 2011;13:667–675.
South ST, Lee C, Lamb AN, Higgins AW, Kearney HM . Working group for the American College of Medical Genetics and Genomics (ACMG) Laboratory Quality Assurance Committee. ACMG Standards and Guidelines for constitutional cytogenomic microarray analysis, including postnatal and prenatal applications: revision 2013. Genet Med 2013;15:901–909.
Yaron Y, Jani J, Schmid M, Oepkes D . Current status of testing for microdeletion syndromes and rare autosomal trisomies using cell-free DNA technology. Obstet Gynecol 2015;126:1095–1099.
Yatsenko SA, Peters DG, Saller DN et al. Maternal cell-free DNA-based screening for fetal microdeletion and the importance of careful diagnostic follow-up. Genet Med 2015;17:836–838.
Bianchi DW, Parsa S, Bhatt S et al. Fetal sex chromosome testing by maternal plasma DNA sequencing: clinical laboratory experience and biology. Obstet Gynecol 2015;125:375–382.
Devaney SA, Palomaki GE, Scott JA, Bianchi DW . Noninvasive fetal sex determination using cell-free fetal DNA. A systematic review and meta-analysis. JAMA 2011;306:627–636.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
A.M.C., Y.M.A., H.M.K., J.H.T. and J.M.M. are clinical laboratory directors at their respective institutions and perform the assays described herein as a clinical service. The other authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Cherry, A., Akkari, Y., Barr, K. et al. Diagnostic cytogenetic testing following positive noninvasive prenatal screening results: a clinical laboratory practice resource of the American College of Medical Genetics and Genomics (ACMG). Genet Med 19, 845–850 (2017). https://doi.org/10.1038/gim.2017.91
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.91
Keywords
This article is cited by
-
Positive predictive value estimates for noninvasive prenatal testing from data of a prenatal diagnosis laboratory and literature review
Molecular Cytogenetics (2022)
-
Performance of cell-free DNA sequencing-based non-invasive prenatal testing: experience on 36,456 singleton and multiple pregnancies
BMC Medical Genomics (2021)
-
Quality assurance checklist and additional considerations for canine clinical genetic testing laboratories: a follow-up to the published standards and guidelines
Human Genetics (2019)