Abstract
Purpose
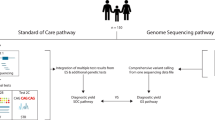
We sought to determine the diagnostic yield of whole-exome sequencing (WES) combined with phenotype-driven analysis of variants in patients with suspected genetic disorders.
Methods
WES was performed on a cohort of 51 patients presenting dysmorphisms with or without neurodevelopmental disorders of undetermined etiology. For each patient, a clinical geneticist reviewed the phenotypes and used the phenotype-driven analysis software PhenoVar (http://phenovar.med.usherbrooke.ca/) to analyze WES variants. The prioritized list of potential diagnoses returned was reviewed by the clinical geneticist, who selected candidate variants to be confirmed by segregation analysis. Conventional analysis of the individual variants was performed in parallel. The resulting candidate variants were subsequently reviewed by the same geneticist, to identify any additional potential diagnoses.
Results
A molecular diagnosis was identified in 35% of the patients using the conventional analysis, and 17 of these 18 diagnoses were independently identified using PhenoVar. The only diagnosis initially missed by PhenoVar was rescued when the optional “minimal phenotypic cutoff” filter was omitted. PhenoVar reduced by half the number of potential diagnoses per patient compared with the conventional analysis.
Conclusion
Phenotype-driven software prioritizes WES variants, provides an efficient diagnostic aid to clinical geneticists and laboratories, and should be incorporated in clinical practice.
Similar content being viewed by others
Introduction
Genetic disorders represent a significant health burden in developed countries.1 The incidence of genetic disorders is estimated at 5.32% in newborns, based on a follow-up period of 25 years.
In general genetics clinics, the traditional diagnostic studies, including array comparative genome hybridization, single-gene or targeted multiple-gene panel sequencing, and biochemical tests, yield a diagnosis in only 46% of the cases.2 Phenotypic variability and genetic heterogeneity still pose significant challenges to obtaining a molecular diagnosis, particularly for groups of disorders with similar or overlapping phenotypes, such as intellectual disability. Gene panels for a single clinical indication often vary from one laboratory to another. Adding to this complexity, the number of genes responsible for human disorders is expanding steadily every month, as only a fraction (15%) of the known protein-coding genes in the human genome are associated with diseases so far.3 Thus, negative results obtained using a gene panel for a given clinical indication would likely require the physician to order sequencing of additional genes or update panels as knowledge evolves. A genomic-based test (whole-exome or whole-genome sequencing) can overcome these limitations and potentially increase the molecular diagnostic yield.
Multiple studies have demonstrated the effectiveness of exome sequencing to unravel the molecular cause in patients with suspected genetic disorders. Among these, several studies addressed the etiologies of neurodevelopmental disorders. Soden et al.4 sequenced the exome of patients with neurodevelopmental disorders and found a diagnosis in 45 of 100 families. A large study by Wright et al.,5 aiming to decipher developmental disorders using trios, yielded a diagnosis in 27% of cases, of 1,133 children. Yang et al.6 found a molecular diagnosis in 25% of their heterogeneous cohort of 2,000 patients, but the diagnostic yield increased to 36% when considering only those having a neurologic disorder. Retterer et al.7 sequenced the exomes of 3,040 patients with suspected genetic disorders and found a definitive diagnosis in 29% of cases. Diagnostic yield was shown to vary according to the patient’s phenotype. The group with the highest diagnostic rate was patients who had hearing deficiencies (55%), followed by those who had visual deficiencies (47%) and the group with musculoskeletal system involvement (40%).
Routine utilization of exome sequencing in the clinical setting still faces challenges related to interpretation of a large number of candidate variants. Once variants of low quality were removed, Yang et al.6 identified a mean of 875 variants per patient to be analyzed. To perform the analysis of such variants, several software programs have been designed. A small but growing number of bioinformatics tools have been designed to incorporate the patient’s phenotype in the algorithm to identify the causal mutation(s), including our tool, PhenoVar.8 Each of these tools has its own particularities, while they generally share some common features. Most use the Human Phenotype Ontology (HPO) database. Exomiser,9 for example, uses the Mouse Phenotype Ontology and the Zebrafish Phenotype Ontology database in addition to HPO to link the phenotype to a disease. eXtasy is another tool that uses HPO to relate the phenotype to a disease.10 However, eXtasy can only perform prioritization on nonsynonymous variants. Another tool used to prioritize variants is Phevor.11 Besides HPO, it also uses the Mammalian Phenotype Ontology, the Disease Ontology, and the Gene Ontology databases, which allows this tool to not be restricted to known disease-associated genes. Phevor is a Web-based tool, but does not support the standard VCF format file and requires VAAST simple files. Other programs rely only on phenotypic traits (HPO terms) to prioritize certain genes and do not require any genotypic data, such as Phenolyser12 and Phenomizer.13 However, it should be noted that an automated Phenolyzer analysis pipeline has been implemented in the wANNOVAR server to facilitate its use with sequencing data.12 Our software, PhenoVar, uses the HPO and OMIM databases to determine the gene–phenotype and phenotype–disease correlation, and to prioritize variants.8 It accepts files in standard VCF format, and includes various variant quality filters and classification of variants according to predictions of their pathogenicity, to improve diagnostic performance. PhenoVar is a Web-based tool usable by clinicians that focuses on known disease-associated genes.
Here, we report a comparison of the performance of the phenotype-based tool PhenoVar versus the conventional analysis of the individual variants identified in a cohort of patients with suspected genetic disorders.
Materials and methods
Recruitment of the patients
A total of 70 patients with unknown diagnoses, followed in the genetic clinics at the University Health Center of Sherbrooke from 2013 to 2016, were proposed for inclusion in the project (Supplementary Figure S1 online). An expert committee, composed of two additional and independent medical geneticists, reviewed each submitted case to evaluate whether study criteria were met (see Supplementary Figure S2), and proposed additional genetic testing (single gene or gene panel) when applicable. Of the 70 patients considered, 9 did not meet the criteria and diagnoses were found for 3 patients following suggested additional testing. Genetic counseling was then provided to the remaining 58 patients. Of these, 7 patients (or parents of patients) declined to participate in the project. We therefore sequenced the exomes of 51 patients (single unrelated probands): 21 females and 30 males (Table 1). At the time of sequencing, 21 patients were less than 5 years old, 25 were between 5 and 17 years old, and 5 were aged 18 years or older. All patients had negative or inconclusive results from previous investigations. These investigations included comparative genome hybridization microarray (96%), karyotype (43%), metabolic workup (78%), single gene or gene panel (84%), imaging (82%), methylation analysis (33%), fluorescence in situ hybridization (10%), and other molecular tests for a specific disease (31%). Blood samples of the patients, the patient’s parents, and their siblings if applicable, were collected. Patients were followed on a yearly basis after the exome sequencing was performed, if results were negative. The study has been approved by the institutional ethics review board of Université de Sherbrooke (project 12–167). All participants or their legal tutors provided written consent.
Exome sequencing
Exome sequencing was performed at the McGill University and Génome Québec Innovation Centre (Montreal, Canada) and Fulgent (Temple City, CA). DNA libraries were prepared for each patient (TruSeq; Illumina), followed by target enrichment (Agilent SureSelect All Exon kit v4 or v5 or Illumina Truseq Exome) and sequenced on a HiSeq 2000 (Illumina) with 100-bp paired-end protocol or HiSeq 4000 (Illumina) with 150-bp paired-end protocol. Median coverage per sample ranged from 86 × to 423 ×, with an overall average coverage of 190 ×. All but 2 of the 51 exomes had a median coverage of more than 100 ×.
Bioinformatics analyses
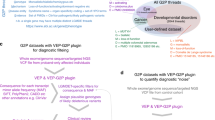
We analyzed the sequencing data using a Linux-based bioinformatics pipeline based on the one developed by the McGill University and Génome Québec Innovation Centre (https://bitbucket.org/mugqic/mugqic_pipelines) as previously described.14 Briefly, (i) raw reads were trimmed using Trimmomatic15 (version 0.32); (ii) sequence alignment was performed with Burrows–Wheeler Aligner16 (version 0.7.10); (iii) genetic variations (single-nucleotide polymorphisms and indels) were called with haplotypeCaller using the Genome Analysis Toolkit17 (version 3.2.2) with prior local realignment, base recalibration, and removal of polymerase chain reaction duplicates using Picard (version 1.123, http://broadinstitute.github.io/picard/); (iv) gene annotation was performed with SnpEff/SnpSift18 (version 3.6, including SIFT, Polyphen2, MutationTaster predictions) with an additional in-house script to annotate variants present in the ClinVar19 database; and (v) a filtering process removed variations outside targeted sequences, with population frequency >1% (dbSNP 138 and ExAC 0.3 (ref.20)), genotype quality less than Q30 or present in three or more local controls sequenced on the same platform (Figure 1). Coverage depth was calculated using BED Tools.21
Filtered variant lists obtained from the bioinformatics pipeline were then interpreted through two parallel approaches: using our previously described software PhenoVar8 and using conventional analysis (Figure 1).
Phenotype-driven analysis of variants, using PhenoVar
The conception, algorithm, and use of PhenoVar (http://phenovar-dev.udes.genap.ca) have been described in detail.8 Briefly, the clinician inputs in PhenoVar the patient’s list of variants (VCF file format) and selects three phenotypic traits or more, using the HPO nomenclature. PhenoVar automatically prioritizes diagnoses for validation based on both the phenotypic and genomic information of a proband. It calculates a patient-specific diagnostic score for each OMIM entry with known molecular basis. The diagnostic score assigned to a given syndrome is the sum of its phenotypic and genotypic weights, the latter having a larger impact. For each syndrome listed in the HPO database the phenotypic weight is determined by calculating the similarity between the proband and the different patients available in a local database (Phenobase). Phenobase includes patients simulated using HPO and real patients. The genotypic weight for each syndrome corresponds to the highest predicted pathogenicity of any variant(s) present in the corresponding associated gene(s). The variants are sorted into three categories that are assigned a different score (from high to low): (i) known disease-causing (ClinVar19) and likely pathogenic variants (e.g., splice-site donor and acceptor, nonsense, frameshift variants), (ii) variants of uncertain clinical significance (e.g., missense, in-frame deletion/insertion), (iii) likely benign and benign variants (untranslated regions, synonymous or intronic variants, unless they are reported as pathogenic or likely pathogenic in ClinVar). The different syndromes are then ranked according to their diagnostic score. Syndromes for which the phenotypic score is below a predetermined cutoff value, and therefore considered unrelated to the patient’s phenotype, are removed from the list if this “minimal phenotypic cutoff” option is selected. When there is no phenotypic trait in common between the patient investigated and the syndrome definition in the HPO database, the phenotypic score will usually be below the minimum phenotypic cutoff value. However, other factors will also influence the phenotypic score, such as a syndrome’s phenotypic trait frequencies, the presence of one or more traits in the patient that have not been reported in the syndrome, and whether a syndrome is defined by a very large number of traits.8 This cutoff is the default option that was used in the present study. The clinical geneticist then reviews the short list of potential diagnoses and selects candidates for confirmation by segregation analysis using Sanger sequencing (parents and additional available relatives when appropriate).
In version 2.0, additional modifications have been incorporated into the original version of PhenoVar.8 Genes with two variants of uncertain significance causing recessive disorders have increased genotypic weight and are thus prioritized on the list of potential diagnoses. Custom filters enable the user to filter out variants according to a desired sequencing quality score (QUAL and GQ scores), conservation and evolutionary constraint (PhasConst100way, genomic evolutionary rate profiling22), or variant frequency cutoffs specific to the disorder’s mode of inheritance. This last filter uses a conservative approach when more than one mode of inheritance is reported, with the highest frequency cutoff being selected.
Conventional manual analysis of variants
Variants located in genes with known disease associations according to the OMIM database were first selected from the VCF file, originating from the McGill University and Génome Québec Innovation Centre pipeline described above. Then, single heterozygous variants in recessive genes and variants with population frequencies above 0.01% in dominant genes were filtered out. However, single heterozygous variants found within genes causing recessive disorders and known to be likely pathogenic or pathogenic in ClinVar or Human Gene Mutation Database public databases, or predicted to cause a loss of function, were included in the analysis. Manual review of read depth was performed for those genes to rule out deletion/duplication. In addition, missense variants that were predicted to be tolerated using bioinformatics tools (SIFT,23 PolyPhen2 (ref.24) and MutationTaster2 (ref.25)), and involving amino acid changes that were not conserved between species, were filtered out. Variants predicted to cause loss of function (frameshifts, splice-site donors/acceptors, and nonsenses) or variants known to be disease-causing (ClinVar or public Human Gene Mutation Database) were prioritized in the final list to be reviewed by the clinical geneticist, after the PhenoVar analysis was completed, in order to select additional candidates that could have been missed by PhenoVar.
Results
Whole-exome sequencing was performed on patients with a suspected genetic disorder, but with unknown diagnosis. Most of the patients presented with behavioral and cognitive involvement (84%), a defect in the nervous system with or without malformations (59%), and/or craniofacial dysmorphisms (53%) (Figure 2). On average (mean ± SD), 527 ± 70 rare variants were observed per patient following filtration, ranging from 412 to 746 variants. Of these, 125 ± 16 variants (range: 93–170) were present in genes known to be associated with Mendelian disorders.
Among the 51 patients sequenced, we identified a diagnosis in 18 using conventional manual analysis, giving a diagnostic yield of 35%. Putative diagnoses identified are listed in Table 2. One of these 18 cases (patient EX0014 (ref.26)) was negative on the initial analysis in mid-2013, but reanalysis of the genomic data on follow-up clinic a year later led to the diagnosis of Schaaf–Yang syndrome, the molecular basis of which was first reported in late 2013.27 No phenotypes were associated with a diagnostic yield significantly higher than the mean of 35%, although some involved systems were associated with a lower diagnostic yield: renal anomalies (0%), skin–hair–nails (16%), gastrointestinal system (22%), and limb anomalies (24%) (Figure 2). Most causal mutations were inherited, with de novo mutations identified in 7/18 diagnosed patients (39%) (Supplementary Figure S3 and Table 2).
In comparison to manual analysis, the analysis performed with the phenotype-driven software independently identified 17 of the 18 diagnoses, one initially being missed with PhenoVar (case EX0022). The phenotype of this patient consisted of intrauterine growth restriction with severe microcephaly, which has been associated with LIG4 deficiency, but this association was first reported only a few months prior to our exome analysis.28 These phenotypic traits were not yet incorporated in the HPO database at the time of analysis. This resulted in a phenotypic score below the minimal cutoff, and thus LIG4 mutations were classified by PhenoVar as unrelated to the patient’s phenotype and were filtered out. Following the manual analysis that uncovered the diagnosis, we were able to visualize the causative variants in PhenoVar by deselecting the optional filter that removes diagnoses not reaching the “minimal phenotypic cutoff” score. However, this had a significant impact on the number of diagnoses to review. When the minimal phenotypic cutoff was selected, only 17 potential diagnoses were listed by PhenoVar, not including LIG4 syndrome. Deselecting the cutoff option led to 136 potential diagnoses, but LIG4 was visualized this time, in eleventh position.
We sought to compare our results with those obtained from another phenotype-driven software based on the HPO database, and to investigate whether this limitation could be overcome by a different algorithm not relying on a minimum phenotypic cutoff. We decided on Exomiser, because it is popular, freely available online, and it enabled us to use directly the VCF file (http://www.sanger.ac.uk/science/tools/exomiser, accessed January 2017). We limited our analysis to the first prioritized 100 potential diagnoses listed by Exomiser. By way of comparison, our manual analysis usually required the clinician to review about 34 potential diagnoses per patient (ranging from 26 to 45), while with PhenoVar this number dropped to 15 on average (ranging from 1 to 26). The diagnosis of the missed case, EX0022, was not included among the first 100 diagnoses listed by Exomiser. Furthermore, when we compared the 18 diagnoses made with the conventional analysis, only 13 of these (72%) were found in the first 100 diagnoses listed by Exomiser. In addition, 89% of the 18 diagnoses were found in the top 10 ranks of PhenoVar whereas only 56% were found in the top 10 ranks of Exomiser (Figure 3).
Discussion
Several studies have revealed the potential of exome sequencing by proving that it can help find diagnoses where other traditional approaches have failed.,6,7,29 In our cohort of 51 patients, most of whom (84%) presented with dysmorphisms and/or neurodevelopmental disorders, a total of 18 diagnoses were found, representing a global diagnostic yield of 35%. All patients had undergone extensive workup prior to exome sequencing. This high yield is in agreement with other studies involving cohorts with a large proportion of patients having neurodevelopmental disorders: Retterer et al.7 and Soden et al.4 found a molecular diagnosis in 29% and 45% of their patients, respectively. In addition, the proportion of diagnosed patients in our study with a dominant de novo mutation is similar to those studies, with 7 of 18 (39%), compared with 44% in Retterer et al.’s7 study. We did not observe particular phenotypes significantly associated with higher diagnostic yield within our cohort compared with others.7 This might be related to the relatively small sample size and to the fact that our cohort was more homogeneous.
The reported diagnostic yield in our study might be underestimated owing to methodological limitations. On average, four single heterozygous likely pathogenic or pathogenic variants were found per patient, in genes known to cause recessive disorders. Limited sensitivity of next-generation sequencing deletion/duplication analysis or low coverage in GC-rich or deep intronic regions might have prevented the finding of a second mutation to support causality, and contributed to decreased diagnostic yield. The large number of missense variations of uncertain significance also implies the use of bioinformatic predictions on gene function to balance the amount of time analyzing each case, but potentially at the expense of decreased clinical sensitivity.
Because exome sequencing identifies a large number of variations, we hypothesized that phenotype-driven analysis might facilitate the integration of exome sequencing as a more routine test in the clinic, by providing an alternative tool that reduces time of analysis in the laboratory and enables clinicians to interact directly with genomic data. Indeed, PhenoVar was able to reduce by about half the number of potential diagnoses per patient (mean of 15 vs. 34) in comparison with our manual approach. This decreased number of diagnoses implies a twofold reduction in review time by the clinical geneticist. In addition, PhenoVar can also decrease the time spent on analyzing the variants before producing the list of potential diagnoses. This task is completed in less than 2 min by PhenoVar. In our hands, this translated to roughly 15 min compared with 90 min per patient spent in total for the variants analysis and the review of candidate conditions. Because time spent on variants analysis may vary significantly from one laboratory to another according to the staff’s experience, or because of the incorporation of additional bioinformatic scripts in the manual analysis, the time savings by the use of PhenoVar would also be variable. Moreover, reanalysis of negative exomes puts a burden on laboratories,30 whereas enabling the clinician to reanalyze the exome data with PhenoVar at the time of follow-up in the clinic provided the diagnosis for patient EX0014 (ref.26).
However, in comparison to the conventional manual analysis, our phenotype-driven software PhenoVar missed one diagnosis, which suggested a limitation to phenotype-driven analysis alone. As shown by our comparison between PhenoVar and Exomiser, this was not only related to our use of a phenotypic cutoff, below which disorders are considered not related to the patient phenotype, because Exomiser also missed this case. In both cases, this was caused by the absence of the relevant phenotypic traits in the HPO database. Phenotype database completeness and accurate choice of phenotypic trait are both critical; limitations in these aspects can lead to false negative outcomes in phenotype-driven analysis.
There is a risk that the phenotype database (in our case HPO) does not yet contain a particular phenotype linked to a gene/disorder, and thus it will be ignored or not prioritized depending on the algorithm used. Although the HPO and OMIM databases have different frequencies of updates for adding genes and/or phenotypes, it might take months before a specific entry is updated following a publication that redefines the phenotypic spectrum of a genetic condition. This is not unexpected, given the burden of reviewing the literature in detail and the high discovery rate of new variants causing diseases. When updating database entries, the selection of the appropriate terms is critical and requires specialized clinical expertise, which is not always readily available. Phenotype-driven tools such as PhenoVar will likely need to access a clinical grade phenotypic database to improve performance. In the case of PhenoVar, the limitation associated with delayed updates may sometimes be overcome by deselecting the “minimum phenotypic cutoff.” Diagnoses with likely pathogenic or known pathogenic variants will be then prioritized over variants of uncertain significance by the algorithm, because the genotype has more weight on the final diagnostic score than does the phenotype. While one could therefore question the utility of the minimal phenotypic cutoff, performing an initial analysis omitting the cutoff has the significant disadvantage of increasing the number of potential diagnoses listed for manual review. As mentioned above for LIG4 deficiency, when the minimal phenotypic cutoff was removed, 136 potential diagnoses were listed compared with only 17 when this cutoff was in use. Moreover, in the case of causative missense variants that are classified as being of uncertain significance because they have not been yet reported in databases, the correct diagnosis is found at a lower rank when the minimal phenotypic cutoff is omitted. This is secondary to the retention of diagnoses unrelated to the patient’s phenotype but included because of likely pathogenic or pathogenic variants, which are prioritized over variants of uncertain significance. However, these will usually represent carrier states or incidental findings. The strategy adopted by our laboratory therefore is to perform a second analysis without the minimum phenotypic cutoff, if the initial analysis yields negative results, and to examine all disorders caused by likely pathogenic and known pathogenic variants.
The phenotypic traits or HPO terms chosen by the clinician are critical in the comparison of the patient against the phenotypic criteria of the various syndromes included in the databases or in our case, patient descriptions included in Phenobase. Our algorithm accepts terms that are closely related to the actual traits listed in the syndrome definition (following the HPO nomenclature), but misidentification of one or more phenotypic traits might lead to failure to recognize the phenotype as corresponding to the causal syndrome. It is however possible to retry the analysis with modifications to the descriptive terms entered and/or with addition of further phenotypic traits, if the diagnosis is not identified at the first pass.
Some additional improvements are desirable to facilitate adoption of phenotype-driven analysis in both laboratories and clinics. An important potential source of improvement of PhenoVar’s performance is the possibility to expand the number of real patients included in Phenobase, which serves to determine the similarity between a given syndrome and the tested patient. This would certainly help to better capture the phenotypic diversity of the syndromes. The PhenoVar algorithm would then be less reliant on the completeness of the HPO database, which is currently used to simulate a minimal number of patients affected by a given syndrome who are then included in Phenobase. Different strategies could be used, such as addition of well-described patients from the literature, or through a collaborative effort of clinicians using PhenoVar subsequently contributing cases following confirmation of the correct diagnosis. However, these approaches might still remain limited. PhenomeCentral, a collaborative collection of patients (phenotypes and genotypes), has been used successfully in the research setting to match patients with potential similar but yet-undefined conditions.31 The various phenotype-driven software could certainly benefit from a similar clinical resource but with patients with known diagnoses. Finally, the Web interface of PhenoVar has been created to favor utilization by clinicians, but an adapted version that could be incorporated in bioinformatic pipelines could help to facilitate adoption by laboratories. Currently, PhenoVar is compatible with VCF produced by Genome Analysis Toolkit and annotated with SnpEff/SnpSift.
In conclusion, analytic approaches using phenotype-driven software to prioritize whole-exome sequencing variants provide an efficient diagnostic aid to clinical geneticists and laboratories, and should be incorporated in clinical practice.
References
Verma IC & Puri RD. Global burden of genetic disease and the role of genetic screening. Semin Fetal Neonatal Med 2015;20:354–363.
Shashi V et al. The utility of the traditional medical genetics diagnostic evaluation in the context of next-generation sequencing for undiagnosed genetic disorders. Genet Med 2014;16:176–182.
Chong JX et al. The genetic basis of Mendelian phenotypes: discoveries, challenges, and opportunities. Am J Hum Genet 2015;97:199–215.
Soden SE et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med 2014;6:265ra168.
Wright CF et al. Genetic diagnosis of developmental disorders in the DDD study: a scalable analysis of genome-wide research data. Lancet 2015;385:1305–1314.
Yang Y et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 2014;312:1870.
Retterer K et al. Clinical application of whole-exome sequencing across clinical indications. Genet Med 2015;18:1–9.
Trakadis YJ et al. PhenoVar: a phenotype-driven approach in clinical genomics for the diagnosis of polymalformative syndromes. BMC Med Genomics 2014;7:22.
Robinson PN et al. Improved exome prioritization of disease genes through cross-species phenotype comparison. Genome Res 2014;24:340–348.
Smedley D & Robinson PN. Phenotype-driven strategies for exome prioritization of human Mendelian disease genes. Genome Med 2015;7:81.
Singleton MV et al. Phevor combines multiple biomedical ontologies for accurate identification of disease-causing alleles in single individuals and small nuclear families. Am J Hum Genet 2014;94:599–610.
Yang H, Robinson PN & Wang K. Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat Methods 2015;12:841–843.
Köhler S et al. Clinical diagnostics in human genetics with semantic similarity searches in ontologies. Am J Hum Genet 2009;85:457–464.
Lévesque S et al. Diagnosis of late-onset Pompe disease and other muscle disorders by next-generation sequencing. Orphanet J Rare Dis 2016;11:8.
Bolger AM, Lohse M & Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014;30:2114–2120.
Pabinger S et al. A survey of tools for variant analysis of next-generation genome sequencing data. Brief Bioinform 2014;15:256–278.
DePristo MA et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 2011;43:491–498.
Cingolani P et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118 ; iso-2; iso-3. Fly (Austin) 2012;6:80–92.
Landrum MJ et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res 2016;44:D862–D868.
Lek M et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016;536:285–291.
Quinlan AR & Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010;26:841–842.
Cooper GM et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res 2005;15:901–913.
Ng PC & Henikoff S. Predicting deleterious amino acid substitutions. Genome Res 2001;11:863–874.
Adzhubei IA et al. A method and server for predicting damaging missense mutations. Nat Methods 2010;7:248–249.
Schwarz JM, Cooper DN, Schuelke M & Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 2014;11:361–362.
Fountain MD et al. The phenotypic spectrum of Schaaf-Yang syndrome: 18 new affected individuals from 14 families. Genet Med 2017;19:45–52.
Schaaf CP et al. Truncating mutations of MAGEL2 cause Prader-Willi phenotypes and autism. Nat Genet 2013;45:1405–1408.
Murray JE et al. Extreme growth failure is a common presentation of ligase IV deficiency. Hum Mutat 2014;35:76–85.
Thevenon J et al. Diagnostic odyssey in severe neurodevelopmental disorders: toward clinical whole-exome sequencing as a first-line diagnostic test. Clin Genet 2016;89:700–707.
O’Daniel JM et al. A survey of current practices for genomic sequencing test interpretation and reporting processes in US laboratories. Genet Med 2017;19:575–582.
Buske OJ et al. PhenomeCentral: a portal for phenotypic and genotypic matchmaking of patients with rare genetic diseases. Hum Mutat 2015;36:931–940.
Acknowledgments
The study was supported by institutional funds of the Université de Sherbrooke, La Fondation du Grand Défi Pierre Lavoie, and La Fondation des Étoiles. We are thankful to Génome Québec and Fulgent for the exome sequencing. Moreover, we thank Marie Edmont and Laura Dempsey-Nunez for the genetic counseling. We are also grateful to the patients and their families for their participation in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Thuriot, F., Buote, C., Gravel, E. et al. Clinical validity of phenotype-driven analysis software PhenoVar as a diagnostic aid for clinical geneticists in the interpretation of whole-exome sequencing data. Genet Med 20, 942–949 (2018). https://doi.org/10.1038/gim.2017.239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.239
Keywords
This article is cited by
-
Best practices for the interpretation and reporting of clinical whole genome sequencing
npj Genomic Medicine (2022)
-
Powerful use of automated prioritization of candidate variants in genetic hearing loss with extreme etiologic heterogeneity
Scientific Reports (2021)