Abstract
Purpose
We aimed to determine the frequency of RET mosaicism in Hirschsprung disease (HSCR), test whether it has been underestimated, and to assess its contribution to HSCR risk.
Methods
Targeted exome sequencing (n = 83) and RET single-gene screening (n = 69) were performed. Amplicon-based deep sequencing was applied on multiple tissue samples. TA cloning and sequencing were conducted for validation.
Results
We identified eight de novo mutations in 152 patients (5.2%), of which six were pathogenic mosaic mutations. Two of these patients were somatic mosaics, with mutations detected in blood, colon, and saliva (mutant allele frequency: 35–44%). In addition, germ-line mosaicism was identified in four clinically unaffected subjects, each with an affected child, in multiple tissues (mutant allele frequency: 1–28%).
Conclusion
Somatic mutations of the RET gene are underrecognized in HSCR. Molecular investigation of the parents of patients with seemingly sporadic mutations is essential to determine recurrence risk in these families.
Similar content being viewed by others
Introduction
Hirschsprung disease (HSCR, MIM 142623) was first clinically described in 1888, but its characteristic pathological change, an absence of enteric ganglia along a variable length of the intestine, was not described until several decades later.1 This developmental disorder is the main genetic cause of functional intestinal obstructions in neonates and has a population incidence of 1/5,000 live births.2 HSCR occurs as an isolated trait in 70% of patients, and is frequently associated with a chromosomal abnormality (12%) or additional congenital anomalies (18%).3 Although all Mendelian modes of inheritance have been described in syndromic HSCR, isolated HSCR stands as a model for complex disorders with low, sex-dependent penetrance and variable clinical expressivity.4 Genetic studies have identified rare coding variants in 14 genes (RET, EDNRB, EDN3, ECE1, GDNF, NRTN, SOX10, PHOX2B, ZEB2, TCF4, KBP, L1CAM, SEMA3C, and SEMA3D) that together explain ~10% of cases.4,5,6 Among these, RET is the major gene because (i) it contains >80% of all known disease-causing mutations,7 (ii) it harbors an excess of loss-of-function (LOF) de novo mutations (DNMs),8 and (iii) both targeted exome sequencing (TES) and whole-exome sequencing have revealed a statistically significant excess of deleterious variants at RET.
Surgical treatment of HSCR, developed in 1948, has evidently decreased its mortality and morbidity, and, thereby, uncovered its familial transmission. However, the reasons for its high heritability and phenotypic variation are still unclear, despite large-scale efforts in searching for additional genes,8 regulatory elements,9 and related molecules,10,11 and, finally, nongenetic factors.12,13 During the past several years, germ-line and somatic mosaicism have emerged as important factors that contribute to phenotypic variability and risk in genetic disorders. Somatic mosaicism has been implicated in >30 monogenic disorders that show variable expressivity.14,15,16,17 The first case of somatic mosaicism for a RET mutation was reported in 2014, in a healthy mother who had two offspring both affected by HSCR.18 Second, by analyzing 125 families with HSCR and manually inspecting the sequence electropherograms of 33 families with RET mutations, Müller et al.19 discovered genetic mosaicism of a frameshift mutation in an asymptomatic father (1/33, 3.03%). Third, by comparing the homozygous status of two polymorphisms in RET on DNA extracted from 53 paraffin-embedded tissue samples, Moore and Zaahl concluded that somatic mutations exist in aganglionic tissue and may promote local aganglionosis through deregulated receptor activity.20 The objective of this present work was to determine the true frequency of mosaicism in HSCR in independent newly ascertained samples, test whether it has been underestimated, and to assess its contribution to HSCR risk.
Materials and methods
Subjects and genetic screening
This study was approved by the Ethics Committee of the Capital Institute of Pediatrics (reference SHERLL 2013039). Participants or their parents provided informed, written consent for the genetic studies. Eighty-three of 152 patients (all isolated HSCR except for one, diagnosed based on surgical reports and pathological examination) initially chosen for TES were from our previous project (under review), including 51/32 male/female; 32/19/32 S-HSCR/L-HSCR/TCA (S, short-segment; L, long-segment; TCA, total colonic aganglionosis). Blood DNA from an additional 69 patients (67/2 were male/female; 65/2/2 were S-HSCR/L-HSCR/TCA) were subjected to screening of all exons, intron–exon boundaries, and 10-bp flanking sequences of RET (primer sequences available on request). Sanger sequencing was performed (forward and reverse) with a Big Dye Terminator v3.1 Kit (Applied Biosystems, Foster City, CA) on an ABI 3130XL automated sequencer. Saliva, colon, sperm, and hair DNA was extracted using the TIANamp Micro DNA Kit (Tiangen Biotech, Beijing, China).
Amplicon-based deep sequencing and TA-cloning validation
Specific amplicons containing the variant were generated for amplicon-based deep sequencing (ADS) performed on an Ion Torrent Personal Genome Machine (Life Technologies, Carlsbad, CA) as described previously.14 For TA-cloning validation, polymerase chain reaction (PCR) products were purified and cloned into the Trans109 Chemically Competent Cell (TransGen Biotech, Beijing, China) using the pMD18-T Vector Cloning Kit (Takara Biomedical Technology, Beijing, China).
Results
Identification of LOF de novo mutations in RET in isolated HSCR probands
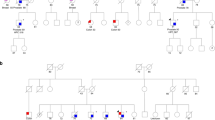
TES identified nine RET variants among 83 patients (10.8%): seven LOF mutations (c.254G>A [p.Trp85X], c.754G>T [p.Glu252X], c.789C>G [p.Tyr263X], c.2308C>T [p.Arg770X], c.2333delT [p.Val778Alafs*1], c.2578C>T [p.Gln860X], and c.2802-2A>G), one pathogenic missense mutation (c.229C>T [p.Arg77Cys], family 6),21 and one possibly pathogenic in-frame insertion (c.200insTCC [p.Arg67insLeu], family 2) (Figure Figure 1). The last variant does not belong with the LOF mutations, but was still suspected to be pathogenic on the basis of its absence from the unaffected sibling and absence from large control populations from the 1000 Genomes Project, and the National Heart, Lung, and Blood Institute and Exome Aggregation Consortium exome sequencing projects. None of the LOF mutations or the possibly pathogenic mutation have been reported in the literature. The male patient (II-1) in family 5 had a nonidentical twin brother (II-2) who was symptom-free and wild-type for the mutation p.Gln860X. In addition, the affected boy (II-1) in family 7 also had a younger unaffected brother who was wild-type. We obtained parental DNA from families 1–8, in which Sanger sequencing indicated that six of these cases (families 1, 2, 5, 6, 7, and 8) were high-confidence candidate de novo (75%). In contrast, the proband’s father in family 3 and the proband’s mother in family 4 both showed somatic mosaicism, each with a small mutant allele peak on the dideoxy-sequence traces (Figure Figure 2a).
(a) Domain organization and identified mutations in RET (GenBank: NP_066124). Variants detected in Hirschsprung disease (HSCR) families by targeted exome sequencing are shown above and those revealed by RET Sanger sequencing are displayed below. (b) Evolutionary conservation of the RET sequence flanking the altered Arg67, Arg77, Thr278, Arg417, and Asp571 amino acids. (c) Three-dimensional representation of the crystal structure of human RET (PDB: 4UX8, residues 29–508). The mutated residues are highlighted in red (Arg67), green (Arg77), purple (Thr278), and orange (Arg417). CD1-4, cadherin domain1-4 (amino acids 168–272); CYS, cysteine-rich domain (amino acids 517–635); SP, signal peptide (amino acids 1–24); PDB, protein data bank; TES, targeted exome sequencing; TK, tyrosine kinase domain (amino acids 724–1,016); TM, transmembrane domain (amino acids 636–657).
(a) Dideoxy-sequence traces for the nine families with loss-of-function or possibly pathogenic mutations detected in RET by targeted exome sequencing. Parental DNA was available for families 1–8, and Sanger sequencing indicated that six of them (families 1, 2, 5, 6, 7, and 8) were high-confidence candidate de novo. In family 3, a small proportion of the mutant c.789C>G allele was suspected to be present in the proband’s father, based on both the presence of a small C peak and the reduced relative height of the normal G peak (reverse sequencing). In family 4, a small proportion of the mutant c.254G>A allele was suspected to be present in the proband’s mother, based on both the presence of a small A peak and a normal sized G peak. (b) Integrative Genomics Viewer visualization of the amplicon-based deep sequencing results on six individuals with mosaic RET mutations. The family member identification number, sample type, variant information, alternate-allele read counts, and total coverage are listed on the left.
Under the LGDbroad (likely gene-disrupted) mutation prioritizing criteria (including stop-gain, splice region (±5) variants, frameshift indels, in-frame indels, and non-synonymous variants predicted to be damaging by at least three bioinformatics tools of five, i.e., SIFT, PolyPhen, MetalR, CADD, and M-Cap), our further screening in a cohort of 69 cases with milder clinical expressivity (mainly S-HSCR patients) revealed variants in six patients (6/69, 8.7%, Figure Figure 1 and Supplementary Table S1 online): one nonsense mutation (c.3148C>T [p.Arg1050X] in one family), three missense variants (c.833C>A [p.Thr278Asn] in two families, c.1250G>A [p.Arg417His] in one family, and c.1711G>A [p.Asp571Asn] in one family), and one splice region variant (c.2392+5G>A in one family). Parental DNA was available for two families (p.Asp571Asn and p.Arg1050X) and the inheritance pattern investigation excluded de novo events (data not shown). Therefore, in 10 families (8 from TES and 2 from RET Sanger sequencing) with RET pathogenic or likely gene-disrupted variants and detectable parental DNA, 6 (families 1, 2, 5, 6, 7, and 8 in TES) were DNMs, 2 (families 3 and 4 in TES) demonstrated parental mosaicism, and 2 (p.Asp571Asn and p.Arg1050X from RET Sanger sequencing) were segregating variants.
ADS revealed mosaic RET mutations
Next, we focused on the six high-confidence candidate DNMs and developed ADS to test their fidelity. We first designed primers (sequences in Supplementary Table S2) for amplifying each of the mutations that were identified by TES. Using blood DNA of all available family members, we found clear evidence for mosaic RET mutations in 4 of 6 pedigrees. Two affected patients (in families 1 and 2) were suspected to be somatic mosaics with mutant allele frequencies of 39% and 35% respectively. Surprisingly, extremely low-level mosaicism in one of the parents was detected in two additional families (5 and 6, mutant allele frequencies 2% and 1%), where the mutant peak was entirety invisible on Sanger sequencing. Patients in families 7 and 8 were the only two germ-line de novo mutation carriers. Somatic mosaicism was confirmed in one of the healthy parents in families 3 and 4, with the mutant allele discovered at 14% and 28%. Copy-number changes at these sites were excluded by performing quantitative PCR using 5 pairs of primers covering exons 2, 3, 4, 13, and 17 in RET (data not shown).
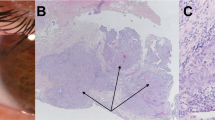
To verify the above results, multiple tissue samples were collected from the six individuals. The second-round ADS validated the nature of the mosaic mutations (Table 1, Figure Figure 2b, and Supplementary Table S3). By comparing four different sequencing techniques, Acuna-Hidalgo et al.22 showed that ADS is the most precise technique for identifying true heterozygosity, with an allelic ratio of 48.2 ± 4.4% (average ± SD). In the current study, two de novo mutations showed a significant deviation from the expected ratio for true germ-line heterozygosity, strongly suggesting a post-zygotic origin (Table 1). The male patient in family 1 (sample ID HSCR0116) was diagnosed with TCA soon after birth and surgically treated at 11 months of age. Deep sequencing confirmed a 39% frequency for c.754G>T in the blood sample and showed that the variant was present at 39% in his saliva and 44% in his colon (formalin-fixed paraffin-embedded (FFPE) sections). In the female patient (sample ID HSCR0127) in family 2, also affected by TCA, the in-frame insertion variant c.200insTCC was discovered at a frequency of 35% in blood, 39% in saliva, and 38% in colon (FFPE). By contrast, the average mutant allelic ratio of all germ-line heterozygous probands in families 3–9 was 49.5% (ranging from 45 to 52%, Supplementary Table S3), except for HSCR0146 in family 4. Considering the relatively high level (28%) of the mutant allele for c.254G>A in his mother’s blood and saliva, the patient was reasonably defined as heterozygous initially. However, when we tested his blood and FFPE samples by ADS (round 1), both alternative-allele frequencies showed statistically significant deviations from 50%, at 63% and 22% (Supplementary Tables S3–S4 and Supplementary Figure S1). To exclude technical artifacts resulting from biased allele amplification during PCR, we then generated a second independent amplicon with different PCR primers to resequence the mutation by ADS (round 2, Supplementary Table S4 and Supplementary Figure S1). This analysis confirmed a statistical coincidence with true germ-line heterozygosity in the allelic ratio for two of four samples (51% in hair, 46% in saliva, 58% in blood, and 15% in FFPE colon), suggesting that it is crucial to distinguish biologically relevant allele imbalances from technical artifacts, especially in clinical genetic practice.
TA-cloning validation
To validate these findings with an independent test, we set out to confirm the presence of variants by Sanger sequencing of individual clones after TA cloning the amplicons. For the variants in families 1–4, the presence of reference and alternative alleles was confirmed for all four sites with at least two, and in most cases three or more, independent clones. The results on allele frequency from TA cloning were compatible with ADS for the four individuals (Table 2).
Discussion
In the current study, we surveyed 8 de novo HSCR families and found somatic mosaicism in 75% of cases, in either the patient or an asymptomatic parent. All of the six mosaic mutations we identified showed strong evidence of pathogenicity. Four are predicted to be null alleles, one has been reported previously, and one inserts an amino acid at a highly conserved site, is absent from the unaffected sibling, and has never been reported previously by the public National Heart, Lung, and Blood Institute or Exome Aggregation Consortium exome sequencing projects. Two mutations were tested and confirmed to be present as mosaic events with allelic ratios between 35 and 44% in the patients. Translating these allelic ratios into percentages of cells carrying the mutation predicted that the mutations must be present in 70–88% of the cells in multiple tissues (blood, saliva, and colon). In addition, 4/8 mutations were actually inherited as a consequence of low-level mosaicism in one of the parents. Mosaicism has been reported in several Mendelian disorders, including Marfan syndrome,23 Duchenne muscular dystrophy,24 hemophilia,25 and ornithine transcarbamylase deficiency.26 However, the high proportion of mosaicism identified in HSCR, as reported here, is unprecedented. Considering that we had high-coverage allele information on only 8/152 families and that Sanger sequencing reveals only a significant degree of mosaicism (over ~10%), we estimate that many more of the de novo families may carry somatic mosaicism for the disease allele that largely goes undetected using the standard diagnostic technique, now and previously.
The frequency of mosaicism depends on the particular disorder, tissue of origin, or selective pressure. Human gastrulation is thought to occur at ~day 16. The presence of a variant in both leukocyte, salivary (mesoderm), and colonic (endoderm) DNA (as in the probands in our families 1 and 2) would imply the appearance of this variant before embryonic day 16, early enough to be potentially also present in his or her germ cells and thus transmissible to the next generation. For autosomal-dominant diseases, the recurrence risk in parents of real de novo patients is considered to be very low, whereas germ-line disease carriers have a 50% probability of having affected offspring. According to Table 1, since asymptomatic mosaic carriers may have as much as 56% of cells carrying the disease allele, such carriers may have an up to 28% risk that their offspring will be affected. In contrast, the mosaic HSCR patient in family 2 may have as little as a 35% risk of having affected offspring (assuming the mosaicism is equally present in her germ line). In short, the phenotype of mosaic patients’ offspring can possibly be underestimated, whereas the recurrence risk in the symptomatic mosaic individuals’ offspring will be overestimated. Detailed analysis of somatic and germ-line mosaicism in de novo HSCR families, consequently, will be required to obtain more accurate figures for genetic counseling.
Evidently, sufficient sequencing coverage is required for reliably identifying mosaic mutations. Increased sequencing coverage steadily decreases the standard deviation of the allelic ratio, which effectively reduces technical variation and allows for better discrimination between true heterozygosity and mosaicism. Acuna-Hidalgo’s model suggests that at least 100-fold coverage is required for distinguishing mosaic mutations with allelic ratios <40% from germ-line mutations with 95% probability. On the other hand, the analysis for parental mosaicism for de novo mutations identified in a proband requires a different approach, and at least 140-fold coverage is needed for detecting low-level mosaicism of ≥5% with ≥95% probability.22 Although next-generation sequencing provides a new diagnostic tool for identifying genetic mosaicism, it can easily be missed as biased allele calls that are filtered out in the bioinformatics workflow. Moreover, the prevailing technologies often use pooled DNA isolated from multiple cells, so they produce data that reflect the overall average of the genomes of the constituent cells. Recent single-cell sequencing studies indeed reveal widespread mosaicism in apparently normal tissues.27 Thus, revision of current diagnostic laboratory protocols is needed to increase the detection rate of mosaic cases.28 In the study by Rahbari et al.,29 although only 1.3% of all DNMs recurred among siblings, this proportion increased to 24% for DNMs that were mosaic in >1% of parental blood cells and 50% for DNMs that were mosaic in >6% of parental blood cells. If this is also the case for HSCR, deep sequencing (with enough sensitivity) of parental blood for pathogenic DNMs seen in children would be highly recommended, and should enable meaningful stratification of families into a substantial majority with a <1% recurrence risk and a small minority with a recurrence risk that could be at least an order of magnitude higher.
Conclusion
Taking these findings together, we hereby propose the following. First, pathogenic mosaics of RET variants could be reliably identified by Sanger sequencing because of the necessary prevalence (of the mutant allele) needed to reach the threshold of pathogenicity, and likewise, would theoretically have an extremely low chance of being “incidentally” detected in healthy control persons (without affected offspring). The advantage of using targeted high-coverage sequencing lies in distinguishing post-zygotic mutations from germ-line mutations. Second, we defend the thesis that HSCR is a dosage-dependent complex disease. The genetic background, or in most cases RET gene function, may define the “boundary” between clinically affected and unaffected individuals, as we found in the current study. Finally, our research discovered mosaicism of an etiologic RET mutation as high as 28% in leukocyte- and saliva-derived DNA of a male patient’s healthy mother, indicating that, presumably, there is a threshold under which the mutation does not place a fatal burden on the biology of the enteric nervous system; and furthermore, at the level of the whole organism, tissue-to-tissue genetic variations may contribute to a mosaic phenotype that might not clearly follow the Mendelian rules of inheritance.
References
Amiel J, Sproat-Emison E, Garcia-Barcelo M et al, Hirschsprung disease, associated syndromes and genetics: a review. J Med Genet 2008; 45: 1–14.
Moore SWAdvances in understanding functional variations in the Hirschsprung disease spectrum (variant Hirschsprung disease). Pediatr Surg Int 2017; 33: 285–298.
Garver KL, Law JC, Garver BHirschsprung disease: a genetic studyClin Genet 1985; 28: 503–508.
Alves MM, Sribudiani Y, Brouwer RW et al, Contribution of rare and common variants determine complex diseases—Hirschsprung disease as a model. Dev Biol 2013; 382: 320–329.
McCallion AS, Chakravarti AEDNRB/EDN3 and Hirschsprung disease type II. Pigment Cell Res 2001; 14: 161–169.
Jiang Q, Arnold S, Heanue T et al, Functional loss of semaphorin 3C and/or semaphorin 3D and their epistatic interaction with RET are critical to Hirschsprung disease liability. Am J Hum Genet 2015; 96: 581–596.
Emison ES, Garcia-Barcelo M, Grice EA et al, Differential contributions of rare and common, coding and noncoding RET mutations to multifactorial Hirschsprung disease liability. Am J Hum Genet 2010; 87: 60–74.
Gui H, Schriemer D, Cheng WW et al, Whole exome sequencing coupled with unbiased functional analysis reveals new Hirschsprung disease genes. Genome Biol 2017; 18: 48.
Chatterjee S, Kapoor A, Akiyama JA et al, Enhancer variants synergistically drive dysfunction of a gene regulatory network in Hirschsprung disease. Cell 2016; 167: 355–368 e310.
Lei H, Li H, Xie H, Du C, Xia Y, Tang WRole of MiR-215 in Hirschsprung’s disease pathogenesis by targeting SIGLEC-8. Cell Physiol Biochem 2016; 40: 1646–1655.
Shen Z, Du C, Zang R et al, Microarray expression profiling of dysregulated long non-coding RNAs in Hirschsprung’s disease reveals their potential role in molecular diagnosis. Neurogastroenterol Motil 2016; 28: 266–273.
Schill EM, Lake JI, Tusheva OA et al, Ibuprofen slows migration and inhibits bowel colonization by enteric nervous system precursors in zebrafish, chick and mouse. Dev Biol 2016; 409: 473–488.
Lake JI, Tusheva OA, Graham BL, Heuckeroth ROHirschsprung-like disease is exacerbated by reduced de novo GMP synthesis. J Clin Invest 2013; 123: 4875–4887.
Twigg SR, Hufnagel RB, Miller KA et al, A recurrent mosaic mutation in SMO, encoding the hedgehog signal transducer smoothened, is the major cause of Curry-Jones syndrome. Am J Hum Genet 2016; 98: 1256–1265.
Couto JA, Huang AY, Konczyk DJ et al, Somatic MAP2K1 mutations are associated with extracranial arteriovenous malformation. Am J Hum Genet 2017; 100: 546–554.
Jansen AM, Crobach S, Geurts-Giele WR et al, Distinct patterns of somatic mosaicism in the APC gene in neoplasms from patients with unexplained adenomatous polyposis. Gastroenterology 2017; 152: 546–549 e543.
Jamuar SS, Lam AT, Kircher M et al, Somatic mutations in cerebral cortical malformations. N Engl J Med 2014; 371: 733–743.
Aminoso C, Garcia-Minaur S, Martinez L et al, Recurrence of Hirschsprung disease due to maternal mosaicism of a novel RET gene mutation. Clin Genet 2014; 85: 401–402.
Muller CM, Haase MG, Kemnitz I, Fitze GGenetic mosaicism of a frameshift mutation in the RET gene in a family with Hirschsprung disease. Gene 2014; 541: 51–54.
Moore SW, Zaahl MGTissue specific somatic mutations and aganglionosis in Hirschsprung’s disease. J Pediatr Surg 2014; 49: 258–261.
Munnes M, Fanaei S, Schmitz B, Muiznieks I, Holschneider AM, Doerfler WFamilial form of Hirschsprung disease: nucleotide sequence studies reveal point mutations in the RET proto-oncogene in two of six families but not in other candidate genes. Am J Med Genet 2000; 94: 19–27.
Acuna-Hidalgo R, Bo T, Kwint MP et al, Post-zygotic point mutations are an underrecognized source of de novo genomic variation. Am J Hum Genet 2015; 97: 67–74.
Rantamäki T, Kaitila I, Syvänen AC, Lukka M, Peltonen LRecurrence of Marfan syndrome as a result of parental germ-line mosaicism for an FBN1 mutation. Am J Hum Genet 1999; 64: 993–1001.
Taniguchi-Ikeda M, Takeshima Y, Lee T et al, Next-generation sequencing discloses a nonsense mutation in the dystrophin gene from long preserved dried umbilical cord and low-level somatic mosaicism in the proband mother. J Hum Genet 2016; 61: 351–355.
Leuer M, Oldenburg J, Lavergne JM et al, Somatic mosaicism in hemophilia A: a fairly common event. Am J Hum Genet 2001; 69: 75–87.
Komaki S, Matsuura T, Oyanagi K et al, Familial lethal inheritance of a mutated paternal gene in females causing X-linked ornithine transcarbamylase (OTC) deficiency. Am J Med Genet 1997; 69: 177–181.
Cai X, Evrony GD, Lehmann HS et al, Single-cell, genome-wide sequencing identifies clonal somatic copy-number variation in the human brain. Cell Rep 2014; 8: 1280–1289.
Gajecka MUnrevealed mosaicism in the next-generation sequencing era. Mol Genet Genomics 2016; 291: 513–530.
Rahbari R, Wuster A, Lindsay SJ et al, Timing, rates and spectra of human germline mutation. Nat Genet 2016; 48: 126–133.
Acknowledgments
This work was made possible by a grant from the Chinese Academy of Medical Sciences Initiative for Innovative Medicine (CAMS-I2M). This study was supported by the National Natural Science Foundation of China (81300266 and 81771620), the Beijing Natural Science Foundation (7142029), the Beijing Excellent Scientist Fund (2013D003034000007), the Beijing Nova Program (Z151100000315091 and Z171100001117125), and the Basic Foundation of the Capital Institute of Pediatrics (FX-2016-02) to Q.J. Q.L. was supported by the Beijing Natural Science Foundation (7154185). X.-L.C. was supported by the National Natural Science Foundation of China (81370708 and 31671310) and the Beijing Natural Science Foundation (7162029). L.L. was supported by grants from the Public Welfare Industry Research Special Foundation of China (201402007) and a Clinical Medicine Development Project of the Beijing Hospital Administration Bureau (ZYLX201306). F.Z. was supported by the National Natural Science Foundation of China (31625015). A.C. was supported by grants from the National Institutes of Health (HD28088). We are grateful to the affected individuals and their family members whose cooperation made this study possible. We thank Qixi Wu, Xiaoxu Yang, and Liping Wei from the National Institute of Biological Sciences for assistance with Bayesian-based mosaic validation, and acknowledge Ashish Kapoor (Johns Hopkins University School of Medicine) and Xue Zhang (Chinese Academy of Medical Sciences) for helpful discussions.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Jiang, Q., Liu, F., Miao, C. et al. RET somatic mutations are underrecognized in Hirschsprung disease. Genet Med 20, 770–777 (2018). https://doi.org/10.1038/gim.2017.178
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.178
This article is cited by
-
Hirschsprung’s disease: m6A methylase VIRMA suppresses cell migration and proliferation by regulating GSK3β
Pediatric Research (2024)
-
RET signaling in breast cancer therapeutic resistance and metastasis
Breast Cancer Research (2023)
-
RET compound inheritance in Chinese patients with Hirschsprung disease: lack of penetrance from insufficient gene dysfunction
Human Genetics (2021)
-
Sequence characterization of RET in 117 Chinese Hirschsprung disease families identifies a large burden of de novo and parental mosaic mutations
Orphanet Journal of Rare Diseases (2019)
-
Do RET somatic mutations play a role in Hirschsprung disease?
Genetics in Medicine (2018)