Abstract
Background
Prostate cancer (PrCa) is one of the most hereditable human cancers, however, only a small fraction of patients has been shown to carry deleterious variants in known cancer predisposition genes.
Methods
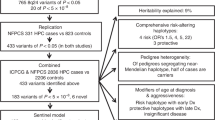
Whole-exome sequencing was performed in multiple affected members of 45 PrCa families to select the best candidate genes behind part of the PrCa missing hereditability. Recurrently mutated genes were prioritised, and further investigated by targeted next-generation sequencing in the whole early-onset and/or familial PrCa series of 462 patients.
Results
PRUNE2 stood out from our analysis when also considering the available data on its association with PrCa development. Ten germline pathogenic/likely pathogenic variants in the PRUNE2 gene were identified in 13 patients. The most frequent variant was found in three unrelated patients and identical-by-descent analysis revealed that the haplotype associated with the variant is shared by all the variant carriers, supporting the existence of a common ancestor.
Discussion
This is the first report of pathogenic/likely pathogenic germline variants in PRUNE2 in PrCa patients, namely in those with early-onset/familial disease. Importantly, PRUNE2 was the most frequently mutated gene in the whole series, with a deleterious germline variant identified in 2.8% of the patients, representing a novel prostate cancer predisposition gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data supporting the results reported in this paper can be found at: https://hive.biochemistry.gwu.edu/biomuta/proteinview/P38936 and https://portal.gdc.cancer.gov/.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J Clin. 2021. https://doi.org/10.3322/caac.21660
Brandão A, Paulo P, Teixeira M. Hereditary predisposition to prostate cancer: from genetics to clinical implications. Int J Mol Sci. 2020;21:1–23.
Bruner D, Moore D, Parlanti A, Dorgan JPE. Relative risk of prostate cancer for men with affected relatives: systematic review and meta-analysis. Int J Cancer. 2003;107:797–803.
Brandt A, Bermejo J, Sundquist J, Hemminki K. Age at diagnosis and age at death in familial prostate cancer. Oncologist. 2009;14:1209–17.
Verhage BAJ, Baffoe-Bonnie AB, Baglietto L, Smith DS, Bailey-Wilson JE, Beaty TH, et al. Autosomal dominant inheritance of prostate cancer: a confirmatory study. Urology. 2001;57:97–101.
Verhage BAJ, Aben KKH, Witjes JA, Straatman H, Schalken JA, Kiemeney LALM. Site-specific familial aggregation of prostate cancer. Int J Cancer. 2004;109:611–7.
Vietri MT, D’elia G, Caliendo G, Resse M, Casamassimi A, Passariello L, et al. Hereditary prostate cancer: genes related, target therapy and prevention. Int J Mol Sci. 2021;22; https://doi.org/10.3390/ijms22073753.
Raghallaigh HN, Eeles R. Genetic predisposition to prostate cancer: an update. Fam Cancer. 2021. https://doi.org/10.1007/S10689-021-00227-3
Ewing CM, Ray AM, Lange EM, Zuhlke KA, Robbins CM, Tembe WD, et al. Germline mutations in HOXB13 and prostate-cancer risk. N. Engl J Med. 2012;366:141–9.
Lin X, Qu L, Chen Z, Xu C, Ye D, Shao Q, et al. A novel germline mutation in HOXB13 is associated with prostate cancer risk in Chinese men. Prostate. 2013;73:169–75.
Maia S, Cardoso M, Pinto P, Pinheiro M, Santos C, Peixoto A, et al. Identification of two novel HOXB13 germline mutations in Portuguese prostate cancer patients. PLoS ONE. 2015;10:e0132728.
Grindedal EM, Møller P, Eeles R, Stormorken AT, Bowitz-Lothe IM, Landrø SM, et al. Germ-line mutations in mismatch repair genes associated with prostate cancer. Cancer Epidemiol Biomark Prev. 2009;18:2460–7.
Castro E, Eeles R. The role of BRCA1 and BRCA2 in prostate cancer. Asian J Androl. 2012;14:409–14.
Maia S, Cardoso M, Paulo P, Pinheiro M, Pinto P, Santos C, et al. The role of germline mutations in the BRCA1/2 and mismatch repair genes in men ascertained for early-onset and/or familial prostate cancer. Fam Cancer. 2016;15:111–21.
Paulo P, Maia S, Pinto C, Pinto P, Monteiro A, Peixoto A, et al. Targeted next generation sequencing identifies functionally deleterious germline mutations in novel genes in early-onset/familial prostate cancer. PLoS Genet. 2018;14; https://doi.org/10.1371/journal.pgen.1007355.
Pritchard CC, Mateo J, Walsh MF, de Sarkar N, Abida W, Beltran H, et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N. Engl J Med. 2016;375:443–53.
Southey MC, Goldgar DE, Winqvist R, Pylkäs K, Couch F, Tischkowitz M, et al. PALB2, CHEK2 and ATM rare variants and cancer risk: data from COGS. J Med Genet. 2016;53:800.
Leongamornlert D, Saunders E, Wakerell S, Whitmore I, Dadaev T, Cieza-Borrella C, et al. Germline DNA repair gene mutations in young-onset prostate cancer cases in the UK: evidence for a more extensive genetic panel. Eur Urol. 2019;76:329–37.
Peixoto A, Santos C, Pinheiro M, Pinto P, Soares MJ, Rocha P, et al. International distribution and age estimation of the Portuguese BRCA2 c.156_157insAlu founder mutation. Breast Cancer Res Treat. 2011;127:671–9.
Peixoto A, Santos C, Pinto P, Pinheiro M, Rocha P, Pinto C, et al. The role of targeted BRCA1/BRCA2 mutation analysis in hereditary breast/ovarian cancer families of Portuguese ancestry. Clin Genet. 2015;88:41–48.
Pinheiro M, Pinto C, Peixoto A, Veiga I, Mesquita B, Henrique R, et al. The MSH2 c.388_389del mutation shows a founder effect in Portuguese Lynch syndrome families. Clin Genet. 2013;84:244–50.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. 2013; http://github.com/lh3/bwa (Accessed 28 Jul 2021).
Garrison E, Marth G. Haplotype-based variant detection from short-read sequencing. 2012; https://arxiv.org/abs/1207.3907v2 (Accessed 28 Jul 2021).
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297.
DePristo MA, Banks E, Poplin RE, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Wei Z, Wang W, Hu P, Lyon GJ, Hakonarson H. SNVer: a statistical tool for variant calling in analysis of pooled or individual next-generation sequencing data. Nucleic Acids Res. 2011;39; https://doi.org/10.1093/NAR/GKR599.
Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009;25:2283–5.
Wang K, Li M, Hakonarson H ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38; https://doi.org/10.1093/NAR/GKQ603.
Li Q, Wang K. InterVar: clinical interpretation of genetic variants by the 2015 ACMG-AMP guidelines. Am J Hum Genet. 2017;100:267–80.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinforma. 2012;13:134.
Browning SR, Browning BL. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet. 2007;81:1084–97.
Browning BL, Browning SR. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics. 2013;194:459–71.
Bandelt HJ, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evolution. 1999;16:37–48.
Leigh JW, Bryant D. POPART: full-feature software for haplotype network construction. Methods Ecol Evolution. 2015;6:1110–6.
Brandão A, Paulo P, Maia S, Pinheiro M, Peixoto A, Cardoso M, et al. The CHEK2 variant C.349A>G is associated with prostate cancer risk and carriers share a common ancestor. Cancers. 2020;12:1–17.
Abramowicz A, Gos M. Splicing mutations in human genetic disorders: examples, detection, and confirmation. J Appl Genet. 2018;59:253–68.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361–2.
Desmet FO, Hamroun D, Lalande M, Collod-Bëroud G, Claustres M, Béroud C. Human splicing finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37; https://doi.org/10.1093/nar/gkp215.
Yeo G, Burge CB. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol. 2004;11:377–94.
Salameh A, Lee AK, Cardó-Vila M, Nunes DN, Efstathiou E, Staquicini FI, et al. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc Natl Acad Sci USA. 2015;112:1–6.
Machida T, Fujita T, Ooo M, Ohira M, Isogai E, Mihara M, et al. Increased expression of proapoptotic BMCC1, a novel gene with the BNIP2 and Cdc42GAP homology (BCH) domain, is associated with favorable prognosis in human neuroblastomas. Oncogene. 2006;25:1931–42.
Clarke RA, Zhao Z, Guo A-Y, Roper K, Teng L, Fang Z-M, et al. New genomic structure for prostate cancer specific gene PCA3 within BMCC1: implications for prostate cancer detection and progression. PLoS ONE 2009;4:e4995.
Yu W, McPherson JR, Stevenson M, van Eijk R, Heng HL, Newey P, et al. Whole-exome sequencing studies of parathyroid carcinomas reveal novel PRUNE2 mutations, distinctive mutational spectra related to APOBEC-catalyzed DNA mutagenesis and mutational enrichment in kinases associated with cell migration and invasion. J Clin Endocrinol Metab. 2015;100:E360–E364.
Harms PW, Vats P, Verhaegen ME, Robinson DR, Wu YM, Dhanasekaran SM, et al. The distinctive mutational spectra of polyomavirus-negative Merkel cell carcinoma. Cancer Res. 2015;75:3720–7.
Alsadoun N, MacGrogan G, Truntzer C, Lacroix-Triki M, Bedgedjian I, Koeb MH, et al. Solid papillary carcinoma with reverse polarity of the breast harbors specific morphologic, immunohistochemical and molecular profile in comparison with other benign or malignant papillary lesions of the breast: a comparative study of 9 additional cases. Mod Pathol. 2018;31:1367–80.
Pinto P, Peixoto A, Santos C, Rocha P, Pinto C, Pinheiro M, et al. Analysis of founder mutations in rare tumors associated with hereditary breast/ovarian cancer reveals a novel association of BRCA2 mutations with ampulla of vater carcinomas. PLoS ONE. 2016;11:e0161438.
Laitinen VH, Wahlfors T, Saaristo L, Rantapero T, Pelttari LM, Kilpivaara O, et al. HOXB13 G84E mutation in Finland: population-based analysis of prostate, breast, and colorectal cancer risk. Cancer Epidemiol Biomark Prev. 2013;22:452–60.
Varon R, Seemanova E, Chrzanowska K, Hnateyko O, Piekutowska-Abramczuk D, Krajewska-Walasek M, et al. Clinical ascertainment of Nijmegen breakage syndrome (NBS) and prevalence of the major mutation, 657del5, in three Slav populations. Eur J Hum Genet. 2000;8:900–2.
Bancroft EK, Page EC, Castro E, Lilja H, Vickers A, Sjoberg D, et al. Targeted prostate cancer screening in BRCA1 and BRCA2 mutation carriers: results from the initial screening round of the IMPACT study. Eur Urol. 2014;66:489–99.
Castro E, Goh C, Olmos D, Saunders E, Leongamornlert D, Tymrakiewicz M, et al. Germline BRCA mutations are associated with higher risk of nodal involvement, distant metastasis, and poor survival outcomes in prostate cancer. J Clin Oncol. 2013;31:1748–57.
Mateo J, Carreira S, Sandhu S, Miranda S, Mossop H, Perez-Lopez R, et al. DNA-repair defects and olaparib in metastatic prostate cancer. N Engl J Med. 2015;373:1697–708.
Dingerdissen HM, Torcivia-Rodriguez J, Hu Y, Chang TC, Mazumder R, Kahsay R. BioMuta and BioXpress: mutation and expression knowledgebases for cancer biomarker discovery. Nucleic Acids Res. 2018;46:D1128–36.
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 2021;49:D412–9.
Funding
This work was supported by IPO-Porto Research Center (CI-IPOP-16-2012) and by Fundação para a Ciência e a Tecnologia (FCT; PTDC/DTP-PIC/1308/2014—POCI-01-0145-FEDER-016889). The following authors were funded by FCT with scholarships or research contracts: MC (SFRH/BD/116557/2016), SM (SFRH/BD/71397/2010), AB (PTDC/DTP-PIC/1308/2014—POCI-01-0145-FEDER-016889; POCI-01-FEDER-028245; UIDP/DTP/00776/2020; 2021.03835.CEECIND), and PP (PEst-OE/SAU/UI0776/2014; UID/DTP/00776/2013/POCI-01-0145-FEDER-006868; CEECINST/00091/2018). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Luis Carvajal-Carmona receives funding from The Auburn Community Cancer Endowed Chair on Basic Science and from the National Cancer Institute of the National Institutes of Health (grant P30CA093373). The opinions expressed in this article are the author’s own and do not reflect the view of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conception and design: MT and LGC-C. Data acquisition: MC, SM, PP, AB, RS and NB. Analysis and interpretation of the data: MC, SM, PP, PL and AB. Drafting of the manuscript: MC. Critical revision of the manuscript: MT, PP, LGC-C and AB. Statistical analysis: AB and PL. Funding obtention: MT and LGC-C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Institutional Ethics Committee of the Portuguese Oncology Institute-Porto (approval number: 38.010) and performed in accordance with the Declaration of Helsinki. Written consent was obtained from all participants.
Consent to publish
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cardoso, M., Maia, S., Brandão, A. et al. Exome sequencing of affected duos and trios uncovers PRUNE2 as a novel prostate cancer predisposition gene. Br J Cancer 128, 1077–1085 (2023). https://doi.org/10.1038/s41416-022-02125-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-02125-6