Abstract
Purpose
Recognizing individuals with inherited diseases can be difficult because signs and symptoms often overlap those of common medical conditions. Focusing on inborn errors of metabolism (IEMs), we present a method that brings the knowledge of highly specialized experts to professionals involved in early diagnoses. We introduce IEMbase, an online expert-curated IEM knowledge base combined with a prototype diagnosis support (mini-expert) system.
Methods
Disease-characterizing profiles of specific biochemical markers and clinical symptoms were extracted from an expert-compiled IEM database. A mini-expert system algorithm was developed using cosine similarity and semantic similarity. The system was evaluated using 190 retrospective cases with established diagnoses, collected from 15 different metabolic centers.
Results
IEMbase provides 530 well-defined IEM profiles and matches a user-provided phenotypic profile to a list of candidate diagnoses/genes. The mini-expert system matched 62% of the retrospective cases to the exact diagnosis and 86% of the cases to a correct diagnosis within the top five candidates. The use of biochemical features in IEM annotations resulted in 41% more exact phenotype matches than clinical features alone.
Conclusion
IEMbase offers a central IEM knowledge repository for many genetic diagnostic centers and clinical communities seeking support in the diagnosis of IEMs.
Similar content being viewed by others
Introduction
Identification of rare genetic disorders has been greatly improved by the advent of genome-wide sequencing. The new technology has expanded our knowledge of rare disease genetics and enhanced our ability to diagnose new patients.1, 2 However, the diagnosis of rare genetic disorders remains a challenge. Misdiagnoses and delayed diagnoses are often3 due to nonspecificity and heterogeneity of signs and symptoms, rarity of conditions, and also limited access to the knowledge of highly specialized experts.2, 4, 5 Inborn errors of metabolism (IEMs) exemplify these challenges: early signs and symptoms are nonspecific6 and insufficiently recognized.7 For example, in a survey of 34 junior pediatric doctors regarding their confidence and knowledge in the acute management of three IEMs—glutaric aciduria type I (MIM 231670), medium-chain acyl Co-A dehydrogenase deficiency (MIM 201450), and maple syrup urine disease (MIM 248600)—only five respondents were able to identify the correct treatment steps for the former two, while only two respondents identified the correct steps for the latter.7 Moreover, more than 22 respondents indicated a low level of confidence in their knowledge.7
The knowledge gap between IEM specialists and other clinicians involved in IEM diagnoses is concerning, given the amenability to targeted treatments for an increasing number of IEMs; a delayed diagnosis can lead to irreversible organ damage or even death. Moreover, this disparity is widening with the explosive amount of knowledge generated by multi-omics technology.2 Such a divide stands in contrast to the historic efforts by the IEM clinical and research community toward early recognition through the creation and use of diagnostic tests, such as population newborn screening. Thus, a potential solution may be found in the rich disease knowledge base established by the IEM community, dating back to Archibald Garrod’s study on alkaptonuria in 1902.8 This compiled knowledge base has, however, lagged behind other fields in the transition to digital form, as much of the work occurred before modern data systems came into existence and therefore the information was stuck on paper. Aspects have been incorporated into large-scale rare-disease databases.9, 10 However, these databases aim to provide an overview of many kinds of individual disorders, and are not designed to guide clinicians in the diagnostic process. Therefore, digital translation and standardization of the IEM community knowledge base are urgently needed to bridge the knowledge gap.
Thus, we created IEMbase, an online application that combines the IEM community knowledge base with a prototype mini-expert system. The expert-compiled knowledge base provides clinical, biochemical, and genetic profiles of 530 known IEMs. The mini-expert system accepts a list of biochemical and clinical phenotypes from users, compares the input phenotypic profile against IEMs in the knowledge base using cosine similarity and semantic similarity, and returns a list of matching IEM diagnoses. With the resulting list, users can generate differential diagnosis charts, biochemical test panels, and targeted gene panels in order to pursue concurrent biochemical and genetic/genomic investigations for a rapid diagnosis. IEMbase aims to renew the existing IEM community knowledge base for the modern age, creating a global resource to facilitate the collection and dissemination of high-quality clinical knowledge for advanced recognition of IEMs.
Materials and methods
Knowledge base compilation
IEMbase was compiled by extracting 530 disease-characterizing profiles from a nascent disease database, which was previously compiled by more than 100 IEM experts to produce a textbook guide on IEM classification.11 Table 1 shows an example of an extracted IEM profile. Each IEM profile consisted of known disorder names, disorder abbreviations, causal gene information, a MIM number, and a list of associated biochemical markers and clinical symptoms. Additionally, the list of biomarkers/symptoms was annotated with information regarding onset, severity/pathological level, and whether the biomarker/symptom is characteristic of the associated IEM. The onset information was organized in five categories (neonatal: birth to 1 month, infant: 1–18 months, childhood: 1.5–11 years, adolescence: 11–16 years, and adulthood: >16 years). The pathological levels of biochemical markers were denoted by up/down/no arrows and the severities of clinical symptoms were denoted by plus/minus signs. The presence or absence of phenotypic characteristics was indicated by yes/no.
The extracted profiles were manually reviewed for consistency and then were imported into IEMbase as three PostgreSQL tables, each representing the type of annotation used in the profiles: disorders, biochemical/clinical phenotypes, or disorder-phenotype associations (Supplementary Figure S10 online). In total, the tables contained 530 disorders, 2,323 biochemical/clinical phenotypes, and 8,465 disorder-phenotype associations.
Additional annotations were created within each IEM profile. One was the amenability of individual IEMs to treatment, which was manually annotated based on previous literature12, 13 and denoted by yes/no/unknown categories. Another was the prevalence of IEMs as reported in literature or clinical resources.9, 10, 11, 13 The last was a list of links to relevant entries in external databases, such as UniProt,14 NCBI Gene,15 GeneCards,16 Kyoto Encyclopedia of Genes and Genomes,17 National Institutes of Health Genetic Testing Registry,18 and GeneReviews.19 The links were created for interoperability with existing systems and were created using a BioMart ID conversion tool,20 as well as URL rules specified on the resource websites.18, 19
The compiled knowledge base was assigned a version number of 1.0.0. This initial version was used for both the methods and the results described herein. Since the initial compilation, IEMbase has been regularly updated with new information. Thus, the version number has been incremented to indicate such updates.
Mapping to structured vocabulary
A known strategy for matching user-provided phenotypic profiles to diseases is to exploit semantic relationships between phenotypic features, which are defined by a structured vocabulary.21 The phenotype vocabulary in IEMbase was not structured, but a structure could be imposed based on a compatible external vocabulary. Therefore, the following four standard medical vocabularies were assessed for their compatibility with IEMbase: Human Phenotype Ontology (HPO),22 Medical Subject Headings (MeSH),23 Systematized Nomenclature of Medicine–Clinical Terms (SNOMED CT),24 and International Classification of Diseases, 10th revision (ICD10).25
During compatibility assessment, HPO OBO file (2016-04-01 release), MeSH ASCII file (2016 version), SNOMED CT RF2 files (2016 versions), and ICD10 XML file (2016 version) were used. The assessment proceeded in three steps. First, unique IDs and medical terms were extracted from IEMbase (version 1.0.0) and the four vocabularies. For SNOMED CT, extraction was restricted to only the terms categorized under “Clinical finding” and “Substance,” to minimize false mapping. The OntoCAT R package26 was used to parse HPO terms. The standard string library and Nokogiri gem in Ruby programming language were used to parse all others. Second, all extracted terms were normalized using the Norm program included in the SPECIALIST Lexical Tools.27 String normalization removed the differences in alphabetic case, singular or plural variants, punctuations, stop words, and word order. Finally, all IEMbase terms were compared against all terms in each vocabulary. Only the exact matches were recorded as compatible mappings.
The initial compatibility assessment revealed that no single vocabulary could completely cover both the biochemical and the clinical phenotypes in IEMbase (Table 2). It also revealed that the most compatible vocabulary was different for biochemical (SNOMED CT) and clinical phenotypes (HPO) (Table 2). Therefore, the assessment was adjusted to consider the two phenotype categories separately. Once adjusted, two additional biochemical vocabularies were added: Chemical Entities of Biological Interest (ChEBI; OBO file; 2016-04-01 release)28 and Logical Observation Identifiers Names and Codes (LOINC; CSV file; version 2.56).29
Based on the adjusted assessment (Table 2), clinical phenotypes were mapped to the most compatible vocabulary, HPO. A medical expert manually reviewed exact matches identified during the compatibility assessment and manually mapped unmatched clinical phenotypes to HPO terms. In total, 1,193 of 1,200 clinical phenotypes were mapped to HPO. The mapped HPO terms and their ancestor/descendant HPO terms were extracted using the OntoCAT R package and were then written into IEMbase as PostgreSQL tables. For biochemical phenotypes, we allowed matches to terms in any of four vocabularies: SNOMED CT, MeSH, LOINC, and ChEBI. However, manual review of unmatched phenotypes revealed that these terms were highly specialized and thus not present in the vocabularies. Therefore, we implemented an alternative strategy for assessing user-supplied biochemical phenotypes and abandoned the established biochemical vocabularies.
Phenotype-matching algorithm for mini-expert system
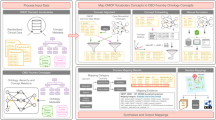
The mini-expert system of IEMbase accepts a list of biochemical and clinical phenotypes as input. The system then employs a two-step algorithm that compares the input phenotypic profile against every IEM profile in IEMbase (Figure 1).
Mini-expert algorithm flowchart. Users enter a list of biochemical/clinical phenotypes into IEMbase’s mini-expert system. The system’s phenotype-matching algorithm first divides the input list into biochemical and clinical categories. The algorithm then ranks the disorders in IEMbase by comparing the biochemical profile of each disorder against the input biochemical profile, using cosine similarity. Subsequently, the algorithm breaks ties in the ranked list by comparing the clinical profiles, using semantic similarity.
First, the algorithm ranks IEMs by assessing only biochemical phenotypes, using cosine similarity. Cosine similarity30 is defined as the cosine of two vectors, TFIDFQ and TFIDFD, which represent the input profile Q and an IEM profile D from IEMbase. The vectors consist of term frequency–inverse document frequency (tfidf) scores30 defined as follows:

tf(d, D) represents the occurrence of biochemical phenotype d in D, expressed as 0 or 1. idf(d, D) represents the specificity of d to D, defined as:

Using the above definitions, the algorithm computes tfidf scores for all d in D and all biochemical phenotypes q in Q. Individual tfidf scores are subsequently multiplied by a score for matching the pathological level (i.e., elevated/normal/decreased), which is −1 if the levels of d and q do not match or 1 if they match. The algorithm then computes the cosine of vector TFIDFQ and vector TFIDFD:

The cosine similarity scores are further multiplied by decay factors defined based on severity/characteristics (sc) scores for disorder D:

λ is a decay constant defined between 0.0 and 1.0. dist is a Euclidean distance between a vector of sc scores for disorder D and a vector of maximum possible sc scores. The vector of sc scores for D consists of sc scores for individual phenotypes d in D that match an input phenotype q in Q. The sc score for individual d is defined as follows:

s(d, D)is the severity score of d ranging from 1 to 3, based on the severity annotation of d. c(d, D) is the characteristic score of d assigned either 1 or 2, based on whether d is characteristic of D.
After the initial ranking of IEMs by biochemical phenotypes, the algorithm breaks ties in this ranking by assessing clinical phenotypes using semantic similarity that is computed based on the work of Kohler et al.21 The similarity between two clinical phenotypes, p and p′, is computed as the information content (IC) of their most informative common ancestor (MICAp,p′) in the HPO. IC is a measure of concreteness of a phenotype p in the HPO. It is defined as:

The similarity between input profile Q and an IEM profile D is computed by averaging the best match scores for clinical phenotypes q in Q:

nq is the number of q in Q. The best match score for each q is defined as EquationSource math mrow miI miC mo( miM miI miC miA miq mo, msub mid mrow mib mie mis mit mo) , where dbest is a clinical phenotype in D whose common ancestor with q has the highest IC and the highest severity score. The similarity score is then multiplied by a decay factor as in biochemical similarity.
Software framework details
IEMbase data is stored in a PostgreSQL database. The front-end user interface was developed using an Angular.js framework. The back-end system was developed in a Ruby on Rails framework.
Code availability
IEMbase is freely available online (http://iembase.org/app) and upon request through an application programming interface. Computer code used for performance evaluation is available upon request.
Mini-expert system case study
To demonstrate a potential use case scenario of the mini-expert system, we used a case of a delayed diagnosis of hyperornithinemia–hyperammonemia–homocitrullinuria syndrome. Case details are described in the Results section.
Performance evaluation of mini-expert system
To evaluate the performance of IEMbase’s mini-expert system, 190 retrospective cases were collected from 15 different metabolic centers. For each case, the contributors provided the final diagnosis and biochemical/clinical information. These cases were collected using an online form, which restricted the contributors to providing the case information using only the disorder and phenotype vocabularies in IEMbase.
Each evaluation case was matched to potential diagnoses using the mini-expert system. The system’s performance was compared against three phenotype-matching algorithms, each of which uses cosine similarity, with or without semantic similarity, and also with or without severity and characteristic scores.
In addition, the system performance was compared using only biochemical phenotypes, and only clinical phenotypes of retrospective cases. For each retrospective case, the phenotypes were separated into biochemical and clinical categories before each category was evaluated with the mini-expert system. Eighteen cases with phenotypes only in either category, were excluded from this paired comparison (n = 172).
We also tested whether the number of phenotypes specified for each case correlated with the rank of correct diagnoses, in order to assess if some cases ranked better than others because more phenotypes were provided for them.
The above evaluations were conducted using version 1.0.0 of IEMbase. Difference in performance was statistically tested using the Mann-Whitney-U test implemented by wilcox.test in R (version 3.3.1). The correlation test was performed using Spearman’s rank correlation test, implemented by cor.test in R. All plots were generated using the ggplot2 R package.
Results
Overview and walkthrough of IEMbase
We developed IEMbase as an online application which combines a comprehensive IEM knowledge base with a diagnosis support (mini-expert) system. IEMbase curates expert-provided information on 530 IEMs, their treatability and genetics, as well as associated biochemical/clinical phenotypes with detailed annotations on the onset/severity/pathological level of the phenotypes. The application is freely available and can be accessed at http://www.iembase.org/app, or from a link on the project overview website (http://www.iembase.org). IEMbase is also available through an application-programming interface for integration into other computational systems. Application-programming interface access is available upon request.
A detailed walkthrough of the application is presented in the Supplementary Materials and Methods.
Applying mini-expert system in clinical settings
We demonstrate the utility of IEMbase’s mini-expert system using a case of a delayed hyperornithinemia–hyperammonemia–homocitrullinuria (HHH) syndrome diagnosis. A girl 2 years and 8 months of age had shown inconspicuous psychomotor development. Following an upper respiratory tract infection, she developed recurrent vomiting, while refusing feeding but drinking occasionally. She was slightly lethargic. Over the following weeks she never fully recovered and continued to undergo episodes of postprandial vomiting, lethargy, and apparent seizures reminiscent of absences. Laboratory tests revealed hyperammonemia (260 μmol/L) together with the constellation of acute liver failure (ASAT 130 U/l, ALAT 233 U/l, ALP 267 U/l, Quick 10%, INR 4.87, aPTT 52 sec.). Plasma amino acids demonstrated high to normal glutamine, elevated ornithine, and low citrulline and arginine, all as abnormalities. Orotic acid was highly elevated in urine. Homocitrulline was specifically tested for but could not be identified in plasma or urine. With a presumptive diagnosis of ornithine transcarbamylase deficiency, the patient was referred to a metabolic center and treated, accordingly, with protein restriction and ammonia scavengers. Over the following months, there were several similar episodes, usually triggered by minor intercurrent infections. Molecular analysis of ornithine transcarbamylase was negative.
When the constellation of symptoms was entered into the IEMbase’s mini-expert system (Supplementary Table S1), hyperornithinemia–hyperammonemia–homocitrullinuria syndrome was suggested as the most likely disease candidate, while ornithine transcarbamylase deficiency was listed as the second probable disease candidate. Indeed, molecular analysis of SLC25A15 identified biallelic variants in the gene, confirming the diagnosis of hyperornithinemia–hyperammonemia–homocitrullinuria syndrome and enabling targeted treatment.
Mini-expert system performance evaluation
IEMbase’s mini-expert system matched 62% of cases to exact diagnoses, 86% of cases within the top five candidate disorders, and 90% of cases within the top ten. The performance comparison between the mini-expert system algorithm (combined + weighted) and three other phenotype-matching algorithms (combined + unweighted, cosine + weighted, cosine + unweighted) is shown in Table 3 and Supplementary Figure S11. There was no significant difference in performance between the mini-expert algorithm and the alternative phenotype-matching algorithms. Cases that were ranked out of the top 20 tended to have entries of unspecific biochemical markers, such as “Acylcarnitines, all” or “Amino acids, all.” Refer to Supplementary Table S2 for an overview of the cases and their ranks. Refer to Supplementary Table S3 for more information about the cases that were ranked out of the top 20.
The system performance using only biochemical queries was significantly better than using only clinical queries (P < 2.2e-16; Figure 2 and Supplementary Table S4). Using only biochemical phenotypes, 60% of cases were matched to exact diagnoses, 83% of cases within the top five candidate disorders, and 89% of cases within the top ten. The success rate of biochemical phenotypes plateaued after 90%, as the number of assessed candidates increased, reflecting 13 cases which failed to produce candidates owing to insufficient/unspecific biochemical information and/or the system’s inability to recognize similar biochemical phenotypes. As an example of the latter, the current implementation fails to recognize “Acylcarnitines, all” and “Long-chain acylcarnitine” as related phenotypes. Using only clinical phenotypes, only 19% of cases were matched to exact diagnoses, 38% of cases within the top five candidate disorders, and 49% of cases within the top ten.
Mini-expert system performance using only biochemical/clinical information. The system performance when using only biochemical phenotypes was compared with that when using only clinical phenotypes of 172 retrospective cases. Percentage success N measures % of cases whose actual diagnoses ranked within the top N ranks. The system performance when using only biochemical phenotypes was significantly better than that when using only clinical phenotypes (P < 2.2e-16; Mann-Whitney-U).
There was no significant correlation between the rank of correct diagnoses and the number of provided phenotypes (P = 0.69; Supplementary Figure S12).
Discussion
Although disease databases for IEMs have been developed in the past,31, 32, 33 they were either based on individual case reports31, 32 or more focused on specific symptoms, such as intellectual disability.33 Large-scale rare disease databases currently available for a general clinical audience9, 10 do cover a wide range of rare diseases, but by their nature do not provide the depth of information found in specialized expert knowledge bases. IEMbase is designed to fill this gap, by combining a central knowledge repository with a basic diagnostic support system. This design allows simultaneous collection of the current expert knowledge and its dissemination to the broader clinical community. In addition, it leads to further improvement of the mini-expert system as the depth of knowledge is compiled. Curated knowledge bases are intended to surpass the capacity of any single expert. IEMbase is therefore of utility for all those involved in IEM diagnoses: pediatricians, internists, neurologists, geneticists, and metabolic specialists. As our case study demonstrates, the utility of IEMbase can also be extended to established metabolic centers and biochemical genetics laboratories to help broaden the array of potential differential diagnoses—specifically to include lesser-known diseases when their constellations overlap with typical presentations of better-known diseases.
The evaluation of the mini-expert system revealed that phenotype-matching performance is significantly higher with the use of biochemical phenotypes than that of clinical phenotypes. This probably reflects two influences: (i) many clinical features of IEMs are not specific, while biochemical alterations are frequently so6, 34 and (ii) the IEM community has made intense efforts toward both disease-specific biomarker discovery and the annotation of biochemical phenotypes.11, 12, 34, 35 The second point draws upon a hundred years of IEM community efforts, leading to a depth and breadth of biochemical annotations that constitute a phenome space well suited to research of assisted diagnostic methods. Furthermore, the uniting of biochemical annotations with genetic and clinical annotations aligns with the imminent shift in investigative paradigm, where multi-omics technology allows holistic investigation into an individual’s genome, epigenome, transcriptome, proteome, metabolome, and phenome.34 Extrapolating from our experience, the knowledge bases of other clinical communities may hold untapped high-quality offline information which could be renewed in a similar way to that held in IEMbase.
Owing to a lack of compatible structured vocabulary for biochemical phenotypes in IEMbase, the current mini-expert system algorithm uses a nonsemantic information retrieval metric (tfidf-cosine similarity) to compare biochemical phenotypes. We recognize that this approach is not robust when matching imprecise terms. For example, the use of tfidf-cosine similarity will not take into account the fact that neopterin and biopterin belong to the same group of pterins. The use of structured vocabulary and semantic similarity can mitigate this shortcoming. Therefore, we plan to contribute our biochemical vocabulary to existing ontologies as we make updates to our system.
Biochemical test/gene panel suggestions that are provided with the output of the mini-expert system are currently restricted to basic information (e.g., gene names or chemical test panels), as detailed specification will require future contributions from the expert community. We anticipate that such improvements will be introduced over time as a result of community outreach efforts such as those described below.
For the long-term viability of IEMbase, continuous contribution from the expert community is crucial, especially with the large number of novel IEMs and phenotypes now being revealed with the use of multi-omics technologies. Therefore, we will periodically reach out to the IEM community for knowledge contribution, in addition to assembling an expert panel, which will regularly review and update the knowledge base. To encourage adoption among the new generation of clinicians, we plan to develop a mobile version of the application and a training module.
In summary, IEMbase is a web application intended to provide the clinical community with a comprehensive IEM knowledge base and a tool to facilitate early and accurate diagnoses of IEMs. Its knowledge base features expert-curated clinical resources on 530 IEMs. Its mini-expert system empowers clinicians and complements their workflow with suggested diagnoses, differential diagnosis charts, biochemical test panels, and gene panels. The multitude of suggestions enables clinicians to initiate concurrent biochemical and genetic evaluations, where the former can help focus the latter for rapid diagnosis, especially in clinical exome/genome interpretations. We believe that the power of IEMbase comes from the community of experts who contribute their knowledge for the greater benefit of the broader clinical community and as such, the value of community science should be recognized as a key component of digital medicine in the 21st century.
References
Tarailo-Graovac M, Shyr C, Ross CJ et al. Exome sequencing and the management of neurometabolic disorders. N Engl J Med 2016;374:2246–2255.
Chong JX, Buckingham KJ, Jhangiani SN et al. The genetic basis of Mendelian phenotypes: discoveries, challenges, and opportunities. Am J Hum Genet 2015;97:199–215.
Survey of the delay in diagnosis for 8 rare diseases in Europe (“EurordisCare2”). Eurodis - Rare Diseases Europe. http://www.eurordis.org/IMG/pdf/Fact_Sheet_Eurordiscare2.pdf. Accessed 1 November 2016.
Shashi V, McConkie-Rosell A, Rosell B et al. The utility of the traditional medical genetics diagnostic evaluation in the context of next-generation sequencing for undiagnosed genetic disorders. Genet Med 2014;16:176–182.
Rare diseases: understanding this public health priority. Eurodis—Rare Diseases Europe. http://www.eurordis.org/sites/default/files/publications/princeps_document-EN.pdf. Accessed 1 November 2016.
Leonard JV, Morris AAM . Diagnosis and early management of inborn errors of metabolism presenting around the time of birth. Acta Paediatr 2006;95:6–14.
Hawkes CP, Walsh A, O’Sullivan S, Crushell E . Doctors’ knowledge of the acute management of Inborn Errors of Metabolism. Acta Paediatr 2011;100:461–463.
Garrod A . The incidence of alkaptonuria: a study in chemical individuality. The Lancet 1902;160:1616–1620.
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA . Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 2005;33:D514–517.
Rath A, Olry A, Dhombres F, Brandt MM, Urbero B, Ayme S . Representation of rare diseases in health information systems: the Orphanet approach to serve a wide range of end users. Hum Mutat 2012;33:803–808.
Blau N, Duran M, Gibson KM, Dionisi-Vici C (eds). Physician’s guide to the diagnosis, treatment, and follow-up of inherited metabolic diseases. Heidelberg, Germany: Springer, 2014.
van Karnebeek CDM, Shevell M, Zschocke J, Moeschler JB, Stockler S . The metabolic evaluation of the child with an intellectual developmental disorder: diagnostic algorithm for identification of treatable causes and new digital resource. Mol Genet Metab 2014;111:428–438.
Zschocke J, Hoffmann GF . Vademecum metabolicum: diagnosis and treatment of inborn errors of metabolism. Friedrichsdorf, Germany: Schattauer, 2011.
The UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res 2015;43:D204–D212.
Brown GR, Hem V, Katz KS et al. Gene: a gene-centered information resource at NCBI. Nucleic Acids Res 2015;43:D36–D42.
Stelzer G, Rosen N, Plaschkes I et al. The GeneCards suite: From gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 2016;54:1.30.1–1.30.33.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M . KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 2016;44:D457–462.
Rubinstein WS, Maglott DR, Lee JM et al. The NIH genetic testing registry: a new, centralized database of genetic tests to enable access to comprehensive information and improve transparency. Nucleic Acids Res 2013;41:D925–D935.
Pagon RA, Adam MP, Ardinger HH et al, (eds) GeneReviews. Seattle: University of Washington, Seattle, 1993–2017. https://www.ncbi.nlm.nih.gov/books/NBK1116/. Accessed February 27, 2017.
Smedley D, Haider S, Durinck S et al. The BioMart community portal: an innovative alternative to large, centralized data repositories. Nucleic Acids Res 2015;43:W589–W598.
Köhler S, Schulz MH, Krawitz P et al. Clinical diagnostics in human genetics with semantic similarity searches in ontologies. Am J Hum Genet 2009;85:457–464.
Köhler S, Doelken SC, Mungall CJ et al. The Human Phenotype Ontology project: linking molecular biology and disease through phenotype data. Nucleic Acids Res 2014;42:D966–974.
Rogers FB . Medical subject headings. Bull Med Libr Assoc 1963;51:114–116.
SNOMED-CT (Systematized Nomenclature of Medicine—Clinical Terms). Copenhagen: International Health Terminology Standards Development Organisation, 2016. http://www.ihtsdo.org/snomed-ct. Accessed October 18, 2016.
ICD-10 (International Classification of Diseases–10). World Health Organization, 2016. http://www.who.int/classifications/icd/en/. Accessed October 18, 2016.
Kurbatova N, Adamusiak T, Kurnosov P, Swertz MA, Kapushesky M . ontoCAT: an R package for ontology traversal and search. Bioinformatics 2011;27:2468–2470.
SPECIALIST Lexical Tools [computer program] Release 2015. Bethesda, MD: Lexical Systems Group, 2015.
Hastings J, de Matos P, Dekker A et al. The ChEBI reference database and ontology for biologically relevant chemistry: enhancements for 2013. Nucleic Acids Res 2013;41:D456–463.
McDonald CJ, Huff SM, Suico JG et al. LOINC, a universal standard for identifying laboratory observations: a 5-year update. Clin Chem 2003;49:624–633.
Salton G, Wong A, Yang CS . A vector space model for automatic indexing. Commun ACM 1975;18:613–620.
Frauendienst-Egger G, Trefz FK . www.metagene.de: online knowledge base for inborn errors of metabolism. J Inherit Metab Dis 2006;29(suppl 1):84.
Töpel T, Scheible D, Trefz F, Hofestädt R . RAMEDIS: a comprehensive information system for variations and corresponding phenotypes of rare metabolic diseases. Hum Mutat 2010;31:E1081–1088.
van Karnebeek CDM, Houben RFA, Lafek M, Giannasi W, Stockler S . The treatable intellectual disability APP www.treatable-id.org: a digital tool to enhance diagnosis & care for rare diseases. Orphanet J Rare Dis 2012;7:47.
Tebani A, Afonso C, Marret S, Bekri S . Omics-based strategies in precision medicine: toward a paradigm shift in inborn errors of metabolism investigations. Int J Mol Sci 2016;17:E1555.
Sahoo S, Franzson L, Jonsson JJ, Thiele I . A compendium of inborn errors of metabolism mapped onto the human metabolic network. Mol Biosyst 2012;8:2545–2558.
Acknowledgements
We sincerely thank all validators who contributed retrospective cases (Carlos Ferreira, USA; Saumya Jamuar, Singapore; Stefan Kölker, Germany; Ylmaz Yildiz, Turkey; Luis Umana, USA; Saikat Santra, UK; Consuelo Duran, Argentina; Bryce Medelsohn, USA; Somesh Kumar, India; G.F.H., Germany, N.B., Germany; Himanshu Goel, Australia; Dorothea Haas, Germany; Susan Brooks, USA; Burcu Öztürk-Hismi, Turkey; C.D.M.v.K., Amsterdam & Canada; Christian Staufer, Germany; Stanley Korman, Israel; Friedrich K. Trefz, Germany); Georg Frauendienst-Egger for review and revision of clinical terminology mappings to HPO; Dora Pak for research management support; Miroslav Hatas for system support. C.D.M.v.K. is recipient of the Michael Smith Foundation for Health Research Scholar Award. J.J.Y.L. is recipient of the Jan M. Friedman Studentship from BC Children’s Hospital Foundation. This work is part of the RD-CONNECT initiative and was supported by the FP7-HEALTH-2012-INNOVATION-1 EU grant 305444 (to NB), funding from the Canadian Institutes of Health Research (to C.D.M.v.K. and W.W.W.), funding from Genome Canada/Genome BC/CIHR Large Scale Applied Research Grant ABC4DE project Ref 174CDE (to W.W.W.), and funding from the Dietmar Hopp Foundation (to G.F.H. and N.B.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary material is linked to the online version of the paper at
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Lee, J., Wasserman, W., Hoffmann, G. et al. Knowledge base and mini-expert platform for the diagnosis of inborn errors of metabolism. Genet Med 20, 151–158 (2018). https://doi.org/10.1038/gim.2017.108
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.108
Keywords
This article is cited by
-
Extending inherited metabolic disorder diagnostics with biomarker interaction visualizations
Orphanet Journal of Rare Diseases (2023)
-
Frequency of carriers for rare metabolic diseases in a Brazilian cohort of 320 patients
Molecular Biology Reports (2022)
-
Rare and common genetic determinants of metabolic individuality and their effects on human health
Nature Medicine (2022)
-
A cross-platform approach identifies genetic regulators of human metabolism and health
Nature Genetics (2021)