Abstract
Purpose:
Identifying women at high risk for breast cancer can trigger a personal program of annual mammograms and magnetic resonance imaging scans for early detection, prophylactic surgery, or chemoprevention to reduce the risk of cancer. Yet, current strategies to identify high-risk mutations based on sequencing panels of genes have significant false-positive and false-negative results, suggesting the need for alternative approaches.
Methods:
Flow-variant assays (FVAs) that assess the effects of mutations in the double-strand break (DSB) repair genetic pathway in lymphoblastoid cells in response to treatment with radiomimetic agents were assessed for sensitivity, specificity, and accuracy both alone and as part of a logistic regression classification score. In turn, these assays were validated in circulating B cells and applied to individuals with personal and/or family history of breast and/or ovarian cancer.
Results:
A three-FVA classification score based on logistic regression had 95% accuracy. Individuals from a breast cancer–positive cohort with affected family members had high-risk FVA classification scores.
Conclusion:
Application of a classification score based on multiple FVAs could represent an alternative to panel sequencing for identifying women at high risk for cancer.
Genet Med advance online publication 16 March 2017
Similar content being viewed by others
Introduction
Approximately 10% of incident cases of breast cancer occur in women from high-risk families with multiple affected relatives, some with ovarian cancer or other primaries and often with earlier ages of onset. Germ-line mutations in BRCA1 and BRCA2 can be detected for 20–40% of these cases.1,2 Moderate- to high-penetrance mutations in other genes also contribute to heritable breast cancer risk.3,4 Along with BRCA1 and BRCA2, many of these phenocopy genes play a role in the repair of double-strand breaks (DSBs) in DNA.5 The identification of these genes has led to the development of panel-based sequencing tests for which annotation of nontruncating genetic variants represents a challenge for interpretation. To meet this challenge of interpreting variants of uncertain significance (VUS), we developed molecular phenotyping flow-variant assays (FVAs) that assess the biological effects of heterozygous mutations in genes in the DSB repair pathway by measuring the response cultured or circulating cells exposed to radiomimetic agents.6 These methods have been benchmarked against various classes of variants that were curated rigorously to represent mutations, benign variants, or VUS.
Here, we extend the range of DSB repair FVAs by developing new assays, extending their performance to circulating B cells, and developing classification scores that combine results for multiple assays. In the process, we demonstrate how these scores might be used as risk predictors in subjects with and without breast or ovarian cancer who did not harbor mutations. We also applied these assays to primary B cells isolated from the blood of these subjects to demonstrate how these FVAs could be converted into a real-time clinical test.
Materials and Methods
Subjects
All subjects were recruited at Albert Einstein College of Medicine under approved institutional review board protocols. Informed consent was obtained from all subjects. Of 29 subjects, 20 had breast and/or ovarian cancer and affected relatives (BOC-positive) and 9 did not have a personal history of breast or ovarian cancer but had affected relatives (BOC-negative). All subjects tested negative for mutations in BRCA1 and BRCA2 at commercial laboratories. IBIS and BOADICEA scores were used to predict 5-year risk and lifetime risk for these subjects based on age, personal and family history, and ethnicity, and these were not different between the two groups.7,8 One BRCA2 mutation–positive case was included as a positive control ( Table 1 ).
EBV-immortalized lymphoblastoid cell lines (LCLs) were created for newly recruited subjects by the Molecular Cytogenetics Core at Albert Einstein College of Medicine.9 In addition, B cells were isolated from 7 ml of EDTA-anticoagulated whole blood from subjects using the MACSxpress B Cell Isolation Kit (Miltenyi Biotec, San Diego, CA, Cat: 130-098-190) , a MACSmix Tube Rotator (Miltenyi Biotec, Cat: 130-090-753), and a MACSxpress Separator (Miltenyi Biotec, Cat: 130-098-308) following the manufacturer’s guidelines.
Whole-genome sequencing and analysis
DNA was extracted from 5 ml of blood using the Gentra Puregene Blood Kit (Qiagen, Germantown, MD 158389). Whole-genome sequencing (WGS) was performed on unamplified, high-molecular weight, genomic DNA (3–6 μg) from BOC-positive individuals at the New York Genome Center (NYGC) using the TruSeq DNA Nano Prep kit (Illumina, San Diego, CA) on an Illumina HiSeq 2500 with 2 × 150-bp paired-end reads at 30× coverage. After removal of adapters from the raw sequence reads, the trimmed reads were aligned to the human reference genome (build GRCh37/Hg19) by using the Burrows–Wheeler Aligner.10 The Genome Analysis ToolKit (GATK) Broad Institute, Cambridge, MA was used to perform local realignments, indel realignment, and base recalibration.11 GATK’s Unified Genotyper was used to call the variants in all samples to detect missense, frameshift, and splice site variants, thereby forming a single variant caller file and annotating for quality metrics. Variants were prioritized against a unified gene panel that was based on their known function in the DSB pathway and described in published studies about genes involved in familial breast and ovarian cancer (Supplementary Table S1 online).12,13,14,15 Known common variants with a global minor allele frequency >5% were removed. The final annotation was based on American College of Medical Genetics and Genomics (ACMG) guidelines.16 The variant annotation was performed by literature review as well as reports in ClinVar (http://www.ncbi.nlm.nih.gov/clinvar/) and the Breast Cancer Information Core Database (http://lgdfm3.ncifcrf.gov/bic/BIC.html).
Whole-exome sequencing and analysis
Whole-exome sequencing (WES) was performed on unamplified, high-molecular weight, genomic DNA (3–6 μg) from BOC-negative individuals at Admera Health using xGen Exome Research Panel v1.0 on an Illumina platform with 2 × 150-bp paired-end reads at 30× coverage. The downstream analysis was similar to that used for WGS analysis.
IBIS score and BOADICEA
The IBIS Risk Evaluation Tool was used to estimate the 5-year and lifetime risks for developing breast cancer based on personal and family history (http://www.ems-trials.org/riskevaluator).7 The Breast and Ovarian Analysis of Disease Incidence and Carrier Estimation Algorithm (BOADICEA) was also used to estimate 5-year and lifetime risks for developing breast cancer based on personal and family history (http://ccge.medschl.cam.ac.uk/boadicea/boadicea-model).8
Cell lines and mutations
A total of 36 LCLs were purchased from the Coriell Institute Human Genetic Cell Repository (Camden, NJ) from individuals with different sets of variants and included controls from the 1000 Genome Project (GM19740, GM11995, GM11894, GM19044, GM19371, GM19379, GM19461, GM20771, HG00096, HG00097, HG00099, HG00100, HG00102, HG01083), known BRCA1 pathogenic mutations (GM14097 p.Cys61Gly, GM14090 c.66_67delAG, GM13711 p.Ser1040Asn, GM13713 p.Glu1250Ter, GM14637 p.Arg1443Ter, GM13710 p.Arg1443Gly, GM13708 p.Tyr1563Ter, GM14092 p.Val1713Ala), and BRCA2 (GM14805 p.Trp194Ter, GM14626 p.Lys3326Ter, GM14170 c.5946delT, GM14622 c.6275_6276delTT, GM14624 c.5722_5723delCT, GM14639 p.Ser2067Hisfs, GM14788 p.Asp252Valfs), ATM (GM01525 p.Arg2136Ter, GM03334 p.Trp2638Ter), FANCC (GM20731 c.456 + 4A>T), FANCD2 (GM16633 p.Arg1236His, GM16756 p.Arg1236His), FANCF (GM16757 p.Gln6Ter), and NBS1 (GM15813 p.Lys219Asnfs) mutations (Supplementary Table S2 online). Sequences of the cell lines derived from the 1000 Genomes Project were analyzed for mutations in other genes reported as moderately to highly penetrant for breast cancer when mutated.
Antibodies
For FVAs, the following antibodies were used for native or phosphorylated forms of proteins: BRCA1 (Thermo Fisher, Waltham, MA, Cat: PA5-17512), BRCA2 (Abnova, Taipei, Taiwan, Cat: H00000675), total p53 (R&D systems, Minneapolis, Minnesota, Cat: AF1355), and phospho p53 (R&D AF1043).These antibodies were specific to their targets, as judged by single bands on western blot analysis. All of the antibodies were conjugated with fluorochromes (Innova Bio lightning-link, Cambridge, United Kingdom, Cat: FITC: 322-0010, PECy7: 762-0010, APCCy7: 765-0010, and PECy5.5: 761-0010) as described previously.6
FVAs
The general approach for the analyses included cell culture of isolated B cells as well as LCLs with and without radiomimetic agents.6 The cell culture was followed by cell fixation, permeabilization, and binding with labeled antibodies within cells or cell lysis, as described previously.6 The quantified BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio were compared for the BRCA1 mutant, phenocopy mutant, and control LCLs as well as for the BOC-positive and BOC-negative B cells.
Flow cytometry was performed using BD FACSCanto II (Becton, Dickinson and Company, Franklin Lakes, NJ) equipped with blue (488 nm) and red (633 nm) lasers. The results were analyzed using FlowJo data analysis software (FlowJo, Ashland, OR). The mean of nine replicates was calculated for each individual sample in each assay. Boxplots and Mann-Whitney tests were performed to determine whether the differences in the BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio were significant between cell lines (controls, BRCA1 mutants, and phenocopy mutants) and between the BOC-positive and BOC-negative groups.
Classifiers
Classifiers were built for BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio FVAs using half of each set via random selection to distinguish BRCA1 mutants and phenocopy mutants from controls. They were then applied to the other half to calculate sensitivity, specificity, and accuracy (weighted means of sensitivity and specificity). The classification scores are weighted sums of the BRCA1 nuclear localization, BRCA2 nuclear localization, and phospho p53/total p53 ratio logistic values by their regression coefficients in a fitted multivariable logistic model. The classification score model was then applied to the patient samples.
Based on published data, the values of the FVAs are tightly distributed within the groups to be compared and the between-group means are well-separated. To compare the FVA scores between the control group and the phenocopy group with a sample size of 14 and 15, there is more than 90% power to detect a mean difference of 1.3 common standard deviations using a two-sided two-sample t-test at the 5% alpha level. Similarly, to compare the FVA-based scores between the BOC-negative group and BOC-positive group, for a sample size of 9 and 20, we would have more than 85% power to detect a mean difference of 1.3 common standard deviations using a two-sample t-test at the 5% alpha level.
Results
BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio and the multivariable classification score are reduced in mutant BRCA1 and phenocopy gene LCLs
We have shown previously that BRCA1 nuclear localization and phospho p53/total p53 ratio were reduced in BRCA1, BRCA2, FANCC, and NBS1 mutation carriers after treatment with a cocktail of radiomimetic agents (diepoxybutane, bleomycin, and mitomycin C). In the current study, BRCA2 and BRCA1 nuclear localization was reduced for BRCA1 and BRCA2 mutation carriers as well as for mutation carriers in a broader variety of genes ( Figure 1 ). Eight mutations in BRCA1, seven mutations in BRCA2, two mutations in ATM, and single mutations in FANCC, FANCF, FANCD2, and NBS1 were assessed in Coriell Institute LCLs for both assays. The BRCA1 mutant cell lines showed reduction in BRCA2 nuclear localization (Mann-Whitney test, P = 4 × 10−4) as well as in BRCA1 nuclear localization (P = 6.3 × 10−6). The same effects were observed in the mutant cell lines for the other genes, such as reduction in BRCA2 nuclear localization (P = 6 × 10−5) and BRCA1 nuclear localization (P = 1.4 × 10−5), with both suggesting phenocopies for BRCA1 mutations. The BRCA1 mutant and phenocopy cell lines also showed a reduced phospho p53/total p53 ratio compared with controls ( Figure 1 ; P = 4.4 × 10−5 and P = 1.2 × 10−7, respectively).
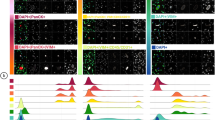
Nuclear localization of BRCA1 and BRCA2 and phospho p53/total p53 ratio are reduced in BRCA1 mutation–bearing and phenocopy lymphoblastoid cell lines compared with controls. Box plots compare standardized nuclear localization of BRCA1 and BRCA2. Phospho p53/total p53 ratio is measured by DCW. P-values of the pairwise comparisons of the various mutation groups relative to controls by the Mann-Whitney test are shown.
To determine sensitivity and specificity of these assays, a fixed threshold was selected. Using “0” as the threshold, the outliers observed for each assay included four for BRCA1 nuclear localization (one control (GM19379) and three phenocopies (GM14624, GM14788 and GM03334)), three for BRCA2 nuclear localization (one each in control (GM11995), BRCA1 mutant (GM14092), and phenocopy (GM14788)), and four for the phospho p53/total p53 ratio (two controls (GM111995 and GM11894), one BRCA1 mutant (GM13710), and one phenocopy (GM16756)). Thus, the sensitivity and specificity for the individual BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio assays ranged from 83 to 91% and 86 to 93%, respectively ( Table 2 ). The accuracy based on the weighted mean of sensitivity and specificity for individual assays ranged from 86 to 89%. When the classification score was calculated by combining these three assays, the sensitivity, specificity, and accuracy increased to 91, 100, and 95%, respectively.
BRCA1 and BRCA2 nuclear localization and phospho p53/total p53 ratio and the multivariable classification score are lower in B cells from BOC-positive individuals even when a mutation cannot be identified
After demonstrating anticipated results for positive and negative control cell lines, FVAs were performed on matched B cells and LCLs from BRCA1 and BRCA2 mutation-negative subjects. These paired cell types showed correlations with BRCA1 nuclear localization (r2= 0.85, Supplementary Figure S1 online) and with phospho p53/total p53 ratio (r2= 0.70, Supplementary Figure S1 online). When the two groups were compared, the BRCA1 nuclear localization, BRCA2 nuclear localization, and phospho p53/total p53 ratio were lower in the BOC-positive group compared with subjects in the BOC-negative group (P = 0.004, P = 0.035, and P = 0.001, respectively, Figure 2 ). When classification scores based on the logistic regression coefficients from the LCL analysis were applied to the B cell groups, the scores were lower for the BOC-positive compared with the BOC-negative group (P = 2.9 × 10−7, Figure 3 ).
Nuclear localization of BRCA1 and BRCA2 is reduced in BOC-positive cases in B cells compared with BOC-negative controls. Box plots compare standardized nuclear localization of BRCA1 and BRCA2. Phospho-p53/total p53 ratio is measured by DCW. P-values of the pairwise comparisons of BOC-positive relative to BOC-negative groups by the Mann-Whitney test are shown.
Classification scores for mutation-bearing and phenocopies cell lines are higher in controls and BOC-negative patients compared with BOC-positive patients. P-values of the pairwise comparisons of the various mutation groups relative to controls and BOC-positive group relative to the BOC-negative group by the Mann-Whitney test are shown.
In the BOC-negative group, two distinct clusters were apparent, with some subjects showing reduced BRCA1 and BRCA2 nuclear localization and reduced phospho p53/total p53 ratio, although not overlapping among all three assays. The observation of two groups was also apparent when classification scores were calculated, with one cluster having lower classification scores that were similar to those of the BOC-positive group ( Figure 3 ). The IBIS and BOADICEA scores calculated for 5-year and lifetime risk for all patients did not show any concordance with the classification scores (although these were correlated with each other), suggesting that classification scores might represent an alternative for identifying those at high risk ( Table 2 ).
WGS and WES analyses did not identify additional mutations in the BOC-positive and BOC-negative cohorts, respectively
WGS was performed for 13 subjects from the BRCA1 and BRCA2 mutation–negative BOC-positive cohort. After alignment and variant calling, and excluding those with a global minor allele frequency of >5% in dbSNP137, 59 variants were identified. When annotated for potential protein truncation or splice site disruption, a single FANCD2 splice site donor deletion was identified in all subjects that was prevalent in the TAGC cohort, suggesting a prevalent benign variant. An additional 15 nonsynonymous missense variants were classified as VUS following the ACMG scoring system (Supplementary Table S3 online). WES using an analysis pipeline that was similar to the WGS analysis was performed for nine subjects from the BOC-negative cohort. Within this group, five nonsynonymous missense mutations were identified as VUS that were not pathogenic (Supplementary Table S4 online).
Discussion
Previously, we and others suggested that cancer gene panel sequencing represents one way of quantifying breast cancer risk. These panels have been widely accepted, even though the clinical utility for genes other than BRCA1 and BRCA2 is unclear.3,4,17 When sequencing was applied to large numbers of women at high risk, mutations were commonly found in BRCA1, BRCA2, and other genes.12,13,14,15,18,19 In up to 40% of women, a VUS was identified; commonly, two or more VUS were observed, especially for those of Asian or African-American ethnicity. Adding genes into sequencing panels increased the likelihood of identifying VUS while minimally increasing the likelihood of finding mutations.18 Interpreting these VUS represents a particular challenge for genetic counseling. Informatics approaches (Align-GVGD, http://agvgd.iarc.fr/) and low- or high-throughput functional assays in yeast or transfected cell lines have been developed for curating variants in BRCA1 and BRCA2, but not for the other genes.20,21,22 In addition to the issue of prevalent VUS, most sequencing methods are biased in scope and depth because certain genomic regions can be difficult to capture, amplify, or assemble. As a result, finished sequences comprise less than the whole of the desired region and important functional variants may be missed. Furthermore, the magnitude of the risk was not always established for mutations in genes other than BRCA1 and BRCA2.3,4 Thus, new methods for assessment of breast cancer risk seem warranted.
Assessing breast cancer risks accurately has clinical utility. Established guidelines exist for increased surveillance, early detection, and risk reduction for women who are at increased risk for cancer due to having a known cancer syndrome, a strong family history, or significant personal medical history. The National Comprehensive Cancer Network (NCCN) and the American Cancer Society (ACS) provide recommendations for women at increased risk for breast cancer based on a number of different factors (http://www.cancer.org/cancer/breastcancer/moreinformation/breastcancerearlydetection).23 For lower penetrance mutations that lack established management guidelines, the implications for clinical care are less clear.24
Although the sample size in the current study was modest, analysis of larger cohorts, including affected and unaffected family members, should demonstrate that phenotypes in FVAs are inherited in a Mendelian fashion. The preliminary results with the BOC-negative group suggest that it might comprise two separate Mendelizing cohorts: one high-risk and the other low-risk. When applied to family cohorts, it will be possible to calculate hazard ratios by decade for the siblings of probands whose B cells demonstrate impairment of BRCA1 and BRCA2 nuclear localization and p53 phosphorylation in response to treatment with radiomimetic agents, thereby improving genetic counseling. Such studies will also demonstrate whether FVAs and resulting classification scores demonstrate significant heterogeneity with loss of accuracy, an issue that could be addressed by adding FVAs for other functions in the DSB repair pathway.
Conclusion
FVA methods are a highly plausible alternative to panel sequencing for identifying functionally important alterations in the DSB repair pathway. A direct functional test at the protein level queries whether key biological functions—BRCA1 and BRCA2 nuclear localization and p53 phosphorylation—are being altered. As shown in this study, tests based on multiple, direct, functional analyses appear to be more sensitive and specific for identifying genetic risks and identify those at high risk, even when a mutation cannot be identified by sequencing. These assays lend themselves to ease of adoption in the clinical and research laboratory environments with minimal change of equipment and workflow because they use common reagents and readily available devices. Analysis can be performed for multiple targets and the results from multiple assays can be combined into an accurate risk prediction score that accounts for the heterogeneity of individual assays.
Disclosure
J.L. and H.O. own shares in Morgan and Mendel Genomics, Inc., which has licensed flow-variant assay technology from Albert Einstein College of Medicine. Albert Einstein College of Medicine has applied for a patent for this application (J.L., A.P., and H.O. are inventors). The other authors declared no conflict of interest.
References
Hopper JL, Southey MC, Dite GS, et al. Population-based estimate of the average age-specific cumulative risk of breast cancer for a defined set of protein-truncating mutations in BRCA1 and BRCA2. Australian Breast Cancer Family Study. Cancer Epidemiol Biomarkers Prev 1999;8:741–747.
Tonin P, Weber B, Offit K, et al. Frequency of recurrent BRCA1 and BRCA2 mutations in Ashkenazi Jewish breast cancer families. Nat Med 1996;2:1179–1183.
Hiraki S, Rinella ES, Schnabel F, Oratz R, Ostrer H. Cancer risk assessment using genetic panel testing: considerations for clinical application. J Genet Couns 2014;23:604–617.
Easton DF, Pharoah PD, Antoniou AC, et al. Gene-panel sequencing and the prediction of breast-cancer risk. N Engl J Med 2015;372:2243–2257.
Ceccaldi R, Sarangi P, D’Andrea AD. The Fanconi anaemia pathway: new players and new functions. Nat Rev Mol Cell Biol 2016;17:337–349.
Loke J, Pearlman A, Upadhyay K, Tesfa L, Shao Y, Ostrer H. Functional variant analyses (FVAs) predict pathogenicity in the BRCA1 DNA double-strand break repair pathway. Hum Mol Genet 2015;24:3030–3037.
Tyrer J, Duffy SW, Cuzick J. A breast cancer prediction model incorporating familial and personal risk factors. Stat Med 2004;23:1111–1130.
Lee AJ, Cunningham AP, Kuchenbaecker KB, Mavaddat N, Easton DF, Antoniou AC ; Consortium of Investigators of Modifiers of BRCA1/2; Breast Cancer Association Consortium. BOADICEA breast cancer risk prediction model: updates to cancer incidences, tumour pathology and web interface. Br J Cancer 2014;110:535–545.
Chen C, Johnston TD, Jeon H, Gedaly R, McHugh P, Ranjan D. Cyclosporine promotes epstein-barr virus-infected human B-cell transformation assayed by three correlated assay methods. Transplant Proc 2009;41:366–370.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010;20:1297–1303.
LaDuca H, Stuenkel AJ, Dolinsky JS, et al. Utilization of multigene panels in hereditary cancer predisposition testing: analysis of more than 2,000 patients. Genet Med 2014;16:830–837.
Kurian AW, Hare EE, Mills MA, et al. Clinical evaluation of a multiple-gene sequencing panel for hereditary cancer risk assessment. J Clin Oncol 2014;32:2001–2009.
Tung N, Battelli C, Allen B, et al. Frequency of mutations in individuals with breast cancer referred for BRCA1 and BRCA2 testing using next-generation sequencing with a 25-gene panel. Cancer 2015;121:25–33.
Lincoln SE, Kobayashi Y, Anderson MJ, et al. A systematic comparison of traditional and multigene panel testing for hereditary breast and ovarian cancer genes in more than 1000 patients. J Mol Diagn 2015;17:533–544.
Richards S, Aziz N, Bale S, et al.; ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Walsh T, Casadei S, Lee MK, et al. Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing. Proc Natl Acad Sci USA 2011;108:18032–18037.
Maxwell KN, Hart SN, Vijai J, et al. Evaluation of ACMG-guideline-based variant classification of cancer susceptibility and non-cancer-associated genes in families affected by breast cancer. Am J Hum Genet 2016;98:801–817.
Susswein LR, Marshall ML, Nusbaum R, et al. Pathogenic and likely pathogenic variant prevalence among the first 10,000 patients referred for next-generation cancer panel testing. Genet Med 2016;18:823–832.
Lindor NM, Guidugli L, Wang X, et al. A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat 2012;33:8–21.
Carvalho MA, Couch FJ, Monteiro AN. Functional assays for BRCA1 and BRCA2. Int J Biochem Cell Biol 2007;39:298–310.
Starita LM, Young DL, Islam M, et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 2015;200:413–422.
Bevers TB, Ward JH, Arun BK, et al. Breast cancer risk reduction, version 2.2015. J Natl Compr Canc Netw 2015;13:880–915.
Robson ME, Storm CD, Weitzel J, Wollins DS, Offit K ; American Society of Clinical Oncology. American Society of Clinical Oncology policy statement update: genetic and genomic testing for cancer susceptibility. J Clin Oncol 2010;28:893–901.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figures and Table
(DOCX 139 kb)
Rights and permissions
About this article
Cite this article
Syeda, M., Upadhyay, K., Loke, J. et al. Prediction of breast cancer risk based on flow-variant analysis of circulating peripheral blood B cells. Genet Med 19, 1071–1077 (2017). https://doi.org/10.1038/gim.2016.222
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2016.222
Keywords
This article is cited by
-
Imbalances in cellular immunological parameters in blood predetermine tumor onset in a natural mouse model of breast cancer
Cancer Immunology, Immunotherapy (2019)