Abstract
Purpose:
A combination of oligonucleotide and single-nucleotide polymorphism probes on the same array platform can detect copy-number abnormalities and copy-neutral aberrations such as uniparental disomy and long stretches of homozygosity. The single-nucleotide polymorphism probe density in commercially available platforms varies widely, which may affect the detection of copy-neutral abnormalities.
Methods:
We evaluated the ability of array platforms with low (Oxford Gene Technology CytoSure ISCA uniparental disomy), mid-range (Agilent custom array), and high (Affymetrix CytoScan HD) single-nucleotide polymorphism probe density to detect copy-number variation, mosaicism, uniparental isodisomy, and absence of heterozygosity in 50 clinical samples.
Results:
All platforms reliably detected copy-number variation, mosaicism, and uniparental isodisomy; however, absence-of-heterozygosity detection varied significantly. The low-density array called absence-of-heterozygosity regions not confirmed by the other platforms and also overestimated the length of true absence-of-heterozygosity regions. Furthermore, the low- and mid-density platforms failed to detect some small absence-of-heterozygosity regions that were identified by the high-density platform.
Conclusion:
Variation in single-nucleotide polymorphism density can lead to major discrepancies in the detection of and confidence in copy-neutral abnormalities. Although suitable for uniparental disomy detection, copy-number plus single-nucleotide polymorphism arrays with 30,000 or fewer unique single-nucleotide polymorphism probes miscall absence-of-heterozygosity regions due to identity by descent.
Genet Med 15 9, 706–712.
Similar content being viewed by others
Main
The use of microarrays as a first-tier test for detecting copy-number aberrations in children with developmental delay, intellectual disability, multiple congenital anomalies, and autism has vastly improved the diagnostic yield in clinical cytogenetic laboratories.1,2,3 Although oligonucleotide array comparative genome hybridization platforms detect whole-chromosome aneuploidies and submicroscopic deletions and duplications, they miss copy-neutral aberrations such as uniparental disomy (UPD) and absence of heterozygosity (AOH). By contrast, single-nucleotide polymorphism (SNP) microarrays detect copy-neutral aberrations, but due to lack of coverage in “SNP deserts,” they may overlook some copy-number alterations.4 Combining both probe types on a single array, referred to here as copy number plus SNP (CN+SNP), has the advantage of identifying copy-number variations (CNVs) and copy-neutral aberrations in the same test.
Because CN+SNP arrays detect copy-number changes and allele genotypes in a single platform, they can provide an internal confirmation of CNVs that may eliminate the need for secondary confirmatory testing such as quantitative PCR, multiplex ligation-dependent probe amplification, or fluorescence in situ hybridization;5 however, such internal confirmation requires independent analysis of oligonucleotide and SNP probes. Not all platforms analyze SNP and oligonucleotide probes independently, so evaluating the probe chemistry and aberration-detection algorithms of various platforms is critical.
Array comparative genome hybridization and SNP microarrays are more sensitive to mosaicism than traditional 20-cell G-banded chromosome analysis, as they can detect mosaicism levels as low as 5%.6,7,8,9 This is important in clinical laboratories because ~1% of all cases are mosaic.9 Furthermore, G-band analysis is limited to cultured cells and may be biased by growth failure and/or lack of proliferation of a subset of cells in culture. By contrast, microarrays can analyze cultured and uncultured samples, thereby uncovering some mosaic abnormalities that escape detection by standard G-band analysis.9,10,11 Moreover, the combination of allele genotypes (SNP probes) with copy-number data (oligonucleotide probes) in CN+SNP arrays can improve sensitivity and confidence in the detection of mosaicism.
SNP arrays also detect regions of excessive homozygosity in constitutional samples, known as AOH. Regions of AOH can point to UPD of single chromosomes or identity by descent of multiple genomic regions consistent with parental consanguinity. UPD is the inheritance of two of the same chromosome from one parent, and there are at least three types of UPD: heterodisomy, isodisomy, and segmental isodisomy.5 Heterodisomy is the inheritance of two biallelic copies of the same chromosome from one parent, so it cannot be detected by SNP analysis without parental studies. Detection of UPD via SNP arrays is thus limited to isodisomy and segmental isodisomy and must be confirmed by another molecular method.5
Elevated AOH levels can also indicate an increased risk for recessive disorders due to regions of the genome identical by descent. Recently, SNP arrays have been applied to homozygosity mapping, also known as autozygosity mapping, to search for genes involved in recessive disease (reviewed in ref. 5). In the clinical laboratory, homozygosity mapping may shorten the time to diagnosis for offspring of a consanguineous mating with a phenotype suggestive of a recessive disorder.12,13 Sequencing genes within regions of AOH could uncover homozygous mutations responsible for patient phenotypes. However, as costs drop in the future, whole-genome sequencing will likely replace homozygosity mapping for recessive disease testing.
The growing demand for CN+SNP microarrays has spawned a number of commercially available platforms that vary in the density of oligonucleotide and SNP probes. Some platforms require specific equipment and software that may force laboratories to commit to one platform over the other based on the availability of costly scanners and/or software licenses. Here, we compared three leading microarray platforms, Affymetrix CytoScan HD (Affymetrix, Santa Clara, CA) (high SNP density), a custom Agilent array (Agilent, Santa Clara, CA) (mid SNP density), and Oxford Gene Technology (OGT) Cytosure ISCA UPD (OGT, Begbroke, Oxfordshire, UK) (low SNP density), to detect CNV, mosaicism, UPD, and long stretches of AOH.
Materials and Methods
Subjects
Fifty DNA samples extracted from peripheral blood were obtained from Emory Genetics Laboratory and Boston Children’s Hospital Genetic Diagnostic Laboratory. One sample from an unaffected, healthy parent was used as a normal control. Forty patients self-reported consanguinity or had initial results suggestive of consanguinity from the low-density (LD) array. Five of these 40 samples also carried CNVs. In addition, we evaluated two UPD samples previously analyzed via DNA methylation, three mosaic samples identified by previous G-banded chromosome analysis, and four samples with pathogenic CNVs previously identified on the LD array. The ethnicities of most patients were not provided to the laboratories.
Microarray hybridization and analysis
We performed microarray analysis of isolated genomic DNA with three different commercially available microarray platforms. Microarray hybridization and analysis were performed according to the manufacturer’s instructions, unless otherwise noted.
Affymetrix (HD). The Affymetrix CytoScan HD array (Affymetrix, catalog no. 901835) has the highest SNP density. This 1 × 2.6 M platform contains 750,000 unique SNP and 1.9 million oligonucleotide probes, with a mean backbone spacing of one oligonucleotide probe every 2 kb and one oligonucleotide probe every 400 basepairs (bp) in targeted regions. The mean spacing of SNP probes is 200 per megabase (Mb). All probes are 25 bp long. Each SNP is targeted by six probes, three for each allele. Approximately 90% of SNPs have a minor allele frequency of 0.05 or greater, based on data from the 1000 Genomes Project.
The high-density (HD) assay was run according to the manufacturer’s protocol. Briefly, 250 ng of patient DNA was digested with Nsp1, amplified with TITANIUM Taq DNA polymerase (Clontech, Mountain View, CA; catalog no. 639240), fragmented with Affymetrix fragmentation reagent, and labeled with biotin end-labeled nucleotides (Affymetrix Reagent Kit Bundle, catalog no. 901835). The DNA was hybridized to the microarray for 16 hours, washed on the GeneChip Fluidics Station 450 (Affymetrix), stained with Affymetrix GeneChip Stain Reagents, and scanned on the GeneChip Scanner 3000 7G (Affymetrix).
Data analysis was performed using Chromosome Analysis Suite software version 1.2.2 (Affymetrix). Hybridization of patient DNA to oligonucleotide and SNP probes was independently compared against a previously analyzed cohort of normal samples to call CNVs and allele genotypes. Chromosome Analysis Suite uses the Bayesian robust linear modeling using Mahalanobis distance perfect match algorithm to call SNP genotypes.14 CNVs require a 25-probe call. The percentage mosaicism of whole-chromosome aneuploidies was determined using the average log2 ratio of the entire chromosome. AOH regions required a 50-probe call of at least 3 Mb. The following regions were not included in the AOH calculation due to segmental duplications in the area: chr1:145,693,382–151,094,545; chr11: 46,031,567–51,563,636; and chr16: 31,451,325–35,220,544 (GRCh37/hg19 build of the Human Genome Assembly).
Agilent Technologies (MD). The custom Aligent CGH+SNP 4 × 180 K array is a mid-density (MD) array with 150,000 oligonucleotide and 30,000 SNP probes, with one oligonucleotide probe every 20 kb and 10 SNP probes per Mb, designed by Boston Children’s Hospital Genetic Diagnostic Laboratory (AMADID no. 036518). This custom design combines dense oligonucleotide coverage in clinically relevant loci with lower coverage in nongenic regions. All probes are 60 bp long, and there is one probe per SNP. The 30,000 SNPs were selected from 60,000 SNPs empirically tested by Agilent. SNPs were evenly spaced throughout the genome, and most SNPs have a minor allele frequency of 0.1 or greater.
Array experiments were performed according to the manufacturer’s protocol with sex-matched controls; control DNA was provided in the SureTag DNA Complete Labeling Kit (Agilent, catalog no. 5190-4240). Briefly, 1–1.5 µg of patient and sex-matched control DNA was digested with AluI and RsaI and labeled with Cy3- and Cy5-dUTP using Exo-Klenow (Agilent). The DNA was purified with an Amicon 30-kDa centrifugal filter (Millipore, Billerica, MA; catalog no. UFC5003BK) and hybridized to the array for 40 hours. The microarray was washed according to wash procedure B (including stabilization and drying solution) and scanned using the Agilent High-Resolution Microarray Scanner model G2505C at 3-µm resolution.
Data were processed with feature extraction for cyto software and analyzed with Agilent CytoGenomics version 1.5. For each SNP site, the allele-specific copy number algorithm calculates an expectation value for the copy number of the uncut SNP allele using data from neighboring oligonucleotide probes. SNP calls are determined from the log2 ratios of the sample signal versus the signal from a genotyped internal reference. Because the reference genotype is known, the raw log2 ratios are “reference adjusted” to values that reflect a hypothetical allele-specific copy number of two for the reference at all SNP sites.
CNV detection required a five-probe call with a minimum absolute average log2 ratio of 0.25. Mosaicism detection required a five-probe call with a minimum absolute average log2 ratio of 0.15. Percentage mosaicism was calculated using the clonal fraction algorithm, which separates out the major aberrant clone from other clones in the sample and calculates a percentage mosaicism based on the log2 ratio.15 AOH regions were detected using the loss of heterozygosity (LOH) algorithm at the default threshold of 6.0, unless otherwise noted.
Oxford Gene Technology (LD). The OGT CytoSure ISCA UPD array 4 × 180 K platform (OGT, catalog no. 020050) has 6,186 unique SNP and 137,100 oligonucleotide probes, with one oligonucleotide probe every 25 kb and higher oligonucleotide density in targeted regions determined by the International Standards for Cytogenomic Array consortium.16 This LD array has an average of three SNP probes every Mb. All probes are 60 bp long, and each SNP has three identical probes per allele. The SNPs have a minor allele frequency of 0.4–0.5.
Array experiments were performed according to the manufacturer’s protocol with some modifications. Briefly, 900 ng of patient and sex-matched control DNA was sonicated to a length between 500 bp and 2 kb. The sex-matched control DNA was pooled from five normal males or females. DNA samples were labeled with Cy3- or Cy5-deoxycytidine triphosphate using Exo-Klenow (OGT Cytosure Labeling Kit, catalog no. 020020), purified with a multiscreen PCR plate (Millipore, catalog no. MSNU03050), and hybridized to the microarray at 65 °C for 40 hours. The microarray was washed using the Little Dipper microarray processor (SciGene, Sunnyvale, CA) and scanned with the Agilent High-Resolution Microarray Scanner Model G2505B at 3-µm resolution.
Data analysis was performed using Feature Extraction version 9.5 (Agilent) and CytoSure Interpret Software version 3.4.8 (OGT). CNV detection required a four-probe call; the log2 ratio threshold is +0.3 for a gain and −0.5 for a loss. The log2 ratio threshold for a mosaic gain or loss is ±0.1. AOH regions required a LOH score ≥50, which is calculated by the CytoSure Interpret Software algorithm. A higher LOH score correlates with greater confidence in the AOH region. For samples C1–C2 and U1–U11, autosomal AOH calls were also detected using LOH score cutoffs of 100, 200, 300, and 400. False-positive and false-negative AOH regions were identified by comparison to the HD array data. The total lengths of false-positive and false-negative AOH regions were summed for each sample at the various LOH scores and divided by the total autosomal genomic length to calculate the percentage autosomal false-positive or false-negative AOH for each sample. These values were averaged for samples U1–U11 and C1–C2 (Figure 4).
For all three CN+SNP platforms, the percentage autosomal AOH was determined by summing AOH regions and dividing by total autosomal genomic length, excluding regions without probe coverage (centromeres, telomeres, and acrocentric chromosome arms).5 Whole-genome views of AOH regions were generated with the National Center for Biotechnology Information Genome Decoration Page (http://www.ncbi.nlm.nih.gov/genome/tools/gdp).
Results
CNV detection
We compared the LD, MD, and HD arrays for their ability to detect nine known CNVs ( Figure 1a ). Three CNVs were 300–800 kb, four CNVs were ~100 kb, and two intragenic CNVs were 9 and 16 kb in length. The two smallest CNVs lie in International Standards for Cytogenomic Array–targeted regions16 and were detected only by the LD and MD arrays. The HD array lacked oligonucleotide probes in the 9-kb CNV and had only 20 probes in the 16-kb CNV. Although visual inspection showed an increase in copy number of those 20 probes, there were not enough probes to meet the 25-probe threshold set for HD array analysis. All three platforms detected CNVs >100 kb; however, the MD array did not detect one CNV that was 91–100 kb due to poor probe coverage in the region. As expected, the length of CNVs varied as measured by different platforms based on probe location and density. The mean difference in minimum CNV length was 11, 32, and 45 kb for the LD versus MD, LD versus HD, and MD versus HD platforms, respectively. Only the HD platform independently detects CNVs with oligonucleotide and SNP probes, thereby providing detection and confirmation for CNVs simultaneously ( Figure 1a ).
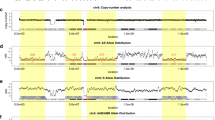
Copy-number variations (CNVs) and uniparental disomy (UPD) are detected on all three platforms. (a) Log2 ratios of oligonucleotide probe signal intensities detect the deletion of 6q16.3 on the low-density (LD) (upper), mid-density (MD) (center), and high-density (HD) (lower) arrays. The allele difference plot from the single-nucleotide polymorphism probes on the HD array (lower) confirms the same CNV. (b) Absence-of-heterozygosity regions detected in two different UPD samples; one with chromosome 7 uniparental segmental isodisomy and the other with chromosome 14 uniparental isodisomy.
Mosaicism detection
The LD, MD, and HD arrays successfully detected mosaicism in three mosaic samples. As measured by G-banded chromosome analysis, the trisomy 8, trisomy 18, and 45,X samples exhibited mosaicism levels of 92, 54, and 47% for the abnormal cell line, respectively. We calculated percentage mosaicism for the MD and HD data using the software packages for each platform (see Materials and Methods). However, the LD software does not calculate percentage mosaicism. Percentage mosaicism values from the MD and HD platforms differed from the G-banded analysis by 6–20% (Supplementary Table S1 online). These differences may be explained by uncultured samples used for the microarrays versus the cultured samples required for G-band analysis.
UPD detection
One of the major goals of CN+SNP platforms is the detection of UPD. We tested two previously characterized UPD samples (UPD7 and UPD14) on the three different platforms. Each array detected AOH regions consistent with whole chromosome 14 and segmental chromosome 7 uniparental isodisomy ( Figure 1b ). The size of AOH regions on chromosome 7 varied among arrays; however, all three platforms detected AOH consistent with segmental isodisomy of chromosome 7.
Long stretches of AOH detection
Forty samples were derived from patients with suspected consanguinity according to self-report or previous LD array–testing results. Twenty-nine of the 40 individuals reported parental consanguinity. Eleven exhibited elevated levels (3–8%) of total autosomal AOH as detected by LD array. We analyzed 14 samples in all three platforms: two from known consanguineous matings (C1–C2), 11 with elevated AOH per the LD array (U1–U11), and one negative control (N). The three arrays detected AOH in the two consanguineous samples and showed no AOH in the negative control, but the results were discrepant for the 11 other samples ( Table 1 ).
Although all three arrays detected AOH in the two consanguineous samples (C1–C2), the AOH calls differed among the arrays. For sample C1, the LD array detected ~12% total autosomal AOH, whereas the MD and HD arrays detected ~8% total autosomal AOH ( Figure 2a ). In sample C2, all three platforms detected ~9% total autosomal AOH ( Figure 2b ). In both samples, there were concordant AOH calls, as well as discordant AOH calls that differed in regions detected and the size of those regions ( Figure 3 ). For samples C1 and C2, the LD array called 14 and 3 AOH regions, respectively, that were not detected by either of the other arrays. These 17 regions were 3–19 Mb, included 10–42 LD SNP probes, and had LOH scores from 50 to 190. The HD and MD arrays included a mean of 1,800 and 90 SNP probes in these regions, respectively. Therefore, we interpret these 17 AOH calls as false positives on the LD array. It is important to point out that false positives detected with the LD array exhibited a range of LOH scores, some higher than those calculated for true AOH positives.
Concordant and discordant absence-of-heterozygosity (AOH) calls. (a) One AOH region in sample C1 detected on all three arrays, (b) one terminal AOH region in sample C2 detected by the mid-density (MD) and high-density (HD) arrays but not the low-density (LD) array, (c) and one terminal AOH region in sample U1 detected by the LD array but not the MD or HD arrays. For each platform, single-nucleotide polymorphism probes are indicated by vertical lines (LD) or dots (MD and HD). AA, AB, and BB alleles are indicated on the left. Gray boxes highlight the AOH calls detected by the software and the number of probes within a call. For LD calls, the loss of heterozygosity score is listed.
In addition, there were three AOH regions in sample C2 that were detected by the MD and HD arrays but not the LD array. These regions ranged from 5 to 12.5 Mb in size, were supported by 71–121 MD and 1,287–2,905 HD SNP probes, but only included ~15–37 SNP probes in the LD array ( Figure 3b ). There were two ~5-Mb AOH regions detected by the HD array that were not detected by the MD or LD array and two ~7-Mb AOH regions detected by the LD and HD arrays but absent from the MD array ( Figure 2 ).
The size of AOH regions is expected to vary slightly between platforms, depending on the location of SNP probes; however, we found large discrepancies in the size of three AOH regions detected by the LD array for C1 and C2 versus the other two arrays ( Figure 2 ). These three regions were 21–28 Mb (49–63 SNP probes and LOH scores of 136–370) as detected by the LD array and only 8–15 Mb as measured by the other arrays, meaning the LD array overestimates the size of AOH stretches in some cases.
Differences in AOH detection were even more pronounced in the 11 samples without known consanguinity (U1–U11). These samples exhibited elevated levels of AOH in the LD array that would suggest relationships between first cousins once removed (3.125%) or first cousins (6.25%) ( Table 1 ).17,18 However, in these samples the MD array detected no AOH regions, and the HD array detected only 0–0.66% autosomal AOH. As an example, the U1 sample exhibits 8.35% autosomal AOH on the LD array and 0% on the MD and HD arrays ( Table 1 and Figure 2c ). In samples U1–U11, the discrepant LD AOH calls ranged from 3 to 26 Mb with 10–56 SNP probes and LOH scores from 50 to 325 ( Figure 3c ). There was a mean of 20 AOH calls (median of 18) per sample detected on the LD array, of which only three regions (~3.5 Mb each) were confirmed by the HD array but not the MD array. These three AOH regions differed in length by 5–11 Mb between LD and HD platforms. There were also seven small AOH regions (3–5 Mb) in the 11 samples that were detected by the HD array but not by the other arrays (Supplementary Table S2 online).
It is possible that raising the LOH threshold in the LD software could lead to more concordant AOH findings among the three platforms. Increasing the LOH score cutoff not only reduced the percentage of the autosomal genome in false-positive AOH calls but also inflated the missed (false-negative) AOH regions. As the LOH cutoff was increased from 50 to 400, the number of AOH calls in samples U1–U11 decreased to levels inconsistent with consanguinity ( Figure 4 ); however, with these more stringent LOH settings, the number of false-negative AOH regions in consanguineous samples C1–C2 rose dramatically. The percentage autosomal AOH in samples C1 and C2 was reduced by 6.8 and 2.8%, respectively, when the LOH score cutoff was increased from 50 to 400 ( Figure 4 ). Therefore, there is a significant trade-off between reducing false positives in regions that are not AOH and increasing false negatives in true stretches of AOH when choosing LD software settings.
False-positive and false-negative absence-of-heterozygosity (AOH) calls detected by the LD software. The percentage of the autosomal genome included in AOH false-positive regions (x-axis) is plotted versus the percentage of the autosomal genome included in AOH false-negative regions (y-axis) for samples C1–C2 and U1–U11. Values for C1–C2 and U1–U11 are average percentage AOH, as described in the Materials and Methods. Loss of heterozygosity (LOH) scores per experiment are indicated. For samples U1–U11, the results were the same at LOH scores of 300 and 400.
MD versus HD array comparison
To better assess the detection of true AOH, we analyzed 27 additional samples (C3–C29) from self-reported consanguineous matings using the MD and HD arrays. All AOH regions detected by the MD array were also detected by the HD array; however, 111 regions detected by the HD array were not detected using MD array default parameters (LOH score = 6.0) (Supplementary Table S3 online). Three of these regions are >10 Mb, 31 regions are 5–10 Mb, and 77 regions are 3–5 Mb. Given the HD SNP probe density in these regions (346–4,056 probes per region), we expect that most are false negatives in the MD platform.
We lowered the stringency of the MD LOH threshold (LOH score = 5.25), which revealed 28 additional AOH regions that were also detected on the HD array. These regions ranged from 3 to 17 Mb (mean = 6.5 Mb). Nevertheless, the MD analysis still failed to detect 83 AOH regions detected by the HD array (3–9 Mb; mean = 4 Mb). Furthermore, at this threshold, the MD array detected 10 AOH regions that were not detected by the HD array (5–16 Mb; mean = 8 Mb); these are likely false positives on the MD platform. Therefore, there is a trade-off between missing some AOH regions and introducing false-positive regions with different stringencies of MD array software.
Discussion
All three CN+SNP arrays were able to detect CNVs >100 kb in size, which is more sensitive than the American College of Medical Genetics and Genomics recommendation of 400 kb.19 Due to differences in platform chemistry and analysis algorithms, only the HD array can detect and confirm CNV calls at the same time ( Figure 1 ). The LD platform independently analyzes the two probe types; however, the software does not distinguish between hemizygous and homozygous genotypes and therefore cannot confirm CNV aberrations with SNP genotypes. The MD software combines data from the oligonucleotide and SNP probes to call genotypes; therefore, the two sets of data are not independent and do not provide internal confirmation. Furthermore, the MD software only displays SNP data from the major clone in the “uncut SNP alleles plot,” which prevents visualization of mosaic genotypes. In our cases, samples had relatively high percentages of mosaic cell lines that were detectable by all three platforms (Supplementary Table S1 online). For lower levels of mosaicism, different arrays may be more or less sensitive, depending on probe coverage and analysis algorithms.
Detection of long stretches of AOH produced the most variable results between arrays. In some cases, AOH regions were detected by one platform but not by the other two, which may represent false positives in the discrepant platform; however, such calls could also represent real, albeit small, regions of AOH that are only detected by platform(s) with dense SNP coverage. It is also possible that patients from ethnic groups underrepresented in SNP studies are heterozygous for fewer SNPs on these platforms, leading to more AOH false positives in certain populations.
Many calls detected on the LD array in samples U1–U11 are likely false positives. Within LD AOH regions, there are abundant heterozygous SNPs on the MD and HD arrays that do not support AOH ( Figure 3c ). Furthermore, in U1–U11, only 3 of 217 AOH calls detected by the LD array were confirmed by the HD array, and for those 3 calls, the HD regions were much smaller than the LD regions. On the basis of these data, we conclude that the LD arrays overestimate the number of AOH regions per genome and the size of individual AOH regions, leading to a major inflation in total autosomal AOH.
It is possible that different LD analysis parameters could improve AOH detection. To investigate this, we increased the stringency of the LOH score in the LD software. We found that increasing the LOH score from 50 in the original analysis to 400 reduced the number of false positives in samples U1–U11 and lowered the percentage of autosomal AOH to levels expected in a normal population, but increasing the LOH score to 400 also decreased percentage AOH in consanguineous samples C1 and C2 by 6.8 and 2.8% of the total autosomal genome, respectively, much lower than their true AOH values ( Figure 4 ). Therefore, when adjusting the LD analysis parameters, one must choose between avoiding false positives and retaining true positives.
Although the MD array performed better than the LD array in AOH detection, there were some AOH regions called by the HD array that were absent from the MD array using our default parameters (Supplementary Table S3 online). To further investigate this, we lowered the stringency of the MD LOH threshold from 6.0 to 5.25 for samples C3–C29. These settings not only improved the detection of previous false-negative regions but also introduced false positives in regions that were not called as AOH in the HD analysis. Therefore, even for the MD array, some small AOH regions may confound detection.
Discrepancies in the size of AOH regions called by the three platforms can lead to differences of up to tens of megabases per AOH call ( Figure 2 ). Minor differences in AOH boundaries are expected when using platforms with different probes, but SNP probe density and/or allele frequency in the LD, MD, and HD arrays can significantly alter the start and stop of AOH regions. Because clinicians currently rely on AOH detection for homozygosity mapping of recessive traits, variation in AOH regions could prompt investigation of genes that are not truly identical by descent. Therefore, the boundaries of AOH should be interpreted with caution, especially when identified by lower-density SNP platforms.
Variation in AOH detection can also affect the interpretation of the degree of parental relationship. Overestimation of total AOH due to false positives or errors in AOH size can erroneously suggest that a given sample exhibits more AOH than that in the general population. Consanguinity is not an uncommon incidental finding in SNP testing,11,20,21,22 and AOH findings are not typically confirmed by a second method, so there is a legitimate risk for reporting false-positive AOH results when using LD arrays. AOH results must therefore be reported with these caveats and should never be considered diagnostic for a recessive condition.
Overall, the LD, MD, and HD arrays are suitable for the detection of CNV, mosaicism, and large regions AOH due to uniparental isodisomy. The MD and the HD arrays detected similar levels of AOH in consanguineous and nonconsanguineous samples, whereas the LD array tended to inflate autosomal AOH. Due to differences in SNP density, some small regions of AOH were detected on the HD array but not the MD array. Absence of these regions from the MD analysis did not significantly alter the percentage autosomal AOH ( Table 1 ) and did not change the interpretation of the patient result. Therefore, MD and HD arrays reliably detect AOH due to identity by descent.
Disclosure
The authors declare no conflict of interest.
References
Miller DT, Adam MP, Aradhya S, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet 2010;86(5):749–764.
Shen Y, Dies KA, Holm IA, et al.; Autism Consortium Clinical Genetics/DNA Diagnostics Collaboration. Clinical genetic testing for patients with autism spectrum disorders. Pediatrics 2010;125:e727–e735.
Coulter ME, Miller DT, Harris DJ, et al. Chromosomal microarray testing influences medical management. Genet Med 2011;13:770–776.
Schaaf CP, Wiszniewska J, Beaudet AL . Copy number and SNP arrays in clinical diagnostics. Annu Rev Genomics Hum Genet 2011;12:25–51.
Kearney HM, Kearney JB, Conlin LK . Diagnostic implications of excessive homozygosity detected by SNP-based microarrays: consanguinity, uniparental disomy, and recessive single-gene mutations. Clin Lab Med 2011;31:595–613, ix.
Hook EB . Exclusion of chromosomal mosaicism: tables of 90%, 95% and 99% confidence limits and comments on use. Am J Hum Genet 1977;29:94–97.
Ballif BC, Rorem EA, Sundin K, et al. Detection of low-level mosaicism by array CGH in routine diagnostic specimens. Am J Med Genet A 2006;140:2757–2767.
Valli R, Marletta C, Pressato B, et al. Comparative genomic hybridization on microarray (a-CGH) in constitutional and acquired mosaicism may detect as low as 8% abnormal cells. Mol Cytogenet 2011;4:13.
Conlin LK, Thiel BD, Bonnemann CG, et al. Mechanisms of mosaicism, chimerism and uniparental disomy identified by single nucleotide polymorphism array analysis. Hum Mol Genet 2010;19:1263–1275.
Cheung SW, Shaw CA, Scott DA, et al. Microarray-based CGH detects chromosomal mosaicism not revealed by conventional cytogenetics. Am J Med Genet A 2007;143A:1679–1686.
Bruno DL, White SM, Ganesamoorthy D, et al. Pathogenic aberrations revealed exclusively by single nucleotide polymorphism (SNP) genotyping data in 5000 samples tested by molecular karyotyping. J Med Genet 2011;48:831–839.
Alkuraya FS . Homozygosity mapping: one more tool in the clinical geneticist’s toolbox. Genet Med 2010;12:236–239.
Lam CW, Lau KC, Tong SF . Microarrays for personalized genomic medicine. Adv Clin Chem 2010;52:1–18.
Lamy P, Grove J, Wiuf C . A review of software for microarray genotyping. Hum Genomics 2011;5:304–309.
Curry B, Ghosh J, Troup C . Normalization of array CG data. In: Stafford P (ed). Methods in Microarray Normalization. CRC Press: Boca Raton, FL, 2008:233–244.
Baldwin EL, Lee JY, Blake DM, et al. Enhanced detection of clinically relevant genomic imbalances using a targeted plus whole genome oligonucleotide microarray. Genet Med 2008;10:415–429.
Wright S . Coefficients of inbreeding and relationship. Am Nat 1922;56:330–338.
Wright S, McPhbe H . An approximate method of calculating coefficients of inbreeding and relationships from livestock pedigrees. J Agric Res 1925;31:377–383.
Kearney HM, South ST, Wolff DJ, Lamb A, Hamosh A, Rao KW ; Working Group of the American College of Medical Genetics. American College of Medical Genetics recommendations for the design and performance expectations for clinical genomic copy number microarrays intended for use in the postnatal setting for detection of constitutional abnormalities. Genet Med 2011;13:676–679.
Bruno DL, Ganesamoorthy D, Schoumans J, et al. Detection of cryptic pathogenic copy number variations and constitutional loss of heterozygosity using high resolution SNP microarray analysis in 117 patients referred for cytogenetic analysis and impact on clinical practice. J Med Genet 2009;46:123–131.
Schaaf CP, Scott DA, Wiszniewska J, Beaudet AL . Identification of incestuous parental relationships by SNP-based DNA microarrays. Lancet 2011;377:555–556.
Sund KL, Zimmerman SL, Thomas C, et al. Regions of homozygosity identified by SNP microarray analysis aid in the diagnosis of autosomal recessive disease and incidentally detect parental blood relationships. Genet Med 2013;15:70–78.
Acknowledgements
We thank Hutton Kearney and Madhuri Hegde for scientific expertise and the Emory Genetics Laboratory and the Boston Children’s Hospital staff for technical support. We also thank Cheryl Strauss for editorial assistance.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Table S1
(DOC 28 kb)
Supplementary Table S2
(DOC 34 kb)
Supplementary Table S3
(DOC 149 kb)
Rights and permissions
About this article
Cite this article
Mason-Suares, H., Kim, W., Grimmett, L. et al. Density matters: comparison of array platforms for detection of copy-number variation and copy-neutral abnormalities. Genet Med 15, 706–712 (2013). https://doi.org/10.1038/gim.2013.36
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2013.36