Abstract
Purpose:
Major advances have been made in our understanding and clinical application of genetic testing in hypertrophic cardiomyopathy. Determining pathogenicity of a single-nucleotide variant remains a major clinical challenge. This study sought to reassess single-nucleotide variant classification in hypertrophic cardiomyopathy probands.
Methods:
Consecutive probands with hypertrophic cardiomyopathy with a reported pathogenic mutation or variation of uncertain significance were included. Family and medical history were obtained. Each single-nucleotide variant was reassessed by a panel of four reviewers for pathogenicity based on established criteria together with updated cosegregation data and current population-based allele frequencies.
Results:
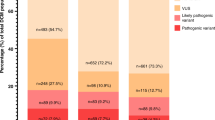
From 2000 to 2012, a total of 136 unrelated hypertrophic cardiomyopathy probands had genetic testing, of which 63 (46%) carried at least one pathogenic mutation. MYBPC3 (n = 34; 47%) and MYH7 (n = 23; 32%) gene variants together accounted for 79%. Five variants in six probands (10%) were reclassified: two variation of uncertain significance were upgraded to pathogenic, one variation of uncertain significance and one pathogenic variant were downgraded to benign, and one pathogenic variant (found in two families) was downgraded to variation of uncertain significance. None of the reclassifications had any adverse clinical consequences.
Conclusion:
Given the rapid growth of genetic information available in both disease and normal populations, periodic reassessment of single-nucleotide variant data is essential in hypertrophic cardiomyopathy.
Genet Med 2014:16(4):286–293.
Similar content being viewed by others
Introduction
Hypertrophic cardiomyopathy (HCM) is a primary disorder of the myocardium characterized by unexplained left ventricular hypertrophy.1,2 HCM is the most common genetic cardiovascular disorder affecting 0.2% (1 in 500) of the general population.3 The clinical course of HCM shows marked heterogeneity, and can lead to progressive heart failure and sudden cardiac death.4 HCM is inherited in an autosomal dominant pattern and displays extreme genetic heterogeneity and variable degree of penetrance. Widespread application of genetic testing in HCM over the last two decades has lead to the identification of more than 1,400 mutations in approximately 13 genes, encoding proteins of the sarcomere, Z-disc and calcium regulation with varying evidence of pathogenicity.5,6,7 A subset of HCM patients (5%) are found to harbor more than one pathogenic mutation (i.e., homozygous or compound heterozygous), which may confer a more severe form of disease with a higher incidence of adverse outcomes including heart failure and sudden death.8,9,10
Genetic testing in HCM is part of routine clinical practice, and identification of a pathogenic mutation allows predictive genetic testing to be performed in at-risk relatives. Overall, the mutation detection rate for HCM genetic testing is 50–60%, although some variation exists in different cohorts.7,11,12,13 Determining whether a single-nucleotide variant (SNV) is a pathogenic disease-causing change or a benign variant of no clinical significance remains a key step in the genetic testing process. Furthermore, in some cases, the SNV is classified as a variant of uncertain significance (VUS) when data to determine pathogenicity are insufficient.
Determining pathogenicity has typically involved bringing together various forms of information about the specific SNV, to either support or refute its pathogenic role. These pathogenic factors include whether the SNV causes an amino acid substitution, is at a conserved site among species, results in a significant functional change in the protein, and is absent or rare in the normal population. Cosegregation data among families are also important, although this information is not available at the time of the first testing of the proband. Importantly, incorrect determination of pathogenicity can lead to inappropriate inclusion or exclusion of asymptomatic family members from clinical surveillance, and in the most serious setting may mean relatives are falsely reassured that they will not develop HCM.
Recent advances in next-generation sequencing techniques have facilitated the availability of large amounts of whole genome and exome data in normal populations. These technologies have also enabled testing of panels of genes in HCM.14 The increase in available genetic information and the greater number of HCM families undergoing comprehensive genetic testing may alter the classification of pathogenicity of previously described HCM mutations. This study sought to reassess SNV classification in HCM probands in the setting of recent availability of population genetic information, in a specialized multidisciplinary clinic setting.
Materials and Methods
Patient cohort
Patients referred to the HCM Centre at Royal Prince Alfred Hospital (Sydney, Australia) were clinically evaluated with full clinical history and physical examination, 12-lead electrocardiography, and 2D and M-mode transthoracic echocardiography, as previously described.5 A maximal left ventricular wall thickness of ≥15 mm on echocardiography in adults in the absence of other loading conditions was used as the primary diagnostic criterion for HCM.1 This study was carried out in strict accordance with the Sydney South West Area Health Service Human Ethics standards.
Genetic analysis
Probands who had undergone genetic testing were included in the study cohort. In those where a pathogenic mutation or VUS was identified, updated clinical and predictive genetic testing results for first-degree relatives were collected during follow-up, and an updated pedigree for each family was constructed. Genetic testing was performed in a clinical setting utilizing both research and a number of different commercial genetic testing laboratories. The panel of genes screened and the method of genetic testing varied across testing laboratories. However, all patients were screened for a panel of 7–10 of the most common HCM-associated genes (MYBPC3, MYH7, TNNI3, TNNT2, TPM, MYL2, MYL3, ACTC, ACTN2, and TCAP). The number of genes included on testing panels has increased over time to encompass genes that have been associated with HCM (CSRP3, MYOZ2, NEXN, PLN, and TTC), and genes causing HCM phenocopies, such as Fabry disease (GLA), Danon disease (LAMP2), and glycogen storage disease (PRKAG2).
Pathogenicity of variants
The pathogenic status of each variant was reassessed into three broad categories: Pathogenic mutation, VUS and benign variant. A variant was considered pathogenic based on established criteria: (i) a missense variant with an amino acid change at a highly conserved position among species, (ii) a variant altering protein structure and function e.g., insertions and deletions causing a reading frameshift, and nonsense and splice-site mutations, (iii) previously reported as a cause of HCM, (iv) cosegregation with disease in the family, and (v) absence of the variant or in <1% among unrelated and ethnically matched controls.5,7,15 VUS were those in which the effect on the protein function was not clear and disease-risk association with HCM was not known. Benign variants were considered synonymous variants that did not alter the amino acid sequence, and missense variants that were SNVs at a frequency of >1% in normal control populations.
Each variant represented an alternate allele from the reference human genome sequence, version GRCh37/hg19. The variants were evaluated for conservation level, including genomic evolutionary rate profiling,16 PhastCons,17 and PhyloP scores. Pathogenicity prediction programs that assess the functional significance of amino acid substitutions were used, including Polyphen 2,18 Grantham score,19 and Sorting intolerant from tolerant20 score. All variants were annotated using SeattleSeq single-nucleotide polymorphism annotation (http://snp.gs.washington.edu/SeattleSeqAnnotation137/), and searched in the published literature for pathogenicity and possible functional studies using PubMed and Human Genome Mutation Database public release version (http://www.hgmd.cf.ac.uk/ac/index.php).21 Occurrence and frequency was sought in the 1000 Genomes Project data,22 dbSNP build 137, and the National Heart, Lung and Blood Institute Exome Sequencing Project (ESP) version.0.0.15 (http://evs.gs.washington.edu/EVS/). A minor allele frequency of <1% was used to define a rare variant. The variants were also analyzed using Alamut software version 2.2 (Interactive Biosoftware, Rouen, France), an integrated platform to look at different predictors for pathogenicity of genetic variants.
Independent evaluation of pathogenicity
A panel of four members (J.D., J.I., R.B., and C.S.) performed an independent review of all variants using current and updated cosegregation data for each family. The panel of reviewers consisted of cardiologist, junior medical doctor, genetic counselor, and human geneticist. The four reviewers analyzed all variants in a blinded manner, and a scoring system was used based on how many reviewers were in agreement, i.e., if four reviewers all agreed, then the score was 4.
Results
Clinical and genetic characteristics of the probands
A total of 136 unrelated HCM probands underwent genetic testing. Sixty-three (46%) probands were reported to have a pathogenic mutation or VUS, of which 37 (59%) were males. The mean age at diagnosis in probands with a positive genetic result was 35 ± 17 years (range 4–68 years) and the mean left ventricular wall thickness was 22 ± 6 mm. The distribution of all reported variants in the HCM probands and initial classification are summarized in Table 1 . There were a total of 72 variations in the 63 probands, representing 54 unique variants, of which 48 were initially classified as pathogenic and 6 as VUS. The variants were mostly distributed across the sarcomere protein genes MYBPC3 (n = 34; 47%) and MYH7 (n = 23; 32%). The most common type of variant identified in MYBPC3 was missense (n = 15; 65%), followed by splice-site substitutions (n = 3; 13%), and 4 (12%) were reported as a VUS. By contrast, all MYH7 variants were missense (n = 18; 100%) and all classified as pathogenic. Multiple pathogenic mutations were reported in six probands (4%) of which five had two mutations (double mutations or compound heterozygote) and one carried a previously described triple mutation (see Supplementary Table S1 online).23
Variant reclassification
A total of 54 unique variants across seven genes were identified in the entire study cohort, of which some of the variants were represented more than once among clinically unrelated probands, e.g., MYBPC3–Arg502Trp variant was present in seven probands (11%), representing a frequently encountered pathogenic variant in the HCM population. Of variants with a discordant pathogenicity status reported by the 4 independent reviewers, 2 variants had a score of 2, and 13 variants scored 3 (see Supplementary Table S2 online for complete details). Each variant with a discordant classification of pathogenicity following review was further assessed by the combined panel to determine a consensus in the pathogenicity status, leading to the reclassification of five variants in six (10%) HCM families ( Figure 1 and Table 2 ).
Family pedigree of probands with a change in pathogenicity status. Circles = females, squares = males. Arrow = proband. Diagonal line = deceased. Black symbols = clinically affected. Open symbols = unknown clinical status. “N” = clinically normal. The results of genetic testing are denoted below the individual as +/− for heterozygous, +/+ for homozygous, and −/− for absence of the variant.
Two variants, Arg162TRp in TNNI3 and an intronic variation, IVS12-9G>A in MYBPC3 were initially reported as VUS but were upgraded to pathogenic mutations. Arg326Gln in MYBPC3 was initially classified as pathogenic in two unrelated probands and subsequently downgraded to a VUS following reassessment. Arg816Gln in MYBPC3 was initially reported as a pathogenic mutation but was reclassified as a benign variant by all the four reviewers, after evaluation of the family revealed a clinically affected HCM individual who did not carry the variant. An MYL3 c.*89G>A substitution in the 3′ untranslated region was initially reported as a VUS and reclassified as benign by three reviewers.
Cosegregation and predictive genetic testing of first-degree relatives
First-degree relatives of all 63 probands were clinically screened. Of a total of 321 living first-degree relatives, 131 (41%) underwent clinical assessment ( Table 3 ). A clinical diagnosis of HCM was made in 56 (43%) members, whereas 70 members (53%) were clinically unaffected. A diagnosis of borderline HCM (left ventricular wall thickness of 11–13 mm) was made in five (4%) of the living first-degree relatives. Pretest genetic counseling and predictive testing of first-degree relatives was performed as part of routine clinical practice. Overall, 84 (64%) first-degree relatives underwent predictive genetic testing, equating to 26% of the total living first-degree relatives. There were 21 (48%) relatives identified as clinically unaffected who had a positive-predictive gene test result (genotype-positive phenotype-negative), and confirmatory genetic testing was performed in 35 (63%) of the affected living first-degree relatives identifying the genetic variant in all but one (Arg816Gln), a pick-up rate of 97%.
Discussion
HCM is the most common genetic heart disease showing marked clinical and genetic heterogeneity.24 Accurately determining the pathogenicity of identified variants remains an important clinical challenge that can impact on diagnosis and allocation of at-risk family members who require regular clinical follow-up. In this study, five variants in six HCM probands (10%) were reclassified based on the most currently available genetic and functional information. The reclassification of variants in these six families did not result in any significant clinical implications; however, the potential to exclude individuals from clinical surveillance based on a predictive test using a misclassified pathogenic mutation remains an important consideration. The findings, coupled with the rapidly growing and readily available genomic data in human populations, support the notion of periodic reassessment of SNV data in HCM, as well as other inherited heart diseases.
The need to reassess, and in some cases reclassify, SNV data have been based largely on the enormous impact of high-throughput sequencing methods, which have provided extensive catalogs of human variation, such as the NHLBI GO ESP, and the 1000 Genomes Project. As a direct result, many variants previously described as novel and disease-causing have been found to be present in these databases indicating that some may actually be rare benign variants. Recent studies in two disease populations, familial long QT syndrome25 and familial dilated cardiomyopathy,26 identified a higher than expected prevalence of variants in the ESP data. Most recently, up to 14% of all previously HCM-associated variants and 18% of all dilated cardiomyopathy and arrhythmogenic right ventricular cardiomyopathy-associated variants were reported to be present in ESP.27 The 1000 Genomes database was also found to carry higher than expected prevalence of sarcomere gene mutations linked to HCM.28
On the basis of the growth in available human genome data, coupled with increasing knowledge of disease mechanisms, and availability of newer in silico prediction programs, reevaluation of genetic findings over the last decade in clinical settings is of importance. Six HCM families (of 63) had SNV data reclassified in our HCM cohort over a 12-year period. The clinical implications of the reclassification of variants in these six HCM families are briefly summarized in the following section.
Proband RK1 was identified to be homozygous for the Arg162Trp variant in TNNI3, which was initially reported as a VUS. RK1, a child of consanguineous parents, was diagnosed with HCM at the age of 17 years. Her brother had a resuscitated cardiac arrest, had clinical HCM, and was also found to be homozygous for Arg162Trp. Both clinically unaffected parents and two clinically unaffected siblings were found to be heterozygous for Arg162Trp. The Arg162Trp missense variant is located in the troponin domain with a prediction favoring pathogenic mutation on Polyphen2 and Sorting intolerant from tolerant and has a moderate Grantham score (101). Functional studies show that the variant can alter calcium sensitivity of the myocardium and thus may lead to HCM.29,30 This variant was therefore reclassified as a pathogenic recessive variant and represents a rare example of recessive inheritance of HCM.10
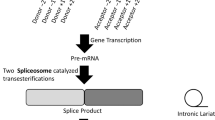
Proband US1, with mild HCM, was found to be heterozygous for a G>A substitution in intron 11 of MYBPC3, 9 bases downstream of exon 12. This previously reported variant creates a new acceptor splice site (Human splicing finder score = 87.2) stronger than the wild type (human splicing finder score = 81.8), resulting in complete skipping of exon 12 and a truncated protein.31 This intronic variant was originally reported as VUS; but on subsequent predictive testing, it was found to be present in the affected mother and also the two affected siblings and was subsequently upgraded to a pathogenic variant. Proband PI1, with severe HCM, had two reported variants on genetic testing, i.e., Glu123Gln in TNNI3 and c.*89G>A in MYL3, which were classified as a pathogenic mutation and a VUS, respectively, at diagnosis. The c.*89G>A variant is a substitution located in the 3′ untranslated region, 89 residues downstream of the termination codon, and has now been identified to have a minor allele frequency of 20% in the 1000 Genomes Project data, thus representing a common polymorphism. The currently available evidence for this variant therefore resulted in a reclassification from VUS to benign variant.
Proband NU2, with moderate HCM, was found to be heterozygous for the Arg816Gln variant in MYBPC3 and was classified as a pathogenic variant at the time of diagnosis. This novel missense variant at a conserved position was predicted to be pathogenic by Polyphen2 and Sorting intolerant from tolerant. However, an affected sibling with clinical HCM did not carry the Arg816Gln variant on predictive testing, which was repeated twice at different testing facilities to ensure no blood collection and genotyping error. Therefore, this variant was downgraded to a benign variant, and has not been used in any subsequent predictive testing. Arg816Gln variant was identified in two probands, AI1 and O1. In both the probands, the variant was initially classified as pathogenic, having been described in HCM families previously. Reassessment of this variant with current genomic data in both ESP and 100 Genomes showed a minor allele frequency of up to 0.38%, and was subsequently reclassified as a VUS. There was no impact on predictive testing or clinical surveillance in either family.
While not available at the time of proband genetic testing, and therefore at the time of variant classification, cosegregation of genetic data in the family setting is important in either supporting or excluding a pathogenic role during follow-up. The genetic testing conducted in this study was in the setting of a multidisciplinary clinic, where both clinical and genetic evaluation (of probands and at-risk relatives) is performed as part of a single clinic.32,33 Even with the best efforts at our multidisciplinary clinic, we were able to clinically screen only 131 (41%) of the relatives. The importance of cosegregation data in determining the pathogenicity of a variant was highlighted in a number of cases in this study, including the reclassification of one variant from pathogenic to benign based on the lack of segregation in a clearly affected individual with HCM (family NU). Predictive testing in HCM has also identified a number of individuals who carry the genotype but have no phenotype features of HCM. These so-called “genotype-positive phenotype-negative” patients represent a new subgroup of HCM patients which have arisen directly by the increase in volume of predictive genetic testing in HCM.34,35
On the basis of the current literature, the explosion of available genomic data in human populations, and the findings from our study, a proposed model of evaluation of genetic testing in HCM is summarized in Figure 2 , with a strong emphasis on periodic reevaluation of variants. In practical terms, determining pathogenicity of an SNV includes demonstration that the variant is novel or of rare frequency in human genome and exome databases, that the variant changes the amino acid which is highly conserved or results in a significant disruption of the protein (e.g., frameshift or nonsense mutations), and that the SNV has been previously reported as disease causing. Other predictive programs such as Polyphen or Sorting intolerant from tolerant, and other tools such as Grantham and genomic evolutionary rate profiling scores, can add supportive information for pathogenicity, as can the availability of functional data and animal models of disease. Once an SNV is classified in the proband, cosegregation studies in families are important in further supporting pathogenicity. This approach, while focused on HCM, is directly relevant to all inherited autosomal dominant heart diseases.36
Model of genetic testing and variant classification in hypertrophic cardiomyopathy (HCM) in the setting of a specialized multidisciplinary clinic. Following proband genetic testing of HCM patients attending a specialized cardiac genetic clinic, variants will be classified as radical mutations, missense variants, variation of uncertain significance (VUS), or a synonymous benign variant. Periodic reevaluation of all variants will allow incorporation of new information and appropriate management of first-degree relatives. CMR, cardiac magnetic resonance imaging; ECG, electrocardiogram.
The difficulties surrounding determining pathogenicity is likely to increase as more variants are identified with newer next-generation sequencing platforms, rather than smaller panel-based genetic testing. More VUSs will be identified. The clinical utility of a VUS is limited, and cannot be used confidently in the setting of predictive testing, although the variant may have some influence on disease pathogenesis and outcomes. The complexities in determining variant pathogenicity in the setting of genetic testing supports the role of specialized cardiac genetic clinics, where interpretation of either a positive or negative result is done in a comprehensive manner by a team of trained cardiologists, geneticists, and genetic counselors.33,37,38,39
Study limitations
This study describes the clinical experience of a multidisciplinary cardiac clinic in the application of genetic testing in HCM. Given the issues relating to high cost of genetic testing in Australia, uptake of genetic testing in HCM families has been low. The panel of genes tested and method of testing varied across testing facilities. The testing was done based on the best available technology at the time, and reflects the evolving nature and technologies of genetic testing in HCM over a period of 12 years.
Conclusion
Genetic testing plays a key role in the diagnosis and evaluation of at-risk family members in HCM. Accurately determining the pathogenicity of variants identified through genetic testing is of paramount importance in how the test is subsequently implemented in the setting of predictive testing. With the rapid growth of genetic information in human populations and the expansion of genetic testing from gene panels to whole-genome approaches, periodic reassessment of variants is essential in the ongoing management of families with HCM. This reassessment is best undertaken in the setting of a multidisciplinary clinic, where decisions can be made collectively by the attending cardiologist, geneticist, laboratory scientist, and the genetic counselor.
Disclosure
The authors declare no conflict of interest.
Change history
26 April 2018
When this article was published, the Supplementary Material was omitted. The files are now provided in the online version of the article. The publisher regrets the error.
References
Gersh BJ, Maron BJ, Bonow RO, et al.; American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines; American Association for Thoracic Surgery; American Society of Echocardiography; American Society of Nuclear Cardiology; Heart Failure Society of America; Heart Rhythm Society; Society for Cardiovascular Angiography and Interventions; Society of Thoracic Surgeons. 2011 ACCF/AHA guideline for the diagnosis and treatment of hypertrophic cardiomyopathy: executive summary: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. Circulation 2011;124:2761–2796.
Seidman JG, Seidman C . The genetic basis for cardiomyopathy: from mutation identification to mechanistic paradigms. Cell 2001;104:557–567.
Maron BJ, Gardin JM, Flack JM, Gidding SS, Kurosaki TT, Bild DE . Prevalence of hypertrophic cardiomyopathy in a general population of young adults. Echocardiographic analysis of 4111 subjects in the CARDIA Study. Coronary Artery Risk Development in (Young) Adults. Circulation 1995;92:785–789.
Maron BJ . Sudden death in young athletes. Lessons from the Hank Gathers affair. N Engl J Med 1993;329:55–57.
Chiu C, Bagnall RD, Ingles J, et al. Mutations in alpha-actinin-2 cause hypertrophic cardiomyopathy: a genome-wide analysis. J Am Coll Cardiol 2010;55:1127–1135.
Richard P, Charron P, Carrier L, et al.; EUROGENE Heart Failure Project. Hypertrophic cardiomyopathy: distribution of disease genes, spectrum of mutations, and implications for a molecular diagnosis strategy. Circulation 2003;107:2227–2232.
Maron BJ, Maron MS, Semsarian C . Genetics of hypertrophic cardiomyopathy after 20 years: clinical perspectives. J Am Coll Cardiol 2012;60:705–715.
Ingles J, Doolan A, Chiu C, Seidman J, Seidman C, Semsarian C . Compound and double mutations in patients with hypertrophic cardiomyopathy: implications for genetic testing and counselling. J Med Genet 2005;42:e59.
Maron BJ, Maron MS, Semsarian C . Double or compound sarcomere mutations in hypertrophic cardiomyopathy: a potential link to sudden death in the absence of conventional risk factors. Heart Rhythm 2012;9:57–63.
Gray B, Yeates L, Medi C, Ingles J, Semsarian C . Homozygous mutation in the cardiac troponin I gene: clinical heterogeneity in hypertrophic cardiomyopathy. Int J Cardiol 2012; e-pub ahead of print 24 December 2012.
Mörner S, Richard P, Kazzam E, et al. Identification of the genotypes causing hypertrophic cardiomyopathy in northern Sweden. J Mol Cell Cardiol 2003;35:841–849.
Erdmann J, Daehmlow S, Wischke S, et al. Mutation spectrum in a large cohort of unrelated consecutive patients with hypertrophic cardiomyopathy. Clin Genet 2003;64:339–349.
Van Driest SL, Ommen SR, Tajik AJ, Gersh BJ, Ackerman MJ . Yield of genetic testing in hypertrophic cardiomyopathy. Mayo Clin Proc 2005;80:739–744.
Meder B, Haas J, Keller A, et al. Targeted next-generation sequencing for the molecular genetic diagnostics of cardiomyopathies. Circ Cardiovasc Genet 2011;4:110–122.
Richards CS, Bale S, Bellissimo DB, et al.; Molecular Subcommittee of the ACMG Laboratory Quality Assurance Committee. ACMG recommendations for standards for interpretation and reporting of sequence variations: revisions 2007. Genet Med 2008;10:294–300.
Cooper GM, Goode DL, Ng SB, et al. Single-nucleotide evolutionary constraint scores highlight disease-causing mutations. Nat Methods 2010;7:250–251.
Siepel A, Bejerano G, Pedersen JS, et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 2005;15:1034–1050.
Adzhubei IA, Schmidt S, Peshkin L, et al. A method and server for predicting damaging missense mutations. Nat Methods 2010;7:248–249.
Grantham R . Amino acid difference formula to help explain protein evolution. Science 1974;185:862–864.
Ng PC, Henikoff S . Predicting deleterious amino acid substitutions. Genome Res 2001;11:863–874.
Stenson PD, Mort M, Ball EV, et al. The Human Gene Mutation Database: 2008 update. Genome Med 2009;1:13.
Abecasis GR, Auton A, Brooks LD, et al. An integrated map of genetic variation from 1,092 human genomes. Nature 2012;491:56–65
Ingles J, Sarina T, Yeates L, et al. Clinical predictors of genetic testing outcomes in hypertrophic cardiomyopathy. Genet Med 2013; 15: 972–977.
Maron BJ . Hypertrophic cardiomyopathy: a systematic review. JAMA 2002;287:1308–1320.
Refsgaard L, Holst AG, Sadjadieh G, Haunsø S, Nielsen JB, Olesen MS . High prevalence of genetic variants previously associated with LQT syndrome in new exome data. Eur J Hum Genet 2012;20:905–908.
Norton N, Robertson PD, Rieder MJ, et al.; National Heart, Lung and Blood Institute GO Exome Sequencing Project. Evaluating pathogenicity of rare variants from dilated cardiomyopathy in the exome era. Circ Cardiovasc Genet 2012;5:167–174.
Andreasen C, Nielsen JB, Refsgaard L, et al. New population-based exome data are questioning the pathogenicity of previously cardiomyopathy-associated genetic variants. Eur J Hum Genet 2013;21:918–928.
Golbus JR, Puckelwartz MJ, Fahrenbach JP, Dellefave-Castillo LM, Wolfgeher D, McNally EM . Population-based variation in cardiomyopathy genes. Circ Cardiovasc Genet 2012;5:391–399.
Takahashi-Yanaga F, Morimoto S, Harada K, et al. Functional consequences of the mutations in human cardiac troponin I gene found in familial hypertrophic cardiomyopathy. J Mol Cell Cardiol 2001;33:2095–2107.
Elliott K, Watkins H, Redwood CS . Altered regulatory properties of human cardiac troponin I mutants that cause hypertrophic cardiomyopathy. J Biol Chem 2000;275:22069–22074.
Rodríguez-García MI, Monserrat L, Ortiz M, et al. Screening mutations in myosin binding protein C3 gene in a cohort of patients with Hypertrophic Cardiomyopathy. BMC Med Genet 2010;11:67.
Ingles J, Lind JM, Phongsavan P, Semsarian C . Psychosocial impact of specialized cardiac genetic clinics for hypertrophic cardiomyopathy. Genet Med 2008;10:117–120.
Ingles J, Yeates L, Semsarian C . The emerging role of the cardiac genetic counselor. Heart Rhythm 2011;8:1958–1962.
Maron BJ, Yeates L, Semsarian C . Clinical challenges of genotype positive (+)-phenotype negative (-) family members in hypertrophic cardiomyopathy. Am J Cardiol 2011;107:604–608.
Maron BJ, Semsarian C . Emergence of gene mutation carriers and the expanding disease spectrum of hypertrophic cardiomyopathy. Eur Heart J 2010;31:1551–1553.
Ackerman MJ, Priori SG, Willems S, et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies this document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Heart Rhythm 2011;8:1308–1339.
Ingles J, Semsarian C . Sudden cardiac death in the young: a clinical genetic approach. Intern Med J 2007;37:32–37.
Hershberger RE, Lindenfeld J, Mestroni L, Seidman CE, Taylor MR, Towbin JA ; Heart Failure Society of America. Genetic evaluation of cardiomyopathy–a Heart Failure Society of America practice guideline. J Card Fail 2009;15:83–97.
Christiaans I, Birnie E, Bonsel GJ, Wilde AA, van Langen IM . Uptake of genetic counselling and predictive DNA testing in hypertrophic cardiomyopathy. Eur J Hum Genet 2008;16:1201–1207.
Acknowledgements
J.D. has received an Australian Prime Minister’s Endeavour Award Scholarship. J.I. is the recipient of an cofunded National Heart Foundation and National Health and Medical Research Council (NHMRC) Early Career Fellowship. C.S. is the recipient of a NHMRC Practitioner Fellowship. This study was also supported, in part, by an NHMRC project grant.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Das K, J., Ingles, J., Bagnall, R. et al. Determining pathogenicity of genetic variants in hypertrophic cardiomyopathy: importance of periodic reassessment. Genet Med 16, 286–293 (2014). https://doi.org/10.1038/gim.2013.138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2013.138
Keywords
This article is cited by
-
Protein Thermodynamic Destabilization in the Assessment of Pathogenicity of a Variant of Uncertain Significance in Cardiac Myosin Binding Protein C
Journal of Cardiovascular Translational Research (2020)
-
Development of a communication aid for explaining hypertrophic cardiomyopathy genetic test results
Pilot and Feasibility Studies (2017)