Abstract
Purpose: The advent of molecular cytogenetic technologies has altered the means by which new microdeletion syndromes are identified. Whereas the cytogenetic basis of microdeletion syndromes has traditionally depended on the serendipitous ascertainment of a patient with established clinical features and a chromosomal rearrangement visible by G-banding, comparative genomic hybridization using microarrays has enabled the identification of novel, recurrent imbalances in patients with mental retardation and apparently nonspecific features. Compared with the “phenotype-first” approach of traditional cytogenetics, array-based comparative genomic hybridization has enabled the detection of novel genomic disorders using a “genotype-first” approach. We report as an illustrative example the characterization of a novel microdeletion syndrome of 1q41q42.
Methods: We tested more than 10,000 patients with developmental disabilities by array-based comparative genomic hybridization using our targeted microarray. High-resolution microarray analysis was performed using oligonucleotide microarrays for patients in whom deletions of 1q41q42 were identified. Fluorescence in situ hybridization was performed to confirm all 1q deletions in the patients and to exclude deletions or other chromosomal rearrangements in the parents.
Results: Seven cases were found with de novo deletions of 1q41q42. The smallest region of overlap is 1.17 Mb and encompasses five genes, including DISP1, a gene involved in the sonic hedgehog signaling pathway, the deletion of which has been implicated in holoprosencephaly in mice. Although none of these patients showed frank holoprosencephaly, many had other midline defects (cleft palate, diaphragmatic hernia), seizures, and mental retardation or developmental delay. Dysmorphic features are present in all patients at varying degrees. Some patients showed more severe phenotypes and carry the clinical diagnosis of Fryns syndrome.
Conclusions: This new microdeletion syndrome with its variable clinical presentation may be responsible for a proportion of Fryns syndrome patients and adds to the increasing number of new syndromes identified with array-based comparative genomic hybridization. The genotype-first approach to identifying recurrent chromosome abnormalities is contrasted with the traditional phenotype-first approach. Targeting developmental pathways in a functional approach to diagnostics may lead to the identification of additional microdeletion syndromes.
Similar content being viewed by others
Main
In 1963, Black and Carter1 posited that the “elfin”-like facies characteristic of two clinical entities, supravalvular aortic stenosis and infantile hypercalcemia, suggested that the two disorders may be related. Nine years later, Beuren2 demonstrated that supravalvular aortic stenosis and idiopathic infantile hypercalcemia, were, in fact, components of the same disorder, which might present with or without mental retardation. However, the genetic basis of the disorder, now commonly referred to as Williams-Beuren syndrome, remained unknown for almost 30 years. During those three decades, numerous loci were erroneously suggested as causative for the phenotype based on single case reports.3–8 It was not until the finding of a translocation segregating with the supravalvular aortic stenosis phenotype and cloning of the breakpoints that the cytogenetic basis for Williams-Beuren syndrome was uncovered.9
Historically, patients with microdeletion syndromes were identified by a constellation of key clinical features that had evolved from the ascertainment of large collections of individuals with similar abnormalities. The cytogenetic bases of these syndromes was not known. The advent of G-banding and other chromosomal banding techniques in the 1970s allowed the identification of the alternating light- and dark-staining bands comprising each human chromosome, facilitating the detection of aneuploidies and large structural rearrangements such as deletions, duplications, and translocations. Thus, the cytogenetic basis of many syndromes was delineated. For example, although Langer-Giedion syndrome (LGS) was characterized by a recognizable phenotype as early as 1969,10 its cytogenetic basis was unknown until high-resolution banding identified interstitial deletions in the long arm of chromosome 8 in patients with LGS11,12; the location of the deletion was subsequently determined to be 8q24.1.13,14 Mapping of the deletion breakpoints in a cohort of LGS patients showed that 75% have cytogenetically detectable deletions of 8q24.1.15
The identification of patients with unbalanced translocations may facilitate the detection of submicroscopic deletions. This approach requires that the deleterious effects of each component chromosome of the unbalanced translocation be parsed. The association of DiGeorge syndrome with an interstitial deletion of the long arm of chromosome 22 was suggested by de la Chapelle and colleagues16 following the identification of an unbalanced translocation resulting in a deleted chromosome 22 and comparison with published reports of monosomy 22. High-resolution G-banding and fluorescence in situ hybridization (FISH) in a large collection of patients later confirmed this hypothesis; more than 95% of patients have deletions of 22q11.2.17,18
Much like those with unbalanced translocations, patients with balanced translocations have helped identify novel microdeletions; typically, these deletions were too small to be easily visualized under the light microscope and necessitated the serendipitous ascertainment of a patient with a translocation segregating the deletion and recognizable clinical features. For example, Shaffer et al.19 reported two family members with mental retardation and multiple congenital anomalies including multiple exostoses, enlarged parietal foramina, and craniofacial dysostosis. Karyotyping of the mother revealed an insertional translocation between chromosomes 11 and 13. Affected individuals inherited a deletion of 11p11.2 following malsegregation of the insertional translocation.19 Additional patients were subsequently described with del(11)(p11.2p12) and multiple exostoses,20,21 confirming that this constellation of features constituted a syndrome. In another example, three cases of Rubinstein-Taybi syndrome (RTS), a disorder consisting of mental retardation, characteristic facial features, and broad thumbs and toes, presented with balanced translocations involving chromosome 16 and 2, 7, or 20.22 The breakpoint on chromosome 16 occurred in band 16p13.3, suggesting this as a location for the RTS locus. Subsequent analysis using a cosmid mapping to 16p13.3 of a cohort of 24 RTS patients showed microdeletions of this region in six cases, confirming that a substantial proportion of RTS result from microdeletions.22
In 1995, Flint et al.23 developed a method for simultaneously interrogating all the unique human subtelomeres using FISH. These probe sets24,25 could be used to detect submicroscopic chromosomal abnormalities in patients with idiopathic mental retardation but without features suggestive of a particular syndrome. Some of the abnormalities initially detected with subtelomere FISH have since been identified in a sufficiently large number of patients that the abnormality could be conclusively linked to the phenotype, which could in turn be delineated by comparison of clinical features among affected individuals. For example, one of the first syndromes discovered using subtelomere FISH resulted from deletion of 22q13.326; once a common cause was established among a collection of patients, the clinical features could be delineated.27 Other subtelomeric abnormalities have been identified in only one or a few patients and cannot be considered recurrent until a consistent collection of clinical features is delineated.
The use of subtelomere FISH panels has illustrated that, in the absence of specific clinical features suggestive of a syndrome, patients with mental retardation can be “screened” for a novel chromosomal abnormality. This development represented a shift from the traditional “phenotype-first” approach explained earlier, wherein a set of patients was grouped based on shared clinical features, to a “genotype-first” approach by which individuals can be characterized first by a common cytogenetic aberration and then as more patients with the same abnormality are ascertained, a clinical presentation can be delineated.
The development of comparative genomic hybridization (CGH), particularly CGH using microarrays (array CGH), broadened the scope and resolution at which the genotypes of patients with idiopathic mental retardation could be assayed. In contrast to subtelomere FISH, microarrays may be constructed with contiguous or noncontiguous coverage of the entire genome or with consideration of well-known microdeletion and microduplication syndrome loci; thus, the principles of subtelomere FISH panels could be applied to a much larger proportion of the genome. For example, in screening patients with unexplained overgrowth syndrome by whole-genome array CGH, Redon et al.28 identified two individuals with de novo interstitial deletions of 9q22.32-q22.33. Although the breakpoints differed in each patient, the similarity of clinical features—both individuals presented with macrocephaly, overgrowth, trigonocephaly, and hyperactivity in addition to a constellation of distinct facial features—led the authors to suggest that deletion of 9q22.3 was a novel microdeletion syndrome.
In another example, three simultaneously published reports detailed the identification of a microdeletion syndrome encompassing 17q21.3.29–31 In one of these reports, Sharp et al.30 constructed a microarray targeted to potential “rearrangement hotspots” that, because of their genomic architecture containing segmental duplications, were likely to be affected by genomic instability through nonallelic homologous recombination.32 Through the screening of 290 children with idiopathic mental retardation with or without associated congenital anomalies, Sharp et al.30 identified de novo microdeletions of 17q21.31 in four individuals. In addition, the authors identified one patient with a de novo microdeletion of 15q24; three more patients with submicroscopic deletions of this region were subsequently identified by array CGH using either targeted or whole-genome BAC or oligonucleotide microarrays.33
In contrast to the Sharp et al.30 publication, which targeted regions associated with segmental duplications, Ballif et al.34 targeted the pericentromeric regions of the genome. These regions are inherently difficult to assess by chromosome analysis and are known to harbor microdeletions, including Williams, DiGeorge, and Prader-Willi syndromes. Array CGH analysis identified inherited copy-number variants (benign polymorphisms) and de novo deletions and duplications with potential clinical significance, including a recurrent de novo interstitial deletion of 16p11.2p12.2 in four patients. FISH and whole-genome microarray analysis of these four deletions showed that the breakpoints cluster at complex segmental duplications that flank the deletion region consistent with nonallelic homologous recombination mediating these rearrangements. The common clinical features of these patients suggested that deletion of 16p11.2p12.2 constitutes a novel microdeletion syndrome.
We have taken a directed, functional approach to identify individuals with novel microdeletions. In developing a targeted microarray for use with CGH,35,36 we included regions commonly rearranged in chromosome abnormalities including microdeletion syndromes and the subtelomeric regions. In addition, to increase our coverage over the genome in a deliberate manner, we included genes in important developmental pathways, such as PTCH, ZIC2, and DISP1, which are involved in the sonic hedgehog (SHH) signaling pathway, the disruption of which has been implicated in holoprosencephaly (HPE). This microarray has been used to identify chromosome abnormalities in patients with developmental delay, mental retardation, seizures, and other developmental abnormalities36,37 and has led to the identification of recurrent de novo deletions of 1q41q42 that include the DISP1 locus.
MATERIALS AND METHODS
Subjects
At the time of this writing, more than 10,000 peripheral blood samples have been analyzed by our diagnostic laboratory (Signature Genomic Laboratories, LLC, Spokane, WA) by array CGH using the SignatureChip.37 These samples were received from referring physicians (medical geneticists, pediatric neurologists, neonatologists, and general pediatricians) from the United States and abroad. The indications for referral were the same as those received for routine cytogenetic testing including developmental delay, mental retardation, dysmorphic features, failure to thrive, seizures, other developmental disabilities, and physical anomalies. Array CGH results for our first 1500 consecutive cases were recently published36 and the results for a total case volume of 8789 are in press.37
Array CGH and FISH
All samples were tested by CGH with the SignatureChip diagnostic microarray using previously published methods.35,36 There have been four versions of our microarray, but all have included bacterial artificial chromosomes (BACs) that contain the HPE or HPE-candidate loci of interest to this report. For this study, FISH was performed with published methods38 using BAC RP11-139E20 containing the DISP1 gene to confirm all 1q deletions in the subjects and to exclude deletions or other chromosomal rearrangements in the parents.
High-resolution microarray studies
Informed consent using a Washington State University Institutional Review Board–approved protocol and consent form was obtained through the referring physicians from parents of children in whom 1q41q42 deletions were identified, except for Subject 2 for whom informed consent was obtained using a Children's Hospital of Philadelphia Institutional Review Board–approved protocol and consent form. DNA samples were then sent to Children's Hospital of Philadelphia for high-resolution analysis. The microarray experiments were performed using the Affymetrix GeneChip 250K Sty chip (Affymetrix, Santa Clara, CA), as previously described.39
RESULTS
Approximately 7% of samples submitted for array CGH in our laboratory were found to have clinically relevant chromosome abnormalities.36,37 Of these abnormal results, seven cases were identified with microdeletions of 1q41q42 that included the DISP1 locus (Fig. 1; Table 1). In all cases, FISH was used to confirm the presence of a heterozygous deletion of the region. Parental analyses in all families demonstrated two copies of the DISP1 locus; thus, these deletions of 1q41q42 are de novo in all tested cases. All seven deletions were identified in our laboratory, although one case (Subject 4) was previously published as a surviving Fryns syndrome patient with a normal karyotype.40 The clinical features of another case (Subject 7) were published after deletion identification.41
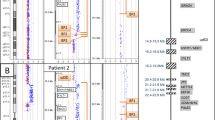
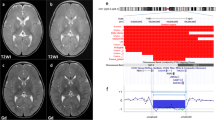
Results for chromosome 1 using a targeted microarray. Each clone on the plot is arranged along the x-axis according to its location on the chromosome with the most distal 1p telomeric clones on the left and the most distal/telomeric 1q clones on the right. The dark blue line represents the control:subject fluorescence intensity ratios for each clone, whereas the pink line represents the fluorescence intensity ratios obtained from a second hybridization in which the dyes have been reversed (subject:control). (A) Normal plot for chromosome 1. Note that all data points are at a log2 ratio of zero. (B) Plot for Subject 2 showing a deletion from BAC RP11-1031M6 through BAC RP11-61M2. (C) Plot for Subject 3 showing a deletion from BAC RP11-1148E24 through BAC RP11-61M2. (D) Plot for Subject 1 showing a deletion from BAC RP11-208F18 through RP11-61M2. For each of the plots B, C, and D, the deletion is identified as a mirror image deviation from a log2 ratio of zero to a log2 ratio of 0.3 to −0.3.
The 1q41q42 deletions were analyzed further using the Affymetrix 250K SNP array platform to (1) determine the full extent of the deletions at a higher resolution and (2) define the smallest region of overlap (SRO) to identify candidate genes. Table 1 and Figure 2 show the results of this higher resolution analysis. Subject 6 has the largest deletion, which is ∼9.07 Mb in size. The deletion in Subject 3 is the smallest at 2.72 Mb. Subject 7 was not tested by high-resolution analysis.
Results for chromosome 1 using a high-density oligonucleotide microarray. Copy number data for a selected region of chromosome 1 (210,000,000–230,000,000, NCBI build 35) is shown for Subject 1(A), Subject 2 (B), and Subject 3 (C). The plots show the signal intensity ratio (log2 ratio) of each probe on the Affymetrix 250K Sty chips resulting from analysis with CNAG software.42 Dots represent raw log2 ratio values for each SNP. Lines represent copy number inferences based on local mean analysis for five consecutive single nucleotide polymorphisms (SNPs). Heterozygous SNP calls are shown as bars below the chromosome. For probes that are a normal copy number, the signal intensity ratio of the subject versus controls is expected to be 1 and log2 ratio should be around 0.0 (log21 = 0). The deletions detected in Subjects 1 to 3 based on log2 ratio are underlined. Loss of copy number due to deletion in the subjects results in a negative log2 ratio (mean log2 ratio ∼−0.5).
The SRO between the seven deletions is ∼1.17 Mb (Chr1:219486921–220657758) (Fig. 3). There are five genes within the SRO (Table 2), based on the UCSC known genes track (http://genome.ucsc.edu; NCBI build35, May 2004), and four of these five genes have known functions.
Breakpoint locations for subjects with deletions of 1q41q42. The chromosome bands of the 1q42 region are shown with the distance from the 1p telomere in Mb along the top (based on the UCSC May 2004 draft of the human genome). Yellow bars indicate the deleted regions for each patient based on the array CGH data of Table 1. Gray bars for Subject 7 indicate the regions containing the breakpoints based on the data in the publication by Kantarci et al.41 The region shaded in gray indicates the smallest region of overlap (SRO) defined by the breakpoints in these subjects and contains the DISP1 locus.
The common clinical features found in our seven subjects are shown in Table 3. Figure 4 shows a composite of five subjects identified to have deletions of 1q41q42. Although none of the subjects has frank HPE, they show some common clinical features including significant developmental delay and distinct facial dysmorphism (frontal bossing, deep-set eyes, broad nasal tip, depressed nasal bridge, anteverted nares). Some subjects showed coarse facies in infancy, microcephaly, cleft palate, clubfeet, seizures, and short stature. Two subjects (Subjects 4 and 7) had diaphragmatic hernia and lung hypoplasia and the clinical diagnosis of Fryns syndrome.40,41
DISCUSSION
Traditional cytogenetic approaches to syndrome detection rely on the acumen—and, with many rare novel syndromes, luck—of the physician and cytogeneticist. Because of the relative rarity of some of these novel syndromes, it is difficult for one physician to ascertain sufficient patients with similar phenotypes to draw meaningful conclusions about a pattern of concurrent clinical features. This assumes that the clinical features are distinctive enough to establish a phenotypic relationship among patients. Thus, the physician or cytogeneticist is limited by human subjectivity in discerning subtle gradations of clinical presentations. The phenotypic overlap between distinct syndromes may also confound diagnoses; many cases of monosomy 1p36, for example, were initially diagnosed as Prader-Willi syndrome because of shared manifestations.43 Furthermore, basing the clinical picture on a single index case may also mask unappreciated or unknown variability in the phenotype.
High-resolution molecular cytogenetic technologies such as array CGH have altered the means by which new microdeletion syndromes are discovered by introducing an objective means of collecting a cohort of patients. Thus, a common phenotype among a group of patients may be delineated only after a common genotype has been isolated. The subjects in our study show common clinical features including significant developmental delay and distinct facial dysmorphism (frontal bossing, deep-set eyes, broad nasal tip, depressed nasal bridge, anteverted nares) (Fig. 4). Some subjects showed coarse facies in infancy, microcephaly, cleft palate, clubfeet, seizures, and short stature. Two subjects (Subjects 4 and 7) had diaphragmatic hernia and lung hypoplasia and the clinical diagnosis of Fryns syndrome.40,41
Deletions of 1q41q42 have been infrequently reported in the literature41,44–46; these cases showed a similar constellation of clinical features as the cases reported here, such as congenital diaphragmatic hernia41,44 and possible Fryns syndrome.41 The patient reported by Slavotinek et al.44 had a 10-Mb deletion of 1q41q42 that includes our SRO and the DISP1 gene (Table 1). This patient had diaphragmatic hernia, pulmonary hypoplasia, heart defects, bilateral cleft lip and palate, facial dysmorphism, and clubfeet.44 Given the midline defects found in Fryns syndrome (diaphragmatic hernia, orofacial clefting), limb malformations (nail and finger hypoplasia), central nervous system anomalies (seizures, agenesis of the corpus callosum), and dysmorphic features (coarse facies, anteverted nares, hypertelorism)47 and the findings of 1q41q42 abnormalities in some Fryns patients (Patient 4 in this study40; Subject 7 in this study27,41), we propose that deletion of 1q41q42 has a variable presentation, with the extreme end of the spectrum demonstrating a Fryns syndrome phenotype.
The common features found in these cases support our hypothesis that deletion of 1q41q42 is a new microdeletion syndrome. However, in extrapolating a common set of features from a small cohort of patients, one must be wary of confirmation bias, by which characteristics that confirm a hypothetical phenotype are selected over outliers. The recent report of a novel genomic disorder on 2p15 illustrates the inherent difficulty in delineating a phenotype in a small sample of cases.48 To illustrate, Subject 4 in our cohort has a Fryns syndrome phenotype but also one of the largest deletions (∼8.5 Mb) among our subjects (Fig. 3). The clinical variability that we observed may be attributed to either variable penetrance due to haploinsufficiency of a single gene in the SRO, modifiers within the larger deletion or elsewhere in the genome, or haploinsufficiency of genes outside the SRO, shared by these Fryns syndrome subjects. Repeated occurrences of the same alteration will be necessary to clarify the phenotype.
Although our study illustrates the difficulty in establishing a common phenotype in a small cohort of patients after the identification of a common genotype, it also highlights the inherent limitations of traditional phenotype-first cytogenetic disease identification. Whereas previous investigations of HPE candidate loci focused on patients with varying degrees of HPE and apparently normal karyotypes,49–51 we tested individuals with a variety of phenotypes who did not necessarily present with HPE. The role that these genes play in the SHH signaling pathway, however, suggested that they have critical functions in development and are logical targets for investigation. Surprisingly, although many of our patients had other midline defects (cleft palate, diaphragmatic hernia), seizures, distinct dysmorphic features, and mental retardation or developmental delay, none had frank HPE, as might be expected from deletion of genes involved in the SHH pathway. However, because none of these patients has HPE, subjects such as these would not have been enrolled in any of the previous HPE screening studies,49–51 preventing their identification using traditional means.
Although we have five candidate genes in the SRO (Table 2), we propose that haploinsufficiency of DISP1 is a reasonable cause for some of the developmental defects found in this deletion syndrome. DISP1 is essential in the SHH signaling pathway52,53 and belongs to a family of 12-pass transmembrane proteins that all have a sterol-sensing domain consisting of about 180 amino acids that form five membrane-spanning domains.54 This sterol-sensing domain is shared with other family members including Ptch1 and several proteins that regulate cholesterol biosynthesis or trafficking.55–57 A mouse knockout of Disp1 shows forebrain defects consistent with HPE.58 The function of DISP1 is conserved from Drosophila to mouse, making it likely to be important for normal development of the brain and limbs.57 Given the evidence that DISP1 is involved in the SHH signaling pathway, haploinsufficiency through mutations or deletions may be expected to cause HPE or other malformations such as midline defects or limb abnormalities. Cognitive deficits, developmental delay, or mental retardation would also be expected.
We have used a functional approach, using knowledge of important developmental pathways, such as the SHH signaling pathway, to target developmentally important genes in children with developmental delay, mental retardation, seizure, and dysmorphism. Our analysis of more than 10,000 clinical cases yielded seven patients with deletion of 1q41q42, a region containing the DISP1 gene. The variable phenotypes suggest that deletion of the SRO is sufficient for the developmental delay, mental retardation, seizures, and dysmorphic features. Larger deletions result in cleft palate, clubfeet, and, in two cases, diaphragmatic hernia. Identification and analysis of additional cases of deletion 1q41q42 or mutations in DISP1 should prove useful for delineating the genes responsible for the various features of this syndrome and will help define whether this is a contiguous gene syndrome or a single gene disorder with variability in phenotypic presentation.
SUMMARY
History shows that banded chromosomes have been important in the identification of the chromosomal basis of some syndromes (e.g., Prader-Willi syndrome,59,60 Williams-Beuren syndrome 9,61 DiGeorge syndrome16,62) and in the identification of new syndromes (e.g., Potocki-Shaffer syndrome,19 monosomy 1p3663). However, it took many decades to discover the etiology of most well-known microdeletion syndromes. Since the advent of array CGH as a routine research and diagnostic tool in the past several years, numerous new syndromes have been identified,28–31,33,34,48,64 and the molecular basis for some known syndromes has been uncovered.65–68 The discovery of novel deletions in multiple individuals allows for delineation of critical regions in which to search for genes causing the features of the syndrome. In some cases, true contiguous gene syndromes are identified (e.g., Potocki-Shaffer syndrome,20,21 LGS69), whereas in others, a single gene results in the complex phenotype (e.g., Alagille syndrome70). We anticipate that large-scale molecular testing methods such as array CGH will continue to play an important role in identifying new microdeletion syndromes and in delineating the critical regions in which to search for phenotype-contributing genes.
References
Black JA, Carter RE . association between aortic stenosis and facies of severe infantile hypercalcaemia. Lancet 1963; 91: 745–749.
Beuren AJ . Supravalvular aortic stenosis: a complex syndrome with and without mental retardation. Birth Defects Orig Artic Ser 1972; 8: 45–56.
Miles JH, Michalski KA . Familial 15q12 duplication associated with Williams phenotype. Am J Hum Genet 1983; 35: 144A.
Kaplan LC, Wharton R, Elias E, Mandell F, et al. Clinical heterogeneity associated with deletions in the long arm of chromosome 15: report of 3 new cases and their possible genetic significance. Am J Med Genet 1987; 28: 45–53.
Jefferson RD, Burn J, Gaunt KL, Hunter S, et al. A terminal deletion of the long arm of chromosome 4 [46, XX, del(4)(q33)] in an infant with phenotypic features of Williams syndrome. J Med Genet 1986; 23: 474–477.
Bzduch V, Lukacova M . Interstitial deletion of the long arm of chromosome 6(q22.2q23) in a boy with phenotypic features of Williams syndrome. Clin Genet 1989; 35: 230–231.
Colley A, Thakker Y, Ward H, Donnai D . Unbalanced 13;18 translocation and Williams syndrome. J Med Genet 1992; 29: 63–65.
Tupler R, Maraschio P, Gerardo A, Mainieri R, et al. A complex chromosome rearrangement with 10 breakpoints: tentative assignment of the locus for Williams syndrome to 4q33-q35.1. J Med Genet 1992; 29: 253–255.
Ewart AK, Morris CA, Atkinson D, Jin W, et al. Hemizygosity at the elastin locus in a developmental disorder, Williams syndrome. Nat Genet 1993; 5: 11–16.
Hall BD, Langer LO, Giedion A, Smith DW, et al. Langer-Giedion syndrome. Birth Defects Orig Artic Ser 1974; 10: 147–164.
Buhler EM, Buhler UK, Stalder GR, Jani L, et al. Chromosome deletion and multiple cartilaginous exostoses. Eur J Pediatr 1980; 133: 163–166.
Pfeiffer RA . Langer-Giedion syndrome and additional congenital malformations with interstitial deletion of the long arm of chromosome 8 46, XY, del 8 (q13-22). Clin Genet 1980; 18: 142–146.
Langer LO Jr, Krassikoff N, Laxova R, Scheer-Williams M, et al. The tricho-rhino-phalangeal syndrome with exostoses (or Langer-Giedion syndrome): four additional patients without mental retardation and review of the literature. Am J Med Genet 1984; 19: 81–112.
Bowen P, Biederman B, Hoo JJ . The critical segment for the Langer-Giedion syndrome: 8q24.11-q24.12. Ann Genet 1985; 28: 224–227.
Ludecke HJ, Johnson C, Wagner MJ, Wells DE, et al. Molecular definition of the shortest region of deletion overlap in the Langer-Giedion syndrome. Am J Hum Genet 1991; 49: 1197–1206.
de la Chapelle A, Herva R, Koivisto M, Aula P . A deletion in chromosome 22 can cause DiGeorge syndrome. Hum Genet 1981; 57: 253–256.
Carey AH, Kelly D, Halford S, Wadey R, et al. Molecular genetic study of the frequency of monosomy 22q11 in DiGeorge syndrome. Am J Hum Genet 1992; 51: 964–970.
Driscoll DA, Budarf ML, Emanuel BS . A genetic etiology for DiGeorge syndrome: consistent deletions and microdeletions of 22q11. Am J Hum Genet 1992; 50: 924–933.
Shaffer LG, Hecht JT, Ledbetter DH, Greenberg F . Familial interstitial deletion 11(p11.12p12) associated with parietal foramina, brachymicrocephaly, and mental retardation. Am J Med Genet 1993; 45: 581–583.
Bartsch O, Wuyts W, Van Hul W, Hecht JT, et al. Delineation of a contiguous gene syndrome with multiple exostoses, enlarged parietal foramina, craniofacial dysostosis, and mental retardation, caused by deletions in the short arm of chromosome 11. Am J Hum Genet 1996; 58: 734–742.
Potocki L, Shaffer LG . Interstitial deletion of 11(p11.2p12): a newly described contiguous gene deletion syndrome involving the gene for hereditary multiple exostoses (EXT2). Am J Med Genet 1996; 62: 319–325.
Breuning MH, Dauwerse HG, Fugazza G, Saris JJ, et al. Rubinstein-Taybi syndrome caused by submicroscopic deletions within 16p13.3. Am J Hum Genet 1993; 52: 249–254.
Flint J, Wilkie AO, Buckle VJ, Winter RM, et al. The detection of subtelomeric chromosomal rearrangements in idiopathic mental retardation. Nat Genet 1995; 9: 132–140.
A complete set of human telomeric probes and their clinical application. National Institutes of Health and Institute of Molecular Medicine collaboration. Nat Genet 1996; 14: 86–89.
Knight SJ, Horsley SW, Regan R, Lawrie NM, et al. Development and clinical application of an innovative fluorescence in situ hybridization technique which detects submicroscopic rearrangements involving telomeres. Eur J Hum Genet 1997; 5: 1–8.
Ning Y, Rosenberg M, Biesecker LG, Ledbetter DH . Isolation of the human chromosome 22q telomere and its application to detection of cryptic chromosomal abnormalities. Hum Genet 1996; 97: 765–769.
Phelan MC, Rogers RC, Saul RA, Stapleton GA, et al. 22q13 deletion syndrome. Am J Med Genet 2001; 101: 91–99.
Redon R, Baujat G, Sanlaville D, Le Merrer M, et al. Interstitial 9q22.3 microdeletion: clinical and molecular characterisation of a newly recognised overgrowth syndrome. Eur J Hum Genet 2006; 14: 759–767.
Koolen DA, Vissers LE, Pfundt R, de Leeuw N, et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet 2006; 38: 999–1001.
Sharp AJ, Hansen S, Selzer RR, Cheng Z, et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 2006; 38: 1038–1042.
Shaw-Smith C, Pittman AM, Willatt L, Martin H, et al. Microdeletion encompassing MAPT at chromosome 17q21.3 is associated with developmental delay and learning disability. Nat Genet 2006; 38: 1032–1037.
Stankiewicz P, Lupski JR . Genome architecture, rearrangements and genomic disorders. Trends Genet 2002; 18: 74–82.
Sharp AJ, Selzer RR, Veltman JA, Gimelli S, et al. Characterization of a recurrent 15q24 microdeletion syndrome. Hum Mol Genet 2007; 16: 567–572.
Ballif BC, Hornor SA, Jenkins E, Madan-Khetarpal S, et al. Discovery of a previously unrecognized microdeletion syndrome of 16p11.2p12.2. Nat Genet. In press.
Bejjani BA, Saleki R, Ballif BC, Rorem EA, et al. Use of targeted array-based CGH for the clinical diagnosis of chromosomal imbalance: is less more?. Am J Med Genet A 2005; 134: 259–267.
Shaffer LG, Kashork CD, Saleki R, Rorem E, et al. Targeted genomic microarray analysis for identification of chromosome abnormalities in 1500 consecutive clinical cases. J Pediatr 2006; 149: 98–102.
Shaffer LG, Bejjani BA, Torchia B, Kirkpatrick S, et al. The identification of microdeletion syndromes and other chromosome abnormalities: cytogenetic methods of the past, new technologies for the future. Am J Med Genet C. In press.
Shaffer LG, McCaskill C, Han JY, Choo KH, et al. Molecular characterization of de novo secondary trisomy 13. Am J Hum Genet 1994; 55: 968–974.
Ming JE, Geiger E, James AC, Ciprero KL, et al. Rapid detection of submicroscopic chromosomal rearrangements in children with multiple congenital anomalies using high density oligonucleotide arrays. Hum Mutat 2006; 27: 467–473.
Van Hove JL, Spiridigliozzi GA, Heinz R, McConkie-Rosell A, et al. Fryns syndrome survivors and neurologic outcome. Am J Med Genet 1995; 59: 334–340.
Kantarci S, Casavant D, Prada C, Russell M, et al. Findings from aCGH in patients with congenital diaphragmatic hernia (CDH): a possible locus for Fryns syndrome. Am J Med Genet A 2006; 140: 17–23.
Nannya Y, Sanada M, Nakazaki K, Hosoya N, et al. A robust algorithm for copy number detection using high-density oligonucleotide single nucleotide polymorphism genotyping arrays. Cancer Res 2005; 65: 6071–6079.
Eugster EA, Berry SA, Hirsch B . Mosaicism for deletion 1p36.33 in a patient with obesity and hyperphagia. Am J Med Genet 1997; 70: 409–412.
Slavotinek AM, Moshrefi A, Davis R, Leeth E, et al. Array comparative genomic hybridization in patients with congenital diaphragmatic hernia: mapping of four CDH-critical regions and sequencing of candidate genes at 15q26.1-15q26.2. Eur J Hum Genet 2006; 14: 999–1008.
Smith SA, Martin KE, Dodd KL, Young ID . Severe microphthalmia, diaphragmatic hernia and Fallot's tetralogy associated with a chromosome 1;15 translocation. Clin Dysmorphol 1994; 3: 287–291.
Youssoufian H, Chance P, Tuck-Muller CM, Jabs EW . Association of a new chromosomal deletion [del(1)(q32q42)] with diaphragmatic hernia: assignment of a human ferritin gene. Hum Genet 1988; 78: 267–270.
Slavotinek AM . Fryns syndrome: a review of the phenotype and diagnostic guidelines. Am J Med Genet A 2004; 124: 427–433.
Rajcan-Separovic E, Harvard C, Liu X, McGillivray B, et al. Clinical and molecular cytogenetic characterisation of a newly recognised microdeletion syndrome involving 2p15-16.1. J Med Genet 2007; 44: 269–276.
Bendavid C, Haddad BR, Griffin A, Huizing M, et al. Multicolour FISH and quantitative PCR can detect submicroscopic deletions in holoprosencephaly patients with a normal karyotype. J Med Genet 2006; 43: 496–500.
Bendavid C, Dubourg C, Gicquel I, Pasquier L, et al. Molecular evaluation of foetuses with holoprosencephaly shows high incidence of microdeletions in the HPE genes. Hum Genet 2006; 119: 1–8.
Dubourg C, Lazaro L, Pasquier L, Bendavid C, et al. Molecular screening of SHH, ZIC2, SIX3, and TGIF genes in patients with features of holoprosencephaly spectrum: mutation review and genotype-phenotype correlations. Hum Mutat 2004; 24: 43–51.
Burke R, Nellen D, Bellotto M, Hafen E, et al. Dispatched, a novel sterol-sensing domain protein dedicated to the release of cholesterol-modified hedgehog from signaling cells. Cell 1999; 99: 803–815.
Tian H, Tenzen T, McMahon AP . Dose dependency of Disp1 and genetic interaction between Disp1 and other hedgehog signaling components in the mouse. Development 2004; 131: 4021–4033.
Cohen MM Jr . The hedgehog signaling network. Am J Med Genet A 2003; 123: 5–28.
Carstea ED, Morris JA, Coleman KG, Loftus SK, et al. Niemann-Pick C1 disease gene: homology to mediators of cholesterol homeostasis. Science 1997; 277: 228–231.
Hua X, Sakai J, Brown MS, Goldstein JL . Regulated cleavage of sterol regulatory element binding proteins requires sequences on both sides of the endoplasmic reticulum membrane. J Biol Chem 1996; 271: 10379–10384.
Tian H, Jeong J, Harfe BD, Tabin CJ, et al. Mouse Disp1 is required in sonic hedgehog-expressing cells for paracrine activity of the cholesterol-modified ligand. Development 2005; 132: 133–142.
Ma Y, Erkner A, Gong R, Yao S, et al. Hedgehog-mediated patterning of the mammalian embryo requires transporter-like function of dispatched. Cell 2002; 111: 63–75.
Ledbetter DH, Mascarello JT, Riccardi VM, Harper VD, et al. Chromosome 15 abnormalities and the Prader-Willi syndrome: a follow-up report of 40 cases. Am J Hum Genet 1982; 34: 278–285.
Ledbetter DH, Riccardi VM, Airhart SD, Strobel RJ, et al. Deletions of chromosome 15 as a cause of the Prader-Willi syndrome. N Engl J Med 1981; 304: 325–329.
Curran ME, Atkinson DL, Ewart AK, Morris CA, et al. The elastin gene is disrupted by a translocation associated with supravalvular aortic stenosis. Cell 1993; 73: 159–168.
Kelley RI, Zackai EH, Emanuel BS, Kistenmacher M, et al. The association of the DiGeorge anomalad with partial monosomy of chromosome 22. J Pediatr 1982; 101: 197–200.
Shapira SK, McCaskill C, Northrup H, Spikes AS, et al. Chromosome 1p36 deletions: the clinical phenotype and molecular characterization of a common newly delineated syndrome. Am J Hum Genet 1997; 61: 642–650.
Balciuniene J, Feng N, Iyadurai K, Hirsch B, et al. Recurrent 10q22-q23 deletions: a genomic disorder on 10q associated with cognitive and behavioral abnormalities. Am J Hum Genet 2007; 80: 938–947.
Klopocki E, Schulze H, Strauss G, Ott CE, et al. Complex inheritance pattern resembling autosomal recessive inheritance involving a microdeletion in thrombocytopenia-absent radius syndrome. Am J Hum Genet 2007; 80: 232–240.
Zweier C, Peippo MM, Hoyer J, Sousa S, et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt-Hopkins syndrome). Am J Hum Genet 2007; 80: 994–1001.
Vissers LE, van Ravenswaaij CM, Admiraal R, Hurst JA, et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet 2004; 36: 955–957.
Shieh JT, Aradhya S, Novelli A, Manning MA, et al. Nablus mask-like facial syndrome is caused by a microdeletion of 8q detected by array-based comparative genomic hybridization. Am J Med Genet A 2006; 140: 1267–1273.
Hou J, Parrish J, Ludecke HJ, Sapru M, et al. A 4-megabase YAC contig that spans the Langer-Giedion syndrome region on human chromosome 8q24.1: use in refining the location of the trichorhinophalangeal syndrome and multiple exostoses genes (TRPS1 and EXT1). Genomics 1995; 29: 87–97.
Oda T, Elkahloun AG, Pike BL, Okajima K, et al. Mutations in the human Jagged1 gene are responsible for Alagille syndrome. Nat Genet 1997; 16: 235–242.
Acknowledgements
The authors thank Sara Minier, Kyle Sundin, and Emily Rorem (Signature Genomic Laboratories) for their assistance with Figure 1. We dedicate this manuscript to Mary Curtis and thank her for contributions to the field of genetics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: L.G. Shaffer and B.A. Bejjani have ownership, receive consulting fees, and sit on the Members' Board of Signature Genomic Laboratories, LLC. The other authors report no conflict of interest.
Rights and permissions
About this article
Cite this article
Shaffer, L., Theisen, A., Bejjani, B. et al. The discovery of microdeletion syndromes in the post-genomic era: review of the methodology and characterization of a new 1q41q42 microdeletion syndrome. Genet Med 9, 607–616 (2007). https://doi.org/10.1097/GIM.0b013e3181484b49
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181484b49
Keywords
This article is cited by
-
Sushi domain-containing protein 4 binds to epithelial growth factor receptor and initiates autophagy in an EGFR phosphorylation independent manner
Journal of Experimental & Clinical Cancer Research (2022)
-
Mosaic trisomy of chromosome 1q in human brain tissue associates with unilateral polymicrogyria, very early-onset focal epilepsy, and severe developmental delay
Acta Neuropathologica (2020)
-
The hedgehog pathway and ocular developmental anomalies
Human Genetics (2019)
-
Identification of novel genomic imbalances in Saudi patients with congenital heart disease
Molecular Cytogenetics (2018)
-
Psychomotor retardation with a 1q42.11–q42.12 deletion
Hereditas (2017)