Abstract
Purpose: Autism is a chronic neurodevelopmental disorder characterized by deficits in reciprocal social interaction, language and communication, and by the presence of stereotypical behaviors. The disorder is a complex genetic trait with no known predisposing genes. We report the results of a pilot project to screen for aberrations in the gene-rich subtelomeric chromosomal regions of a cohort of children with autism.
Methods: For our pilot project, we used a multiprobe system that includes probes for the subtelomeric regions of all human chromosomes. We assessed the subtelomeric regions of chromosomes from 10 children with a diagnosis of autism.
Results: The screen identified one child with an apparent deletion of the subtelomeric region of chromosome 2q; nine children and pooled control samples yielded normal results. The deletion in our patient was confirmed with two other subtelomeric probes and a targeted cytogenetic study revealed a subtle difference in appearance for one chromosome 2 homologue.
Conclusion: There have been several reports of children with dysmorphic features, autistic behaviors, and 2q deletions detectable with standard cytogenetic techniques. It may be that the distal region of chromosome 2q harbors a gene or genes that may predispose to autism.
Similar content being viewed by others
Main
Autism (OMIM: 209850) is a chronic neurodevelopmental disorder characterized by pervasive deficits in social relating, language and communication, and by the presence of stereotyped behaviors. This disorder has an approximate prevalence rate of 2 to 5 cases in 10,000 children.1 Although the diagnosis is primarily clinical, several tools exist to gather the necessary history and behavioral observations to determine whether criteria from the American Psychiatric Association's Diagnostic and Statistical Manual of Mental Disorders, 4th edition (DSM-IV),2 are fulfilled. Gold standard assessment tools include the Autism Diagnostic Interview-Revised (ADI-R)3 and the Autism Diagnostic Observation Scale-Generic (ADOS-G).4
For most children with autism, the underlying cause leading to their stereotypic behaviors is unknown5 and laboratory analyses (hematological, biochemical, and cytogenetic) do not offer assistance.6 Although extensive studies have not revealed a definitive biological marker for autism, the disorder has been shown to be a complex genetic disease. A variety of genetic factors, including chromosome abnormalities, have been implicated in the pathogenesis of autism.7 Abnormalities in chromosome 15 and the sex chromosomes are the most common findings in studies of children with autistic disorder;6 however, aberrations in the majority of individual chromosomes have been reported.7
Family and twin studies support the involvement of numerous genes involved in autism.8,9 Recently, genome screens have identified several areas with potentially significant linkage.10–15 These studies report the strongest evidence for an autism susceptibility locus on 7q. In addition, regions on 2q, 16p, and 19p were found to be suggestive of linkage by at least two genome scans.15,16 Buxbaum et al.14 presented evidence suggestive of linkage to distal 2q, particularly in patients with autism and delayed onset of phrase speech.
Over 70% of individuals with autism have comorbid cognitive impairment.17 Chromosomal abnormalities are known to be one of the most common causes of mental retardation and several studies have reported using a novel technique to screen for cryptic, submicroscopic chromosomal aberrations in patients with idiopathic mental retardation.18–20 Deletions and rearrangements in the subtelomeric regions were detected in a significant proportion of individuals with mental impairment, with aberration detection rates ranging from 3% to 23%.21–23 These studies suggest that rearrangements in the clinically important subtelomeres may be relatively common and that they may serve as markers for the underlying causes of idiopathic mental retardation.
The subtelomeric regions of human chromosomes are specialized units of chromosome-specific DNA just proximal to the simple tandem telomeric repeats delineating the physical ends of the human chromosomes. A high concentration of genes and several candidate genes for recognizable syndromes are known to be present in the subtelomeric regions.24 To determine whether cryptic rearrangements and deletions in the subtelomeric region represent a significant source of pathology in autism, the use of a screening assay is of great utility. Researchers at the Institute of Molecular Medicine in Oxford, England, have developed an easy-to-use device that allows for simultaneous detection of all subtelomeric sequences on a single microscope slide.18 Thus, a large number of individuals may be analyzed relatively quickly and inexpensively. We used this technique for a pilot screening study of 10 children with a diagnosis of autism.
MATERIALS AND METHODS
Patients and controls
Children with a diagnosis of autism were ascertained from the Developmental Disorders and Evaluation Clinic at the Medical University of South Carolina. All patients had previous normal routine cytogenetic studies. Diagnostic procedures for each child varied slightly, but each received, at minimum, an evaluation by an experienced diagnostician using the Childhood Autism Rating Scale (CARS)25 and review of criteria for autistic disorder from the DSM-IV.2 Two of the children, including the subject with the deletion, received a more comprehensive evaluation from an autism diagnostic team. The team was composed of a developmental pediatrician, psychologist, educational specialist, speech/language therapist, occupational therapist, and social worker. Tools used included the ADI-R,3 Pre-Linguistic Autism Diagnostic Schedule (PL-ADOS),26 CARS,25 and review of the DSM-IV2 criteria. These two children were evaluated in 1997 before the use of the updated Autism Diagnostic Observation Schedule-Generic.4
The PL-ADOS26 consists of a set of structured situations that allow systematic observation and rating of a variety of behavioral characteristics that may be indicative of autism in preschool children. The patient with the deletion met criteria for autistic disorder on the PL-ADOS26 in both areas: Social/Communication score of 16 (cut-off for autism = 12) and Restricted/Repetitive Behaviors score of 4 (cut-off for autism = 2).
The ADI-R3 is a lengthy developmental interview, which is completed with the parent or other caregiver. Responses to questions are then scored in four areas of concern relative to autism. Significant ratings must be obtained in all four areas to indicate the presence of autism. This patient met criteria in all four areas: Qualitative Impairment in Reciprocal Social Interaction, Qualitative Impairment in Communication, Repetitive Behaviors and Stereotyped Patterns, and Abnormal Development Before 36 months. The CARS25 is a 15-item behavioral rating scale developed to identify a person with autism. The patient received a total score of 33.5 (cut-off for autism = 30), which is in the mild-moderate range of autism. Review of DSM-IV2 criteria found the patient to meet criteria for a diagnosis of autistic disorder (299.00).
The study was approved by the Institutional Review Board for Human Research, and informed consents were obtained from parents or guardians of all 10 children.
Controls were selected from a pool of anonymized cytogenetically normal individuals' cell pellets. Two pooled control samples, one made up of five normal 46,XX females and the other of five normal 46,XY males, were tested.
Case report of patient with 2q deletion
The patient with the deletion of 2q has been diagnosed with autistic disorder, moderate mental retardation, and disruptive behaviors. Originally seen in Developmental Pediatrics at age 28 months for concerns with receptive and expressive language delays, the patient was found to have autistic behaviors as well as global developmental delays. CARS done at 2 years 5 months fell into the mild-moderate range of autism with a score of 33.5. The diagnosis of autistic disorder was further confirmed at 3 years 11 months with the administration of the PL-ADOS and the ADI-R. He was found to meet criteria for autistic disorder on both the PL-ADOS and the ADI-R as well as by DSM-IV criteria for autism. Physical exams have revealed normal growth parameters, and there are no discernible dysmorphic features. Family history was unremarkable for developmental or psychiatric disorders on the maternal side. The patient's father was not involved in his care, and mother was unaware of his family history.
The patient has been maintained in a self-contained special education class since age 3 years. He has had difficulty with hyperkinetic, impulsive, and aggressive behaviors in the school setting but not at home. He has had problems in the past with sleep disturbance, but this condition has resolved.
Multiprobe FISH
Peripheral blood cell samples were cultured using standard protocol. Prepared cell pellets were stored in 3:1 methanol:acetic acid at 0 to 6°C for no longer than 6 months before testing, some samples were stored for longer than a month in the freezer at −20°C.
The Cytocell Chromoprobe Multiprobe-T System (Banbury, England) was used for slide preparation, hybridization, and detection following the manufacturer's recommended protocol. However, NP40 was substituted for Tween 20 for the stringent washes, and the overnight hybridization was done in an open, water-filled, plastic container incubating in a 37°C oven instead of a 37°C waterbath.
During the course of the study, the manufacturer switched from an indirect labeling system to directly labeled probes. Two patient samples and the second run of each pooled control sample were assayed using the new devices. In addition, the 2q deletion was confirmed with the indirect device.
In most instances, five metaphases were scored for each section on the slide, and the observed number and color of each signal were recorded. A chromosome was considered normal if at least two cells from the section had the correct number of signals, i.e., two red and two green signals (Fig. 1), or two red signals for the acrocentric chromosomes. Scoring criteria were not met for several chromosome-specific areas on a minority of patient devices. In these cases, single chromosome telomere coverslips were used to complete the patient analysis.
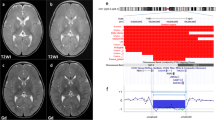
Examples of analysis of chromosome 2 subtelomeres in a normal control (A, indirect method; B, direct method) and in a patient with a deletion of 2q (C, indirect method; D, direct method). Green probe signals correspond to the short arm subtelomere, and the q arm subtelomeres reveal red signals. Yellow arrows depict the deleted chromosome.
Confirmation of deletion and FISH with YAC probes
The deletion in our one patient was confirmed using the 2q subtelomeric probe for locus D2S447 (Vysis) according to manufacturer's instructions. To determine the extent of the deletion in this patient, a biotin-labeled 2qYAC probe, 938A7, that is estimated to be approximately 10cM from the 2q telomere was hybridized to a slide from the deleted patient as described in Kumar et al. 1998.27
RESULTS
We screened the subtelomeres of 10 children with a diagnosis of autism and two pooled control samples. The control data (Table 1) was used to estimate variability in the testing methodology. Each control pooled sample was tested initially using the indirect detection devices. Several cells were missing one or several signals (no cells had a pattern consistent with additional signals), indicating decreased hybridization, decreased detection efficiency, or both, with some probes. The “abnormal” signal patterns seen in a percentage of control cells were assumed to reflect technical artifact and not true biologic deletions because (1) in a subsequent analysis using the direct detection devices, all normal patterns were reported for the same pooled control samples (Table 1); and (2) similar variability was seen in the autistic patient samples. Thus, scoring criteria for the patient samples (at least two cells with a normal signal pattern indicates a normal result) were adopted to account for the technical variability.
Nine of the 10 patient samples demonstrated normal signal patterns in 2 or more metaphases in each of the 23 chromosome-specific areas (Fig. 1 A,B). Thus, there were no abnormalities detectable by the Cytocell Chromoprobe Multiprobe-T System in these individuals.
Insufficient results were obtained for seven subtelomere hybridizations (2 -#13; 1 each of #14, #15, #16, #19, #20). In these squares, no cell or only one cell revealed signal because of an air bubble inhibiting hybridization or other technical problem. Thus, each of these subtelomere probe hybridizations was repeated using a single coverslip with the appropriate probe. This single coverslip analysis yielded normal results for each probe studied.
Cells from one patient exhibited an abnormality - the absence of one red signal on chromosome 2q. Signal patterns for the remaining 22 chromosomes were normal, with no chromosome appearing to have three signals. A total of 10 metaphases gave a signal pattern consistent with a deletion of the subtelomeric region of 2q at locus D2S2986 (Fig. 1 C) for this patient using the indirect probe methodology. To confirm the abnormal signal pattern, this patient's sample was also tested using the direct detection device and with the Vysis 2q probe (D2S447). Ten metaphase cells from each assay gave the same abnormal pattern with these FISH probes (Fig. 1 D). The patient's blood was re-cultured for chromosome studies with a targeted analysis of 2q. Although no length discrepancy was noted, the 2q regions exhibited differing banding patterns suggestive of a subtle abnormality. Each of the apparent abnormal chromosome 2s terminated in a blunted dark band (Fig. 2).
A peripheral blood sample was obtained from this patient's mother (father was unavailable) to look for a balanced rearrangement and to rule out the possibility that the abnormal pattern reflected a polymorphism segregating in this patient's family. The mother's study yielded a normal high-resolution karyotype and a normal subtelomere study.
To further define the deletion of 2q, a YAC from distal 2q was hybridized to the patient's cells. YAC 938A7 revealed signals on both chromosome 2s, indicating that the deletion was distal to the marker D2S338 (data not shown).
DISCUSSION
Ten percent (1 of 10) of the unselected patients in our pilot project had abnormal results. This finding suggests that the subtelomeres may be involved in the pathogenesis of a significant percentage of autism cases. This finding is similar to recent reports of an association of subtelomeric deletions/rearrangements and idiopathic mental retardation.18,19 In individuals with mental retardation of unknown cause, deletions and rearrangements in the subtelomeric regions were detected in 3% to 23%.21–23 These studies suggest that rearrangements in the clinically important subtelomeres may be relatively common and that they may reveal underlying causes of multiple disorders.
There have been several reports of phenotypically abnormal children with 2q deletions, detectable by routine cytogenetics, and a diagnosis of autism, or behaviors consistent with this diagnosis.28 The patients with the most distal deletions tend to have mental retardation, frontal bossing, macrocephaly, a depressed nasal bridge, and social and communication abnormalities with repetitive behaviors.28,29 Ghaziuddin and Burmeister29 hypothesized that patients with deletions of 2q37 may form a distinct subgroup within the spectrum of autistic disorders and that a gene within 2q37, when deleted, may contribute to the etiology of autism. The deletion in our nondysmorphic patient with autism is smaller than those previously reported, suggesting that his deletion may contain the gene strictly responsible for the behavioral phenotype. At least 25 genes have been localized to the possible critical region, several of which are specifically expressed in the brain or nervous system (OMIM Gene Maps).
Whole genome screens have suggested several chromosomal regions that are potentially associated with a susceptibility gene for autism.10–15 Three studies revealed positive linkage to 2q10,11,15 and a third study demonstrated linkage in distal 2q of a subset of patients with autism and delayed onset of phrase speech.14 Genomic scans are limited by the number of loci that are assessed, therefore, not all areas may be equally represented. It is important to note that telomeric regions may have increased meiotic recombination and may be under-represented in these types of analyses. Thus, the FISH approach is an important correlative study in the search for susceptibility genes.
Subtelomeres are specialized units of chromosome-specific DNA just proximal to the simple tandem telomeric repeats delineating the physical ends of the human chromosomes. A high concentration of genes and several candidate genes for recognizable syndromes are known to be present in the subtelomeric regions.24 However, some subtelomeric DNA is noncoding, polymorphic with no clinical significance, or both.30 In particular, the subtelomeric region of chromosome 2q has been shown to exhibit a chromosome length polymorphism31 and a 2q polymorphism detectable using the Cytocell indirect probe (PAC 1011O17) has been reported.32 (Note that this reference32 reports that the direct 2q telomere probe [P1 210E14] for the generation 2 Cytocell device revealed normal results for the patients with the polymorphism.) Therefore, deletions and rearrangements of subtelomeric sequences may not always indicate a clinically relevant abnormality. When an abnormality is identified in a patient with an aberrant phenotype, family studies should also be performed to evaluate the inheritance pattern and to rule out a clinically insignificant polymorphism. It is possible that the deletion of 2q observed in our patient was inherited from his father and may represent a polymorphism. However, confirmation of our patient's abnormality with the generation 2 Cytocell direct detection device and the more proximal Vysis probe strongly suggests that this aberration does not represent a polymorphism. In addition, the cytogenetic analysis also suggested a subtle difference between the two chromosome 2 homologs (Fig. 2). It seems most likely that the patient's FISH results are due to the significant loss of genetic material.
The simultaneous detection of all subtelomeric sequences on a single microscope slide allowed for an efficient screen of these regions in our autistic patients. However, it is important to note that results may vary from cell to cell, and strict scoring criteria may need to be developed. As evidenced by our control data using the indirect detection (Table 1), technical artifact may give false-positive or -negative results. The direct method provides cleaner data and may not need as strict a scoring system.
To determine the frequency of 2q and other subtelomeric abnormalities, additional patients with autistic disorder need to be analyzed. It may be useful to screen families with multiple affected individuals, dysmorphic patients with autism, or both, to increase the likelihood of detecting chromosomal abnormalities. Once identified, abnormalities in the subtelomeres may lead to the identification of genes important in the etiology of autism and a better understanding of this disorder.
References
Klin A, Volkmar FR . Autism and other pervasive developmental disorders. In: Goldstein S, Reynolds CR, editors. Handbook of neurodevelopmental and genetic disorders in children. New York: Guilford Press, 1999; 257.
American Psychiatric Association ( 1994) Diagnostic and Statistical Manual of Mental Disorders, 4th edition (DSM-IV). Washington, DC: American Psychiatric Association.
Lord C, Rutter M, Le Couteur A . Autism diagnostic interview-revised: a revised version of a diagnostic interview for caregivers of individuals with possible pervasive developmental disorders. J Autism Dev Disord 1994; 24: 659–685.
Lord C, Rutter M, DiLavore P . Autism diagnostic observation schedule-generic. Chicago, IL: University of Chicago, Dept. of Psychiatry, 1998.
Rutter M, Bailey A, Bolton P, Le Couteur A . Autism and known medical conditions. J Child Psychol Psychiatry 1994; 35: 311–322.
Schroer RJ, Phelan MC, Michaelis RC, Crawford EC, Skinner SA, Cuccaro M, Simensen RJ, Bishop J, Skinner C, Fender D, Stevenson RE . Autism and maternally derived aberrations of chromosome 15q. Am J Med Genet 1998; 76: 327–336.
Weidmer-Mikhail E, Sheldon S, Ghaziuddin M . Chromosomes in autism and related pervasive developmental disorders: a cytogenetic study. J Intellect Disabil Res 1998; 42: 8–12.
Bailey A, Phillips W, Rutter M . Autism: towards an integration of clinical, genetic, neuropsychological, and neurobiological perspectives. J Child Psychol Psychiatry 1996; 37: 89–126.
Szatmari P, Jones MB, Zwaigenbaum L, MacLean JE . Genetics of autism: overview and new directions. J Autism Dev Dis 1998; 28: 351–368.
International Molecular Genetic Study of Autism Consortium. A full genome screen for autism with evidence for linkage to a region on chromosome 7q. Hum Mol Genet 1998; 7: 571–578.
Philippe A, Martinez M, Guilloud-Bataille M, Gillberg C, Rastam M, Sponheim E, Coleman M, Zappella M, Aschauer H, van Malldergerme L . Genome-wide scan for autism susceptibility genes. Paris Autism Research International Sibpair Study. Hum Mol Genet 1999; 8: 805–812.
Barrett S, Beck JC, Bernier R, Bisson E, Braun TA, Casavant TL, Childress D, Folstein SE, Garcia M, Gardiner MB, Gilman S, Haines JL, Hopkins K, Landa R, Meyer NH, Mullane JA, Nishimura DY, Palmer P, Piven J, Purdy J, Santangelo SL, Searby C, Sheffield V, Singleton J, Slager S, et al. An autosomal genomic screen for autism. Collaborative linkage study of autism. Am J Med Genet 1999; 88: 609–615.
Risch N, Spiker D, Lotspeich L, Nouri N, Hinds D, Hallmayer J, Kalaydjieva L, McCague P, Dimiceli S, Pitts T, Nguyen L, Yang J, Harper C, Thorpe D, Vermeer S, Young H, Hebert J, Lin A, Ferguson J, Chiotti C, Wiese-Slater S, Rogers T, Salmon B, Nicholas P, Myers RM, et al. A genomic screen of autism: evidence for a multilocus etiology. Am J Hum Genet 1999; 65: 493–507.
Buxbaum JD, Silverman JM, Smith CJ, Kilifarski M, Reichert J, Hollander E, Lawlor BA, Fitzgerald M, Greenberg DA, Davis KL . Evidence for a susceptibility gene for autism on chromosome 2 and for genetic heterogeneity. Am J Hum Genet 2001; 68: 1514–1520.
International Molecular Genetic Study of Autism Consortium. A genomewide screen for autism: strong evidence for linkage to chromosomes 2q, 7q, and 16p. Am J Hum Genet 2001; 69: 570–581.
Lamb JA, Moore J, Bailey A, Monaco AP . Autism: recent molecular genetic advances. Hum Mol Genet 2000; 9: 861–868.
Mauk JE . Autism and pervasive developmental disorders. Pediatr Clin North Am 1993; 40: 567–578.
Knight S, Horsley S, Regan R, Lawrie N, Maher E, Cardy D, Flint J, Kearney L . Development and clinical application of an innovative fluorescence in situ hybridization technique which detects submicroscopic rearrangements involving telomeres. Eur J Hum Genet 1997; 5: 1–8.
Ghaffari S, Boyd E, Tolmie JL, Crow Y, Trainer A, Connor M . A new strategy for cryptic telomeric translocation screening in patients with idiopathic mental retardation. J Med Genet 1998; 35: 225–233.
Knight SJ, Regan R, Nicod A, Horsley SW, Kearney L, Homfray T, Winter RM, Bolton P, Flint J . Subtle chromosomal rearrangements in children with unexplained mental retardation. Lancet 1999; 354: 1676–1681.
Flint J, Wilkie A, Buckle VJ, Winter RM, Holland AJ, McDermid HE . The detection of subtelomeric chromosomal rearrangements in idiopathic mental retardation. Nat Genet 1995; 9: 132–140.
Voit G, Gosset P, Fert S, Prieur M, Turleau C, Raoul O, DeBlois MC, Lyonnet S, Munnich A, Vekemans M . Cryptic subtelomeric rearrangements detected by FISH in mentally retarded and dysmorphic patients. Am J Hum Genet Suppl 1998; 63: A44.
Vorsanova SG, Kolotii AD, Sharonin VO, Soloview IV, Yurov YB . FISH analysis of microaberrations at telomeric and subtelomeric regions in chromosomes of children with mental retardation. Am J Hum Genet Suppl (Suppl) 1998; 63: A873.
National Institutes of Health, and Institute of Molecular Medicine Collaborative Group. A complete set of human telomeric probes and their clinical application. Nat Genet 1996; 14: 86–89.
Schopler E, Reichler RJ, Renner BR . The Childhood Autism Rating Scale. Los Angeles, CA: Western Psychological Services, l998.
DiLavore P, Lord C, Rutter M . The Pre-Linguistic Autism Diagnostic Observation Schedule. J Autism Dev Disord 1995; 24: 355–379.
Kumar A, Becker LA, Depinet TW, Haren JM, Kurtz CL, Robin NH, Cassidy SB, Wolff DJ, Schwartz S . Molecular characterization and delineation of subtle deletions in de novo “balanced” chromosomal rearrangements. Hum Genet 1998; 103: 173–178.
Conrad B, Dewald G, Christensen E, Lopez M, Higgins J, Pierpont ME . Clinical phenotype associated with terminal 2q37 deletion. Clin Genet 1995; 48: 134–139.
Ghaziuddin M, Burmeister M . Deletion of chromosome 2q37 and autism: a distinct subtype?. J Autism Dev Disord 1999; 29: 259–263.
Trask BJ, Friedman C, Martin-Gallardo A, Rowen L, Akinbami C, Blankenship J, Collins C, Giorgi D, Iadonato S, Johnson F, Kuo WL, Massa H, Morrish T, Naylor S, Nguyen OT, Rouquier S, Smith T, Wong DJ, Youngblom J, van den Engh G . Members of the olfactory receptor gene family are contained in large blocks of DNA duplicated polymorphically near the ends of human chromosomes. Hum Mol Genet 1998; 7: 13–26.
Macina RA, Negorev DG, Spais C, Ruthig LA, Hu X-L, Riethman HC . Sequence organization of the human chromosome 2q telomere. Hum Mol Genet 1994; 3: 1847–1853.
Ballif BC, Kashork CD, Shaffer LG . The promise and pitfalls of telomere region-specific probes [letter]. Am J Hum Genet 2000; 67: 1356–1359.
Acknowledgements
The labeled YAC probe was a kind gift from Dr. Stuart Schwartz. Appreciation is expressed to Donna Jacobs and Tara Ellingham for expert technical assistance.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wolff, D., Clifton, K., Karr, C. et al. Pilot assessment of the subtelomeric regions of children with autism: Detection of a 2q deletion. Genet Med 4, 10–14 (2002). https://doi.org/10.1097/00125817-200201000-00002
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/00125817-200201000-00002
Keywords
This article is cited by
-
Phenotypic variant of Brachydactyly-mental retardation syndrome in a family with an inherited interstitial 2q37.3 microdeletion including HDAC4
European Journal of Human Genetics (2013)
-
The genetics of autistic disorders and its clinical relevance: a review of the literature
Molecular Psychiatry (2007)
-
Systematic Screening for Subtelomeric Anomalies in a Clinical Sample of Autism
Journal of Autism and Developmental Disorders (2007)
-
Autism and cytogenetic abnormalities: Solving autism one chromosome at a time
Current Psychiatry Reports (2007)
-
Genetics of autism spectrum disorder
European Journal of Human Genetics (2006)