Abstract
Purpose
To identify genetic defects in a large family with open angle glaucoma (OAG) and microcornea.
Methods
Genomic DNA was prepared from leukocytes of 15 individuals from three generations of a Chinese family, including seven individuals with OAG and microcornea, one with microcornea alone, and seven healthy individuals. Whole exome sequencing was performed on genomic DNA of the proband. Candidate variants were obtained through multiple steps of bioinformatics analysis and validated by Sanger sequencing and segregation analysis.
Results
Exome sequencing detected a candidate variant in GJA1, a novel truncation mutation (c.791_792delAA, p.K264Ifs*43). This mutation was present in all seven individuals with OAG and microcornea and the individual with microcornea alone, but not in the seven unaffected relatives in the family. It was not present in 1394 alleles from 505 unrelated controls without glaucoma and 192 normal controls. Extraocular signs were not observed in seven out of the eight individuals; only one was affected with dental enamel hypoplasia and syndactyly.
Conclusions
A novel truncation mutation in GJA1 is associated with OAG and microcornea in a Chinese family. This suggests that GJA1 should be included as a candidate gene for glaucoma.
Similar content being viewed by others
Introduction
Glaucoma is a neurodegenerative disease characterised by typical visual dysfunction and structural damage to the optic nerve.1 The prevalence of primary open angle glaucoma (OAG) was 2.6% in Chinese with age over 40 years.2 It is widely accepted that reduced drainage of aqueous humor is the main cause for elevated intraocular pressure (IOP), which is associated with retina ganglion cell loss.3 Both genetic and environmental factors are involved in the development of glaucoma.3 A few genes are associated with primary OAG,4, 5, 6, 7, 8 but mutations in these genes only account for 5–7.8% of cases.9, 10 The molecular basis of OAG in most patients has yet to be identified.
We sampled individuals from a large Chinese family with OAG and microcornea. In this study, the genetic defect in the family was investigated by whole exome sequencing and a novel truncation mutation in GJA1 was identified.
Materials and methods
The proband from a Chinese family with OAG and microcornea was collected from the Zhongshan Ophthalmic Center as part of an ongoing program to identify the genes responsible for glaucoma. An additional 14 family members were collected for segregation analysis thereafter. Written informed consent was obtained from the participants or their guardians before the study. Genomic DNA was prepared from the peripheral venous leukocytes of 15 subjects from the family by using procedures described in a previous study,11 including seven individuals with OAG and microcornea, one individual with microcornea alone, and seven unaffected relatives. The diagnostic criteria for OAG is as follows:12 IOP over 21 mm Hg; glaucomatous optic neuropathy (vertical cup-to-disc ratio ≥0.7, asymmetry ≥0.2, rim width ≤0.1 or disc hemorrhage); glaucomatous visual field defect meeting the Hodapp–Parrish–Anderson criteria for early defect;13 and open anterior chamber angle on gonioscopy (Shaffer grades III or IV). Microcornea refers to a horizontal corneal diameter <10 mm.14 The clinical data of four patients (II:1, II:6, II:8, and III:6) were in consensus with the diagnostic criteria of OAG and microcornea; one patient (III:3) reached the diagnostic criteria of microcornea alone, and the other three patients (II:3, III:4, and III:5) were diagnosed as OAG and microcornea by the same ophthalmologist, although detailed clinical data were not available. Our study was consistent with the tenets of the Declaration of Helsinki and was approved by the Institutional Review Board of the Zhongshan Ophthalmic Center. We certified that all applicable institutional and governmental regulations concerning the ethical use of human volunteers were followed during this research.
Genomic DNA from the proband was initially analyzed by whole exome sequencing using a procedure similar to that described in our previous study.9 The variants detected in proband III:4 were filtered using the following steps: (1) variations predicted to affect the coding residues or mRNA splicing were selected; (2) after comparison with the 1000 Genomes Project database, variants with a minor allele frequency (MAF) <0.01 were selected; (3) variants in genes associated with ocular diseases were selected; (4) the remaining variants were compared with ethnicity-matched regional controls (195 patients with high myopia and 310 patients with hereditary retinal diseases but not glaucoma based on our unpublished data of whole exome sequencing on patients with hereditary eye diseases) to exclude ethnicity-specific polymorphisms (MAF≥0.01); (5) effects of the remaining variants on coding residues of the remaining variants were analyzed using Polyphen-2 and SIFT;15, 16 (6) the remaining variants predicted to have damaging effects on coding sequence and to affect splicing were confirmed by Sanger sequencing and validated in family members, as well as in 192 normal controls. A pair of primers used to confirm the novel variant in GJA1 were designed using the Primer3 online tool (http://frodo.wi.mit.edu/primer3/):17 GJA1-Forward 5′-aaaagagatccctgcccaca-3′ and GJA1-Reverse 5′-aggctgttgagtaccacctc-3′. The polymerase chain reaction was used to amplify the target variants, and the amplicons were analyzed with an ABI BigDye Terminator cycle sequencing kit v3.1 (Applied Biosystems, Foster City, CA, USA) on an ABI3130 Genetic Analyzer (Applied Biosystems) as described in a previous study.9
Results
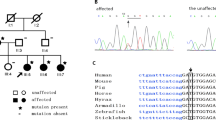
Overall, 74 740 single nuclear variants, insertions, and deletions were captured by whole exome sequencing on the proband. After filtering the whole exome sequencing data, one novel candidate heterozygous variant (c.791_792delAA, p.K264Ifs*43) in GJA1 was identified and subsequently confirmed in the proband by Sanger sequencing. This variant was neither present in databases (1000 Genomes Project and Exome Variant Server) nor in 1394 unrelated ethnicity-matched control alleles (192 normal controls, 195 patients with high myopia, and 310 patients with hereditary retinal diseases based on our unpublished data of whole exome sequencing on patients with hereditary eye diseases). It was present in the seven individuals with OAG and microcornea and the one with microcornea alone but absent in the seven unaffected relatives (Figure 1). By using this mutation as a marker, two-point linkage analysis yielded a maximum lod score of 2.709 at θ=0, theoretically the maximum lod score for this type of family.
The GJA1 mutation and the pedigree. (a) The mutation cosegregated with OAG and microcornea. Under each individual, + indicates a wild-type allele, and M indicates a mutant allele c.791_792delAA. Black symbols indicate individuals with OAG and microcornea; the black-and-white symbol indicates the individual with microcornea alone. (b) Sequencing chromatography demonstrated a heterozygous c.791_792delAA mutation in GJA1.
Clinical data for the family members participating in this study are listed in Table 1. The c.791_792delAA mutation was present in all eight individuals with microcornea, in whom the cornea diameter ranged from 9.0 to 9.5 mm. Of these eight individuals, seven had OAG, as diagnosed based on increased IOP, degenerative changes to the optic disc and the retina ganglion cell layers, and/or typical glaucomatous visual field defects (Figure 2). The individual with microcornea alone had no detectable signs of glaucoma when examined at the age of 29 years, but was at risk of developing glaucoma. Of these eight individuals, none had a remnant pupillary membrane, porous and spongy iris tissue, aniridia, or posterior embryotoxon. Systemic examination of the eight individuals identified syndactyly, camptodactyly, and dental enamel hypoplasia in one individual (III:6) and palmoplantar keratodermas in another individual (II:8). None of the eight individuals were observed to have paralysis, mental retardation, congenital cardiovascular defects, or hearing loss.
Discussion
In this study, we report on a large Chinese family with OAG and microcornea. For these conditions, glaucoma is the most prominent sign that brings the individuals to the hospital for treatment. Except for OAG and microcornea, most patients in the studied family do not have other noticeable ocular or systemic anomalies. A novel truncation mutation (c.791_792delAA, p.K264Ifs*43) in GJA1 that cosegregates with the disease has been identified in this family. This mutation is predicted to result in complete loss of function of the coded protein by the mutant allele and is absent in 1394 ethnicity-matched control alleles.
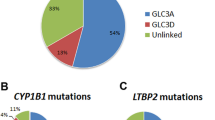
Previously, based on the HGMD database (http://www.hgmd.org/, accessed on 22 May 2014), 92 mutations have been identified in GJA1, including 82 missense mutations, two nonsense mutations, four in-frame insertion/deletion mutations, three frameshift mutations, and one splicing mutation.18 Of the 92 mutations, 71 are associated with oculodentodigital dysplasia (ODDD); 13 are associated with heart malformations; four are associated with deafness; two are associated with sudden infant death syndrome; and one is associated with craniometaphyseal dysplasia (Supplementary Table S1).19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57 Glaucoma has been recorded in 33 of 161 patients with ODDD and GJA1 mutations but in none of the 33 patients with other conditions and GJA1 mutations.23, 26, 27, 28, 29, 30, 31, 37, 40, 46, 47 These glaucoma-associated mutations distribute randomly in the functional domains of GJA1. It total, glaucoma is reported in ~17% of patients (33/194) with GJA1 mutations. Both OAG and angle closure glaucoma have been reported.37 In nearly half of the previous reported 33 patients with glaucoma (48%, 16/33), the age at diagnosis of glaucoma was less than 35 years old (Supplementary Table S1).23, 26, 27, 28, 29, 40, 46, 47 In the present study, OAG is present in seven of the eight individuals (87.5%) with the GJA1 mutation, which is a significantly higher association than that reported in previous studies.
Here we identify a novel truncation mutation in GJA1 which is associated with OAG and microcornea in a Chinese family. The results suggest that GJA1 mutations may frequently associate with OAG and, therefore, it should be considered as a causative gene for glaucoma.

References
Foster PJ, Buhrmann R, Quigley HA, Johnson GJ . The definition and classification of glaucoma in prevalence surveys. Br J Ophthalmol 2002; 86 (2): 238–242.
Wang YX, Xu L, Yang H, Jonas JB . Prevalence of glaucoma in North China: the Beijing Eye Study. Am J Ophthalmol 2010; 150 (6): 917–924.
Quigley HA . Glaucoma. Lancet 2011; 377 (9774): 1367–1377.
Monemi S, Spaeth G, DaSilva A, Popinchalk S, Ilitchev E, Liebmann J et al. Identification of a novel adult-onset primary open-angle glaucoma (POAG) gene on 5q22.1. Hum Mol Genet 2005; 14 (6): 725–733.
Stone EM, Fingert JH, Alward WL, Nguyen TD, Polansky JR, Sunden SL et al. Identification of a gene that causes primary open angle glaucoma. Science 1997; 275 (5300): 668–670.
Rezaie T, Child A, Hitchings R, Brice G, Miller L, Coca-Prados M et al. Adult-onset primary open-angle glaucoma caused by mutations in optineurin. Science 2002; 295 (5557): 1077–1079.
Pasutto F, Matsumoto T, Mardin CY, Sticht H, Brandstatter JH, Michels-Rautenstrauss K et al. Heterozygous NTF4 mutations impairing neurotrophin-4 signaling in patients with primary open-angle glaucoma. Am J Hum Genet 2009; 85 (4): 447–456.
Melki R, Colomb E, Lefort N, Brezin AP, Garchon HJ . CYP1B1 mutations in French patients with early-onset primary open-angle glaucoma. J Med Genet 2004; 41 (9): 647–651.
Huang X, Li M, Guo X, Li S, Xiao X, Jia X et al. Mutation analysis of seven known glaucoma-associated genes in Chinese patients with glaucoma. Invest Ophthalmol Vis Sci 2014; 55 (6): 3594–3602.
Fingert JH . Primary open-angle glaucoma genes. Eye (Lond) 2011; 25 (5): 587–595.
Wang Q, Wang P, Li S, Xiao X, Jia X, Guo X et al. Mitochondrial DNA haplogroup distribution in Chaoshanese with and without myopia. Mol Vis 2010; 16: 303–309.
Fan BJ, Wang DY, Fan DS, Tam PO, Lam DS, Tham CC et al. SNPs and interaction analyses of myocilin, optineurin, and apolipoprotein E in primary open angle glaucoma patients. Mol Vis 2005; 11: 625–631.
Hodapp E, Parrish RK II, Anderson DR . Clinical Decisions in Glaucoma. The CV Mosby Co: St Louis, 1993; 52–61.
Friede R . Surface area of cornea and sclera in embryos and in newborn infants and its relation to megalocornea in adults. Z Augenheilkd 1933; 8: 213.
Kumar P, Henikoff S, Ng PC . Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 2009; 4 (7): 1073–1081.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P et al. A method and server for predicting damaging missense mutations. Nat Methods 2010; 7 (4): 248–249.
Rozen S, Skaletsky H . Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 2000; 132: 365–386.
Stenson PD, Ball EV, Mort M, Phillips AD, Shiel JA, Thomas NS et al. Human Gene Mutation Database (HGMD): 2003 update. Hum Mutat 2003; 21 (6): 577–581.
de la Parra DR, Zenteno JC . A new GJA1 (connexin 43) mutation causing oculodentodigital dysplasia associated to uncommon features. Ophthalmic Genet 2007; 28 (4): 198–202.
Jamsheer A, Badura-Stronka M, Sowinska A, Debicki S, Kiryluk K, Latos-Bielenska A . A severe progressive oculodentodigital dysplasia due to compound heterozygous GJA1 mutation. Clin Genet 2010; 78 (1): 94–97.
Fenwick A, Richardson RJ, Butterworth J, Barron MJ, Dixon MJ . Novel mutations in GJA1 cause oculodentodigital syndrome. J Dent Res 2008; 87 (11): 1021–1026.
Himi M, Fujimaki T, Yokoyama T, Fujiki K, Takizawa T, Murakami A . A case of oculodentodigital dysplasia syndrome with novel GJA1 gene mutation. Jpn J Ophthalmol 2009; 53 (5): 541–545.
Paznekas WA, Karczeski B, Vermeer S, Lowry RB, Delatycki M, Laurence F et al. GJA1 mutations, variants, and connexin 43 dysfunction as it relates to the oculodentodigital dysplasia phenotype. Hum Mutat 2009; 30 (5): 724–733.
Gardner P, Oitmaa E, Messner A, Hoefsloot L, Metspalu A, Schrijver I . Simultaneous multigene mutation detection in patients with sensorineural hearing loss through a novel diagnostic microarray: a new approach for newborn screening follow-up. Pediatrics 2006; 118 (3): 985–994.
Jamsheer A, Wisniewska M, Szpak A, Bugaj G, Krawczynski MR, Budny B et al. A novel GJA1 missense mutation in a Polish child with oculodentodigital dysplasia. J Appl Genet 2009; 50 (3): 297–299.
Gabriel LA, Sachdeva R, Marcotty A, Rockwood EJ, Traboulsi EI . Oculodentodigital dysplasia: new ocular findings and a novel connexin 43 mutation. Arch Ophthalmol 2011; 129 (6): 781–784.
Kelly SC, Ratajczak P, Keller M, Purcell SM, Griffin T, Richard G . A novel GJA 1 mutation in oculo-dento-digital dysplasia with curly hair and hyperkeratosis. Eur J Dermatol 2006; 16 (3): 241–245.
Paznekas WA, Boyadjiev SA, Shapiro RE, Daniels O, Wollnik B, Keegan CE et al. Connexin 43 (GJA1) mutations cause the pleiotropic phenotype of oculodentodigital dysplasia. Am J Hum Genet 2003; 72 (2): 408–418.
Furuta N, Ikeda M, Hirayanagi K, Fujita Y, Amanuma M, Okamoto K . A novel GJA1 mutation in oculodentodigital dysplasia with progressive spastic paraplegia and sensory deficits. Intern Med 2012; 51 (1): 93–98.
Richardson R, Donnai D, Meire F, Dixon MJ . Expression of Gja1 correlates with the phenotype observed in oculodentodigital syndrome/type III syndactyly. J Med Genet 2004; 41 (1): 60–67.
Gladwin A, Donnai D, Metcalfe K, Schrander-Stumpel C, Brueton L, Verloes A et al. Localization of a gene for oculodentodigital syndrome to human chromosome 6q22-q24. Hum Mol Genet 1997; 6 (1): 123–127.
Richardson RJ, Joss S, Tomkin S, Ahmed M, Sheridan E, Dixon MJ . A nonsense mutation in the first transmembrane domain of connexin 43 underlies autosomal recessive oculodentodigital syndrome. J Med Genet 2006; 43 (7): e37.
Kellermayer R, Keller M, Ratajczak P, Richardson E, Harangi F, Merei E et al. Bigenic connexin mutations in a patient with hidrotic ectodermal dysplasia. Eur J Dermatol 2005; 15 (2): 75–79.
Klaver EC, Versluijs GM, Wilders R . Cardiac ion channel mutations in the sudden infant death syndrome. Int J Cardiol 2011; 152 (2): 162–170.
Feller L, Wood NH, Sluiter MD, Noffke C, Raubenheimer EJ, Lemmer J et al. Report of a black South African child with oculodentodigital dysplasia and a novel GJA1 gene mutation. Am J Med Genet A 2008; 146A (10): 1350–1353.
Izumi K, Lippa AM, Wilkens A, Feret HA, McDonald-McGinn DM, Zackai EH . Congenital heart defects in oculodentodigital dysplasia: report of two cases. Am J Med Genet A 2013; 161A (12): 3150–3154.
Vasconcellos JP, Melo MB, Schimiti RB, Bressanim NC, Costa FF, Costa VP . A novel mutation in the GJA1 gene in a family with oculodentodigital dysplasia. Arch Ophthalmol 2005; 123 (10): 1422–1426.
Pizzuti A, Flex E, Mingarelli R, Salpietro C, Zelante L, Dallapiccola B . A homozygous GJA1 gene mutation causes a Hallermann-Streiff/ODDD spectrum phenotype. Hum Mutat 2004; 23 (3): 286.
Honkaniemi J, Kalkkila JP, Koivisto P, Kahara V, Latvala T, Simola K . Letter to the editor: Novel GJA1 mutation in oculodentodigital dysplasia. Am J Med Genet A 2005; 139 (1): 48–49.
Wiest T, Herrmann O, Stogbauer F, Grasshoff U, Enders H, Koch MJ et al. Clinical and genetic variability of oculodentodigital dysplasia. Clin Genet 2006; 70 (1): 71–72.
Kjaer KW, Hansen L, Eiberg H, Leicht P, Opitz JM, Tommerup N . Novel Connexin 43 (GJA1) mutation causes oculo-dento-digital dysplasia with curly hair. Am J Med Genet A 2004; 127A (2): 152–157.
Debeer P, Van Esch H, Huysmans C, Pijkels E, De Smet L, Van de Ven W et al. Novel GJA1 mutations in patients with oculo-dento-digital dysplasia (ODDD). Eur J Med Genet 2005; 48 (4): 377–387.
Brueton LA, Huson SM, Farren B, Winter RM . Oculodentodigital dysplasia and type III syndactyly: separate genetic entities or disease spectrum? J Med Genet 1990; 27 (3): 169–175.
Schrander-Stumpel CT, De Groot-Wijnands JB, De Die-Smulders C, Fryns JP . Type III syndactyly and oculodentodigital dysplasia: a clinical spectrum. Genet Couns 1993; 4 (4): 271–276.
Wang B, Wen Q, Xie X, Liu S, Liu M, Tao Y et al. Mutation analysis of Connexon43 gene in Chinese patients with congenital heart defects. Int J Cardiol 2010; 145 (3): 487–489.
van Es RJ, Wittebol-Post D, Beemer FA . Oculodentodigital dysplasia with mandibular retrognathism and absence of syndactyly: a case report with a novel mutation in the connexin 43 gene. Int J Oral Maxillofac Surg 2007; 36 (9): 858–860.
Vitiello C, D'Adamo P, Gentile F, Vingolo EM, Gasparini P, Banfi S . A novel GJA1 mutation causes oculodentodigital dysplasia without syndactyly. Am J Med Genet A 2005; 133A (1): 58–60.
Shapiro RE, Griffin JW, Stine OC . Evidence for genetic anticipation in the oculodentodigital syndrome. Am J Med Genet 1997; 71 (1): 36–41.
Brice G, Ostergaard P, Jeffery S, Gordon K, Mortimer PS, Mansour S . A novel mutation in GJA1 causing oculodentodigital syndrome and primary lymphoedema in a three generation family. Clin Genet 2013; 84 (4): 378–381.
Vreeburg M, de Zwart-Storm EA, Schouten MI, Nellen RG, Marcus-Soekarman D, Devies M et al. Skin changes in oculo-dento-digital dysplasia are correlated with C-terminal truncations of connexin 43. Am J Med Genet A 2007; 143 (4): 360–363.
Hu Y, Chen IP, de Almeida S, Tiziani V, Do Amaral CM, Gowrishankar K et al. A novel autosomal recessive GJA1 missense mutation linked to Craniometaphyseal dysplasia. PloS One 2013; 8 (8): e73576.
Chen P, Xie LJ, Huang GY, Zhao XQ, Chang C . Mutations of connexin43 in fetuses with congenital heart malformations. Chin Med J 2005; 118 (12): 971–976.
van Steensel MA, Spruijt L, van der Burgt I, Bladergroen RS, Vermeer M, Steijlen PM et al. A 2-bp deletion in the GJA1 gene is associated with oculo-dento-digital dysplasia with palmoplantar keratoderma. Am J Med Genet A 2005; 132A (2): 171–174.
Yang JJ, Huang SH, Chou KH, Liao PJ, Su CC, Li SY . Identification of mutations in members of the connexin gene family as a cause of nonsyndromic deafness in Taiwan. Audiol Neurootol 2007; 12 (3): 198–208.
Britz-Cunningham SH, Shah MM, Zuppan CW, Fletcher WH . Mutations of the Connexin43 gap-junction gene in patients with heart malformations and defects of laterality. N Engl J Med 1995; 332 (20): 1323–1329.
Dasgupta C, Martinez AM, Zuppan CW, Shah MM, Bailey LL, Fletcher WH . Identification of connexin43 (alpha1) gap junction gene mutations in patients with hypoplastic left heart syndrome by denaturing gradient gel electrophoresis (DGGE). Mutat Res 2001; 479 (1-2): 173–186.
Kooshavar D, Tabatabaiefar MA, Farrokhi E, Abolhasani M, Noori-Daloii MR, Hashemzadeh-Chaleshtori M . Digenic inheritance in autosomal recessive non-syndromic hearing loss cases carrying GJB2 heterozygote mutations: assessment of GJB4, GJA1, and GJC3. Int J Pediatr Otorhinolaryngol 2013; 77 (2): 189–193.
Acknowledgements
We thank the patient and his family for their participation. This study was supported by the National Natural Science Foundation of China (81170881, U1201221), Guangdong Department of Science & Technology Translational Medicine Center grant 2011A080300002, and the Fundamental Research Funds of the State Key Laboratory of Ophthalmology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Eye website
Supplementary information
Rights and permissions
About this article
Cite this article
Huang, X., Wang, N., Xiao, X. et al. A novel truncation mutation in GJA1 associated with open angle glaucoma and microcornea in a large Chinese family. Eye 29, 972–977 (2015). https://doi.org/10.1038/eye.2015.74
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/eye.2015.74
This article is cited by
-
Rare mosaic variant of GJA1 in a patient with a neurodevelopmental disorder
Human Genome Variation (2024)
-
P2X7/P2X4 Receptors Mediate Proliferation and Migration of Retinal Microglia in Experimental Glaucoma in Mice
Neuroscience Bulletin (2022)