Abstract
Regulatory T (Treg) cells are a distinct subset of CD4+ T cells. Instead of triggering adaptive immunity, they suppress immune responses. Small numbers of Treg cells reside within lymphoid organs and peripheral tissues, but their contribution to immune tolerance is so significant that defects in Treg cell function cause catastrophic immune disorders. Since they were first discovered 20 years ago, efforts have been made to understand the differences in developmental processes between Treg cells and conventional T cells that determine the ultimate fate of the overall T-cell population. Transcription factor Foxp3 is crucial for Treg cell differentiation, but it is not the whole story. Owing to recent advances in Treg cell research, we are now on the verge of appreciating the comprehensive mechanisms underlying Treg cell generation. Here, we discuss major discoveries, active study topics and remaining questions regarding Treg cell development.
Similar content being viewed by others
Introduction
The human body is defended by an immune system that responds to invading microorganisms. However, excessive or improper immune responses against self-antigens, innocuous antigens present in food, commensal microorganisms or fetal antigens can have detrimental effects; thus, they have to be constrained. Regulatory T (Treg) cells play a major role in restraining immune responses to maintain immune homeostasis. Since Treg cells are involved in many aspects of immune regulation, they have attracted much attention over the past two decades in terms of their basic mechanism(s) of action and their therapeutic potential. Since the discovery of Treg cells, knowledge about their development and differentiation has increased. Here, we briefly summarize established knowledge and describe recent advancements in the study of Treg cell development.
The discovery of Treg cells
Considering the boom in the Treg cell research field at the beginning of the twenty-first century, it is surprising that the earliest evidence of the existence of suppressive T cells goes back to 1969. In Japan, Nishizuka and Sakakura et al.1 found that thymectomizing 3-day-old female mice caused sterility; however, this was not the case for mice aged 7 days or older. The cause of sterility was later found to be a part of autoimmune oophoritis due to a missing cell population derived from the thymus that was produced within the first week after birth.2 Later reports strongly suggested the presence of T-cell-mediated immune tolerance to autoantigens.3 When the likely candidate cell type, reported to express a marker called the I–J determinant, was isolated, T cells with an immunosuppressive phenotype became an active research subject.4 However, despite two decades of study, researchers started to question both the existence of the I-J determinant and that of T cells expressing it. When the I-J determinant was proved to be absent from the expected locus,5 the field of so-called “suppressor T lymphocytes” collapsed. T cells that could induce immune tolerance were later identified as a subset of CD4+ T cells that constantly express the interleukin (IL-2) receptor α-chain CD25.6 This cell type was further characterized by the expression of Foxp3,7, 8 a transcription factor already linked to immune dysregulation, polyendocrinopathy, enteropathy and X-linked syndrome (IPEX).9 A detailed history of Treg cell discovery is documented elsewhere.10
Transcriptional regulation of Foxp3 expression
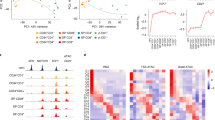
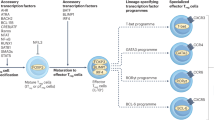
As mentioned above, expression of Foxp3 is a distinctive feature of Treg cells. Ectopic expression of Foxp3 alone can induce a suppressive phenotype in conventional T (Tconv) cells.7 Therefore, stringent regulation of Foxp3 expression is required to maintain homeostasis of T-cell-mediated immune responses. Many years have been dedicated to studying the mechanisms regulating the Foxp3 locus (Figure 1), making Foxp3 one of the most intensively studied genes in recent years.
Schematic diagram of transcriptional regulation of the Foxp3 locus. Regulatory regions of the Foxp3 locus including the promoter CNS1, CNS2, CNS3, and recently discovered CNS0 are shown. Transcription factors (TFs) binding to each regulatory region and the function of each regulatory region are shown.
Regulatory elements of the Foxp3 locus
Comparative genomic approaches involving alignment of human, rat and mouse genomes initially discovered three conserved non-coding sequences (CNSs) on the Foxp3 locus: a promoter and two enhancers that are positioned within the first intron.11, 12, 13 Later, another intronic enhancer, located directly after exon 1, was found (Figure 1).14 The Foxp3 promoter has minimal transcriptional activity, and the mechanism underlying lineage-specific expression of Foxp3 relies heavily on other cis-regulatory elements (Figure 1). Intensive studies of the function of CNSs were undertaken, the most notable being the systemic deletion of each CNS by Rudensky and co-workers.14 They revealed that CNS1 is largely dispensable for thymic Treg (tTreg) cell development, but is necessary for peripheral induction of Treg cells. Its significance was highlighted when deletion of CNS1 markedly reduced the Treg cell population in gut-associated lymphoid tissue. On the other hand, deleting CNS3 causes a severe reduction in thymic output of Treg cells, indicating its importance in tTreg cell generation. CNS2, also known as Treg cell-specific demethylated region, contains CpG islands that are highly demethylated only in functional Treg cells; its demethylation is considered to be the most definitive marker of commitment to the Treg cell lineage.15, 16 Deletion of CNS2 does not change thymic generation of Treg cells; rather, it affects the stability of Foxp3 expression during proliferation. The most recent discovery in the Foxp3 locus is another regulatory element named CNS0, which lies on an intron of the neighboring gene 5′ of the Foxp3 locus (Figure 1).17 It was found in an attempt to localize Treg cell-specific super enhancers using high-throughput chromatin immunoprecipitation sequencing of acetylated histone H3K27.
Transcription factors binding to regulatory elements
Many transcription factors have been studied for their ability to transactivate the Foxp3 gene (Figure 1). Among them is c-Rel. The significance of c-Rel was demonstrated by showing that c-Rel deficiency causes a marked reduction in tTreg cell generation.18 Individual studies suggest different mechanisms for the function of c-Rel during Foxp3 transcription; these include binding and demethylation of CNS2,19 binding to the promoter followed by formation of a c-Rel enhanceosome over the Foxp3 locus18 and binding to CNS3 and triggering Foxp3 induction by T-cell receptor (TCR) and costimulatory signals.14 Foxo family of transcription factors are also involved in regulating Foxp3 induction. Foxo1 and Foxo3 act redundantly on Foxp3 transcription by binding directly to the promoters, CNS1 and CNS3.20, 21 T-cell-specific deletion of both genes in mice halves the tTreg cell population and causes a multifocal inflammatory disorder. It was discovered that not only Foxp3 but also Treg cell-specific genes rely on Foxo transcription factors. Smad3 and NFAT modulate Foxp3 expression by binding to CNS1 upon transforming growth factor-β (TGF-β) and TCR signaling, respectively.22 NFAT also binds to CNS2 and mediates formation of a chromatin loop between the promoter and CNS2 of the Foxp3 locus via a mediator–cohesin complex.23 AP-1 transcription factors also bind to CNS1 and transactivate Foxp3 induction, while signal transducer and activator of transcription 3 (Stat3) binding to the CNS2 region silences Foxp3 transcription.24 Stat5, a protein downstream of IL-2 and other common γ-chain cytokine signaling pathways, targets the Foxp3 locus directly.25 IL-2 signaling and Stat5 binding to CNS2 protect Treg cell identity from other cytokine signals and maintain heritable transcription of Foxp3.26 Nr4a nuclear receptor family members Nr4a1 (also known as Nur77), Nr4a2 and Nr4a3 (the induction of which is proportional to the intensity of the TCR signal) function redundantly to bind the proximal promoter of Foxp3 to induce transactivation and are thought to translate TCR signaling intensity into a T-cell fate decision via Foxp3 induction or by triggering negative selection.27 Recently, chromatin organizer Satb1 was found to bind CNS0 and act as a pioneer factor to activate Treg cell-specific super enhancers of the Foxp3 gene and other Treg cell-related genes such as Ctla4 and Il2ra at the early stages of tTreg cell differentiation.17 Satb1 works by binding to closed chromatin structures and modifies the epigenetic status of the Foxp3 locus to a poised state, thereby allowing other transcription factors to bind to regulatory elements. Since Satb1 acts not only on Treg cell but also on general thymic T-cell development,28 it is unclear how Satb1 is induced and binds to Treg cell-specific super enhancers in a Treg cell-specific manner.
Epigenetic regulation
As a marker of definitive Treg cell lineage commitment, the mechanism underlying CNS2 demethylation has been the center of attention in the quest to understand Treg cell differentiation. The importance of CNS2 demethylation in Treg cell differentiation and function is highlighted by the finding that transcription factors CREB/ATF, NF-κB, Ets-1 and the Runx-Foxp3 complex, all of which induce Foxp3 expression, cannot bind to CNS2 without demethylation,11, 14, 29, 30 which results in loss of Foxp3 expression in Treg cells. Studies using the DNA methyltransferase inhibitor azacytidine and Dnmt1-deficient T cells reveal that the methylation status of the Foxp3 locus is actively maintained in T cells.11, 31 However, the exact machinery responsible for CNS2 demethylation was not identified until recently. In 2013, researchers found that vitamin C is a critical cofactor for Tet activity (Tet hydroxylates 5-methylcytosine to 5-hydroxymethylcytosine, the first step of CpG demethylation).32, 33, 34 This is also true for demethylation of the Foxp3 locus in Treg cells, as Tet2 and Tet3 work redundantly during demethylation of CNS2 in the presence of vitamin C.35, 36 Since supplementation with vitamin C alone demethylates CNS2 in vitro, demethylation of CNS2 might be a spontaneous reaction during normal Treg cell differentiation in vivo, where vitamin C is abundant. This breakthrough could provide a new opportunity for Treg cell therapy because in vitro generation of Treg cells showing stable Foxp3 expression is now more likely.
Permissible histone modification also contributes to Foxp3 expression in Treg cells. In fact, as in the case of in vitro-induced Treg cells, Foxp3 expression can be achieved without CNS2 demethylation when histone modification is permissive to transcription.37 By activating naïve T cells in the presence of TGF-β, Floess et al.15 and Ohkura et al.38 found prominent trimethylation of H3K4 on the promoter and CNS1 regions of the Foxp3 locus, although Foxp3 expression was transient and subsequently fell in the absence of exogenous TGF-β. Trimethylation of H3K4 is strongly correlated with Foxp3 expression, and this modification is only visible on the Foxp3 locus of fully differentiated Treg cells. However, there are other histone modifications on the Foxp3 locus that are more persistent throughout Treg cell development. From as early as the double-negative 1 stage of thymocyte development, CNS3 shows monomethylation of H3K4 (a poised enhancer marker); this modification can be inherited by subsequent T-cell lineages regardless of Foxp3 expression.39 Interestingly, naïve CD4+ T cells lacking Foxp3 expression maintain the monomethylation of H3K4 poised status of the Foxp3 promoter region in a CNS3-dependent manner, underlining the importance of CNS3 in peripheral Treg (pTreg) cell development. This could explain the previously held notion of defective in vitro-induced Treg cell development in CNS3-KO T cells.14 Another prevailing histone modification on the Foxp3 locus is H3K27. In the case of tTreg cell development, H3K27 starts to lose trimethylation status and acquires acetylation on enhancer regions at the pre-Treg cell stage, which implies initiation of enhancer activation.17 Not only internal cues but also environmental signals affect these modifications. The short-chain fatty acid butyrate, produced by the gut microbiota, promotes H3K27 acetylation on the Foxp3 locus by inhibiting histone deacetylase during pTreg cell development.40, 41
FOXP3-independent differentiation of Treg cells
Although the significance of Foxp3 drew much attention, other genes are also regulated in a Treg cell-specific manner. Some of those signature genes are not induced by Foxp3, but are regulated in parallel with the Foxp3 gene. Many different approaches to screening Treg cell-specific genes on a whole-genome scale have been conducted. Some of these genes are independent of Foxp3 expression.42, 43 Foxp3-deletion studies support the notion that, in the absence of Foxp3, a portion of T cells dubbed “wannabe” Treg cells still maintain a Treg cell-like phenotype.44, 45 Epigenetic regulation of Treg cell-signature genes is also independent of Foxp3. Demethylation of Treg cell-signature genes Ctla4, Il2ra, Tnfrsf18 and Ikzf4 is independent of Foxp3 expression, but dependent on TCR stimulation, and is required for the suppressive function of Treg cells.38 Enhancers of signature genes, in this case Ctla4, Il2ra and Ikzf2, are acetylated on H3K27 from the pre-Treg cell stage.17 Based on these studies, it is evident that the developmental process of Treg cells is not entirely dependent on the expression of Foxp3, and that a more comprehensive understanding of Treg cell development is required.
tTreg cell development
Thymocytes undergo positive and negative selection during TCR rearrangement. During these processes, a progenitor Treg cell population emerges that expresses a CD25hi CD4 single-positive phenotype (Figure 2). This population can later induce Foxp3 expression via stimulation by common γ-chain cytokines IL-2 and IL-15.46 Although Foxp3 is an essential transcription factor for Treg cells, this two-step developmental stage implies that Foxp3 expression is not a prerequisite for Treg cell lineage commitment, but rather an event that follows Treg cell differentiation. The main candidate determinant of the fate of thymocytes is TCR activation.
Schematic diagram of Treg cell development. tTreg cells develop in the thymus by two-step process. First, high-affinity tissue-restricted self-antigens presented by medullary thymic epithelial cells (mTECs) or bone-marrow derived antigen-presenting cells (BM-APCs) derive single-positive (SP) T cells into Treg pathway. Second, cytokine IL-2 or IL-15 derives the precursor cells into fully committed tTreg cells. pTreg cells develop in the periphery by environmental antigens such as microbial antigens or food antigens presented by mucosal tissue-resident dendritic cells (DCs). TGF-β, retinoic acids (RAs) and short chain fatty acids (SCFAs) produced in the immunosuppressive environments promote pTreg development.
Before the discovery of Treg cells, clonal deletion of autoreactive T cells was the main hypothesis that explained how T-cell-mediated autoimmunity is prevented, a process termed “central tolerance.” However, the importance of Treg cell function in corresponding “peripheral tolerance” has since been demonstrated, and a paradox became obvious: self-recognizing TCR clones must be negatively selected to prevent autoimmunity; however, they have to survive to generate Treg cells specific for self-antigens. This issue remained a mystery until recently.
A Nur77-GFP transgenic mouse experiment showed that, during thymocyte development, Treg cells receive a stronger TCR signal than Tconv cells.47 Stronger TCR signaling is associated with induction of Treg cell-specific epigenetic changes and gene expression patterns.48 It has long been known that a Treg cell TCR recognizes a self-antigen with high affinity,49 which increases the intensity of subsequent signals in the selection process. But how do Treg cells with self-reactive TCR clones evade deletion? Which is more important for determining Treg cell fate: the availability of autoantigen in the thymus or TCR affinity? Legoux et al.50 and Malhotra et al.51 designed elegant experiments to address this issue. They used readily available strains of Cre and fluorescent protein transgenic mice in which expression of the fluorescent protein is either universal or tissue-specific, depending on the promoter activity of the transgenes. They found that T cells reactive with ubiquitously expressed antigens are likely to be deleted, but clones reactive with tissue-restricted peptides are the main sources of Treg cell generation (Figure 2). In medullary thymic epithelial cells (mTECs), tissue-specific antigens are ectopically induced by the transcription factor Aire, and are presented to developing thymocytes at the thymic medulla at low frequency. Some tissue-specific antigens not induced by Aire are ignored. This is also true for endogenous self-antigens.52 In the absence of Aire, clones that normally differentiate into Treg cells become Tconv cells and induce organ-specific autoimmune diseases.53 From this point of view, it is noteworthy that Aire-dependent expression of tissue-specific genes is unique to each mTEC, or to each individual mouse.54 In addition, peptide processing and presentation by mTECs differs between perinatal and adult mice, thereby generating a distinct TCR repertoire on Treg cells in an age-dependent manner.55 The disparities in Treg cell TCR repertoires caused by altered antigen display might explain different susceptibilities to autoimmune diseases between individuals or at different ages.
Bone marrow-derived antigen-presenting cells (BM-APCs) also participate in T-cell selection in the thymic medulla. For certain TCR clones, tTreg cell development requires Aire-independent antigen presentation by bone marrow-derived antigen-presenting cells.56 Those APCs, primarily dendritic cells and B cells, take up blood-borne antigens and peripheral tissue antigens and then either migrate to present them within the thymus or acquire antigens from mTECs.57, 58 Another mode of B-cell participation is presentation of ectopic self-antigens along with expression of Aire.59 It is not clear whether BM-APCs display non-self-antigens derived from the microbiota or environmental antigens in the thymus, and whether this affects negative selection of thymocytes or tTreg cell differentiation. A fluorophore painted onto the skin of mice was found in thymic dendritic cells, implying that migration of peripheral APCs carrying foreign substances to the thymus is possible.60 A significant proportion of Treg cell TCRs react with non-self-antigens.61 Also, tTreg cells accumulate in neonatal skin to induce tolerance against Staphylococcus epidermidis colonization.62 Thus, it is plausible that tolerance to non-self-antigens could, at least in part, be established by thymic selection.
Still, the difference between TCR signals that bifurcate T-cell fate into Tconv or Treg cells is not well understood. Evidence presented so far supports quantitative differences in signaling, but it is possible that the signals are qualitatively variable. An interesting report shows topological differences in a Treg cell TCR–peptide–MHC docking structure. The report shows a 180° rotation of the binding structure between MHC with an insulin peptide and an in vitro-induced Treg cell TCR.63 Inverted orientation of TCR–peptide–MHC binding was also found in other T cells with specificity for epitopes; however, they showed minimal participation in immune responses against cognate antigens.64 Too few TCR–peptide–MHC complex structures have been studied to verify whether reversed polarity is a common phenomenon in a Treg cell population.
Peripheral Treg cell development
The generation of pTreg cells is even less clear than that of tTreg cells. This is, in part, due to a lack of reliable methods that discriminate pTreg cells from mixed Treg cell populations in peripheral lymphoid tissues. Although the transcription factor Helios,65 or the surface antigen neuropilin1,66, 67 are suggested to be exclusively expressed by tTreg cells, it is unclear whether they are genuine markers for tTreg cells.68, 69, 70 This makes identifying the origin of pTreg cells troublesome, especially in tissues such as gut, lung or skin in which environmental antigens make direct contact with the immune system and in which tTreg and pTreg cells are thought to coexist. Attempts to tackle this issue by direct TCR sequencing have yielded contradicting results. Some studies of intestinal Treg cells show results that favor peripheral differentiation of major Treg cell populations being influenced by colonic commensal microbiota,71 while others support a thymic origin.72 The opposing claims for the origin of major populations of intestinal Treg cells could be derived from the inherent limitations of TCR sequencing because the method uses TCR transgenic mice, which have a relatively small repertoire when compared with that in wild-type mice. In fact, a recent publication notes that it requires a full TCR repertoire to maintain intestinal homeostasis.73 Other studies that focused on sub-populations of intestinal Treg cells suggest that those Treg cells comprise a mixture of various origins and phenotypes.74 The exact mechanisms by which certain clones of T cells are driven into the Treg cell lineage by non-self-antigens remain unclear.
Aside from the origin issue, the existence of pTreg cells and the molecular mechanisms underlying their development are well defined (Figure 2). Treg cells can develop from naïve CD4+Foxp3− T cells in vitro upon TGF-β stimulation.75 Chronic exposure to antigens in small dosages induces a Treg cell population that is indistinguishable from tTreg cells in vivo.76 This population of Treg cells is also found in mice harboring a chronic Leishmania major infection.77 Owing to the nature of peripheral differentiation induced by non-self-antigens, pTreg cells are assumed to be a main component responsible for immunological tolerance to non-pathogenic substances such as environmental antigens or commensal microbiota. Removal of pTreg cells causes dysregulated immune responses in the gastrointestinal tract and airway,78 where the mucosal surface is in direct contact with environmental antigens. Under homeostatic conditions, such organs are favorable to Treg cell generation. Gut-associated lymphoid tissue-residing CD103+ dendritic cells produce TGF-β and retinoic acid, both of which promote de novo Treg cell induction.79, 80, 81 Also, the microbiota affects Treg cell differentiation. Metabolites of the microbiota, such as butyrate or propionate, boost Treg cell differentiation in the colon.40, 41 A recent study found that Treg cell abundancy in epithelial tissue is increased by CD11b+CD103−CX3CR1+ dendritic cells via the TRIF-IFN-β (TIR-domain-containing adapter-inducing interferon-β) pathway.82 These dendritic cells are suppressed by phosphatidylserine exposed on apoptotic epithelial cells, which binds to CD300a on the dendritic cell surface; this implies that apoptosis of epithelial cells indirectly regulates Treg cell differentiation. This finding may help us understand the mechanism by which barrier surfaces strike a balance between immune tolerance and inflammatory responses. Clonal pTreg cells are thought to be induced mostly by commensal microbiota. Recently, experiments using antigen-free mice fed an amino-acid diet revealed that small intestinal lamina propria Treg cells are induced mostly by dietary antigens.83 These pTreg cells are relatively short-lived and disappear fast when the supply of dietary antigen stops. It is possible that the reason for the short lifespan of these Treg cells is their conversion into Foxp3–CD8αα+CD4+ intraepithelial lymphocytes.84 The anti-inflammatory property of intraepithelial lymphocytes means that they complement Treg cell-mediated immune suppression.
Treg cell plasticity
Maintaining Treg cell identity is critically important for immune homeostasis;37, 85, 86, 87 for example, if Treg cells are readily converted to Tconv cells, maintaining homeostasis is impossible. Understanding the precise nature of Treg cell stability is crucial for therapeutic applications that use or modulate Treg cells to treat autoimmune diseases, allergies, graft rejection and tumors.87 Whether Treg cells are phenotypically and functionally stable is a contentious issue.87, 88, 89, 90, 91 Although it was originally thought that Treg cells are quite stable, a number of studies show that some Treg cells lose their identity or are converted into pathogenic effector CD4 T cells under lymphopenic and proinflammatory conditions.92, 93, 94, 95, 96, 97 Yang et al.97 showed that, when in vitro-induced Treg and tTreg cells are treated with IL-6 in combination with IL-1 or IL-23 in vitro, they are induced to express IL-17 and show defective suppressive activity. Duarte et al.96 showed that half of Foxp3+ cells lose Foxp3 expression when adoptively transferred to lymphopenic mice. This is prevented when Foxp3− cells are cotransferred, or IL-2 is injected into the recipient mice. The cells that lost Foxp3 expression produce IL-2 and lose suppressive activity upon secondary transfer to lymphopenic mice. Using bacterial artificial chromosome transgenic mice containing Foxp3-GFP-Cre crossed with Rosa26-YFP mice, Zhou et al.94 showed that a substantial percentage of cells have transient or unstable expression of Foxp3. These “exFoxp3 cells” are more numerous in inflamed tissues under autoimmune conditions. Adoptive transfer of these cells leads to rapid induction of diabetes, suggesting that they have an activated-memory phenotype. Tsuji et al.92 showed that, when Foxp3+ T cells from Foxp3-GFP mice are transferred to T-cell-deficient mice, some cells convert into Foxp3− cells, migrate into the germinal centers of Peyer’s patches and differentiate into T follicular helper cells. Using a Toxoplasma gondii infection model, Oldenhove et al.93 showed that Treg cell numbers decline in infected mice. In these mice, Treg cells also acquire T-bet and IFN-γ expression. These studies suggest that proinflammatory cytokines cause Treg cell instability by downregulating Foxp3. In human patients of several inflammatory diseases including psoriasis, inflammatory bowel disease and rheumatoid arthritis, Treg cells expressing IL-17 (and IFN-γ in some cases) were shown to increase compared with healthy controls.98, 99, 100 Whether these cells maintain suppressive function depends on the context; nevertheless, these results support phenotypic and functional plasticity of human Treg cells.
By contrast, another study showed that Treg cells are very stable and are not easily converted into effector cells.101, 102 Rubtsov et al.101 examined Treg cell stability using tracer mice expressing eGFP-Foxp3-Cre-ER × Rosa26-YFP. In these mice, YFP+ cells, which once expressed Foxp3 during their lifetime, were chased after tamoxifen treatment. YFP+ cells hardly lost GFP (Foxp3) expression under homeostatic conditions or under autoimmune inflammatory conditions, suggesting that Treg cells show remarkable stability.
To resolve this apparent controversy with respect to the plasticity of Treg cells, a few models have been proposed. First is the “heterogeneity model” proposed by Hori et al.103 He proposed that Treg cells consist of heterogeneous populations with different degrees of commitment, including fully committed Treg cells and less committed Treg cells. The model proposes that fully committed Treg cells are stable, but less committed Treg cells are unstable. Another model is the “transient flexibility model” proposed by Piccirillo and co-workers104 In this model, Treg cells are flexible in terms of their phenotype depending on the environment they are in. Under strong inflammatory conditions, Treg cells transiently lose Foxp3 expression and their suppressive properties, but these are recovered after the inflammatory conditions are removed. Thus, further studies should address and resolve the issue. In addition, the key molecular mechanisms that control Treg cell instability under these conditions should be elucidated to fully explain this phenomenon.
Metabolic regulation of Treg vs Th17 cell balance
Another subset of CD4+ T cells, discovered later than Treg cells, are T-helper type 17 (Th17) cells.105, 106 Despite the relatively short history of Th17 cell research, the impact of this subset on many human diseases (from autoimmunity to various infections) has attracted much attention during the past decade. One aspect of Th17 cells that ties them to Treg cells is their similarity in terms of differentiation conditions (such as a requirement of TGF-β in the case of Th17 and pTreg cell differentiation),107, 108 although this is not an absolute necessity for Th17 cells.109, 110 However, differentiation results in extreme differences in immunological activity: Treg cells suppress, while Th17 cells promote, immune responses. Initially, the proinflammatory cytokine IL-6 and subsequent Stat3 signaling were found to dictate the fate of these two subsets. Nowadays, more sophisticated mechanisms underlying T-cell fate decisions are being identified: these include cytokines, cellular metabolic pathways, dietary nutrients and the microbiota. Hundreds of articles have been published about the mechanisms underlying maintenance of the Treg/Th17 balance and its influence on diseases. Here, we review how TCR signaling and T-cell metabolism affect Treg cell differentiation.
Upon activation, naïve T cells undergo major metabolic conversion from oxidative phosphorylation and lipid oxidation to glycolysis to meet energy and material demand due to increased proliferation and cell mass. The metabolic mediator mammalian target of rapamycin (mTOR) is closely involved in this transition. mTOR receives and integrates various signals upon cellular stimulation by the environment and regulates metabolic changes and immune responses.111 The most notable mTOR modulator in T cells is the phosphatidylinositol 3-kinase (PI3K)/Akt pathway. Activated by costimulatory molecules, PI3K activates Akt and subsequently mTORC1. Phosphatase PTEN reverses PI3K activity and suppresses downstream signaling.112 The consequence of PI3K/Akt/mTOR activation is upregulation of glucose transporter Glut1 expression and increased glucose consumption to drive anabolic metabolism in T cells.
mTOR activation is required for effector T-cell development, whereas inhibiting the mTOR pathway promotes Treg cell differentiation. In particular, transcription factor hypoxia-inducible factor 1α (induced by mTOR signaling) promotes glycolysis and Th17 cell differentiation, whereas lack of mTOR113 or hypoxia-inducible factor 1α114, 115 drives cell fate toward Treg cells. Interestingly though, mTORC1 signaling is necessary for proper Treg cell proliferation and function because deletion of Treg cell-specific Raptor, a component of mTORC1, abrogates the suppressive function of Treg cells and eventually leads to inflammatory disorders.116 New evidence suggests that the mTOR pathway is more strictly regulated in Treg cells than in Tconv cells. One way in which Foxp3 regulates mTORC1 signaling is via toll-like receptors.117 The mTOR pathway is necessary for proliferation of Treg cells, and certain environmental cues such as toll-like receptor or leptin118 may enhance mTOR signaling intensity to enrich Treg cells. However, enhanced mTOR signaling also reduces the immunosuppressive function of Treg cells. In parallel with this, uncontrolled Akt signaling in Treg cells due to Treg-specific PTEN deficiency results in increased glycolysis and loss of Foxp3.119, 120 However, it is not mTORC1 activity, but rather mTORC2 activity, that is regulated by PTEN.120 Another recent report shows that inhibiting protein kinase CK2 blocks Th17 development and promotes Treg cell differentiation in mice with experimental autoimmune encephalomyelitis; this is due to a defect in Stat3 phosphorylation.121 Although not shown in that study, Akt is a target of CK2.122 It is not clear whether CK2 activity affects the PI3K/Akt pathway. It is also interesting to note that the same group reported that CK2 is necessary for proper Treg cell function during suppression of type 2 immune responses in the lung;123 this contradicts another study that examined the effect of CK2 on Treg cells.121 Thus, the role of CK2 in Treg cells seems to depend on the context of the immune response, although the exact mechanism(s) is not clear. In Th17 cells, but not Treg cells, glycolysis is linked with de novo fatty acid synthesis by acetyl-CoA carboxylase; this is because Treg cells acquire fatty acids from the environment. Disruption of acetyl-CoA carboxylase results in blockade of Th17 cell differentiation and instead promotes Treg cell differentiation.124
Applications
Understanding the mechanism(s) underlying Treg cell development is a prerequisite for Treg cell-based immunotherapy. Using Treg cells for immunotherapy would enable target-specific immunosuppression; this has clear benefits over nonspecific drug-induced immunosuppression, which can induce side effects such as opportunistic infection and cancer induction. Also, memory Treg cells could provide long-term, hopefully lifelong, tolerance to target antigens, resulting in relatively fewer treatments. Treg cell-mediated immunotherapy could be applied to organ transplantation, autoimmune diseases and allergies, resulting in improved prognoses and fewer side effects. There are more than 60 clinical trials currently registered in the United States examining the utility of Treg cell transfer (or indirect methods of boosting Treg cells (e.g., IL-2)) (https://clinicaltrials.gov) to improve treatment of type 1 diabetes mellitus, graft-versus-host disease, complications arising from organ transplantation, lupus and many other medical conditions. Most of those trials rely on polyclonal Treg cells derived from donors, although such cells are not target-specific. To improve specificity, some trials have utilized donor-alloantigen-reactive Treg cells, which are recipient Treg cells that are activated by, and proliferate in, donor tissue.125 At the preclinical research stage, several methods have been developed to generate targeted Treg cells. The most recent attempt involves application of a chimeric antigen receptor technique to target Treg cells to specific antigens or more broad alloantigens.126, 127
Conclusion
Treg cell research has progressed at an astonishing pace over the past two decades. Appreciation of the mechanisms underlying Treg cell development has led to the first therapeutic applications involving Treg cell induction. Yet, the fundamental question of how Treg cells enter fates different from Tconv cells (even though they arise from a common progenitor T cell) still remains unclear. Future studies of the processes underlying development of Treg cells should focus on comprehensive analyses of the entire TCR repertoire, intra- and intercellular mechanisms that determine Treg cell fate, and distinct features between thymic and peripheral Treg cell development.
References
Nishizuka Y, Sakakura T . Thymus and reproduction: sex-linked dysgenesia of the gonad after neonatal thymectomy in mice. Science 1969; 166: 753–755.
Sakaguchi S, Takahashi T, Nishizuka Y . Study on cellular events in postthymectomy autoimmune oophoritis in mice. I. Requirement of Lyt-1 effector cells for oocytes damage after adoptive transfer. J Exp Med 1982; 156: 1565–1576.
Gershon RK, Kondo K . Cell interactions in the induction of tolerance: the role of thymic lymphocytes. Immunology 1970; 18: 723–737.
Murphy DB, Herzenberg LA, Okumura K, McDevitt HO . A new I subregion (I-J) marked by a locus (Ia-4) controlling surface determinants on suppressor T lymphocytes. J Exp Med 1976; 144: 699–712.
Kronenberg M, Steinmetz M, Kobori J, Kraig E, Kapp JA, Pierce CW et al. RNA transcripts for I-J polypeptides are apparently not encoded between the I-A and I-E subregions of the murine major histocompatibility complex. Proc Natl Acad Sci USA 1983; 80: 5704–5708.
Sakaguchi S, Sakaguchi N, Asano M, Itoh M, Toda M . Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J Immunol 1995; 155: 1151–1164.
Hori S, Nomura T, Sakaguchi S . Control of regulatory T cell development by the transcription factor Foxp3. Science 2003; 299: 1057–1061.
Fontenot JD, Gavin MA, Rudensky AY . Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat Immunol 2003; 4: 330–336.
Bennett CL, Christie J, Ramsdell F, Brunkow ME, Ferguson PJ, Whitesell L et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat Genet 2001; 27: 20–21.
Shevach EM . The resurrection of T cell-mediated suppression. J Immunol 2011; 186: 3805–3807.
Kim HP, Leonard WJ . CREB/ATF-dependent T cell receptor-induced FoxP3 gene expression: a role for DNA methylation. J Exp Med 2007; 204: 1543–1551.
Mantel PY, Ouaked N, Ruckert B, Karagiannidis C, Welz R, Blaser K et al. Molecular mechanisms underlying FOXP3 induction in human T cells. J Immunol 2006; 176: 3593–3602.
Zorn E, Nelson EA, Mohseni M, Porcheray F, Kim H, Litsa D et al. IL-2 regulates FOXP3 expression in human CD4+CD25+ regulatory T cells through a STAT-dependent mechanism and induces the expansion of these cells in vivo. Blood 2006; 108: 1571–1579.
Zheng Y, Josefowicz S, Chaudhry A, Peng XP, Forbush K, Rudensky AY . Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 2010; 463: 808–812.
Floess S, Freyer J, Siewert C, Baron U, Olek S, Polansky J et al. Epigenetic control of the foxp3 locus in regulatory T cells. PLoS Biol 2007; 5: e38.
Wieczorek G, Asemissen A, Model F, Turbachova I, Floess S, Liebenberg V et al. Quantitative DNA methylation analysis of FOXP3 as a new method for counting regulatory T cells in peripheral blood and solid tissue. Cancer Res 2009; 69: 599–608.
Kitagawa Y, Ohkura N, Kidani Y, Vandenbon A, Hirota K, Kawakami R et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat Immunol 2017; 18: 173–183.
Ruan Q, Kameswaran V, Tone Y, Li L, Liou HC, Greene MI et al. Development of Foxp3(+) regulatory t cells is driven by the c-Rel enhanceosome. Immunity 2009; 31: 932–940.
Long M, Park SG, Strickland I, Hayden MS, Ghosh S . Nuclear factor-kappaB modulates regulatory T cell development by directly regulating expression of Foxp3 transcription factor. Immunity 2009; 31: 921–931.
Harada Y, Elly C, Ying G, Paik JH, DePinho RA, Liu YC . Transcription factors Foxo3a and Foxo1 couple the E3 ligase Cbl-b to the induction of Foxp3 expression in induced regulatory T cells. J Exp Med 2010; 207: 1381–1391.
Ouyang W, Beckett O, Ma Q, Paik JH, DePinho RA, Li MO . Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat Immunol 2010; 11: 618–627.
Tone Y, Furuuchi K, Kojima Y, Tykocinski ML, Greene MI, Tone M . Smad3 and NFAT cooperate to induce Foxp3 expression through its enhancer. Nat Immunol 2008; 9: 194–202.
Li X, Liang Y, LeBlanc M, Benner C, Zheng Y . Function of a Foxp3 cis-element in protecting regulatory T cell identity. Cell 2014; 158: 734–748.
Xu L, Kitani A, Stuelten C, McGrady G, Fuss I, Strober W . Positive and negative transcriptional regulation of the Foxp3 gene is mediated by access and binding of the Smad3 protein to enhancer I. Immunity 2010; 33: 313–325.
Yao Z, Kanno Y, Kerenyi M, Stephens G, Durant L, Watford WT et al. Nonredundant roles for Stat5a/b in directly regulating Foxp3. Blood 2007; 109: 4368–4375.
Feng Y, Arvey A, Chinen T, van der Veeken J, Gasteiger G, Rudensky AY . Control of the inheritance of regulatory T cell identity by a cis element in the Foxp3 locus. Cell 2014; 158: 749–763.
Sekiya T, Kashiwagi I, Yoshida R, Fukaya T, Morita R, Kimura A et al. Nr4a receptors are essential for thymic regulatory T cell development and immune homeostasis. Nat Immunol 2013; 14: 230–237.
Alvarez JD, Yasui DH, Niida H, Joh T, Loh DY, Kohwi-Shigematsu T . The MAR-binding protein SATB1 orchestrates temporal and spatial expression of multiple genes during T-cell development. Genes Dev 2000; 14: 521–535.
Polansky JK, Schreiber L, Thelemann C, Ludwig L, Kruger M, Baumgrass R et al. Methylation matters: binding of Ets-1 to the demethylated Foxp3 gene contributes to the stabilization of Foxp3 expression in regulatory T cells. J Mol Med (Berl) 2010; 88: 1029–1040.
Mouly E, Chemin K, Nguyen HV, Chopin M, Mesnard L, Leite-de-Moraes M et al. The Ets-1 transcription factor controls the development and function of natural regulatory T cells. J Exp Med 2010; 207: 2113–2125.
Josefowicz SZ, Wilson CB, Rudensky AY . Cutting edge: TCR stimulation is sufficient for induction of Foxp3 expression in the absence of DNA methyltransferase 1. J Immunol 2009; 182: 6648–6652.
Blaschke K, Ebata KT, Karimi MM, Zepeda-Martinez JA, Goyal P, Mahapatra S et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 2013; 500: 222–226.
Minor EA, Court BL, Young JI, Wang G . Ascorbate induces ten-eleven translocation (Tet) methylcytosine dioxygenase-mediated generation of 5-hydroxymethylcytosine. J Biol Chem 2013; 288: 13669–13674.
Yin R, Mao SQ, Zhao B, Chong Z, Yang Y, Zhao C et al. Ascorbic acid enhances Tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J Am Chem Soc 2013; 135: 10396–10403.
Sasidharan Nair V, Song MH, Oh KI . Vitamin C facilitates demethylation of the Foxp3 enhancer in a Tet-dependent manner. J Immunol 2016; 196: 2119–2131.
Yue X, Trifari S, Aijo T, Tsagaratou A, Pastor WA, Zepeda-Martinez JA et al. Control of Foxp3 stability through modulation of TET activity. J Exp Med 2016; 213: 377–397.
Ohkura N, Kitagawa Y, Sakaguchi S . Development and maintenance of regulatory T cells. Immunity 2013; 38: 414–423.
Ohkura N, Hamaguchi M, Morikawa H, Sugimura K, Tanaka A, Ito Y et al. T cell receptor stimulation-induced epigenetic changes and Foxp3 expression are independent and complementary events required for Treg cell development. Immunity 2012; 37: 785–799.
Feng Y, van der Veeken J, Shugay M, Putintseva EV, Osmanbeyoglu HU, Dikiy S et al. A mechanism for expansion of regulatory T-cell repertoire and its role in self-tolerance. Nature 2015; 528: 132–136.
Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013; 504: 451–455.
Furusawa Y, Obata Y, Fukuda S, Endo TA, Nakato G, Takahashi D et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013; 504: 446–450.
Sugimoto N, Oida T, Hirota K, Nakamura K, Nomura T, Uchiyama T et al. Foxp3-dependent and -independent molecules specific for CD25+CD4+ natural regulatory T cells revealed by DNA microarray analysis. Int Immunol 2006; 18: 1197–1209.
Hill JA, Feuerer M, Tash K, Haxhinasto S, Perez J, Melamed R et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity 2007; 27: 786–800.
Wan YY, Flavell RA . Regulatory T-cell functions are subverted and converted owing to attenuated Foxp3 expression. Nature 2007; 445: 766–770.
Gavin MA, Rasmussen JP, Fontenot JD, Vasta V, Manganiello VC, Beavo JA et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature 2007; 445: 771–775.
Lio CW, Hsieh CS . A two-step process for thymic regulatory T cell development. Immunity 2008; 28: 100–111.
Moran AE, Holzapfel KL, Xing Y, Cunningham NR, Maltzman JS, Punt J et al. T cell receptor signal strength in Treg and iNKT cell development demonstrated by a novel fluorescent reporter mouse. J Exp Med 2011; 208: 1279–1289.
Morikawa H, Sakaguchi S . Genetic and epigenetic basis of Treg cell development and function: from a FoxP3-centered view to an epigenome-defined view of natural Treg cells. Immunol Rev 2014; 259: 192–205.
Jordan MS, Boesteanu A, Reed AJ, Petrone AL, Holenbeck AE, Lerman MA et al. Thymic selection of CD4+CD25+ regulatory T cells induced by an agonist self-peptide. Nat Immunol 2001; 2: 301–306.
Legoux FP, Lim JB, Cauley AW, Dikiy S, Ertelt J, Mariani TJ et al. CD4+ T cell tolerance to tissue-restricted self antigens is mediated by antigen-specific regulatory T cells rather than deletion. Immunity 2015; 43: 896–908.
Malhotra D, Linehan JL, Dileepan T, Lee YJ, Purtha WE, Lu JV et al. Tolerance is established in polyclonal CD4(+) T cells by distinct mechanisms, according to self-peptide expression patterns. Nat Immunol 2016; 17: 187–195.
Kieback E, Hilgenberg E, Stervbo U, Lampropoulou V, Shen P, Bunse M et al. Thymus-derived regulatory T cells are positively selected on natural self-antigen through cognate interactions of high functional avidity. Immunity 2016; 44: 1114–1126.
Malchow S, Leventhal DS, Lee V, Nishi S, Socci ND, Savage PA . Aire enforces immune tolerance by directing autoreactive T cells into the regulatory T cell lineage. Immunity 2016; 44: 1102–1113.
Meredith M, Zemmour D, Mathis D, Benoist C . Aire controls gene expression in the thymic epithelium with ordered stochasticity. Nat Immunol 2015; 16: 942–949.
Yang S, Fujikado N, Kolodin D, Benoist C, Mathis D . Immune tolerance. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 2015; 348: 589–594.
Lin J, Yang L, Silva HM, Trzeciak A, Choi Y, Schwab SR et al. Increased generation of Foxp3(+) regulatory T cells by manipulating antigen presentation in the thymus. Nat Commun 2016; 7: 10562.
Perry JS, Hsieh CS . Development of T-cell tolerance utilizes both cell-autonomous and cooperative presentation of self-antigen. Immunol Rev 2016; 271: 141–155.
Oh J, Shin JS . The role of dendritic cells in central tolerance. Immune Netw 2015; 15: 111–120.
Yamano T, Nedjic J, Hinterberger M, Steinert M, Koser S, Pinto S et al. Thymic B cells are licensed to present self antigens for central T cell tolerance induction. Immunity 2015; 42: 1048–1061.
Bonasio R, Scimone ML, Schaerli P, Grabie N, Lichtman AH, von Andrian UH . Clonal deletion of thymocytes by circulating dendritic cells homing to the thymus. Nat Immunol 2006; 7: 1092–1100.
Pacholczyk R, Kern J, Singh N, Iwashima M, Kraj P, Ignatowicz L . Nonself-antigens are the cognate specificities of Foxp3+ regulatory T cells. Immunity 2007; 27: 493–504.
Scharschmidt TC, Vasquez KS, Truong HA, Gearty SV, Pauli ML, Nosbaum A et al. A wave of regulatory T cells into neonatal skin mediates tolerance to commensal microbes. Immunity 2015; 43: 1011–1021.
Beringer DX, Kleijwegt FS, Wiede F, van der Slik AR, Loh KL, Petersen J et al. T cell receptor reversed polarity recognition of a self-antigen major histocompatibility complex. Nat Immunol 2015; 16: 1153–1161.
Gras S, Chadderton J, Del Campo CM, Farenc C, Wiede F, Josephs TM et al. Reversed T cell receptor docking on a major histocompatibility class I complex limits involvement in the immune response. Immunity 2016; 45: 749–760.
Thornton AM, Korty PE, Tran DQ, Wohlfert EA, Murray PE, Belkaid Y et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J Immunol 2010; 184: 3433–3441.
Weiss JM, Bilate AM, Gobert M, Ding Y, Curotto de Lafaille MA, Parkhurst CN et al. Neuropilin 1 is expressed on thymus-derived natural regulatory T cells, but not mucosa-generated induced Foxp3+ T reg cells. J Exp Med 2012; 209: 1723–42, S1.
Yadav M, Louvet C, Davini D, Gardner JM, Martinez-Llordella M, Bailey-Bucktrout S et al. Neuropilin-1 distinguishes natural and inducible regulatory T cells among regulatory T cell subsets in vivo. J Exp Med 2012; 209: 1713–1722 S1–19.
Szurek E, Cebula A, Wojciech L, Pietrzak M, Rempala G, Kisielow P et al. Differences in expression level of helios and neuropilin-1 do not distinguish thymus-derived from extrathymically-induced CD4+Foxp3+ regulatory T Cells. PLoS ONE 2015; 10: e0141161.
Elkord E . Helios should not be cited as a marker of human thymus-derived Tregs. Commentary: Helios(+) and Helios(−) cells coexist within the natural FOXP3(+) T regulatory cell subset in humans. Front Immunol 2016; 7: 276.
Himmel ME, MacDonald KG, Garcia RV, Steiner TS, Levings MK . Helios+ and Helios− cells coexist within the natural FOXP3+ T regulatory cell subset in humans. J Immunol 2013; 190: 2001–2008.
Lathrop SK, Bloom SM, Rao SM, Nutsch K, Lio CW, Santacruz N et al. Peripheral education of the immune system by colonic commensal microbiota. Nature 2011; 478: 250–254.
Cebula A, Seweryn M, Rempala GA, Pabla SS, McIndoe RA, Denning TL et al. Thymus-derived regulatory T cells contribute to tolerance to commensal microbiota. Nature 2013; 497: 258–262.
Nishio J, Baba M, Atarashi K, Tanoue T, Negishi H, Yanai H et al. Requirement of full TCR repertoire for regulatory T cells to maintain intestinal homeostasis. Proc Natl Acad Sci USA 2015; 112: 12770–12775.
Tanoue T, Atarashi K, Honda K . Development and maintenance of intestinal regulatory T cells. Nat Rev Immunol 2016; 16: 295–309.
Chen W, Jin W, Hardegen N, Lei KJ, Li L, Marinos N et al. Conversion of peripheral CD4+CD25− naive T cells to CD4+CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J Exp Med 2003; 198: 1875–1886.
Apostolou I, von Boehmer H . In vivo instruction of suppressor commitment in naive T cells. J Exp Med 2004; 199: 1401–1408.
Belkaid Y, Piccirillo CA, Mendez S, Shevach EM, Sacks DL . CD4+CD25+ regulatory T cells control Leishmania major persistence and immunity. Nature 2002; 420: 502–507.
Josefowicz SZ, Niec RE, Kim HY, Treuting P, Chinen T, Zheng Y et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature 2012; 482: 395–399.
Sun CM, Hall JA, Blank RB, Bouladoux N, Oukka M, Mora JR et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J Exp Med 2007; 204: 1775–1785.
Benson MJ, Pino-Lagos K, Rosemblatt M, Noelle RJ . All-trans retinoic acid mediates enhanced T reg cell growth, differentiation, and gut homing in the face of high levels of co-stimulation. J Exp Med 2007; 204: 1765–1774.
Coombes JL, Siddiqui KR, Arancibia-Carcamo CV, Hall J, Sun CM, Belkaid Y et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism. J Exp Med 2007; 204: 1757–1764.
Nakahashi-Oda C, Udayanga KG, Nakamura Y, Nakazawa Y, Totsuka N, Miki H et al. Apoptotic epithelial cells control the abundance of Treg cells at barrier surfaces. Nat Immunol 2016; 17: 441–450.
Kim KS, Hong SW, Han D, Yi J, Jung J, Yang BG et al. Dietary antigens limit mucosal immunity by inducing regulatory T cells in the small intestine. Science 2016; 351: 858–863.
Sujino T, London M, Hoytema van Konijnenburg DP, Rendon T, Buch T, Silva HM et al. Tissue adaptation of regulatory and intraepithelial CD4(+) T cells controls gut inflammation. Science 2016; 352: 1581–1586.
Rudensky AY . Regulatory T cells and Foxp3. Immunol Rev 2011; 241: 260–268.
Benoist C, Mathis D . Treg cells, life history, and diversity. Cold Spring Harb Perspect Biol 2012; 4: a007021.
Sakaguchi S, Vignali DA, Rudensky AY, Niec RE, Waldmann H . The plasticity and stability of regulatory T cells. Nat Rev Immunol 2013; 13: 461–467.
Liston A, Piccirillo CA . Developmental plasticity of murine and human Foxp3(+) regulatory T cells. Adv Immunol 2013; 119: 85–106.
Li L, Boussiotis VA . Molecular and functional heterogeneity of T regulatory cells. Clin Immunol 2011; 141: 244–252.
Hori S . Stability of regulatory T-cell lineage. Adv Immunol 2011; 112: 1–24.
Pillai MR, Collison LW, Wang X, Finkelstein D, Rehg JE, Boyd K et al. The plasticity of regulatory T cell function. J Immunol 2011; 187: 4987–4997.
Tsuji M, Komatsu N, Kawamoto S, Suzuki K, Kanagawa O, Honjo T et al. Preferential generation of follicular B helper T cells from Foxp3+ T cells in gut Peyer’s patches. Science 2009; 323: 1488–1492.
Oldenhove G, Bouladoux N, Wohlfert EA, Hall JA, Chou D, Dos Santos L et al. Decrease of Foxp3+ Treg cell number and acquisition of effector cell phenotype during lethal infection. Immunity 2009; 31: 772–786.
Zhou X, Bailey-Bucktrout SL, Jeker LT, Penaranda C, Martinez-Llordella M, Ashby M et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nat Immunol 2009; 10: 1000–1007.
Komatsu N, Mariotti-Ferrandiz ME, Wang Y, Malissen B, Waldmann H, Hori S . Heterogeneity of natural Foxp3+ T cells: a committed regulatory T-cell lineage and an uncommitted minor population retaining plasticity. Proc Natl Acad Sci USA 2009; 106: 1903–1908.
Duarte JH, Zelenay S, Bergman ML, Martins AC, Demengeot J . Natural Treg cells spontaneously differentiate into pathogenic helper cells in lymphopenic conditions. Eur J Immunol 2009; 39: 948–955.
Yang XO, Nurieva R, Martinez GJ, Kang HS, Chung Y, Pappu BP et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity 2008; 29: 44–56.
Hovhannisyan Z, Treatman J, Littman DR, Mayer L . Characterization of interleukin-17-producing regulatory T cells in inflamed intestinal mucosa from patients with inflammatory bowel diseases. Gastroenterology 2011; 140: 957–965.
Voo KS, Wang YH, Santori FR, Boggiano C, Arima K, Bover L et al. Identification of IL-17-producing FOXP3+ regulatory T cells in humans. Proc Natl Acad Sci USA 2009; 106: 4793–4798.
Pesenacker AM, Bending D, Ursu S, Wu Q, Nistala K, Wedderburn LR . CD161 defines the subset of FoxP3+ T cells capable of producing proinflammatory cytokines. Blood 2013; 121: 2647–2658.
Rubtsov YP, Niec RE, Josefowicz S, Li L, Darce J, Mathis D et al. Stability of the regulatory T cell lineage in vivo. Science 2010; 329: 1667–1671.
Miyao T, Floess S, Setoguchi R, Luche H, Fehling HJ, Waldmann H et al. Plasticity of Foxp3(+) T cells reflects promiscuous Foxp3 expression in conventional T cells but not reprogramming of regulatory T cells. Immunity 2012; 36: 262–275.
Hori S . Lineage stability and phenotypic plasticity of Foxp3(+) regulatory T cells. Immunol Rev 2014; 259: 159–172.
Bin Dhuban K, Kornete M, S Mason E, Piccirillo CA . Functional dynamics of Foxp3(+) regulatory T cells in mice and humans. Immunol Rev 2014; 259: 140–158.
Harrington LE, Hatton RD, Mangan PR, Turner H, Murphy TL, Murphy KM et al. Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat Immunol 2005; 6: 1123–1132.
Park H, Li Z, Yang XO, Chang SH, Nurieva R, Wang YH et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat Immunol 2005; 6: 1133–1141.
Bettelli E, Carrier Y, Gao W, Korn T, Strom TB, Oukka M et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 2006; 441: 235–238.
Mangan PR, Harrington LE, O'Quinn DB, Helms WS, Bullard DC, Elson CO et al. Transforming growth factor-beta induces development of the T(H)17 lineage. Nature 2006; 441: 231–234.
Das J, Ren G, Zhang L, Roberts AI, Zhao X, Bothwell AL et al. Transforming growth factor beta is dispensable for the molecular orchestration of Th17 cell differentiation. J Exp Med 2009; 206: 2407–2416.
Ghoreschi K, Laurence A, Yang XP, Tato CM, McGeachy MJ, Konkel JE et al. Generation of pathogenic T(H)17 cells in the absence of TGF-beta signalling. Nature 2010; 467: 967–971.
Waickman AT, Powell JD . m TOR metabolism, and the regulation of T-cell differentiation and function. Immunol Rev 2012; 249: 43–58.
Buck MD, O'Sullivan D, Pearce EL . T cell metabolism drives immunity. J Exp Med 2015; 212: 1345–1360.
Delgoffe GM, Kole TP, Zheng Y, Zarek PE, Matthews KL, Xiao B et al. The mTOR kinase differentially regulates effector and regulatory T cell lineage commitment. Immunity 2009; 30: 832–844.
Shi LZ, Wang R, Huang G, Vogel P, Neale G, Green DR et al. HIF1alpha-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J Exp Med 2011; 208: 1367–1376.
Dang EV, Barbi J, Yang HY, Jinasena D, Yu H, Zheng Y et al. Control of T(H)17/T(reg) balance by hypoxia-inducible factor 1. Cell 2011; 146: 772–784.
Zeng H, Yang K, Cloer C, Neale G, Vogel P, Chi H . mTORC1 couples immune signals and metabolic programming to establish T(reg)-cell function. Nature 2013; 499: 485–490.
Gerriets VA, Kishton RJ, Johnson MO, Cohen S, Siska PJ, Nichols AG et al. Foxp3 and Toll-like receptor signaling balance Treg cell anabolic metabolism for suppression. Nat Immunol 2016; 17: 1459–1466.
Procaccini C, De Rosa V, Galgani M, Abanni L, Cali G, Porcellini A et al. An oscillatory switch in mTOR kinase activity sets regulatory T cell responsiveness. Immunity 2010; 33: 929–941.
Huynh A, DuPage M, Priyadharshini B, Sage PT, Quiros J, Borges CM et al. Control of PI(3) kinase in Treg cells maintains homeostasis and lineage stability. Nat Immunol 2015; 16: 188–196.
Shrestha S, Yang K, Guy C, Vogel P, Neale G, Chi H . Treg cells require the phosphatase PTEN to restrain TH1 and TFH cell responses. Nat Immunol 2015; 16: 178–187.
Ulges A, Witsch EJ, Pramanik G, Klein M, Birkner K, Buhler U et al. Protein kinase CK2 governs the molecular decision between encephalitogenic TH17 cell and Treg cell development. Proc Natl Acad Sci USA 2016; 113: 10145–10150.
Di Maira G, Salvi M, Arrigoni G, Marin O, Sarno S, Brustolon F et al. Protein kinase CK2 phosphorylates and upregulates Akt/PKB. Cell Death Differ 2005; 12: 668–677.
Ulges A, Klein M, Reuter S, Gerlitzki B, Hoffmann M, Grebe N et al. Protein kinase CK2 enables regulatory T cells to suppress excessive TH2 responses in vivo. Nat Immunol 2015; 16: 267–275.
Berod L, Friedrich C, Nandan A, Freitag J, Hagemann S, Harmrolfs K et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat Med 2014; 20: 1327–1333.
Feng G, Chan T, Wood KJ, Bushell A . Donor reactive regulatory T cells. Curr Opin Organ Transplant 2009; 14: 432–438.
Elinav E, Waks T, Eshhar Z . Redirection of regulatory T cells with predetermined specificity for the treatment of experimental colitis in mice. Gastroenterology 2008; 134: 2014–2024.
MacDonald KG, Hoeppli RE, Huang Q, Gillies J, Luciani DS, Orban PC et al. Alloantigen-specific regulatory T cells generated with a chimeric antigen receptor. J Clin Invest 2016; 126: 1413–1424.
Acknowledgements
This work was supported by a National Research Foundation of Korea (NRF) grant funded by Korean government (NRF-2014R1A2A1A11052545 and NRF-2017R1A2B3008621).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher’s note:
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Lee, W., Lee, G. Transcriptional regulation and development of regulatory T cells. Exp Mol Med 50, e456 (2018). https://doi.org/10.1038/emm.2017.313
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2017.313
This article is cited by
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
Mannosylated glycans impair normal T-cell development by reprogramming commitment and repertoire diversity
Cellular & Molecular Immunology (2023)
-
Differential distribution of steroid hormone signaling networks in the human choroid-retinal pigment epithelial complex
BMC Ophthalmology (2022)
-
Regulatory T cells in skeletal muscle repair and regeneration: recent insights
Cell Death & Disease (2022)
-
Reduced frequency of circulating regulatory T cells and their related immunosuppressive mediators in treated celiac patients
Molecular Biology Reports (2022)