Abstract
When we treated rat bone marrow stromal cells (rBMSCs) with neuronal differentiation induction media, typical unfolded protein response (UPR) was observed. BIP/GRP78 protein expression was time-dependently increased, and three branches of UPR were all activated. ATF6 increased the transcription of XBP1 which was successfully spliced by IRE1. PERK was phosphorylated and it was followed by eIF2α phosphorylation. Transcription of two downstream targets of eIF2α, ATF4 and CHOP/GADD153, were transiently up-regulated with the peak level at 24 h. Immunocytochemical study showed clear coexpression of BIP and ATF4 with NeuN and Map2, respectively. UPR was also observed during the neuronal differentiation of mouse embryonic stem (mES) cells. Finally, chemical endoplasmic reticulum (ER) stress inducers, thapsigargin, tunicamycin, and brefeldin A, dose-dependently increased both mRNA and protein expressions of NF-L, and, its expression was specific to BIP-positive rBMSCs. Our results showing the induction of UPR during neuronal differentiations of rBMSCs and mES cells as well as NF-L expression by ER stress inducers strongly suggest the potential role of UPR in neuronal differentiation.
Similar content being viewed by others
Introduction
Adult neurogenesis can occur in the mammalian central nervous system through manipulating endogenous neural progenitors or transplantating cells exogenously. Cell transplantation is one of the strategies for treatment of neural disorders, and many kinds of cells including embryonic and neural stem cells have been considered as candidates for transplantation therapy. Although neural stem cells appear to exist in the adult brain, limited availability of these cells requires exogenous sources to obtain sufficient amount of cells to replace damaged cells in conditions such as Parkinson's disease and stroke (Gage, 2000). Bone marrow stromal cells (BMSCs) arose great interest as a source of autologous stem cells with therapeutic potential for cell replacement therapies in neurodegenerative disorders and traumatic injuries, since they can be easily isolated and expanded from patients without serious ethical and technical issues (Mahmood et al., 2003; Dezawa et al., 2005).
Bone marrow stromal cells (BMSCs) are multipotent cells that can be differentiated into mesoderm-derived cells such as osteoblasts, osteoclasts, chondrocytes, adipocytes and myocytes, and may contribute to the regeneration of tissues depending on the microenvironment (Prockop, 1997). BMSCs can also differentiate into non-mesenchymal cells, such as hepatocytes, cardiomyocytes and alveolar epithelium (Jiang et al., 2002). Interestingly, recent studies demonstrated the differentiation of BMSCs into neuron in vitro (Sanchez-Ramos et al., 2000; Woodbury et al., 2002) as well as in vivo (Shichinohe et al., 2004), although it is still controversial if BMSCs indeed transdifferentiate into neuronal cells (Cogle et al., 2004; Munoz-Elias et al., 2004), or transplanted BMSCs fuse to resident cells expressing BMSC-derived markers (Alvarez-Dolado et al., 2003; Weimann et al., 2003). To support the clinical potential of BMSC transplantation, many attempts to differentiate BMSCs in vitro into neural cells have been made. In this connection, a few results showing the molecular pathways and gene expression patterns specific for in vitro neuronal differentiation of mesenchymal stem cells have been published (Jori et al., 2005; Wang et al., 2007). However, precise mechanism for neuronal differentiation or functional role of BMSCs in the brain still remains to be determined.
The endoplasmic reticulum (ER) is an important intracellular organelle that is responsible for the folding and trafficking of secretory proteins, and the biosynthesis of membrane lipids. Misfolded or unfolded proteins accumulate in the ER under various conditions evoking the ER stress, such as perturbed calcium homeostasis, cellular redox status, and the increased synthesis of secretory proteins. To alleviate the ER stress, eukaryotic cells activate a series of self-defense mechanisms collectively referred to as the unfolded protein response (UPR) which are initiated by three different membrane receptors, PERK, IRE1, and ATF6 (Yoshida, 2007). UPR also mediates apoptosis under severe ER stress. It has been reported that the ER stress is induced in various human neurodegenerative diseases, such as Parkinson's disease, Alzheimer's disease and prion diseases. Interestingly, recent studies suggest that the ER stress is also involved in cellular differentiations, as in erythropoiesis, adipogenesis, chondrogenic and osteogenic differentiations, and eyes and bone developments (Cui et al., 2000; Gass et al., 2002; Pereira et al., 2004; Yang et al., 2005). ER is abundant and well developed in neurons, and recently, Zhang et al. (2007) reported the induction of UPR during the embryonic development of the central nervous system in the mouse.
In this study, we investigated the involvement of the UPR and discussed its possible role in the differentiation into neuronal cells of rat BMSCs (rBMSCs) and mouse embryonic stem (mES) cells.
Results
Treatment of rBMSCs with NIM
rBMSCs cultured in the neuronal induction medium (NIM) displayed typical morphological changes of neuronal cells with stretched neurite-like appearance from as early as 3 h (data not shown). To evaluate the neuronal differentiation at the molecular level, we examined the expression patterns of neuronal markers using quantitative real-time PCR and immunoblot analysis. The mRNA levels of two neuronal markers neurofilament-L (NF-L) and -M (NF-M) (Figure 1A) were time-dependently increased (P < 0.05). Protein expression of a mature neuronal marker NeuN was also increased from 12 h after the induction (Figure 1B). The expression of Tuj1, an immature marker for neuronal cells, was transiently up-regulated until 6 h, and then decreased over time (Figure 1B). Expressions of astrocyte (GFAP) and oligodendrocyte (CNPase) markers were not increased by NIM (Figure 1C). These results indicate that neuron-specific, but not glia-specific, proteins were induced in rBMSCs by NIM in vitro.
Expressions of neuronal markers by the treatment of rBMSCs with NIM. Rat BMSCs were isolated and treated with NIM as described in Methods. (A) The expression of two neuronal markers, NF-L (left) and NF-M (right), were examined using real-time PCR. The relative quantities were normalized to the expression of β-actin. (B) Protein expression of mature (NeuN) and immature (Tuj1) neuronal markers were examined by immunoblot analysis. Alpha-tubulin was used for a loading control. (C) Expressions of makers for astrocyte (GFAP) and oligodendrocyte (CNPase) after 24 h of the induction were examined using RT-PCR. Data are mean ± SEM. *, P < 0.05. con, control; PI, preinduction.
Expression of UPR genes during neuronal differentiation
We next examined the expression of UPR genes in rBMSCs by NIM. The expression of BIP, a well known ER stress marker, was time-dependently increased at both protein and mRNA levels (Figure 2A and B). The precursor ATF6α (90 kDa) protein was immediately induced by NIM reaching the maximum level at 6 h, and its processed active form (50 kDa) was detected from 6 h, reached the maximum at 24 h, and returned to the basal level at 72 h (Figure 2A). NI also activated PERK- mediated signaling pathway. PERK and its downstream target eIF2α were transiently activated as evidenced by the induction of their phosphorylated forms (P-PERK and P-eIF2α) with the peak levels at 6 h and 12 h, respectively (Figure 2A). The level of XBP1 mRNA, a target of ATF6α, was also increased by NIM from 24 h (P < 0.05) (Figure 2C). The induction of IRE1-spliced XBP1 mRNA, XBP1(s), (P < 0.05) suggested the activation of IRE1α by NI (Figure 2D). Transcriptions of ATF4/CREB2 and CHOP/GADD153, two downstream targets of eIF2α, were transiently up-regulated with the maximum increase of fifteen-fold at 24 h (P < 0.05) (Figure 2E and F). Expression of another target molecule of PERK, Nrf2 (Cullinan et al., 2003; Cullinan and Diehl, 2006) increased seven-fold at 24 h (P < 0.05) (Figure 2G). To reveal the mechanisms involved in the transient temporal patterns of downstream signaling pathways of PERK/eIF2α, we examined the expression of GADD34 and P58IPK, two well-known negative feedback regulators for PERK signaling. GADD34 inhibits stress-induced gene expression and promotes recovery from translational inhibition by dephosphorylating eIF2α (Novoa et al., 2001). As we expected, the expression of GADD34 began to increase just after the time when eIF2α phosphorylation reached the maximum, and it was peaked at 72 h with the disappearance of P-eIF2α (P < 0.05) (Figure 2H). P58IPK prevents dimerization and autophosphorylation of PERK (Tan et al., 1998), thus interfering with PERK's ability to phosphorylate eIF2α. As in GADD34, temporal pattern of P58IPK expression showed an inverse correlation with that of P-PERK (P < 0.05) (Figure 2I). These data suggested the induction of typical UPR by NIM treatment in rBMSCs.
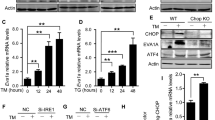
Expression and activation of UPR genes in NIM-treated rBMSCs. Rat BMSCs were isolated and treated with NIM, and the expression of UPR-related genes were examined using immunoblot analysis (A) and real time PCR (B-I). (A) BIP level was time-dependently increased. The precursor ATF6α (90 kDa) and its processed active form (50 kDa) were induced by NIM. PERK and its downstream target eIF2α were transiently activated (P-PERK and P-eIF2α) with the peak levels at 6 and 12 h. (B) The level of BIP messenger RNA was upregulated. (C) and (D) XBP1, a target of ATF6α, and IRE1-spliced XBP1, XBP1(s) mRNAs were increased by NIM. (E-I) Transcriptions of members of PERK-mediated pathway such as ATF4, CHOP, Nrf2, GADD34 and P58IPK were increased during the differentiation period. Note the transient nature of PERK-mediated signaling pathway. Alpha-tubulin was used for a loading control for immunoblot analysis. Data are mean ± SEM. *, P < 0.05. con, control; PI, preinduction.
Co-expression of UPR proteins and neuron-specific proteins
BMSCs are intrinsically heterogeneous population. To investigate whether the UPR is induced specifically in neuronally differentiated cells in response to NIM, we examined the expression of a neuronal marker NeuN and UPR proteins by immunofluorescence microscopy. Figure 3 shows the coimmunostaining of BIP and ATF4 with NeuN and MAP2, respectively, in rBMSCs which were cultured for 72 h in NIM, suggesting the possibility that UPR may induce the expression of neuronspecific proteins or vice versa in rBMSCs.
Co-localization of UPR proteins and neuron-specific proteins. Rat BMSCs were treated with NIM for 72 h, and the co-expression of FITC-labeled neuron-specific proteins (MAP2 and NeuN) (green) with Cy3-labeled UPR proteins (red) such as ATF4 and BIP was examined by immunocytochemical staining and confocal microscope. Scale bars, 50 µm.
Non-apoptotic role of neuronal differentiation-induced ER stress
It has been well-known that severe ER stress can cause apoptosis. To rule out the apoptosis- inducing role of ER stress response, we examined whether the cytotoxicity and apoptosis occurred by NIM treatment (Figure 4). While the proliferation of rBMSCs apparently stopped immediately after the treatment with NIM, their cell number was stably maintained (Figure 4A). FACS analysis also confirmed the lack of significant apoptosis or necrosis even after 4 days of the induction (Figure 4B).
Non-apoptotic role of ER stress induced in NIM-treated rBMSCs. Rat BMSCs were isolated and treated with NIM for the indicated times, and cell viability was examined using CCK-8 (A) and FACS analyses (B). (A) CCK-8 assay: Proliferation of rBMSCs was stopped immediately after the treatment with NIM, but, thereafter, it was stably maintained for 72 h at 70% of the starting level. (B) FACS analysis: No apoptosis or necrosis was observed at 0 (con), 3 (NI 3d) and 4 days (NI 4d) after the neuronal induction. Two-day treatment of adriamycin (10 µg/ml) was used as a positive control. Percentage of apoptotic cells was indicated in each figure. Data are mean ± SEM. *, P < 0.05. NI, neuronal induction.
Induction of UPR during neuronal differentiation of mouse ES cells
To test whether NIM-induced UPR is a common phenomenon, not restricted to rBMSCs, we examined whether similar pattern of UPR also occurred in neuronally-differentiating J1 mouse embryonic stem (mES) cells (Figure 5). Mouse ES cells were got through the stages of embryoid bodies, neuronal precursors and then differentiated for 6 days into neuron-like cells. In this experimental condition, mRNA expression of a neuronal marker NF-L was significantly increased immediately after the induction of differentiation (ND0) (Figure 5B), and NeuN protein level was upregulated from NP stage in a time-dependent manner (Figure 5A). UPR-related gene levels were also increased. P-PERK was temporarily upregulated from EB stage, peaked at NP stage (Figure 5A), and it was followed by the increased mRNA expressions of BIP, ATF4, CHOP and XBP1 (Figure 5B). These findings indicate that the UPR may be commonly induced during neuronal differentiation of stem cells such as mES cells and rBMSCs.
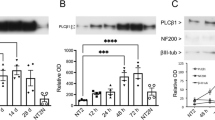
Induction of UPR during neuronal differentiation of mouse ES cells. J1 ES cells were got through the stages of embryoid bodies (EB), neuronal precursors (NP) and differentiated for 6 days (ND0-6) into neuron-like cells, and the expressions of the neuronal markers and UPR genes were examined using immunoblot (A) and real time PCR (B) analyses. (A) NeuN protein level was upregulated from NP stage in a time-dependent manner. P-PERK was temporarily increased from EB stage, and peaked at NP stage. Alpha-tubulin was used for a loading control. (B) Transcription levels of a neuronal marker NF-L and some UPR genes such as BIP, ATF4, CHOP, XBP1 and XBP1(s) were significantly increased from ND0. Data are mean ± SEM. *, P < 0.05. ND, neuronal differentiation.
Expression of a neuronal marker by chemical ER stress inducers
To investigate whether the UPR induction is simply a phenomenon secondary to neuronal differentiation or not, we examined the expression of a neuronal marker NF-L in rBMSCs by ER stress. Treatment of the cells with three well-known ER stress inducers such as thapsigargin, tunicamycin and brefeldin A at the non-cytotoxic concentrations for 48 h significantly induced the expression of NF-L mRNA at dose-dependent manner (Figure 6A, B and C) (P < 0.05). Protein expression of NF-L was also dose-dependently increased by tunicamycin (Figure 6D). BIP, a positive maker for ER stress, was increased by tunicamycin (P < 0.05) (Figure 6E). However, glial markers such as GFAP and CNPase were not increased by ER stress inducers (Figure 6F). In immunocytochemical study, NF-L and BIP were co-expressed in rBMSCs which was treated with tunicamycin (0.25 µg/ml) for 48 h (Figure 6G).
Expressions of NF-L by ER stress inducers in rBMSCs. NF-L expression in rBMSCs which were treated for 2 days with three ER stress inducers such as thapsigargin, tunicamycin or brefeldin A was examined using real time PCR (A-C), immunoblot analysis (D) or immunocytochemistry (G). (A-C) All three ER stress inducers, thapsigargin (A), tunicamycin (B) and brefeldin A (C) increased NF-L expression at dose-dependent manners. (D) Tunicamycin-induced NF-L expression was also observed at the protein level. GAPDH was used for a loading control for immunoblot analysis. (E) Expression of BIP by tunicamycin was examined using real time PCR analysis. (F) RT-PCR result showed no increase of the expressions of glial markers, GFAP and CNPase in rBMSCs which was treated for 2 days with thapsigargin (0.4 µM), tunicamycin (0.25 µg/ml), or brefeldin A (0.25 µg/ml). Beta actin was used as a control. (G) Immunocytochemical staining for BIP (green) and NF-L (red) in rBMSCs which were incubated for 2 days in the absence (con) or presence with tunicamycin (TM, 0.25 µg/ml). DAPI was used to stain nuclei (blue). Scale bars, 50 µm. Data are mean values ± SEM. *, P < 0.05.
Discussion
Stress response is a conserved mechanism that protects cells to respond to a variety of environmental and metabolic conditions. In mammalian cells, ER stress is induced in many physiological and pathological conditions (Yoshida, 2007). Here we presented the evidence for the induction of UPR in rBMSCs by NIM. BIP was induced immediately and time-dependently after the induction, and all three ER stress sensor molecules as well as their downstream signaling pathways were activated (Figure 2). UPR induction was also observed during the neuronal differentiation of mES cells (Figure 5) as well as in PC 12 cells (Cho et al., 2007). These findings strongly indicate that UPR may be a common mechanism that occurs during neuronal differentiation process. However, BMSCs are by nature a heterogeneous with subpopulations of different characteristics and UPR can be occurred ubiquitously in any type of the cells. It is therefore still possible that hematopoietic cells like macrophages or monocytes which are possibly contaminated not to be neuronally differentiated could exhibit UPR in response to NIM. To exclude this possibility, we used the specific passages of rBMSCs in which hematopoietic cells were known to be removed (Pittenger et al., 1999; Woodbury et al., 2000). Our data showing the successful differentiation of rBMSCs into adipocytes also indicate that the cell population used in this study was mesenchymal stem cells (data not shown) (Pittenger et al., 1999; Yeon et al., 2006). Moreover, immunocytochemistry result clearly showed the co-expression of neuron-specific (NeuN and MAP2) and UPR (ATF4 and BIP) proteins (Figure 3).
Insufficient adaptive responses to ER stress result in the initiation of apoptosis, and functions of ER stress-related proteins have been studied primarily from the view of ER-mediated apoptosis. Therefore, it is necessary to examine the proapoptotic role of the UPR which was induced in our experiments. CHOP is a central mediator for ER stress-induced apoptosis, and its expression under the ER stress condition is up-regulated in proportion to the level of apoptotic cell death (Eymin et al., 1997). Reports showing specific activation of ATF6 during apoptosis in myoblasts (Nakanishi et al., 2005) and the delay of proteasome inhibitor-induced apoptosis by the loss of eIF2α phosphorylation (Jiang and Wek, 2005) also suggest their roles in ER stress-induced apoptosis. In our results (Figure 2), although NIM transiently induced CHOP expression, the ATF6 activation as well as the eIF2α phosphorylation, no apoptosis occurred. These findings suggest that the ER stress was induced during the neuronal differentiation of rBMSCs with the potency which was not enough to induce apoptosis. It is also possible that mechanisms to increase cell survival could counteract these pro-apoptotic signals. Among the ER stress proteins, ATF4, XBP1 and Nrf2 have been known to enhance cell survival (Cullinan et al., 2003), and they were increased by NIM in our experiment (Figure 2G). Another possible role of the UPR proteins may be the regulation of neuronal differentiation. It is interesting that most of UPR gene expressions appeared earlier or similar to, but not later, than those of neuronal markers. For example, PERK and ATF6 began to be activated from 3 or 6 h after the induction, respectively. Neuronal maker expressions occurred from 24 or 48 h at which ATF6 and most of PERK-mediated pathways such as CHOP and ATF4 were already inactivated (Figure 2). Although it is not enough to support this idea, our results strongly suggest the possibility that UPR induction may be not an incidental phenomenon secondary to the differentiation, but an active factor for neuronal differentiation. More interestingly, chemical ER stress inducers specifically induced a neuronal marker NF-L expression, and NF-L was coexpressed with an ER stress marker BIP (Figure 6). In accord with this idea, growing evidences suggest potential roles of ER stress-related proteins in other cellular differentiation process. UPR occurs during myoblast differentiation both in vivo and in vitro (Nakanishi et al., 2005; 2007). CHOP regulates the differentiations of erythrocytes, osteocytes, chondrocytes and B cells (Cui et al., 2000; Pereira et al., 2004; Skalet et al., 2005; Yang et al., 2005). XBP1 induces osteogenic and plasma cell differentiations (Iwakoshi et al., 2003), and IRE1 increases lymphopoiesis of B cells (Zhang et al., 2005).
Differentiation into neural cells could be decided based on the changes of morphology, biochemical marker expressions or their excitability by measuring membrane potential. Although a few reports exploited electrophysiological methods (Wislet-Gendebien et al., 2005; Choi et al., 2006; Wenisch et al., 2006), most of the studies have used morphological or biochemical methods for the evaluation of in vitro neuronal differentiation. However, recent findings have cast some serious doubts on the potential role of mesenchymal stem cells to differentiate into neuron, suggesting the morphological changes of neuron-like cells as the phenomenon merely reflecting mechanical reorganization of cytoskeletal elements (Neuhuber et al., 2004; Suon et al., 2004). These findings indicate the caution that transdifferentiation of MSCs into neural cells must be estimated by biochemical rather than morphological parameters. However, the relationship between NIM-induced expression of neuronal markers and neuronal differentiation are still unclear. In our results, NIM induced neuron-like morphological changes immediately after the induction, but the expression of neuronal markers were started from 12 or 24 h after the induction (Figure 1). Moreover, in spite of time-dependent induction of NF-L expression, we could not observe neuron-like morphology in the tunicamycin-treated rBMSCs (Figure 6). These findings suggest the possibility that NIM-induced neuronal marker expression may not be related to the neuronal differentiation of BMSCs. If so, UPR is a candidate factor for the induction of neuronal marker expression by NIM. UPR can be occurred by various situations such as NIM in our experiment, and it is an easy idea that the proteins and the resulting responses induced by UPR may be different accordingly. To confirm the primary role of UPR proteins in the induction of neuronal markers or neuronal differentiation, further studies will be necessary.
Signaling molecules involved in the UPR have been reported to be required during development (Reimold et al., 2000; Urano et al., 2000), and a novel strategy that ER controls developmental pathways by a specific ER retention and release mechanisms has recently emerged (Yamamoto et al., 2005). Roles of UPR during the neuronal development could be indirectly estimated from the studies using developing brain or knock-out mice. Recently, a few reports suggested the role of ER stress during brain development such as the role of XBP1 in neurite outgrowth (Hayashi et al., 2007), ER stress induction during embryonic development of the central nervous system (Zhang et al., 2007), and the critical role of BIP for the development of cerebral cortex (Mimura et al., 2008). Embryonic lethalities were also observed in IRE1α-/-, XBP1-/- or eIF2S51A knock-in mice, suggesting critical role of these genes in the development of mice (Reimold et al., 2000; Urano et al., 2000; Scheuner et al., 2001). However, no obvious abnormalities in brain were found in the mice deficient of Nrf2, PERK, CHOP, ATF4 or ATF6 genes (Chan et al., 1996; Zinszner et al., 1998; Harding et al., 2001; Masuoka and Townes, 2002; Yamamoto et al., 2007). These findings and our results indicate that some specific, but not all, ER stress genes may play a role during neuronal differentiation. To prove causative relationship between UPR proteins and neuronal differentiation, further works using cells deficient of UPR genes, more specifically BIP, IRE1α, XBP1 or eIE2α, will be necessary.
In conclusion, we demonstrated for the first time the specific induction UPR and suggested the possibility of their active roles for the expression of neuronal markers or for the differentiation of rBMSCs and mES into neurons in vitro.
Methods
Isolation and culture of rBMSCs
The rBMSCs were isolated from the femurs of 70-80 g male Sprague-Dawley rats as described previously (Rismanchi et al., 2003). Under sterile conditions, both ends of femurs and tibias were removed, and the bone marrow was flushed out using a needle connected to a syringe filled with PBS supplemented with 2% FBS (GIBCO, Pasley, Scotland, UK). Isolated cells were then diluted and centrifuged through a density gradient (Ficoll-Paque-Plus, Amersham Pharmacia Biotech, Nuckinghamshire, UK) for 30 min at 1,000 g, and the mononuclear cells were collected and resuspended with DMEM (GIBCO) supplemented with 20% FBS. Cells from one rat were plated on a 15 cm culture dish, and incubated at 37℃ with 5% humidified CO2. Within 24 h of harvest, non-adherent hematopoietic cells were removed by vigorous shaking of the culture dishes and media replacement. Adherent rBMSCs were grown to confluency (defined as passage 0), trypsinized, and grown again to confluency (passage 1). To rule out further the contamination of macrophages, cells only at passages 3 to 8 were used for all of the experiments (Pittenger et al., 1999).
Neuronal induction of rBMSCs
Neuronal differentiation of rBMSCs was performed as described previously (Woodbury et al., 2002; Yaghoobi and Mowla, 2006) with modification. Briefly, rBMSCs were retrieved from sub-confluent culture by trypsinization, and plated at the density of 4 × 103 cells/cm2. For preinduction (PI), rBMSCs were grown for 24 h in DMEM supplemented with 20% FBS and 10 ng/ml basic fibroblast growth factor (bFGF, Invitrogen, Carlsbad, CA) (Palmer et al., 1999). For neuronal induction (NI), cells were rinsed twice with PBS and incubated with a neuronal induction media (NIM), DMEM supplemented with 100 µM butylated hydroxyanisole (BHA, Sigma, St Louis, MO), 2% DMSO (Sigma), 25 mM KCl (Sigma), 5 U/ml heparin (Sigma), 10 µM forskolin (Sigma), 10 ng/ml bFGF, 2 mM valproic acid (Sigma), and 1 × B27 (GIBCO). The media was replaced everyday. After the induction, total RNA and protein were prepared from the each culture at the indicated times (0, 1, 3, 6, 12, 24, 48, and 72 h).
Cell culture and neuronal differentiation of mES cells
The mouse blastocyst-derived ES cell line J1 was obtained from ATCC (Rockland, MD), and was propagated and maintained as described previously (Chung et al., 2005). Briefly, undifferentiated mES cells were cultured on gelatin-coated dishes in DMEM supplemented with 2 mM glutamine (Life Technologies, Boone, NC), 0.001% β-mercaptoethanol (Life Technologies), 1 × nonessential amino acids (Life Technologies), 10% donor horse serum (Sigma), and 2000 U/ml human recombinant leukemia inhibitory factor (LIF; Invitrogen).
ES cells were differentiated into embryoid bodies (EB) on non-adherent bacterial dishes (Fisher Scientific, Pittsburgh, PA) for 4 days in LIF-free EB medium containing 10% FBS instead of horse serum. EBs were then plated onto adhesive tissue culture surface (Fisher Scientific), and after 24 h in culture, selection of neuronal precursor cells (NP) was initiated in serum-free medium supplemented with insulin, transferrin, selenium, fibronectin (ITSFn) (Okabe et al., 1996). After 6-10 days of selection, cells were trypsinized and nestin-positive neuronal precursors were plated on 6 cm culture plates pre-coated with poly-L-ornithine/fibronectin in N2 medium (Johe et al., 1996) supplemented with 1 µg/ml laminin (Sigma) and 10 ng/ml bFGF (ND 0). After expansion for 4 days, bFGF was removed to induce differentiation to neuronal phenotypes for 6 days (ND 1-6).
Induction of ER Stress by the chemical inducers
The rBMSCs were cultured in DMEM containing 20% FBS for one day at a density of 1 × 105 cells/cm2, and treated with ER stress inducers, thapsigargin (0 to 0.4 µM; Sigma), tunicamycin (0 to 0.25 µg/ml; Sigma), and brefeldin A (0 to 0.25 µg/ml, Sigma) for 2 days.
Total RNA isolation, RT-PCR and real-time PCR
Total RNA was prepared using TRIzol Reagent (Invitrogen). cDNAs were synthesized by using M-MLV reverse transcriptase (Promega, Madison, WI). The quantitative real time PCR analysis was performed on three independent RNA samples by using Mx3000P Multiplex Quantitative PCR instrument (Stratagene, La Jolla, CA) with the SYBR Green Q-PCR Master Mix (Takara, Shiga, Japan). The relative amount of mRNAs was calculated using the ΔΔCt method with the GAPDH mRNA as an internal control. The primers used for RT-PCR and real time PCR analysis are listed in Supplemental Data Table S1.
Immunoblot analysis
Cells were lysed in RIPA buffer (20 mM Tris-HCl, pH 7.5, 0.1% SDS, 1% Triton X-100, 1% sodium deoxycholate, 150 mM NaCl, 1 mM EDTA, 1% NP-40, and proteinase inhibitor cocktail) and spun for 30 min at 13,200 g to remove debris. Forty micrograms of total protein were separated by 9 or 12% SDS-PAGE and transferred onto a nitrocellulose membrane (Schleicher & Schell, Dassel, Germany). The membranes were blocked with 3% BSA and incubated overnight at 4℃ with the antisera against BIP (1:500; BD Biosciences, San Diego, CA), phosphorylated PERK (1:200; Cell Signaling, Danvers, MA), phosphorylated eIF2α (1:200; Cell Signaling), ATF6 (1:200; Santa Cruz Biotechnology, Santa Cruz, CA), CHOP (1:200; Santa Cruz), XBP1 (1:200; Santa Cruz), NeuN (1:500; Chemicon, Temecula, CA), NF-L (1:200; Santa Cruz), Tuj1 (1:500; Covance, New Jersey, NY), GAPDH (1:1,000; Sigma), and α-tubulin (1:1,000; Sigma). Antibody binding was detected using HRP-conjugated anti-rabbit or anti-mouse IgG (1:2,000; Santa Cruz) and the immunoreactive bands were visualized with ECL method (Amersham Pharmacia Biotech).
Immunocytochemistry
Cells were cultured on poly-D-lysine (50 µg/ml)-coated chambered coverslips (CultureWell, Grace Bio-Labs, Bend, OR) and treated with NIM. After 72 h, NIM-treated or control cells were washed with 0.01 M PBS and then fixed in 2% paraformaldehyde (Sigma) for 15 min on ice. After the fixation, the slides were washed with PBS, blocked for 1 h with 10% normal donkey serum (Sigma) in PBS containing 0.1-0.5% Triton X-100 (Sigma) at room temperature, and then incubated overnight with biotinylated anti-NeuN (1:200), mouse anti-MAP2 (1:200; Chemicon) or mouse anti-NF-L (1:200; Santa Cruz) antibodies at room temperature. The slides were washed with PBS and incubated for 1 h with FITC-conjugated streptavidin or goat anti-mouse IgG (1:1,000; JacksonImmunoResearch, West Grove, PA) at room temperature. For the binding of second primary antibodies, the same procedure used for the first primary antibody was repeated. After the slides were washed with PBS, blocked again for 1 h with same blocking medium, and then incubated overnight with second primary antibodies. Second primary antibodies (1:200) used were mouse anti-BIP (BD Biosciences) and rabbit anti-ATF4 (Santa Cruz) antibodies. After washing, the cells were incubated with Cy3-conjugated goat anti-mouse or anti-rabbit IgG (1:1,000; Jackson ImmunoResearch), and mounted with Vectashield (Vector Laboratories, Burlingame, CA). Fluorescence images were visualized through a Zeiss 510LSM META laser-scanning microscope (Carl Zeiss, Oberkochen, Germany).
Determination of cell survival rates
Cells (2 × 103) were cultured in each well of 96 well tissue culture plates for one day. After the induction of neuronal differentiation or the treatment with ER stress inducers, growth rate was evaluated using the cell counting kit (CCK-8 kit, Dojindo, Kumamoto, Japan) following to the manufacturer's instructions by measuring the absorbance at 450 nm on a microplate reader.
FACS analysis
Apoptotic cells were detected with an annexin V-FITC and propidium iodide (PI) assay kit (Vybrant Apoptosis Assay kit #3, Invitrogen). For FACS analysis, 1 × 105 cells were seeded onto 10 cm plates. After NI for 3 or 4 days, cells were trypsinized, washed with PBS, and resuspended in 100 µl of binding buffer containing annexin V-FITC and PI according to the manufacturer's instructions. After 15 min of incubation, 400 µl of binding buffer was added and the analysis was performed using a FACScan flow cytometer (BD Biosciences), and the percentages of apoptotic cells were analyzed with CellQuest software (BD Biosciences). FACS analysis was repeated at least three times for each sample.
Statistical analysis
All values were expressed as mean ± SEM from at least 3 independent experiments. Data were analyzed using one-way ANOVA between subjects, and post hoc comparisons were made using the Tukey HSD test. Statistical analyses were performed using Graphpad Prism 4 for Windows (Graphpad Prism Software Inc., La Jolla, CA). In all cases, statistical significance was set at P < 0.05.
Abbreviations
- bFGF:
-
basic fibroblast growth factor
- BMSC:
-
bone marrow stromal cells
- EB:
-
embrioid body
- ER:
-
endoplasmic reticulum
- mES cells:
-
mouse embryonic stem cells
- ND:
-
neuronal differentiation
- NF-L:
-
neurofilament-L
- NF-M:
-
neurofilament-M
- NI:
-
neuronal induction
- NIM:
-
neuronal induction medium
- NP:
-
neuronal precursor
- PI:
-
propidium iodide
- UPR:
-
unfolded protein response
References
Alvarez-Dolado M, Pardal R, Garcia-Verdugo JM, Fike JR, Lee HO, Pfeffer K, Lois C, Morrison SJ, Alvarez-Buylla A . Fusion of bone-marrow-derived cells with Purkinje neurons, cardiomyocytes and hepatocytes . Nature 2003 ; 425 : 968 - 973
Chan K, Lu R, Chang JC, Kan YW . NRF2, a member of the NFE2 family of transcription factors, is not essential for murine erythropoiesis, growth, and development . Proc Natl Acad Sci USA 1996 ; 93 : 13943 - 13948
Cho YM, Jang YS, Jang YM, Soe JY, Kim HS, Lee JH, Jeong SW, Kim IK, Kwon OJ . Involvement of endoplasmic reticulum stress response in the neuronal differentiation . Korean J Physiol Pharmacol 2007 ; 11 : 239 - 246
Choi CB, Cho YK, Prakash KV, Jee BK, Han CW, Paik YK, Kim HY, Lee KH, Chung N, Rha HK . Analysis of neuron-like differentiation of human bone marrow mesenchymal stem cells . Biochem Biophys Res Commun 2006 ; 350 : 138 - 146
Chung S, Hedlund E, Hwang M, Kim DW, Shin BS, Hwang DY, Jung Kang U, Isacson O, Kim KS . The homeodomain transcription factor Pitx3 facilitates differentiation of mouse embryonic stem cells into AHD2-expressing dopaminergic neurons . Mol Cell Neurosci 2005 ; 28 : 241 - 252
Cogle CR, Yachnis AT, Laywell ED, Zander DS, Wingard JR, Steindler DA, Scott EW . Bone marrow transdifferentiation in brain after transplantation: a retrospective study . Lancet 2004 ; 363 : 1432 - 1437
Cui K, Coutts M, Stahl J, Sytkowski AJ . Novel interaction between the transcription factor CHOP (GADD153) and the ribosomal protein FTE/S3a modulates erythropoiesis . J Biol Chem 2000 ; 275 : 7591 - 7596
Cullinan SB, Diehl JA . Coordination of ER and oxidative stress signaling: the PERK/Nrf2 signaling pathway . Int J Biochem Cell Biol 2006 ; 38 : 317 - 332
Cullinan SB, Zhang D, Hannink M, Arvisais E, Kaufman RJ, Diehl JA . Nrf2 is a direct PERK substrate and effector of PERK-dependent cell survival . Mol Cell Biol 2003 ; 23 : 7198 - 7209
Dezawa M, Hoshino M, Nabeshima Y, Ide C . Marrow stromal cells: implications in health and disease in the nervous system . Curr Mol Med 2005 ; 5 : 723 - 732
Eymin B, Dubrez L, Allouche M, Solary E . Increased gadd153 messenger RNA level is associated with apoptosis in human leukemic cells treated with etoposide . Cancer Res 1997 ; 57 : 686 - 695
Gage FH . Mammalian neural stem cells . Science 2000 ; 287 : 1433 - 1438
Gass JN, Gifford NM, Brewer JW . Activation of an unfolded protein response during differentiation of antibody- secreting B cells . J Biol Chem 2002 ; 277 : 49047 - 49054
Harding HP, Zeng H, Zhang Y, Jungries R, Chung P, Plesken H, Sabatini DD, Ron D . Diabetes mellitus and exocrine pancreatic dysfunction in perk-/- mice reveals a role for translational control in secretory cell survival . Mol Cell 2001 ; 7 : 1153 - 1163
Hayashi A, Kasahara T, Iwamoto K, Ishiwata M, Kametani M, Kakiuchi C, Furuichi T, Kato T . The role of brain-derived neurotrophic factor (BDNF)-induced XBP1 splicing during brain development . J Biol Chem 2007 ; 282 : 34525 - 34534
Iwakoshi NN, Lee AH, Vallabhajosyula P, Otipoby KL, Rajewsky K, Glimcher LH . Plasma cell differentiation and the unfolded protein response intersect at the transcription factor XBP-1 . Nat Immunol 2003 ; 4 : 321 - 329
Jiang HY, Wek RC . Phosphorylation of the alpha-subunit of the eukaryotic initiation factor-2 (eIF2alpha) reduces protein synthesis and enhances apoptosis in response to proteasome inhibition . J Biol Chem 2005 ; 280 : 14189 - 14202
Jiang Y, Jahagirdar BN, Reinhardt RL, Schwartz RE, Keene CD, Ortiz-Gonzalez XR, Reyes M, Lenvik T, Lund T, Blackstad M, Du J, Aldrich S, Lisberg A, Low WC, Largaespada DA, Verfaillie CM . Pluripotency of mesenchymal stem cells derived from adult marrow . Nature 2002 ; 418 : 41 - 49
Johe KK, Hazel TG, Muller T, Dugich-Djordjevic MM, McKay RD . Single factors direct the differentiation of stem cells from the fetal and adult central nervous system . Genes Dev 1996 ; 10 : 3129 - 3140
Jori FP, Napolitano MA, Melone MA, Cipollaro M, Cascino A, Altucci L, Peluso G, Giordano A, Galderisi U . Molecular pathways involved in neural in vitro differentiation of marrow stromal stem cells . J Cell Biochem 2005 ; 94 : 645 - 655
Mahmood A, Lu D, Lu M, Chopp M . Treatment of traumatic brain injury in adult rats with intravenous administration of human bone marrow stromal cells . Neurosurgery 2003 ; 53 : 697 - 703
Masuoka HC, Townes TM . Targeted disruption of the activating transcription factor 4 gene results in severe fetal anemia in mice . Blood 2002 ; 99 : 736 - 745
Mimura N, Yuasa S, Soma M, Jin H, Kimura K, Goto S, Koseki H, Aoe T . Altered quality control in the endoplasmic reticulum causes cortical dysplasia in knock-in mice expressing a mutant BiP . Mol Cell Biol 2008 ; 28 : 293 - 301
Munoz-Elias G, Marcus AJ, Coyne TM, Woodbury D, Black IB . Adult bone marrow stromal cells in the embryonic brain: engraftment, migration, differentiation, and long-term survival . J Neurosci 2004 ; 24 : 4585 - 4595
Nakanishi K, Sudo T, Morishima N . Endoplasmic reticulum stress signaling transmitted by ATF6 mediates apoptosis during muscle development . J Cell Biol 2005 ; 169 : 555 - 560
Nakanishi K, Dohmae N, Morishima N . Endoplasmic reticulum stress increases myofiber formation in vitro . Faseb J 2007 ; 21 : 2994 - 3003
Neuhuber B, Gallo G, Howard L, Kostura L, Mackay A, Fischer I . Reevaluation of in vitro differentiation protocols for bone marrow stromal cells: disruption of actin cytoskeleton induces rapid morphological changes and mimics neuronal phenotype . J Neurosci Res 2004 ; 77 : 192 - 204
Novoa I, Zeng H, Harding HP, Ron D . Feedback inhibition of the unfolded protein response by GADD34-mediated dephosphorylation of eIF2alpha . J Cell Biol 2001 ; 153 : 1011 - 1022
Okabe S, Forsberg-Nilsson K, Spiro AC, Segal M, McKay RD . Development of neuronal precursor cells and functional postmitotic neurons from embryonic stem cells in vitro . Mech Dev 1996 ; 59 : 89 - 102
Palmer TD, Markakis EA, Willhoite AR, Safar F, Gage FH . Fibroblast growth factor-2 activates a latent neurogenic program in neural stem cells from diverse regions of the adult CNS . J Neurosci 1999 ; 19 : 8487 - 8497
Pereira RC, Delany AM, Canalis E . CCAAT/enhancer binding protein homologous protein (DDIT3) induces osteoblastic cell differentiation . Endocrinology 2004 ; 145 : 1952 - 1960
Pittenger MF, Mackay AM, Beck SC, Jaiswal RK, Douglas R, Mosca JD, Moorman MA, Simonetti DW, Craig S, Marshak DR . Multilineage potential of adult human mesenchymal stem cells . Science 1999 ; 284 : 143 - 147
Prockop DJ . Marrow stromal cells as stem cells for nonhematopoietic tissues . Science 1997 ; 276 : 71 - 74
Reimold AM, Etkin A, Clauss I, Perkins A, Friend DS, Zhang J, Horton HF, Scott A, Orkin SH, Byrne MC, Grusby MJ, Glimcher LH . An essential role in liver development for transcription factor XBP-1 . Genes Dev 2000 ; 14 : 152 - 157
Rismanchi N, Floyd CL, Berman RF, Lyeth BG . Cell death and long-term maintenance of neuron-like state after differentiation of rat bone marrow stromal cells: a comparison of protocols . Brain Res 2003 ; 991 : 46 - 55
Sanchez-Ramos J, Song S, Cardozo-Pelaez F, Hazzi C, Stedeford T, Willing A, Freeman TB, Saporta S, Janssen W, Patel N, Cooper DR, Sanberg PR . Adult bone marrow stromal cells differentiate into neural cells in vitro . Exp Neurol 2000 ; 164 : 247 - 256
Scheuner D, Song B, McEwen E, Liu C, Laybutt R, Gillespie P, Saunders T, Bonner-Weir S, Kaufman RJ . Translational control is required for the unfolded protein response and in vivo glucose homeostasis . Mol Cell 2001 ; 7 : 1165 - 1176
Shichinohe H, Kuroda S, Lee JB, Nishimura G, Yano S, Seki T, Ikeda J, Tamura M, Iwasaki Y . in vivo tracking of bone marrow stromal cells transplanted into mice cerebral infarct by fluorescence optical imaging . Brain Res 2004 ; 13 : 166 - 175
Skalet AH, Isler JA, King LB, Harding HP, Ron D, Monroe JG . Rapid B cell receptor-induced unfolded protein response in nonsecretory B cells correlates with pro- versus antiapoptotic cell fate . J Biol Chem 2005 ; 280 : 39762 - 39771
Suon S, Jin H, Donaldson AE, Caterson EJ, Tuan RS, Deschennes G, Marshall C, Iacovitti L . Transient differentiation of adult human bone marrow cells into neuron-like cells in culture: development of morphological and biochemical traits is mediated by different molecular mechanisms . Stem Cells Dev 2004 ; 13 : 625 - 635
Tan SL, Gale MJ, Katze MG . Double-stranded RNAindependent dimerization of interferon-induced protein kinase PKR and inhibition of dimerization by the cellular P58IPK inhibitor . Mol Cell Biol 1998 ; 18 : 2431 - 2443
Urano F, Bertolotti A, Ron D . IRE1 and efferent signaling from the endoplasmic reticulum . J Cell Sci 2000 ; 113 Pt 21 : 3697 - 3702
Wang TT, Tio M, Lee W, Beerheide W, Udolph G . Neural differentiation of mesenchymal-like stem cells from cord blood is mediated by PKA . Biochem Biophys Res Commun 2007 ; 357 : 1021 - 1027
Weimann JM, Johansson CB, Trejo A, Blau HM . Stable reprogrammed heterokaryons form spontaneously in Purkinje neurons after bone marrow transplant . Nat Cell Biol 2003 ; 5 : 959 - 966
Wenisch S, Trinkaus K, Hild A, Hose D, Heiss C, Alt V, Klisch C, Meissl H, Schnettler R . Immunochemical, ultrastructural and electrophysiological investigations of bone- derived stem cells in the course of neuronal differentiation . Bone 2006 ; 38 : 911 - 921
Wislet-Gendebien S, Hans G, Leprince P, Rigo JM, Moonen G, Rogister B . Plasticity of cultured mesenchymal stem cells: switch from nestin-positive to excitable neuron-like phenotype . Stem Cells 2005 ; 23 : 392 - 402
Woodbury D, Schwarz EJ, Prockop DJ, Black IB . Adult rat and human bone marrow stromal cells differentiate into neurons . J Neurosci Res 2000 ; 61 : 364 - 370
Woodbury D, Reynolds K, Black IB . Adult bone marrow stromal stem cells express germline, ectodermal, endodermal, and mesodermal genes prior to neurogenesis . J Neurosci Res 2002 ; 69 : 908 - 917
Yaghoobi MM, Mowla SJ . Differential gene expression pattern of neurotrophins and their receptors during neuronal differentiation of rat bone marrow stromal cells . Neurosci Lett 2006 ; 397 : 149 - 154
Yamamoto A, Nagano T, Takehara S, Hibi M, Aizawa S . Shisa promotes head formation through the inhibition of receptor protein maturation for the caudalizing factors, Wnt and FGF . Cell 2005 ; 120 : 223 - 235
Yamamoto K, Sato T, Matsui T, Sato M, Okada T, Yoshida H, Harada A, Mori K . Transcriptional induction of mammalian ER quality control proteins is mediated by single or combined action of ATF6alpha and XBP1 . Dev Cell 2007 ; 13 : 365 - 376
Yang L, Carlson SG, McBurney D, Horton WE . Multiple signals induce endoplasmic reticulum stress in both primary and immortalized chondrocytes resulting in loss of differentiation, impaired cell growth, and apoptosis . J Biol Chem 2005 ; 280 : 31156 - 31165
Yeon Lim, Jeun SS, Lee KJ, Oh JH, Kim SM, Park SI, Jeong CH, Kang SG . Multiple stem cell traits of expanded rat bone marrow stromal cells . Exp Neurol 2006 ; 199 : 416 - 426
Yoshida H . ER stress and diseases . The FEBS J 2007 ; 274 : 630 - 658
Zhang K, Wong HN, Song B, Miller CN, Scheuner D, Kaufman RJ . The unfolded protein response sensor IRE1alpha is required at 2 distinct steps in B cell lymphopoiesis . J Clin Invest 2005 ; 115 : 268 - 281
Zhang X, Szabo E, Michalak M, Opas M . Endoplasmic reticulum stress during the embryonic development of the central nervous system in the mouse . Int J Dev Neurosci 2007 ; 25 : 455 - 463
Zinszner H, Kuroda M, Wang X, Batchvarova N, Lightfoot RT, Remotti H, Stevens JL, Ron D . CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum . Genes Dev 1998 ; 12 : 982 - 995
Acknowledgements
This work was supported by a grant R13-2002-005-04002-0 from the MRC of the Korea Science and Engineering Foundation. The authors are grateful to Dr. Ann-Hwee Lee (Harvard School of Public Health, Boston, MA) for his critical comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Experimental & Molecular Medicine website
Supplementary information
Rights and permissions
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Cho, Y., Jang, YS., Jang, YM. et al. Induction of unfolded protein response during neuronal induction of rat bone marrow stromal cells and mouse embryonic stem cells. Exp Mol Med 41, 440–452 (2009). https://doi.org/10.3858/emm.2009.41.6.049
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3858/emm.2009.41.6.049
Keywords
This article is cited by
-
A lifetime of stress: ATF6 in development and homeostasis
Journal of Biomedical Science (2018)
-
In vitro culture expansion impairs chondrogenic differentiation and the therapeutic effect of mesenchymal stem cells by regulating the unfolded protein response
Journal of Biological Engineering (2018)
-
Mechanisms of protein homeostasis (proteostasis) maintain stem cell identity in mammalian pluripotent stem cells
Cellular and Molecular Life Sciences (2018)
-
Novel KDM1A inhibitors induce differentiation and apoptosis of glioma stem cells via unfolded protein response pathway
Oncogene (2017)
-
Neurogenic transdifferentiation of human adipose-derived stem cells? A critical protocol reevaluation with special emphasis on cell proliferation and cell cycle alterations
Histochemistry and Cell Biology (2010)