Abstract
The Bajo, the world’s largest remaining sea nomad group, are scattered across hundreds of recently settled communities in Island Southeast Asia, along the coasts of Indonesia, Malaysia and the Philippines. With a significant role in historical trading, the Bajo lived until recently as nomads, spending their entire lives on houseboats while moving long distances to fish and trade. Along the routes they traveled, the Bajo settled and intermarried with local land-based groups, leading to ‘maritime creolization’, a process whereby Bajo communities retained their culture, but assimilated – and frequently married into – local groups. The origins of the Bajo have remained unclear despite several hypotheses from oral tradition, culture and language, all currently without supporting genetic evidence. Here, we report genome-wide SNP analyses on 73 Bajo individuals from three communities across Indonesia – the Derawan of Northeast Borneo, the Kotabaru of Southeast Borneo and the Kendari of Southeast Sulawesi, with 87 new samples from three populations surrounding the area where these Bajo peoples live. The Bajo likely share a common connection with Southern Sulawesi, but crucially, each Bajo community also exhibits unique genetic contributions from neighboring populations.
Similar content being viewed by others
Introduction
Rapid advances in sea faring technologies in Island Southeast Asia (ISEA) around 5000 years ago created an intricate network of maritime interactions, the leading example being the well-known expansion of Austronesian peoples.1, 2, 3 Triggering inter-continental maritime connections linking ISEA with East Africa and Remote Oceania,2, 4, 5, 6, 7, 8 these contacts drove exchanges of goods, ideas, cultures and people around the Indo-Pacific region.9, 10 Sea-orientated populations, including sea nomads, emerged from this milieu, dominating trade within ISEA for centuries and helping to structure population interactions across Indonesia and beyond. Today, the Indonesian archipelago hosts ~600 ethnic groups,11 of which only a handful are known for their sea-based lifestyles. Some, like the Bugis and Makassar of Southern Sulawesi,12 are maritime inter-regional traders that arose within the framework of regional empires, such as Malay/Hindu Śrīvijaya and Majapahit. However, these groups still have homeland territories on land. Far more extreme are ethnic groups that subsist entirely detached from the land, living their whole lives aboard small boats, and as recently as 40 years ago, living as nomadic seafarers.13, 14, 15
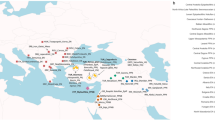
The biggest group, the Bajo (also Bajaw, Bajau or Sama-Bajau),16, 17 number approximately one million people, who today live in numerous scattered hamlets and villages recently created by the Indonesian government along the coasts of the Indonesian archipelago, as well as Sabah in Malaysia, and the Sulu archipelago and South-Western Mindanao in the Southern Philippines.13, 18, 19, 20 The geographical distribution of Bajo communities overlaps large parts of the coral triangle, which contains one of the highest rates of marine biodiversity in the world, thus underpinning the Bajo economy based on exploiting marine resources including fish, tortoise shell and sea cucumber. Within Indonesia, the Bajo presence extends over a wide geographical area (Figure 1). Historically, Bajo were frequently associated with Sulawesi Bugis traders and ship owners12, 17 and were well known for traveling with their families, even for long-distance journeys reaching as far as New Guinea and Australia.21 The Bajo may have mediated westward dispersals into the Indian Ocean, perhaps even having a role in the Indonesian settlement of Madagascar.22
Although some Bajo communities live far apart today, they still have similar social and cultural features, including shared shipbuilding and fishing culture, traditions and myths.17, 23, 24 Their languages belong to a single subfamily, the Sama-Bajau subgroup on the West Malayo-Polynesian branch of the Austronesian language family.25, 26 This subgroup includes at least nine languages,11, 18 with its highest diversity in Sabah (North Borneo) and the Southern Philippines.18 However, the Sama-Bajau languages of Indonesia are poorly documented, and an ongoing survey has identified at least three unrecognized languages (Grangé, personal communication). Some of these languages are mutually unintelligible, suggesting that the Bajo diaspora started centuries ago, fitting with oral tradition. Numerous loanwords indicate that the languages spoken by the Bajo were influenced by neighboring ethnic groups with whom the Bajo interacted and socialized, in a process called ‘maritime creolization’.24 However, the extent of these social interactions on the genetic composition of Bajo communities remains unknown.
The Bajo have no written history, instead relying on oral tradition, especially epic songs, which say little about their early history. Hypotheses about their origins have been drawn from this folklore, as well as linguistic studies and rare records from European sailors from the 16th century onward.16 The Bajo diaspora may have originated in Johor, Malaysia,13 or even Arabia,23 according to oral tradition. Brunei and Southern Sulawesi have also been proposed based on other Bajo stories.23, 27, 28 Linguistic surveys point toward the Sulu archipelago of the Philippines,18 and at an earlier stage, to Southeast Borneo.26 None of these hypotheses have been tested with genetic data.
Here, we undertake a genomic survey to help clarify the history of Bajo sea nomad populations. We present genome-wide analyses from three Bajo communities (n=73; Supplementary Table S1), representing different Sama-Bajaw dialects, together with comparative data from neighboring populations potentially connected historically with the Bajo. Using this large comparative data set, we investigate the genetic origins and history of the Bajo, and characterize the genetic impact of their near-unique lifestyle as some of the world’s last remaining sea nomads.
Materials and methods
Ethics
Biological sampling was conducted by the Eijkman Institute for Molecular Biology, with the assistance of Indonesian Public Health clinic staff, following protocols for the protection of human subjects established by the Eijkman Institute. All samples were collected with informed consent from unrelated individuals. Collection and use of these samples was approved by the Research Ethics Commission at the Eijkman Institute for Molecular Biology, Indonesia.
Samples, data set integration and quality control
Subjects were surveyed for language affiliation, current residence, familial birthplaces and a short genealogy of four generations to establish regional ancestry. A total of 47 saliva samples were collected using the Oragene saliva sampling kit (DNA Genotek Inc., ON, Canada) from two Bajo communities: Derawan (n=18) in coastal Northeastern Borneo, Indonesia and Kotabaru (n=23) in coastal Southeastern Borneo, Indonesia (Figure 1). DNA was extracted using the standard kit protocol. We also added DNA samples from the Samihim in Eastern Borneo (n=25), the Bugis of Southern Sulawesi (n=25), the Mandar of Southern Sulawesi (n=23) and North Maluku individuals from various linguistic groups (n=14) as comparative populations. This sampling strategy is relevant for the statistical tests that are described below, both on population structure and admixture. Genome-wide SNP genotypes were generated using the Illumina Human Omni5 Bead Chip (Illumina Inc., San Diego, CA, USA), which surveys 4 284 426 single-nucleotide markers semi-regularly spaced across the genome. Genotype data from previously published Bajo individuals from the Kendari community of Southern Sulawesi were also included (n=32).29 New genotyping data have been deposited at the European Genome-Phenome Archive (EGA), which is hosted by the EBI and CRG, under accession number EGAS00001002246.
A comparative data set was built from 110 worldwide populations comprising an additional 2256 individuals (Supplementary Table S1). Data quality controls were performed using PLINK v1.9:30 (i) to avoid close relatives, relatedness was measured between all pairs of individuals within each population using an identity-by-descent (IBD) estimation with upper threshold of 0.25 (second-degree relatives); (ii) SNPs that failed the Hardy–Weinberg exact test (P<10−6) were excluded; (iii) samples with an overall call rate <0.99 and individual SNPs with missing rates >0.05 across all samples in each population were excluded. The final data set contains 230 833 SNPs. Genotypes were then phased with SHAPEIT v231 using the 1000 Genomes Project phased data32 as a reference panel and the HapMap phase II genetic map. For specific analyses mentioned below, variants in high linkage disequilibrium (LD) (r2> 0.5; 50 SNP sliding windows) were also pruned, leaving a final data set of 168 368 SNPs.
Population structure
Population structure was evaluated using a suite of different programs, each relying on specific algorithms and types of data, to obtain the most relevant and robust interpretations. A fineSTRUCTURE v2.0733 analysis was performed using 2 × 106 Markov Chain Monte Carlo iterations, discarding the first 106 iterations as ‘burn in’, and sampling from the posterior distribution every 104 iterations following the burn in. This analysis detects shared IBD fragments between each pair of individuals, without self-copying, as calculated with Chromopainter v2.033 to perform a model-based Bayesian clustering of genotypes. From the results, a co-ancestry heat map and dendrogram were built to visualize the number of statistically defined clusters that best describe the data. Principal component analysis (PCA) was performed using the ‘smartpca’ algorithm of EIGENSOFT v6.0.1.34 The runs of homozygosity (ROH) and inbreeding coefficient (FIS) analyses were performed in PLINK v1.9. FST distance calculations were calculated with EIGENSOFT v6.0.1. To ascertain the significance of each pairwise FST value, 10 000 bootstraps were conducted using StAMPP,35 from which probability values were determined.
Population admixture
Admixture scenarios are determined from statistically complex models that rely a priori on the algorithms, and their assumptions, implemented in each program. To compensate for the potential biases of individual methods, we based our interpretations on the convergence of results from multiple different programs and different types of data. ADMIXTURE v1.3036 was used to estimate the genomic ancestry profile of individuals using maximum likelihood for components (K) from K=2 to K=20. Ten replicates were run at each value of K with different random seeds, then merged and assessed for clustering quality using CLUMPP,37 and the cross-validation value was calculated to determine the optimal number of genomic components. To determine the sex bias of admixture for all Bajo communities, unsupervised ADMIXTURE analysis were run on K=2 using both autosomal and X chromosome SNPs using Igorot and PNG highlanders as proxies for East Asian and Papuan ancestry, respectively. Significance tests of the proportion of the Papuan component between the autosomes and X chromosome for all Bajo communities were conducted using the one-tailed Wilcoxon test. Gene flow between populations was first investigated using TreeMix v1.12,38 with blocks of 200 SNPs to account for LD, and migration edges added sequentially until the model explained 99% of the variance. The three-population (f3) test was performed as implemented in ADMIXTOOLS v1.3.39 Haplotype sharing using the Refined IBD algorithm of Beagle v.4.040 was computed to estimate the total number of shared genetic fragments (logarithm of odds ratio >3) between each pair of individuals. Finally, we used Chromopainter v233 and GLOBETROTTER v141 to estimate the ratios and dates of potential admixture events. For all results presented here, we standardized each co-ancestry curve by a ‘NULL’ individual designed to eliminate any spurious LD patterns not attributable to that expected under a genuine admixture event,41 and consistency between each estimated parameter was checked, although we note that results were similar when not performing this standardization. The ‘best-guess’ scenario given by GLOBETROTTER was considered for each target population. Using the parental populations given by GLOBETROTTER, we ran 100 bootstrap iterations to estimate admixture dates, assuming a generation interval of 28 years for all analyses.42 With the parental populations given by GLOBETROTTER, dates of admixture were also estimated using MALDER v1.3.43
Results
We studied genetic variation in three Bajo communities spread across large parts of their geographical range: the Derawan of coastal Northeastern Borneo (n=18, B-DRW), the Kotabaru of coastal Southeastern Borneo (n=23, B-KTBR) and the Kendari of coastal Sulawesi (n=32, B-KDR). To determine the population structure of these three Bajo communities, a PCA was performed using 645 385 overlapping SNPs in just the Bajo (Figure 2). Individuals from the three groups form distinguishable clusters. PC1 (16.2% variance explained) separates the Kotabaru Bajo from the two other groups, whereas PC2 (12.9% variance explained) differentiates the Kendari Bajo from the Derawan Bajo. Interestingly, there is no overlap between the groups presently living in Borneo.
The regional connections of the Bajo were determined from 230 833 overlapping SNPs in 116 surrounding populations. A clear division appears between East Asia/Mainland Southeast Asia and Island Southeast Asia, notably separating Papuan/Eastern Indonesian populations (PC1) from Western Indonesian populations (PC2). All Bajo individuals fall within the Island Southeast Asia cluster, specifically with other Indonesian groups (Supplementary Figure S1). As before, all three Bajo communities still form their own clusters with limited overlap. Most Bajo individuals lie close to populations from Sulawesi, such as the Bugis and Mandar. The Derawan Bajo cluster close to Philippine populations; the Kotabaru Bajo cluster close to Borneo populations; whereas the Kendari Bajo have connections with eastern Indonesia, such as Sumba and North Maluku, and with Papuans.
The PCA results are consistent with fineSTRUCTURE clustering on phased genotype data (Supplementary Figure S2), which shows that all three Bajo communities form a single group, but trend toward their close geographic neighbors. Conversely, pairwise FST values (P≤1 × 10−4 for all FST pairs) suggest that all three Bajo communities have closer genetic ties to their surrounding populations than between themselves (Supplementary Table S2), thus hinting that genetic connections within the Bajo are correspondingly weaker. For instance, the Kendari Bajo have closest genetic distances with Sulawesi Bugis and Mandar; the Kotabaru Bajo with Borneo Banjar and Malay; and the Derawan Bajo with Philippine populations and Borneo Lebbo. Geography, and interactions with local groups, are therefore dominant features in the development of Bajo genetic diversity.
However, all Bajo individuals do share common patterns of genetic ancestry, as revealed by ADMIXTURE analysis (Figure 3,Supplementary Figures S3 and S4). The three Bajo communities have an admixed profile with two major Asian components and a Papuan component, but in varying proportions. The Kendari Bajo have more of the Papuan component (red) than the two Borneo Bajo groups (~20%), in keeping with their location further east. The Asian genetic ancestry is formed by similar components as for other Indonesian groups, with three main contributions: one East Asian (orange), two Austronesian components (pink and yellow) and an indigenous peninsular Malaysia component (cyan), cumulatively summing to 80–90%. The three Bajo communities only differ by relatively minor proportions of genomic ancestry that can be linked to their specific locations: minor Negrito Philippine (Aeta and Batak) components, with white and gray colors respectively, are observed in the Derawan Bajo (~1–2%); and an Indian component (green) is detected in both the Derawan Bajo and Kotabaru Bajo (~6%). Interestingly, this Indian component was not clearly detected in the Kendari Bajo, contra Mörseburg et al.,44 probably because of its very low proportion. We detect sex biased admixture in Kotabaru Bajo and Derawan Bajo (one-tailed Wilcoxon test; P<0.01), but not in Kendari Bajo nor in the Bugis (one-tailed Wilcoxon test; P>0.05). A higher proportion of Papuan X chromosomes relative to the autosomal contribution is also observed (Supplementary Figure S5).
The f3 statistics suggest that the Derawan and Kendari Bajo are admixed (Supplementary Table S3). However, defining the Kotabaru Bajo as a daughter population, all possible surrogate population combinations return positive f3 statistics with Z-scores >−2, indicating no significant gene flow, or recent bottlenecks, or founder effects,39 as also suggested by the Admixture plot at K=20 (Supplementary Figure S3). This is consistent with the ROH and inbreeding coefficient (FIS) analyses, which show higher values compared with the other two Bajo groups (Supplementary Figures S6 and S7).
IBD was used to measure haplotype sharing across the genome. All Bajo communities share longer fragments with each other than with other regional populations (Supplementary Figure S8), suggesting that the Bajo communities did intermarry until their recent land-based resettlement. The highest IBD sharing was observed between the Kendari and Kotabaru Bajo, then between the Kotabaru and Derawan Bajo, with much less between the Derawan and Kendari Bajo, again suggesting that genetic similar does not simply match current geographical location. As also shown by FST distances, high shared IBD between Bajo groups does not exclude sharing with non-Bajo neighbors. Nevertheless, IBD sharing between the Kendari Bajo and Bugis, two sea-based communities currently settled in Sulawesi, is lower than IBD sharing between the Kendari Bajo and other Bajo groups.
Like other analyses, a TreeMix analysis situates the three Bajo communities with eastern Indonesian and Philippine populations (Supplementary Figure S9). The tree supports 16 migration nodes, many showing migration into the Bajo from Papuan clusters. Interestingly, there is Papuan migration into the two Bajo groups on Borneo, as well as the Kendari Bajo, where Papuan contributions were noted by Admixture. The most parsimonious hypothesis is multilayer admixture – from Papuan or Eastern Indonesian groups into the Kendari Bajo, and from there into the other Bajo groups. However, the data cannot exclude a more complex scenario with direct contact between Bajo groups in Borneo and Papuans.
We also inferred admixture scenarios for the three Bajo populations using GLOBETROTTER. This suggests that the Kendari Bajo mixed with surrogates of Sulawesi Bugis and Papuans multiple times. The oldest admixture event occurred around 62 generations ago (1736 years ago, assuming a 28-year generation interval) with 90% and 10% contributions from Sulawesi Bugis and Papuans, respectively, and more recent admixture six generations ago (175 years ago), with admixture just from the Bugis (Figure 4, Supplementary Table S4). In contrast, the Kotabaru Bajo show one admixture event between Indian (5%), Sulawesi Bugis (70%) and Bornean Banjar (25%) sources around 33 generations ago (925 years ago), suggesting that Sulawesi had a major role in shaping the genomes of Kotabaru Bajo individuals. Local populations also contribute to the genomic make-up, highlighting the neighboring Banjar of Borneo as another contributing group. The Derawan Bajo have genomic components from Indian (5%), Filipino (70%) and Malay (25%) sources, dating to around 24 generations ago (675 years ago).
Admixture history of the three Bajo communities inferred with GLOBETROTTER. (a) Admixture of Bugis (South Sulawesi) with multiple populations, including Malay, Filipinos and Papuans, up to 1600 years ago (ya), contemporary to the admixture of pre-Bajo Kendari by Bugis and Papuans around 1750 ya (Supplementary Table S4). (b) The expansion of the Śrīvijaya empire to Southeast Borneo triggered the dispersal of Bajo language, culture and people in many directions, including Southern Sulawesi and the Kendari, who assimilated them into its society. (c) Southern Sulawesi populations subsequently migrated westward to Southeastern Borneo, forming the Kotabaru community by admixing with local Banjar populations, in addition to Indian influences through the reigning Malay empire around 925 ya. Northward migrations formed the Derawan community, which also admixed with local Malay and Filipino groups around 675 ya. The influence of Southern Sulawesi populations (dashed arrows) is observed in both the Kotabaru and Derawan Bajo. (d) Recent interactions between the three Bajo groups were maintained with different intensities (dashed lines).
These results were obtained with significant fit values by excluding other Bajo communities as potential surrogate populations for any given Bajo community. When we allowed all Bajo groups to act as potential surrogate populations, lower fit values were obtained, reflecting uncertain inference of admixture scenarios (Supplementary Table S4). Nonetheless, these runs confirm the earlier GLOBETROTTER results for the Kendari and Kotabaru Bajo, but in a new finding, the Bugis appears to be a surrogate population for the Derawan Bajo, in addition to Malay, Filipinos and Indians. Given the potential role of the Bugis on the genetic make-up of the Bajo, we therefore tested their admixture profile using GLOBETROTTER. The Bugis experienced a multiway admixture event around 57 generations ago (1600 years ago) between Papuans (14%), Filipinos (41%) and Malay (45%), at around the same time as the admixture event with the Kendari Bajo. These admixture events were confirmed using MALDER (Supplementary Table S5).
Discussion
Even among the extraordinary diversity of human lifeways, the entirely sea-based lives of the Bajo – being born, growing up, marrying and dying on the sea – is special. This way of living is unique to Southeast Asia, with the Bajo, Urak Lawoi and Moken being well-known examples.13, 45 However, very little is understood about the genetic structure of these communities. Using genome-wide SNP data, we can reconstruct the genetic background and diversity of the Bajo across three communities with different dialects spanning their geographic range, thus helping to clarify where the Bajo originated and how their society interacted with other groups. Each Bajo community constitutes a homogenous genetic group, with surprisingly little overlap. A common theme is that genetic sharing is greater with neighboring populations than other Bajo groups, although there is a clear shared component of Bajo ancestry. Nevertheless, genetic contributions from these local populations were far from trivial, matching the maritime creolization process observed in their languages.
This admixture seems to have started early. Bajo were never the major ethnic group in the regions where they first lived, but instead seem to have attracted and assimilated people from nearby communities.24 For example, in Kangean, a small archipelago in the Java Sea between Eastern Java and Southern Borneo, where the Bajo language and culture predominates today, ethnicities were historically more numerous.23, 24 Before Bajo migrants arrived, the main island was inhabited by indigenous Madura (East Javanese people), then several waves of migrants spread from Southern Sulawesi (including the Bajo, but also Bugis, Makasar and Mandar). However, the number of Bajo speakers then increased markedly, quickly reaching one-third of the total island population. Over time, non-Bajo speakers adopted Bajo languages and intermarried with the original Bajo. This also occurred elsewhere, with mixing between Bajo and neighboring ‘land owners’ being commonplace.20
Consequently, all Bajo individuals share at least some common genetic background, suggesting that gene flow between these groups occurred until recently, and indeed, may still be ongoing today. Bajo communities maintained contact through sharing of goods, trading, fishing and marriage. Until recently, Bajo trading routes spanned Singapore in the west to New Guinea in the east, and Northeastern Borneo in the north to the Lesser Sunda Islands in the south.24 Records note peaceful contact of Bugis and Bajo with Australian Aborigines along the Northern coast of Australia, where the Bajo harvested trepang (sea cucumber) in shallow near-shore waters,12, 17, 21 but no genetic contact is known. Strikingly, an established and stable Bajo sea trading route connected Southeastern Sulawesi with Southeastern Borneo (including Kotabaru Island), and from there, Kotabaru Island with Northeastern Borneo as far as Brunei, albeit with less intense activity.24 This may explain the very recent admixture seen in the Bajo genomes, best illustrated by long shared IBD regions (Figure 4d).
Despite a complex genetic history involving creolization and multiple admixture events, the genomic data are suggestive of a single population origin for the Bajo, converging on Southern Sulawesi. The oldest estimated admixture event dates to the fourth century CE (Figure 4a, Supplementary Table S4) between ancestral Bugis (90%) and a Papuan group (10%). The two Bajo communities on Borneo appear to have emerged later, around the 12th century for the Kotabaru Bajo and the 14th century for Derawan Bajo, perhaps suggesting that Bajo communities lived in Southern Sulawesi for nearly 800 years before spreading west to Borneo. In contrast, the most recent linguistic studies support an origin of the Bajo language in the Southeast Borneo region, followed by a dispersal up the east coast of Borneo during the 11th century, only later spreading to the Southern Philippines and Northeast Borneo in the 13th–14th centuries.26 The linguistic and genetic evidence are therefore in broad agreement regarding the timing of the Bajo dispersal along the east coast of Borneo, but point to quite different locations for its origin: Southern Sulawesi for the gene pool and Southeast Borneo for the languages.
This apparent contradiction may be reconciled by aspects of recent history, as the expanding influence of the Malay kingdom of Śrīvijaya from the 7th century onward (7–13th centuries)6, 46 heavily modified population structure and interactions in Southeast Borneo, triggering large population movements, such as the likely migration of the Banjar to Madagascar.47, 48 We postulate that similar causes may have also stimulated the dispersal of Bajo speakers from Southeast Borneo, again around the 11th century. The spread of the Bajo culture from Southeast Borneo possibly impacted pre-Bajo groups in Southern Sulawesi, leading to the emergence of the Kendari Bajo (Figure 4b). This Southern Sulawesi community with an incipient Bajo culture then unified the Bajo language and genome by settling other areas, creating communities such as the Kotabaru and Derawan Bajo (11th–14th centuries), likely with sex biased admixture between men from mainland Asia and women from the Bajo ancestral population (Supplementary Figure S5). This sex bias pattern is also observed in other sea nomad populations along coastal Mainland Southeast Asia, such as the Moken sea nomads, who exhibit lower female gene flow from mainland Asian populations.49 Later, admixture with local groups occurred (Figure 4c), as well as ongoing contact between Bajo communities (Figure 4d). A similar process likely impacted other regions where the Bajo culture is common now, such as the Sulu archipelago in the Southern Philippines. Furthermore, Southern Sulawesi was long a center of trading activity during and after the Śrīvijaya Empire,12 reaching its peak during the 16th century.50 The main actors with significant role as traders are the Bugis, Makasar and Bajo, all with Southern Sulawesi connections.13 Therefore, the presence of a Southern Sulawesi genetic background in all Bajo communities may also result from contact directly between these three sea trading groups.
Outside Indonesia, similar admixture behaviors, notably shared long-distance contact with local genetic contributions, has also been observed in other recent diasporas, such as the Romani and the Jewish in Europe. The Romani, who originated in Northwest India, later admixed with local European populations where they settled, yet with relatively modest genetic contributions.51 Similarly, the Jewish diaspora has been traced back to the Levant, but local genetic admixture has been identified in each respective community.52 In both examples, all communities shared a common culture and genetic heritage, but like the Bajo, experienced gene flow from populations surrounding them.
The complexity of the Bajo genomic profile provides a striking reflection of their history, mediated by both migratory and local admixture events, and emphasizing their unique lifestyle had out across multiple geographical scales. Despite speaking Sama-Bajau languages, the Bajo prove to be diverse, encompassing rich genetic inputs from many groups, each distributed differently in the major Bajo communities, but homogenous for individuals within each community. It appears that contact between Bajo groups was a major feature of this history, but countered by strong regional contacts: Papuan influence in the Kendari Bajo; Banjar in the Kotabaru Bajo; and Filipino and Malay in the Derawan Bajo – all with an outsized influence from their Southern Sulawesi origin, possibly obtained by proxy from the Bugis. This genetic structure is in part due to a process of maritime creolization, exhibiting closer genetic connections with neighboring populations than distant Bajo groups. The sea-oriented way of life of the Bajo and their prime role in the maritime trading network placed them in contact with surprisingly diverse populations, including South Asians and Papuans, whose contact left secondary traces in the genomes of the Bajo today. Studies of other Bajo communities, of which there are hundreds scattered across 1300 km of Island Southeast Asia from east to west and 2000 km from north to south, are likely to reveal more nuanced patterns of contact, as well as differential associations with means of subsistence, language, traditions and origin myths. In addition, this may well provide greater insight into the likely genetic histories of other nomadic populations that speak closely related languages, but span wide geographical areas.
References
Solheim WG II, Bulbeck D, Flavel A : Archaeology and Culture in Southeast Asia: Unraveling the Nusantao. Diliman, Quezon City: University of Philippines Press, 2006.
Bellwood P : Prehistory of the Indo-Malaysian Archipelago. Canberra: ANU E Press, 2007.
Bulbeck D : An integrated perspective on the Austronesian diaspora: the switch from cereal agriculture to maritime foraging in the colonisation of Island Southeast Asia. Aust Archaeol 2008; 67: 31–52.
Oppenheimer S, Richards M : Fast trains, slow boats, and the ancestry of the Polynesian islanders. Sci Prog 2001; 84: 157–181.
Kayser M, Lao O, Saar K et al: Genome-wide analysis indicates more Asian than Melanesian ancestry of Polynesians. Am J Hum Genet 2008; 82: 194–198.
Beaujard P : Les Mondes de l’océan Indien. Vol. 1: De lace l’État au Premier Système-Monde Afro-Eurasien (4e millénaire av. J.-C.-6e siècle apr. J.-C.). Paris, France: Armand Collin, 2012.
Beaujard P : Les Mondes de l’ocean Indien. Vol. 2: L’océan Indien, Au Cœur Des Globalisations de l’Ancien Monde (7e-15e siècles). Paris, France: Armand Collin, 2012.
Skoglund P, Posth C, Sirak K et al: Genomic insights into the peopling of the Southwest Pacific. Nature 2016; 538: 510–513.
Chaudhuri KN : Trade and Civilisation in the Indian Ocean: An Economic History from the Rise of Islam to 1750, 1st edn. Cambridge. UK: Cambridge University Press, 1985.
Lawler A : Sailing Sinbad’s seas. Science 2014; 344: 1440–1445.
Lewis MP, Simons GF, Fennig CD, (eds) : Ethnologue: Languages of the World, 19th edn. Texas: SIL International, 2016, Online version:http://www.ethnologue.com.
Pelras C : The Bugis, 1st edn. Oxford, UK: Wiley-Blackwell, 1997.
Sopher DE : The Sea Nomads: A Study of the Maritime Boat People of Southeast Asia. Singapore: National Museum, 1977.
Hogan DW, Pattemore SW : Urak Lawoi’: basic structures and a dictionary Dept. of Linguistics, Research School of Pacific Studies. Australian National University: Canberra, Australia, 1988.
Larish MD . The position of Moken and Moklen within the Austronesian language family. PhD thesis. Ann Arbor: University of Michigan, 2007.
Sather C : The Bajau Laut: Adaptation, History, and Fate in a Maritime Fishing Society of South-Eastern Sabah. Oxford, UK: Oxford University Press, 1997.
Stacey N : Boats to Burn: Bajo Fishing Activity in the Australian Fishing Zone. Canberra, Australia: ANU E Press, 2007.
Pallesen AK: : Culture contact and language convergence. Manila, Philippines: Linguistic Society of the Philippines, 1985.
Verheijen JAJ: : The Sama/Bajau language in the Lesser Sunda islands. Canberra, Australia: Department of Linguistics, Research School of Pacific Studies, Australian National University, 1986.
Nuraini C : The intangible legacy of the Indonesian Bajo. Wacana 2016; 17: 1–18.
Macknight CC : The Voyage to Marege’: Macassan Trepangers in Northern Australia. Melbourne, Australia: Melbourne University Press, 1976.
Kusuma P, Cox MP, Pierron D et al: Mitochondrial DNA and the Y chromosome suggest the settlement of Madagascar by Indonesian sea nomad populations. BMC Genomics 2015; 16: 191.
Nuraini C : Langue et Production de Récits d’une Communauté Bajo des îles Kangean (Indonésie). PhD thesis. La Rochelle: Université de La Rochelle, 2008.
Nagatsu K : Persisting Maritime Frontiers and Multi-Layered Networks in Wallacea. Kyoto: Presented in Asian Core Program Seminar, 2013.
Noorduyn J : A Critical Survey of Studies on the Languages of Sulawesi. Leiden, The Netherlands: KITLV Press, 1991.
Blust R : The linguistic position of Sama-Bajaw. Stud Philipp Lang Cult 2007; 15: 73–114.
Liebner HH: Four oral versions of a story about the origin of the Bajo people of southern Selayar. In: Robinson K, Paeni M (eds): Living through Histories: Culture, History, and Social Life in South Sulawesi. Canberra, Australia: Department of Anthropology, Research School of Pacific and Asian Studies, Australian National University, 1998.
Jubilado RC, Hussin H, Manueli MK : The Sama-Bajaus of Sulu-Sulawesi seas: perspectives from linguistics and culture. J Southeast Asian Stud 2011; 15: 83–95.
Pierron D, Razafindrazaka H, Pagani L et al: Genome-wide evidence of Austronesian–Bantu admixture and cultural reversion in a hunter-gatherer group of Madagascar. Proc Natl Acad Sci USA 2014; 111: 936–941.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ : Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 2015; 4: 7.
Delaneau O, Marchini J, Zagury J-F : A linear complexity phasing method for thousands of genomes. Nat Methods 2012; 9: 179–181.
Delaneau O, Marchini J : Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat Commun 2014; 5: 3934.
Lawson DJ, Hellenthal G, Myers S, Falush D : Inference of population structure using dense haplotype data. PLOS Genet 2012; 8: e1002453.
Patterson N, Price AL, Reich D : Population structure and eigenanalysis. PLoS Genet 2006; 2: e190.
Pembleton LW, Cogan NOI, Forster JW : StAMPP: an R package for calculation of genetic differentiation and structure of mixed‐ploidy level populations. Mol Ecol Resour 2013; 13: 946–952.
Alexander DH, Novembre J, Lange K : Fast model-based estimation of ancestry in unrelated individuals. Genome Res 2009; 19: 1655–1664.
Jakobsson M, Rosenberg NA : CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007; 23: 1801–1806.
Pickrell JK, Pritchard JK : Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 2012; 8: e1002967.
Patterson NJ, Moorjani P, Luo Y et al: Ancient admixture in human history. Genetics 2012; 192: 1065–1093.
Browning SR, Browning BL : Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 2007; 81: 1084–1097.
Hellenthal G, Busby GBJ, Band G et al: A genetic atlas of human admixture history. Science 2014; 343: 747–751.
Fenner JN : Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am J Phys Anthropol 2005; 128: 415–423.
Loh P-R, Lipson M, Patterson N et al: Inferring admixture histories of human populations using linkage disequilibrium. Genetics 2013; 193: 1233–1254.
Mörseburg A, Pagani L, Ricaut F-X et al: Multi-layered population structure in Island Southeast Asians. Eur J Hum Genet 2016; 24: 1605–1611.
Ivanoff J: : Rings of coral: Moken folktales, 1st edn. Bangkok, Thailand: White Lotus Co. Ltd, 2002.
Ras JJ : Hikajat Banjar: A study in Malay Historiography. The Hague: Martinus Nijhoff, 1968.
Brucato N, Kusuma P, Cox MP et al: Malagasy genetic ancestry comes from an historical Malay trading post in Southeast Borneo. Mol Biol Evol 2016; 33: 2396–2400.
Kusuma P, Brucato N, Cox MP et al: Contrasting linguistic and genetic origins of the Asian source populations of Malagasy. Sci Rep 2016; 6: 26066.
Dancause KN, Chan CW, Arunotai NH, Lum JK : Origins of the Moken Sea Gypsies inferred from mitochondrial hypervariable region and whole genome sequences. J Hum Genet 2009; 54: 86–93.
Villiers J : Makassar: the rise and fall of an east Indonesian maritime trading state, 1512–1669. In: Kathririthamby-Wells J, Villiers J (eds): The Southeast Asian Port and Polity: Rise and Demise. Singapore: Singapore University Press, 1990.
Mendizabal I, Lao O, Marigorta UM et al: Reconstructing the population history of European Romani from genome-wide data. Curr Biol 2012; 22: 2342–2349.
Behar DM, Yunusbayev B, Metspalu M et al: The genome-wide structure of the Jewish people. Nature 2010; 466: 238–242.
Acknowledgements
The authors thank Gludhug A Purnomo, Isabella Apriyana and Chelzie C Darusalam from Eijkman Institute for Molecular Biology, Jakarta, Indonesia, for biological sampling, and Laure Tonasso and Stéphanie Schiavinato (University of Toulouse) for laboratory assistance. We acknowledge support from the GenoToul bioinformatics facility of the Genopole Toulouse Midi Pyrénées, France. This research was supported by the French ANR via grant ANR-14-CE31-0013-01 (OCEOADAPTO) to F-XR, the French Ministry of Foreign and European Affairs (French Archaeological Mission in Borneo; MAFBO) to F-XR, a Rutherford Fellowship from the Royal Society of New Zealand (RDF-10-MAU-001) to MPC, the French Embassy in Indonesia through its Cultural and Cooperation Services (Institut Français en Indonésie) to F-XR and the Ministry of Education and Culture of Indonesia to PK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kusuma, P., Brucato, N., Cox, M. et al. The last sea nomads of the Indonesian archipelago: genomic origins and dispersal. Eur J Hum Genet 25, 1004–1010 (2017). https://doi.org/10.1038/ejhg.2017.88
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.88