Abstract
Genetic imputation is a cost-efficient way to improve the power and resolution of genome-wide association (GWA) studies. Current publicly accessible imputation reference panels accurately predict genotypes for common variants with minor allele frequency (MAF)≥5% and low-frequency variants (0.5≤MAF<5%) across diverse populations, but the imputation of rare variation (MAF<0.5%) is still rather limited. In the current study, we evaluate imputation accuracy achieved with reference panels from diverse populations with a population-specific high-coverage (30 ×) whole-genome sequencing (WGS) based reference panel, comprising of 2244 Estonian individuals (0.25% of adult Estonians). Although the Estonian-specific panel contains fewer haplotypes and variants, the imputation confidence and accuracy of imputed low-frequency and rare variants was significantly higher. The results indicate the utility of population-specific reference panels for human genetic studies.
Similar content being viewed by others
Introduction
Genotype imputation is a method for statistically inferring untyped genotypes in a sample of partially genotyped individuals, based on a reference panel of individuals who have been more densely genotyped or sequenced. Imputation methods attempt to identify haplotype sharing between individuals in the sample and in an imputation reference panel (IRP), and use this information to infer the alleles at untyped loci in the sample.1 Imputation allows geneticists to study variants that have not been directly genotyped in a sample and thereby to increase power and resolution of genome-wide association studies (GWAS). Imputation is particularly useful for combining association results across studies that used different genotyping arrays2 and facilitate fine-mapping to localise association signals by considering all genetic variants in a region.
Publicly available IRPs from the International HapMap Project3, 4 and 1000 Genomes Project (1000G)5 have been instrumental to the discovery of thousands of loci affecting diseases and traits in individual GWAS and collaborative meta-analyses. The first wave of studies mostly used the HapMap II IRP, which used microarray-based genotypes from 270 individuals at 3.1 million (M) variants.6, 7, 8, 9, 10 Later studies used IRPs based on the 1000G project, which performed whole-genome sequencing (WGS) on a diverse set of populations, with 2504 individuals and up to 84.4 M variants.11, 12, 13, 14, 15, 16 Although the latter IRP allows robust imputation of common variants (minor allele frequency (MAF)≥5%) and low-frequency variants (0.5≤MAF<5%)5 it has only limited imputation accuracy for rare (MAF<0.5%) variants.17, 18, 19 A recent IRP from Haplotype Reference Consortium (HRC)20 contains even more individuals (N=32 488, mostly with European ancestry) and should therefore enable better imputation of both low-frequency and rare variants in European samples.
Recently, several studies have demonstrated that the use of population-specific IRPs can further improve the imputation accuracy of common and low-frequency variants, and improve the imputation of rarer variants in the relevant population.21, 22, 23, 24 By using an IRP composed of related Dutch individuals, Deelan et al.23 showed that it is possible to substantially improve the completeness and accuracy of imputation of rare variants into a set of Dutch individuals. Gudbjartsson et al used long-range haplotype phasing in combination with imputation to increase imputation accuracy for rare variants down to MAF of 0.1% in the Icelandic population.22 Sidore et al. reported several variants associated with circulating lipid levels in Sardinians that were detected due to accurate imputation achieved by using a Sardinian WGS-based IRP; these authors showed that the variants would not have been identified if the analyses had been based on the 1000G IRP.24 Similar results were obtained in the UK10K project, where the British population-specific IRP combined with 1000G Project reference panel facilitated the discovery of several novel genetic variants associated with medically relevant phenotypes.19, 25, 26
Studies have shown that the genetic structure of European countries correlates closely with their geographic origin.27, 28 The Estonian population, being located in Northeast Europe, is genetically most similar to its neighbouring countries, including Finland, the North-western part of Russia, and other Baltic countries.28, 29, 30 Notwithstanding this overall genetic similarity, the Estonian population still has a substantial proportion of haplotypes that are not expected to be covered by the more diverse IRPs. Moreover, the population-specific differences are expected to increase as allele frequencies decrease.
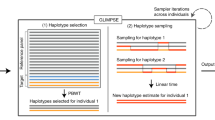
In the current study, we first evaluated two most commonly used phasing algorithms to create population-specific IRP based on high-coverage (30 ×) WGS data from 2244 Estonian individuals. To impute low-frequency and rare variants more accurately in a specific population, one can take two approaches: (i) increase the size of IRPs from diverse populations to capture more reference haplotypes or (ii) employ population-specific IRPs. We assessed the utility of these approaches for improving imputation in Estonian samples by comparing the performance of (i) an Estonian-specific IRP, (ii) the commonly used 1000G IRP, (iii) the much larger HRC IRP and (iv) combinations of these panels.
Materials and methods
Cohort description
2304 geographically distributed individuals (selected randomly by county of birth) from the Estonian Biobank of the Estonian Genome Center, University of Tartu (EGCUT) were selected for WGS. EGCUT is a population-based biobank, containing almost 52 000 samples of the adult population (aged ≥18 years), which closely reflects the age, sex and geographical distribution of the Estonian population. A total of 6394 individuals (selected randomly and not overlapping with WGS data set) from the Estonian Biobank were selected for genotyping using Illumina HumanCoreExome (Illumina, San Diego, CA, USA) array, whereas the subset of 505 of these individuals were also subject to whole-exome sequencing (WES).
WGS and WES sequencing and variant calling
WGS samples followed a PCR-free sample preparation. Libraries sequenced on the Illumina HiSeq X Ten (Illumina, San Diego, CA, USA) with the use of 150 bp paired-end reads to 30 × mean coverage with a median insert size of 400 bp±25%. WES samples DNA was enriched for target sequences (Agilent Technologies, Santa Clara, CA, USA; Human All Exon V5+UTRs) according to manufacturer’s recommendations.
Sequenced reads were aligned to the GRCh37/hg19 human reference genome using BWA-MEM31 v0.7.7. SAMtools32 v1.2 was applied to compress SAM to BAM (samtools view), sort (samtools sort) and index BAM (samtools index) files. PCR duplicates were then marked using Picard (http://broadinstitute.github.io/picard) v1.136 MarkDuplicates.jar. For further BAM improvements, including realignment around known indels and base quality score recalibration, we applied Genome Analysis Toolkit (GATK)33, 34 v3.4 (v3.4-46). Single-sample genotypes were called by GATK HaplotypeCaller algorithm (-ERC GVCF). All gVCF-files were combined (-T CombineGVCFs) and jointly called (-T GenotypeGVCFs).
Quality control
Out of the total 2304 WGS samples submitted for sequencing, 4 samples did not have enough input DNA (<1.2 μg), 7 samples failed in library preparation three times and 9 samples had a contamination rate >10%. Thus, variants of 2284 WGS samples were jointly called. The GATK Variant Quality Score Recalibration was used to filter variants with a truth sensitivity of 99.8%. Also, variants with GATK inbreeding coefficient less than −0.3 were filtered to remove sites with excess heterozygous individuals. Only PASS sites were considered in the further analysis.
The PLINK/SEQ (https://atgu.mgh.harvard.edu/plinkseq) v0.10 i-stats module was used to calculate number of variants (NVAR), number of non-reference (NALT) variants, number of heterozygous (NHET) variants, NHET/NALT ratio, transition/transversion (TITV) ratio per sample and outlier (below or above 3 SD from the population mean) samples were removed. In addition, genotype and phenotype sex concordance was checked for each sample and outliers were removed. The final WGS sample set contained 2244 individuals. The final WES sample set, which passed all quality control filters and was genotyped with Illumina HumanCoreExome array, contained 505 individuals.
Multi-allelic SNVs were removed and we further excluded variants with call rate <0.95, minor allele count ≤2, Hardy–Weinberg equilibrium test P-value<1 × 10−6 and variants in low-complexity regions.35
Genotype array data was filtered sample-wise by excluding on the basis of call rate (<98%), extreme heterozygosity (>mean±3 SD), genotype and phenotype sex discordance, cryptic relatedness (IBD>20%) and outliers from the European descent from the MDS plot in comparison with HapMap reference samples. SNP quality filtering included call rate (<99%), MAF (<1%) and extreme deviation from Hardy–Weinberg equilibrium (P-value<1 × 10−4). Non-autosomal SNPs were excluded from the analysis.
Haplotype phasing
The EGCUT WGS data was phased with SHAPEIT2(ref. 36) (r837), using four computer cores. Pre-phasing of genotype array data was made in similar manner using SHAPEIT2 using four cores. As a separate test for pre-phasing accuracy, we used chromosome 20 sequence of 2244 full genomes, which were filtered beforehand to exclude any non-founder family members and individuals with a genome-wide PI_HAT value above 0.5 (2195 individuals remained) when compared to other individuals in the data set. To assess the efficiency of various approaches to phasing of WGS data, we applied two different tools: SHAPEIT2(ref. 36) and Eagle2.37, 38 Both programs were engaged with the default parameters with varying number of cores (1, 2, 4, 8, 16, 24 and 32). To verify the phasing accuracy for other data sets, the 1000G data was phased using a similar pipeline (1, 8 and 32 cores).
In addition to the regular phasing functionality, the read-aware phasing capability of SHAPEIT2(ref. 39) was also assessed. The first step entailed creating a phase informative read file on the basis of BAM files, using the module ExtractPIRs v1 (r68) with default parameters provided by the authors. After the generation of phase informative reads, the obtained file could be used in a similar fashion to a map file as a reference point for SHAPEIT2 to phase the data sets. Phasing was performed in three parallel runs after which the average run time and accuracy were compared as indicators of phasing quality.
Phasing accuracy was defined as the number of switch errors present in the phased data set. For this, the phased founder genotypes were compared with the non-phased genotypes of their offspring to determine the heredity pattern of heterozygous positions, any shifts in heredity from one parental haplotype to another were counted as switches. Two families with one offspring and two families with two offspring were used to estimate switch error rate in EGCUT sample set, four families with one offspring were used for 1000G sample set. The ratio of switch errors was calculated by dividing the number of haplotype switches to the number of the heterozygous positions where the occurrence of the switch can be reliably determined, after which the results were averaged across the trios.
Genotype imputation
Imputation using EGCUT and 1000G reference panels separately and in combination were performed in High Performance Computing Center, University of Tartu using IMPUTE2 with default parameters. As IMPUTE2 allows to use two-phased reference panels in combination (the ‘imputation with two phased reference panels’ option), we used the EGCUT and 1000G reference panels also together (EGCUT+1000G and 1000G+EGCUT). In case of such panel combining, IMPUTE2 imputes only genotypes for variants that are present in the first (main) panel but in the process, uses additional haplotype information from the second panel to improve the imputation accuracy through larger set of reference haplotypes.40
Imputation with the HRC panel was carried out using IMPUTE2 with default parameters except that the k_hap parameter that was set to 1000.
For all imputation panels, monomorphic SNVs were excluded. No further filtering was performed based on IMPUTE2 info score, but most of the analyses rest on well-imputed (INFO>0.4) and confidently imputed (INFO>0.8) SNVs.
Post-imputation filtering and concordance analyses
The GATK GenotypeConcordance tool was used to calculate imputation accuracy (concordance, non-reference sensitivity and non-reference discordancy) for different imputation panels with WES data for overlapping individuals (N=505) used as the gold standard. Low-complexity regions were filtered out of WES data prior to analysis. PLINK v1.9 was used to convert IMPUTE2 files (imputation output) to VCF format using hard-call threshold 0.9. BCFtools filter option was used to keep genotypes imputed with INFO-value>0.4 and overlapping with WES-target regions. Comparison was performed in three MAF bins (MAF≥5%, 0.5≤MAF<5% and MAF<0.5%) based on WES minor allele frequencies and only well-imputed (INFO>0.4) SNVs were considered. Reference sequence in the concordance analyses was the same for both WGS and WES analysis pipelines.
To assess more stratified imputation accuracy, an additional concordance analysis was run for IRPs for well-imputed (INFO>0.4) variants in WES-based MAF bins of (0, 0.2), (0.2, 0.4), (0.4, 0.6), (0.6, 0.8), (0.8, 1), (1, 2), (2, 3), (3, 4), (4, 5), (5, 10), (10, 20), (20, 30), (30, 40) and (40, 50%).
Functional annotation of variants
We used Variant Effect Predictor41 version 84 to annotate the confidently imputed variants in the 20 345 protein-coding genes in the Ensembl database (with Gencode v19 on GRCh37).
Results
Using high-coverage WGS data of 2244 Estonian individuals from the Estonian Biobank,42 we created a population-specific IRP. After variant calling and rigorous quality control steps (Materials and Methods), we phased the Estonian WGS data and used the resulting Estonian IRP (referred to here as the EGCUT IRP, for the Estonian Genome Center at University of Tartu), together with the 1000G and HRC IRP, to impute genotypes into 6394 Estonians who had been genotyped on microarrays.
Phasing speed and accuracy of multi-threaded haplotype phasing
Haplotype phasing can be a time-consuming process, especially for large WGS-based data sets. We therefore began by evaluating haplotype-phasing algorithms. We compared three different parallel, multi-threaded computational programmes—SHAPEIT2,36 SHAPEIT2-RA (for read-aware)39 and Eagle2(refs. 37, 38) —utilised with different number of processor cores (1, 2, 4, 8, 16, 24 and 32) (Supplementary Figure 1A). These programmes were applied to data from chromosome 20 in the EGCUT samples. Accuracy was assessed by counting the number of haplotype switch errors (Materials and Methods) in four families, for which haplotype phase could be independently determined based on segregation of genetic markers.
While the speed of both SHAPEIT2 and SHAPEIT2-RA increased in proportion to the number of cores used, the speed of Eagle2 increased proportionally up to eight cores but not beyond. Up to this point, Eagle2 was considerably faster than SHAPEIT2, by a factor of roughly 6-fold. The two versions of SHAPEIT2 showed similar accuracy, which was slightly lower for Eagle2 (average haplotype switch error rate of 0.7% with SHAPEIT2 vs 0.81% with Eagle2; Table 1). In all cases, the accuracy did not vary significantly with the number of cores used. To validate that these results were not population-specific, we performed similar analyses with four 1000G family trios (with 1, 8 and 32 cores) and observed similar switch error rates in the corresponding phasing results (Supplementary Table 1). While in our hands, SHAPEIT2 displayed slightly higher accuracy, it did so at the cost of increased computing time, making Eagle2 a viable option for the researchers who require time-efficient phasing of large data sets. However, because the 1000G and HRC IRPs were phased with SHAPEIT, we used this program computer program to phase the EGCUT data (Materials and Methods).
Genotype imputation
To impute genotypes into 6394 Estonian individuals who had been genotyped on Illumina HumanCoreExome microarrays, we used the IMPUTE2 software43, 44 together with three separate IRPs and two combinations of IRPs (Table 2). The first IRP consisted of the 2244 whole-genome sequenced EGCUT individuals; these individuals were selected to be geographically distributed across Estonia and did not overlap with the set of genotyped individuals. The other two were 1000G IRP and the HRC IRP from large diverse populations. The IMPUTE2 software also allows to improve imputation accuracy by using two reference panels simultaneously by pooling haplotype information across both IRPs.40 We used both combinations of the EGCUT and 1000G panels with that option: EGCUT+1000G and 1000G+EGCUT. In such combinations, IMPUTE2 imputes only genotypes for variants that are present in the first (main) IRP while also considering haplotype information from the second IRP to improve the imputation accuracy through larger set of reference haplotypes. Thus, EGCUT+1000G should be viewed as an improvement of the EGCUT reference panel (genotypes observed in the EGCUT panel imputed while considering haplotypes inferred from the EGCUT and 1000G panels) and 1000G+EGCUT should be considered as an improvement of the 1000G panel (genotypes observed in the 1000G panel imputed while considering haplotypes inferred from both panels).
Number of imputed variants
For each IRP, we studied the number of imputed single-nucleotide variants (SNVs) as a function of the imputation confidence estimate—INFO-value—assigned by the IMPUTE2 programme. The INFO-value reflects the information in imputed genotypes relative to the information if only the allele frequency were known.43, 44 We counted the total number of imputed SNVs, the number of ‘well-imputed’ SNVs (INFO>0.4)18 and the number of ‘confidently imputed’ SNVs (INFO>0.8). We also counted the number of imputed SNVs found only with each IRP (Figure 1a).
Number of variants imputed from different IRPs. (a) Number of all shared and panel-specific variants in three distinct reference panels imputed with INFO-value >0.4 (in bold) and >0.8 (given in brackets); (b) Total number of imputed SNVs (bars); the number of SNVs imputed with imputation quality score (INFO-value)>0.4 (coloured) and INFO>0.8 (shaded areas).
Although the number of total variants and well-imputed variants obtained with the larger diverse panels (1000G and HCR) exceeded the corresponding numbers for the population-specific panel, the situation was reversed for confidently imputed SNVs with 12.29 M (75% of total number of imputed SNVs), 10.05 M (48%) and 9.44 M (27%) of SNVs being confidently imputed with the EGCUT, HRC and 1000G panel, respectively (Figure 1b). The combined EGCUT+1000G panel showed almost identical results to EGCUT panel alone, whereas the 1000G+EGCUT panel showed considerable increase in the number of confidently imputed SNVs (by considering additional haplotype information from the population-specific IRP) as compared to the 1000G panel alone. These results indicate that using a population-specific IRP increases the number of confidently imputed variants, due to more similar allele frequencies and greater relatedness between the samples and the IRP. More diverse IRPs have a tendency to employ incorrect allele frequency distribution and also to contain divergent haplotypes, which are not present in the samples (eg., African haplotypes carrying variants that are not polymorphic in non-African populations).
We next stratified these analyses according to the MAFs of the imputed SNVs, dividing them into three groups: common (MAF≥5%), low-frequency (0.5≤MAF<5%) and rare (MAF<0.5%) SNVs. For common variants, the number of imputed SNVs was very similar across the IRPs (Figure 2). For low-frequency variants, the number of well-imputed SNVs was also very similar, whereas the number of confidently imputed SNVs was larger for the population-specific IRP. For rare variants, the results were even more pronounced, 3.48 M (54% of well-imputed rare variants), 2.54 M (33%) and 1.86 M (15%) SNVs were imputed confidently from the EGCUT, HRC and 1000G panels, respectively (Figure 2b,Supplementary Table 2). Notably, the EGCUT panel outperformed the other panels on rare variants despite the fact that the HRC panel contains the largest number of haplotypes (64 976) and the 1000G panel contains the largest number of variants (81 M SNVs on autosomes).
These results show that imputation confidence (measured as INFO-value) decreases substantially as the allele frequency of the imputed variants declines (Supplementary Figure 2). Despite the fact that the larger and more diverse IRPs contained more variants, they contained fewer matching haplotypes than the population-specific panel. As a result, the HRC and 1000G panels yielded genotypes imputed with lower confidence (INFO-value), especially for rare SNVs (Supplementary Figure 3). For the combinations of reference panels, the EGCUT+1000G showed almost identical results in every aspect compared to EGCUT panel alone, while the 1000G+EGCUT panel showed a slight gain for common and low-frequency variants and a substantial gain for rare variants when compared to 1000G panel alone (Figure 2).
Imputation of loss-of-function and missense variants
Loss-of-function (LoF) variants that disrupt protein-coding genes and missense variants that cause amino acid changes are of particular interest because they are potentially clinically relevant. Considering only confidently imputed SNVs (INFO>0.8), we observed that all three reference panels enabled imputation of a similar number of common LoF and missense variants (Figure 3). However, the number of low-frequency LoF variants was higher with the population-specific IRP and the number of rare LoFs was almost twice as high (417, 439 and 730 LoF SNVs with the 1000G, HRC and EGCUT, respectively; Supplementary Table 3) with the population-specific IRP.
Imputation sensitivity and accuracy
Although imputation confidence estimates (such as INFO-values or squared correlations r2)45, 46 are useful for characterising the overall success of the imputation process, high INFO or r2 values do not guarantee that the corresponding genotypes are inferred correctly. Therefore, it is important to directly assess the accuracy of the imputed genotypes. We compared the ‘best guess’ genotypes imputed from the different reference panels to WES data available for a subset of imputed EGCUT individuals (N=505; Supplementary Figure 1B). Treating these WES-based genotype calls as ‘gold standard’, we calculated two metrics for each imputed data set: (i) sensitivity, defined as the proportion of WES-based non-reference (NR) variant calls that were also obtained through imputation process; and (ii) discordancy rate, defined as the proportion of imputed SNVs that had incorrect genotype call.
For well-imputed common SNVs, all of the IRPs gave similarly high sensitivity (88.5–92.4%) (Figure 4a). For low-frequency SNVs, the three panels that included data from the population-specific panel (EGCUT, EGCUT+1000G, and 1000G+EGCUT) yielded in higher sensitivity (~87%) than the more diverse panels (78% and 76% for HRC and 1000G, respectively) (Table 3). For rare SNVs, the proportional difference was even greater (40%, 42% and 49% for 1000G, HRC and EGCUT IRPs, respectively).
Imputation accuracy for common (MAF≥5%), low-frequency (0.5≤MAF<5%) and rare (MAF<0.5%) well-imputed variants (INFO>0.4) imputed from different IRPs. (a) Non-reference (NR) sensitivity—proportion of whole-exome sequencing (WES) based NR variant calls that were also retrieved through imputation process. (b) NR discordancy rate—proportion of NR variants that were retrieved through imputation process but had incorrect genotype calls as compared to the WES genotypes.
Similarly, the population-specific IRP performed better with respect to discordancy rate (Figure 4b). Whereas all three panels had a low discordancy rate for common variants (1.9–3.4%), the EGCUT panel outperformed other panels for low-frequency and rare SNVs (Table 3). Notably, one-quarter (24.7%) of rare SNVs imputed from the 1000G IRP had incorrect genotype calls, whereas the proportion was substantially lower with the EGCUT IRP alone (14.1%) or if it was used in combination with the 1000G panel (13.6% and 14.3% for the EGCUT+1000G and 1000G+EGCUT panels, respectively). Similar results were seen for confidently imputed variants, for which both sensitivity and discordancy rate were better in case of the population-specific reference panel (Supplementary Figure 4, Supplementary Table 4). The better performance is due to a close match between the EGCUT IRP and Estonian samples—owing to the fact that rare variants tend to be more recent and thus more population specific.
We repeated these analyses of imputation accuracy by using finer bins of MAF (Supplementary Figures 5–9). We found that although the overall success of genotype imputation of well-imputed variants decreased steadily with MAF in case of all compared IRPs, imputation accuracy was, especially for rare variants, significantly better in case of the population-specific IRP (Supplementary Figure 7) or if it was used together with the 1000G reference panel (Supplementary Figures 8 and 9).
Discussion
Genotype imputation is a cost-efficient way to improve the power and resolution of GWA studies. Although large IRPs from diverse populations work reasonably well for imputation of common and low-frequency variants, currently available reference panels allow only limited imputation of rare variants.
WGS has become increasingly widespread in recent years and is increasingly used in creating IRPs. The first step in the process of creating an IRP is the correct assignment of polymorphic positions regarding the individual haplotypes. Although the task can be computationally demanding for large data sets, the advent of various phasing algorithms has simplified this task considerably. We compared the performance of the SHAPEIT2 and Eagle2 software, both of which can increase the phasing speed by dividing the phased reference data set into multiple subsets, which are then processed in parallel. Similarly to previously published comparison,38 we found that Eagle2 was considerably faster than SHAPEIT2. However, the decrease in phasing time resulted in a small increase in haplotype switch errors, making SHAPEIT2 a better choice for those aiming at the highest accuracy. Interestingly, we did not observe a difference in phasing accuracy between SHAPEIT2 and SHAPEIT2’s read-aware mode. It is possible that this was due relatively homogeneous nature of our Estonian samples and that the SHAPEIT2 read-aware mode may exhibit advantages for more heterogeneous data sets.
Consistent with previous studies, our results show that population-specific IRPs can improve the genotype imputation, especially for low-frequency and rare variants.21, 22, 23, 24 By being genuinely reflective of the study data set, population-specific IRPs can therefore facilitate discovery of true associations in GWAS and subsequent fine-mapping of causal variants, as demonstrated by others24, 47, 48 and also with the Estonian population-specific reference panel.49
Although the large IRPs from more diverse populations led to the imputation of a larger number of rare SNVs, a large proportion of these genotypes were imputed with low imputation confidence (IMPUTE2 INFO-value). Focusing only on confidently imputed SNVs, the population-specific IRP outperformed the 1000G and HRC IRPs. Although the overall imputation success and accuracy depend on several different factors (including the size of the IRP and the genetic structure of the reference panel and the genotyped sample), these observations are expected to apply to other populations with similar genetic background.
Beyond imputation quality, we also considered sensitivity and discrepancy rate of the imputed genotypes. We found that the population-specific IRP outperformed the large IRPs from diverse populations—a finding that is also in line with other recent imputation accuracy comparisons.50 Using a large IRP that is not well matched in terms of ancestry can thus not only limit the discovery of associations in GWAS as observed previously24 but also introduce variants that are not actually polymorphic in the imputed sample.50
Because short insertion-deletion (indel) variants were not part of the HRC IRP and because calling indel variants is still more error-prone than SNV calling, we did not include indels in our IRP and our comparisons. Once technical limitations related to indel calling and phasing are resolved, indels should be included in all IRPs.
In conclusion, we observe that, although currently publicly accessible large diverse IRPs like 1000G and HRC enable imputation of many low-frequency and rare variants in the Estonian population, most of these variants are imputed with relatively low confidence and furthermore, there is a significant proportion of population-specific variation that cannot be imputed from these panels. Moreover, imputation of low-frequency and rare variants is considerably more accurate with a population-specific reference panel or if one is used in combination with a publicly available reference such as the 1000G panel. Our results also suggest that, given that the population-specific reference panel size (number of haplotypes) is comparable to the 1000G panel size, the previous observations that reference sample size is more important than precise population matching does not apply equally well to all populations and population-specific panels can outperform even an order of magnitude larger but more diverse reference panels.
References
Marchini J, Howie B : Genotype imputation for genome-wide association studies. Nat Rev Genet 2010; 11: 499–511.
Li Y, Willer C, Sanna S, Abecasis G : Genotype imputation. Annu Rev Genomics Hum Genet 2009; 10: 387–406.
HapMap C, Altshuler DM, Gibbs RA et alInternational HapMap 3 Consortium: Integrating common and rare genetic variation in diverse human populations. Nature 2010; 467: 52–58.
International HapMap Consortium International HapMap Consortium, Frazer KA International HapMap Consortium, Ballinger DG et al: A second generation human haplotype map of over 3.1 million SNPs. Nature 2007; 449: 851–861.
1000 Genomes Project Consortium 1000 Genomes Project Consortium, Auton A 1000 Genomes Project Consortium, Brooks LD et al: A global reference for human genetic variation. Nature 2015; 526: 68–74.
Speliotes EK, Willer CJ, Berndt SI et al: Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet 2010; 42: 937–948.
Lango Allen H, Estrada K, Lettre G et al: Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010; 467: 832–838.
Morris AP, Voight BF, Teslovich TM et al: Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet 2012; 44: 981–990.
Stolk L, Perry JR, Chasman DI et al: Meta-analyses identify 13 loci associated with age at menopause and highlight DNA repair and immune pathways. Nat Genet 2012; 44: 260–268.
International Schizophrenia Consortium International Schizophrenia Consortium, Purcell SM International Schizophrenia Consortium, Wray NR et al: Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460: 748–752.
Schizophrenia Working Group of the Psychiatric Genomics Consortium: Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
van Leeuwen EM, Sabo A, Bis JC et al: Meta-analysis of 49 549 individuals imputed with the 1000 Genomes Project reveals an exonic damaging variant in ANGPTL4 determining fasting TG levels. J Med Genet 2016; 53: 441–449.
Soler Artigas M, Wain LV, Miller S et al: Sixteen new lung function signals identified through 1000 Genomes Project reference panel imputation. Nat Commun 2015; 6: 8658.
Genetics of Personality Consortium Genetics of Personality Consortium, de Moor MH Genetics of Personality Consortium, van den Berg SM et al: Meta-analysis of genome-wide association studies for neuroticism, and the polygenic association with major depressive disorder. JAMA Psychiatry 2015; 72: 642–650.
Gormley P, Anttila V, Winsvold BS et al: Meta-analysis of 375,000 individuals identifies 38 susceptibility loci for migraine. Nat Genet 2016; 48: 856–866.
Pickrell JK, Berisa T, Liu JZ, Segurel L, Tung JY, Hinds DA : Detection and interpretation of shared genetic influences on 42 human traits. Nat Genet 2016; 48: 709–717.
Pasaniuc B, Rohland N, McLaren PJ et al: Extremely low-coverage sequencing and imputation increases power for genome-wide association studies. Nat Genet 2012; 44: 631–635.
Zheng HF, Ladouceur M, Greenwood CM, Richards JB : Effect of genome-wide genotyping and reference panels on rare variants imputation. J Genet Genomics 2012; 39: 545–550.
UK10K Consortium UK10K Consortium, Walter K UK10K Consortium, Min JL et al: The UK10K project identifies rare variants in health and disease. Nature 2015; 526: 82–90.
McCarthy S, Das S, Kretzschmar W et al: A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet 2016; 48: 1279–1283.
Pistis G, Porcu E, Vrieze SI et al: Rare variant genotype imputation with thousands of study-specific whole-genome sequences: implications for cost-effective study designs. Eur J Hum Genet 2015; 23: 975–983.
Gudbjartsson DF, Helgason H, Gudjonsson SA et al: Large-scale whole-genome sequencing of the Icelandic population. Nat Genet 2015; 47: 435–444.
Deelen P, Menelaou A, van Leeuwen EM et al: Improved imputation quality of low-frequency and rare variants in European samples using the ‘Genome of The Netherlands’. Eur J Hum Genet 2014; 22: 1321–1326.
Sidore C, Busonero F, Maschio A et al: Genome sequencing elucidates Sardinian genetic architecture and augments association analyses for lipid and blood inflammatory markers. Nat Genet 2015; 47: 1272–1281.
Huang J, Howie B, McCarthy S et al: Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat Commun 2015; 6: 8111.
Timpson NJ, Walter K, Min JL et al: A rare variant in APOC3 is associated with plasma triglyceride and VLDL levels in Europeans. Nat Commun 2014; 5: 4871.
Novembre J, Johnson T, Bryc K et al: Genes mirror geography within Europe. Nature 2008; 456: 98–101.
Nelis M, Esko T, Magi R et al: Genetic structure of Europeans: a view from the North-East. PLoS ONE 2009; 4: e5472.
Kushniarevich A, Utevska O, Chuhryaeva M et al: Genetic heritage of the balto-slavic speaking populations: a synthesis of autosomal, mitochondrial and Y-chromosomal data. PLoS ONE 2015; 10: e0135820.
Khrunin AV, Khokhrin DV, Filippova IN et al: A genome-wide analysis of populations from European Russia reveals a new pole of genetic diversity in northern Europe. PLoS ONE 2013; 8: e58552.
Li H, Durbin R : Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Li H, Handsaker B, Wysoker A et al: The sequence alignment/map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
McKenna A, Hanna M, Banks E et al: The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
Van der Auwera GA, Carneiro MO, Hartl C et al: From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 2013; 43: 11.10.1-33.
Li H : Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 2014; 30: 2843–2851.
Delaneau O, Zagury JF, Marchini J : Improved whole-chromosome phasing for disease and population genetic studies. Nat Methods 2013; 10: 5–6.
Loh PR, Palamara PF, Price AL : Fast and accurate long-range phasing in a UK Biobank cohort. Nat Genet 2016; 48: 811–816.
Loh P-R, Danecek P, Palamara PF et al: Reference-based phasing using the Haplotype Reference Consortium panel. Nat Genet 2016; 48: 1443–1448.
Delaneau O, Howie B, Cox AJ, Zagury JF, Marchini J : Haplotype estimation using sequencing reads. Am J Hum Genet 2013; 93: 687–696.
Howie BN, Donnelly P, Marchini J : A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 2009; 5: e1000529.
McLaren W, Gil L, Hunt SE et al: The ensembl variant effect predictor. Genome Biol 2016; 17: 122.
Leitsalu L, Haller T, Esko T et al: Cohort Profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int J Epidemiol 2015; 44: 1137–1147.
Howie B, Fuchsberger C, Stephens M, Marchini J, Abecasis GR : Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat Genet 2012; 44: 955–959.
Howie B, Marchini J, Stephens M : Genotype imputation with thousands of genomes. G3 2011; 1: 457–470.
Browning BL, Browning SR : Genotype imputation with millions of reference samples. Am J Hum Genet 2016; 98: 116–126.
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR : MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 2010; 34: 816–834.
Steinberg S, Stefansson H, Jonsson T et al: Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat Genet 2015; 47: 445–447.
van Leeuwen EM, Karssen LC, Deelen J et al: Genome of The Netherlands population-specific imputations identify an ABCA6 variant associated with cholesterol levels. Nat Commun 2015; 6: 6065.
Guo MH, Nandakumar SK, Ulirsch JC et al: Comprehensive population-based genome sequencing provides insight into hematopoietic regulatory mechanisms. Proc Natl Acad Sci USA 2017; 114: E327–E336.
Surakka I, Sarin A-P, Ruotsalainen SE et al The rate of false polymorphisms introduced when imputing genotypes from global imputation panels. bioRxiv2016. Available at: https://doi.org/10.110/080770.
Acknowledgements
We would like to acknowledge High Performance Computing Center of University of Tartu. This study was supported by European Union H2020 grants 692145 (ePerMed I/II), 676550, 654248; Estonian Research Council grants IUT20-60, IUT24-6 and SP1GV15007T; European Union through the European Regional Development Fund (Project No. 2014–2020.4.01.15-0012 GENTRANSMED. PP was supported by the Nordic Information for Action eScience Center (NIASC), a Nordic Center of Excellence financed by NordForsk (Project number 62721); and APM is a Wellcome Trust Senior Fellow in Basic Biomedical Science (grant number WT098017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Mitt, M., Kals, M., Pärn, K. et al. Improved imputation accuracy of rare and low-frequency variants using population-specific high-coverage WGS-based imputation reference panel. Eur J Hum Genet 25, 869–876 (2017). https://doi.org/10.1038/ejhg.2017.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.51
This article is cited by
-
Genetic determinants of plasma protein levels in the Estonian population
Scientific Reports (2024)
-
A cost-effective sequencing method for genetic studies combining high-depth whole exome and low-depth whole genome
npj Genomic Medicine (2024)
-
Distinct and shared genetic architectures of gestational diabetes mellitus and type 2 diabetes
Nature Genetics (2024)
-
How local reference panels improve imputation in French populations
Scientific Reports (2024)
-
Genetic drivers of heterogeneity in type 2 diabetes pathophysiology
Nature (2024)