Abstract
Mosaicism, presence of a genetic feature in only a subpopulation of cells, is frequent in de novo genetic diseases. Among large deletions covering the NF1 tumor suppressor gene, the frequency of mosaicism can be as high as 40% in de novo patients. In this study, we demonstrate the high potential of digital PCR in detecting large NF1 deletions and in discriminating mosaic cases. By simultaneously assessing the NF1 gene and a reference gene RPP30, deletions could be unambiguously distinguished from non-deletion samples. Performing the same assay for mixed samples from a DNA with a deletion and a non-deletion DNA, a highly significant linear relation was obtained between the set-up ratio of the two samples and the measured ratio of NF1/RPP30 (P<0.0001), suggesting the high potential of digital PCR in discriminating mosaic deletions. Furthermore, digital PCR detects NF1 allele loss in a tumor specimen that was not detected by loss of heterozygosity analysis using polymorphic markers due to high content of non-tumor cells. Based on the measured ratio of NF1/RPP30, the proportion of the tumor cells in this specimen could be calculated as 25%. Our results demonstrate that dual-probe digital PCR is a simple and effective method for detecting deletions and for discriminating mosaic deletions. Furthermore, this method is sensitive for assigning somatic allele loss in tumor specimen and enables determining proportion of tumor cells.
Similar content being viewed by others
Introduction
At least 5.5% of all pathogenic genetic alterations in human are large genomic deletions or duplications.1 For the autosomal genetic disorder neurofibromatosis type 1 (NF1, MIM# 162200), genomic deletions of 1.2–1.4 Mb covering the NF1 tumor suppressor gene 17q11.2 account for ~5% of all causal genetic alterations.2 Previously, we established a simple and inexpensive method for routine screening of large NF1 deletions by means of genotyping multiple polymorphisms linked to the NF1 gene.2 Combining with subsequent fluorescence in situ hybridization (FISH), we discovered high frequency (40%) mosaicism among NF1 deletions.3 FISH is a sensitive method for discriminating mosaic cases. However, it is laborious, demands special technique and requires fresh blood, which is often not available. In fact, nowadays, FISH has been substituted by multiple ligation-dependent probe amplification (MLPA). However, the semiquantitative method MLPA is less sensitive in discriminating mosaic deletion.4
Somatically, allele loss is a frequent genetic alteration in tumors. However, tumor specimen available for investigations often contain non-tumor cells to various extent. In cases with high proportion of non-tumor cells, polymorphism-based analysis may not have sufficient sensitivity to detect loss of heterozygosity (LOH). Ambiguous cases will usually be concluded as lack of evidence for LOH.
Digital PCR is a recently established technology that enables precise quantification of target sequences.5 In this study, we demonstrate the potential of digital PCR in detecting large NF1 deletions, in discriminating mosaic deletions, in sensitively assigning LOH in tumor specimen with high proportion of non-tumor cells and in determining proportion of tumor cells in such specimen.
Materials and methods
A total of 46 DNA samples were selected from our NF1 patient cohort, among them 24 were confirmed to have large NF1 deletions either by FISH or MLPA.2, 3 Among the 24 deletions, five were confirmed mosaic cases.3 In addition, two specimens of a malignant tumor from a non-deletion patient were included. The two specimens T1 and T2 were obtained from the same tumor at two different operations with a time interval of 1 month. Blood-DNA was originally prepared for genetic diagnosis. Patients gave consent for using the rest of the samples for research. All samples were anonymized for private protection.
LOH analysis for the two tumor specimens was carried out using six intragenic simple repeat polymorphisms of the NF1 gene.2
Dual-probe digital PCR was carried out for counting copy numbers of NF1 and a reference gene RPP30 simultaneously. The validated assays with probes for NF1 (FAM-labeled) and RPP30 (HEX-labeled) were purchased from BioRad (Munich, Germany).
For testing the potential of digital PCR assay in discriminating mosaic deletions, a DNA sample with an NF1 deletion was mixed with a non-deletion sample at 13 different ratios (1:0, 1:2, 1:4, 1:6, 1:8, 1:10, 1:1, 10:1, 8:1, 6:1, 4:1, 2:1 and 0:1) and subjected to the dual-probe digital PCR assay. For a sample without deletion, the NF1/RPP30 ratio should be 1 (both two copies/cell). For a sample with a heterozygous NF1 deletion, this ratio should be 0.5 (one copy of NF1 and two copies of RPP30/cell). In mosaic cases, the proportion of deletion cells varied between 0 and 100% with the corresponding NF1/RPP30 ratio varied between 1 and 0.5.
Raw data of the dual-probe digital PCR assay were analyzed automatically. However, each data set was inspected also manually for clear separation of the positive and negative droplets, and correct setting of the thresholds. Subsequently, the ratio NF1/RPP30 was obtained.
Correlation of set-up proportion of deletion DNA in the mixed sample with the measured ratio NF1/RPP30 was evaluated using linear regression. Proportion of cells with heterozygous deletion of the NF1 allele in a specimen was calculated as: 2 × (1−NF1/RPP30).
Results
Detecting constitutional NF1 deletion
Ratios NF1/RPP30 obtained from dual-probe digital PCR assay unambiguously distinguished 24 NF1 deletion from the 22 samples without deletions (Figure 1).
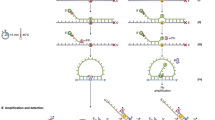
Digital PCR unambiguously discriminates deletion from non-deletion samples. (a) Raw digital PCR data showing numbers of positive droplets of NF1 (blue) and a reference gene RPP30 (green); (b) Corresponding ratios of NF1/RPP30 after correction using Poisson distribution function; (c) Ratios NF1/RPP30 of all samples, among them 22 have confirmed deletions. Orange dots are previously confirmed mosaic deletions. The yellow dot is a deletion case suspected being mosaic. A full color version of this figure is available at the European Journal of Human Genetics journal online.
Potential for discriminating mosaic NF1 deletion
By artificially mixing a deletion DNA with a non-deletion DNA at 13 different ratios, a highly significant linear relation was obtained for these ratios with the measured ratios NF1/RPP30 (P<0.0001, Figure 2). Lineal regression analysis of the data set resulted in the following equation:
Potential of digital PCR for discriminating mosaic deletions. A deletion and a non-deletion sample were mixed at various ratios. The mixed samples were subjected to a dual-probe digital PCR assay, and the resulting ratios NF1/RPP30 (y axis) were plotted against the set-up proportion of the deletion sample (x axis). On the right side, means and SDs from 22 deletion samples and 24 non-deletion samples are given.

The obtained parameters 1.02 and 0.51 were nearly identical to the theoretical ones 1.00 and 0.50, respectively.
Assessing somatic NF1 allele loss
Genotyping intra-NF1 markers revealed LOH in tumor specimen T1 (Figure 3a). Also digital PCR revealed a reduction of the NF1/RPP30 ratio of 0.80, corresponding to 41% of tumor cells with LOH.
NF1 allele loss assessed by polymorphism-genotyping (a) and digital PCR (b, c). In the blood, two peaks are visible indicating heterozygosity of the marker. In tumor specimen T1, one peak is obviously reduced (array), indicating loss of this allele in some cells. By contrast, no allele loss can be assigned for the other tumor specimen T2 that was obtained from later operation of the same tumor. However, digital PCR revealed a slightly reduced NF1 allele in both tumors (b). From the ratios (NF1/RPP30) of 0.80 and 0.87 (c), the tumor content was calculated as 41 and 25% for specimens T1 and T2, respectively.
Interestingly, LOH was not detected in the other specimen T2 that was resected from the same tumor only 1 month later (Figure 3a). However, digital PCR detected a slight reduction in the NF1/RPP30 ratio (0.87). Based on this ratio, the proportion of tumor cells with LOH could be calculated as 25% in this specimen.
Discussion
We demonstrated that dual-probe digital PCR distinguished large NF1 deletions unambiguously from non-deletion cases. Unlike MLPA, digital PCR does not require multiple non-deletion samples as reference and is generally less expensive. Digital PCR has the weakness of examining only one segment and therefore may pick up smaller deletions that do not cover the entire NF1 gene. However, this weakness can be compensated by using two flanking probes for the NF1 gene in the digital PCR assay.
A major strength of digital PCR is its potential in discriminating mosaic cases. For the 24 deletions, mean ratio of NF1/RPP30 was 0.51±0.03, whereas for 22 non-deletions, the mean ratio was 1.00±0.04. Therefore, ratios >0.6 and <0.9 may suggest a deletion in a subpopulation of the cells indicating mosaic deletion. In the present study, we used the BioRad system that has maximal 20 000 nano-droplets for one reaction. An alternative would be the RainDance system (RainDance Technologies, Billerica, MA, USA) that allows 10 000 000 picodroplets for one sample. The 500fold more droplets should further increase the precision and sensitivity in discriminating mosaic deletions.
Interestingly, mosaic NF1 deletions identified in our previous study all had deletions in >90% of blood cells. This fact may reflect a selection effect of our previous screening method that is based on typing polymorphisms. As also demonstrated in the present study, such a method does not have the sensitivity to detect heterozygous deletions in cases where <25% of the cells have such deletions (T2 in Figure 3a). Consequently, mosaic deletions in a small population of affected blood cells may remain undetected.
Similarly, somatic NF1 allele loss in tumors may also have been underestimated in studies based on polymorphism-genotyping. Indeed in the present study, digital PCR detected an allele loss in the specimen T2 that was not detected by marker-based LOH analysis. Furthermore, digital PCR enables calculation of the proportion of tumor cells with defined genetic alteration in the specimen. For many specimens or DNA, especially retrospective ones, information regarding tumor content is often not available. Therefore, digital PCR provides a strategy for recovering this information, which is valuable for designing studies and in interpreting results.
In summary, our results demonstrate that dual-probe digital PCR identifies heterozygous genomic deletions unambiguously, has high potential for discriminating mosaic cases, is sensitive for assigning somatic allele loss in tumor specimen and enables estimation of proportion of tumor cells in study samples.
References
Krawczak M, Cooper DN : The human gene mutation database. Trends Genet 1997; 13: 121–122.
Kluwe L, Siebert R, Gesk S et al: Screening 500 unselected neurofibromatosis 1 patients for deletions of the NF1 gene. Hum Mutat 2004; 23: 111–116.
Kehrer-Sawatzki H, Kluwe L, Sandig C et al: High frequency of mosaicism among patients with neurofibromatosis type 1 (NF1) with microdeletions caused by somatic recombination of the JJAZ1 gene. Am J Hum Genet 2004; 75: 410–423.
van Veghel-Plandsoen MM, Wouters CH, Kromosoeto JN, den Ridder-Klünnen MC, Halley DJ, van den Ouweland AM : Multiplex ligation-depending probe amplification is not suitable for detection of low-grade mosaicism. Eur J Hum Genet 2011; 19: 1009–1012.
Mazaika E, Homsy J : Digital droplet PCR: CNV analysis and other applications. Curr Protoc Hum Genet 2014; 82: 1–13.
Acknowledgements
Mrs Alster is appreciated for her outstanding technical assistance. Also appreciated is Dr Volz and the research group of Dr Dandri for providing the access to their digital PCR device. We further thank Dr Victor Mautner and Dr Hildegard Kehrer-Sawatzki for reading the manuscript and for their suggestions. Funding is in part provided by the Deutsche Krebshilfe (111319).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Kluwe, L. Digital PCR for discriminating mosaic deletions and for determining proportion of tumor cells in specimen. Eur J Hum Genet 24, 1644–1648 (2016). https://doi.org/10.1038/ejhg.2016.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2016.56
This article is cited by
-
Mixed ductal–lobular carcinoma: an analysis of CDH1 DNA copy number variation and mutation
Breast Cancer (2021)
-
High prevalence of SMARCB1 constitutional abnormalities including mosaicism in malignant rhabdoid tumors
European Journal of Human Genetics (2020)