Abstract
The validation and verification of laboratory methods and procedures before their use in clinical testing is essential for providing a safe and useful service to clinicians and patients. This paper outlines the principles of validation and verification in the context of clinical human molecular genetic testing. We describe implementation processes, types of tests and their key validation components, and suggest some relevant statistical approaches that can be used by individual laboratories to ensure that tests are conducted to defined standards.
Similar content being viewed by others
Introduction
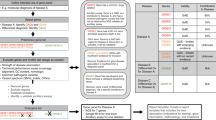
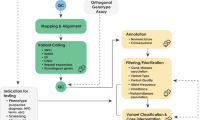
The process of implementing a molecular genetic test for diagnostic use is complex and involves many levels of assessment and validation. The key components of the process, as detailed by the ACCE framework, are analytical validation, clinical validation, clinical utility and consideration of the ethical, legal and social implications of the test.1 After making a decision to set up a diagnostic test, the technology to be used must be chosen and built into a suitable laboratory process. The development stage involves assessment of both the diagnostic and technical use of the process to ensure that the measurements obtained are relevant to the diagnostic question(s) and that the analyte(s) can be unambiguously identified (ie, that there are no confounding factors). The final stage of the laboratory process is to determine whether the performance of the test, in terms of accuracy, meets the required diagnostic standards. Whether this is achieved by performing analytical validation or verification depends on the existence of a suitable performance specification that details the expected accuracy of the test under given conditions. The results of the analytical validation or verification determine whether, and how, the test will be implemented and set the requirements for performance monitoring (ongoing validation) of the test. A simplified process diagram illustrating these concepts is given in Figure 1.
The process of implementing a molecular genetic test for diagnostic use. The shaded arrows represent the two general routes to implementation, depending on the availability of a suitable performance specification: validation (lighter) and verification (darker). Broken arrows represent the situation in which validation or verification fails to meet the specified requirements.
The validation or verification of methods, as defined in Table 1, is a formal requirement for the accreditation of laboratories according to the two major international standards applicable to genetic testing laboratories, ISO 151892 and ISO 17025.3 Although the general requirements are clearly stated (Table 1), the standards provide very little guidance about the detailed requirements or procedures.
To provide more detailed and specific guidance, Eurogentest4 set up a working group comprising clinical and laboratory scientists and experts on quality assurance and statistics from both Europe and the United States. The aims were to develop a framework for validation that could be widely implemented in laboratories to improve the overall quality of genetic testing services while respecting the need for flexibility imposed, for example, by regional requirements and regulations, as well as practical constraints such as test volume and resources. In a recently generated parallel initiative, Jennings et al5 have provided a thorough discussion of FDA regulation, together with a good review of validation procedures. However, specific interpretation of the standards and practical guidance for molecular genetic tests are still lacking. In this paper we propose a generic scheme for the validation and verification of molecular genetic tests for diagnostic use.
Scope
This paper is specifically focused on processes involved in analytical validation and verification of tests in human molecular genetics so as to provide working detail of the first component of the ACCE framework.1 These processes seek to confirm that a particular laboratory process or test delivers reliability that is consistent with the intended diagnostic use. Analytical validation/verification relates only to laboratory processes, and makes no assessment of the manner in which the decision to set up a test is made, as well as the clinical validation, clinical utility or the ethical, legal and social implications of the test.1 In particular, the clinical relevance of the test and the suitability of the chosen measurements with respect to diagnosing a particular genetic disorder are left to professional judgement.
There is much debate about the exact boundary between development and validation, and good cases can be made for different divisions. For the purpose of simplicity, we have defined a definitive boundary placing all concepts that relate to test utility in development (ie, out of scope) and parameters relating to test accuracy in validation (ie, within scope).
These limitations of the scope should not be taken as assigning different levels of importance to the various processes; making clinically useful and appropriate measurements is clearly critical to setting up a valid diagnostic test. For this reason, we have included a brief section outlining the processes involved and important factors that should be considered at the development stage.
Although we are concerned with appropriate use of statistics and sample sizes, this paper is not intended to be a treatise on the subject, but a practical guide for diagnostic molecular geneticists to aid them in designing, performing and reporting suitable validation or verification for the tests they wish to implement. References have been provided in which more complex statistical concepts are involved, but it is recommended that the advice of a statistician be sought in case of doubt. Above all, we seek to promote a pragmatic approach; although validation and verification must be carefully considered, test implementation must also be achievable and not overburdening.
Although there is much literature addressing validation on a more general level,6, 7, 8 we propose a first attempt to identify and organize the components required for validation/verification in the context of molecular genetic diagnostics, and have consequently included some measure of simplification of statistical principles and their interpretation. It is intended that this paper be a starting point for the ongoing development of validation/verification guidelines that will necessarily become more sophisticated as knowledge and experience in the area increase and scenarios that are not adequately covered by this paper are identified. Although these recommendations are aimed primarily at molecular genetic testing, we believe that the principles and concepts are also applicable in the context of cytogenetics.
To help guide the validation process and provide a format for recording validations, a standardized validation pro forma (template) has been provided in Supplementary data. An initial draft of this form was developed from an amalgamation of standard forms used in a range of small and large laboratories that undertake genetic testing. This prototype underwent a field trial to assess its use, as well as ease of use and appropriate amendments made. We recognize that a single format is unlikely to suit all laboratories; hence we recommend that this form should be used as a starting point for the development of a suitable format for local needs.
This paper can be used as detailed explanatory notes for the validation form.
The implementation process
Development
The purpose of development is to establish a testing procedure and broadly show that it is fit for the intended purpose, in terms of what is being tested and of the desired laboratory procedure. This involves defining the analyte(s) to be tested and designing an appropriate methodology, including any assay-specific reagents (eg, primers), controls and a testing workflow. The development process should be used to gain the necessary experience with the test, with the aim of identifying any critical parameters that may affect performance and any necessary control measures and limitations that need to be considered. Examples of critical parameters may include primer design, location of known polymorphisms, the G+C content of the region of interest, fragment length, type of mutations to be detected and location of mutations within fragments. Suitable control measures might include the use of positive, negative and no-template controls, running replicates of the test and a quality scoring system. It therefore follows that the amount of development required would depend on the novelty of the testing procedure, both on a general level (ie, in the literature) and within the laboratory setup of the test. For example, development of a sequencing test for a new gene in a laboratory with extensive experience in sequencing may simply be a case of primer design, whereas setting up an entirely new methodology would require much more extensive investigation.
Assessment of use
Before a test can be validated, it is necessary to establish (a) that the particular measurements are diagnostically useful and (b) that the correct analyte(s), and only the correct analyte(s), are measured. This could involve, for example, ensuring that primers do not overlay known polymorphisms in the primer-binding site and that they are specific to the target of interest. It should be noted that use of a CE-marked kit does not preclude assessment of use; care should still be taken to ensure that the test measures suitable analyte(s) for the intended purpose, as in vitro diagnostic device (IVDD) compliance relates only to technical performance and not to clinical or diagnostic validity. Three other critical concepts that should be considered at this stage are the following:
Selectivity
How good is the method at distinguishing the signal of the target from that of other components? For example, a PCR product run on a denaturing polyacrylamide gel to detect the presence of the CFTR p.Phe508del (p.F508del) mutation associated with cystic fibrosis will also detect the rarer p.Ile507del (p.I507del) mutation, without distinguishing between them. For most genetic tests, selectivity issues are best avoided by careful design (eg, BLAST9 primers to avoid nonspecific amplification) or by applying adapted controls and/or limitations.
Interference
Are there any substances the presence of which in the test can affect the detection of the target sequence? If so, will this cause the reaction to fail or is there a possibility of an incorrect result? For most genetic tests, this is likely to relate to substances that cause a reaction to fail (eg, heparin or ethanol in a DNA sample as a result of the stabilization or extraction procedure). Although failures may not generate false results, there can be issues relating to the use and timeliness of tests if failure rates are too high. In situations in which interference could cause incorrect results, great care needs to be taken to avoid interfering substances, for example, by running a pretest quality check on samples or by including more controls.
Because of their complex nature, multiplex assays are particularly susceptible to interference, which could give rise to incorrect results. Validation and verification of this type of assay can be particularly demanding and is beyond the scope of this paper. The Clinical Laboratory Standards Institute (CLSI) has published a guideline that deals comprehensively with this specialist topic.10
Carryover (cross-contamination)
This relates to residual products from previous or concurrent analyses that may be introduced into an assay (eg, through a contaminated pipette). Stringent procedural precautions should be used as a matter of routine to minimize the risk of such cross-contamination. In particular, physical separation of pre- and post-PCR areas for both reagents and laboratory equipment is critical. Other controls/precautions may include the use of no-template controls and uracil-N-glycosylase treatment.11, 12, 13
Performance specification
Once a suitable test procedure has been established and it is judged that there is sufficient local knowledge of critical parameters, it is necessary to show that
-
a)
test performs to a suitable level of accuracy for the intended purpose: that is, it produces results that can satisfactorily answer the clinical question allowing for uncertainty of measurement; and that
-
b)
this level of accuracy is routinely maintained.
The level of testing required is dependent on the availability of a suitable performance specification. This should define all the test conditions necessary to achieve a particular level of accuracy, together with measurable parameters that can be used to show that this is the case; specifically,
-
a)
an estimate of the test accuracy including measurement uncertainty (eg, confidence limits);
-
b)
control measures required to ensure routine maintenance of accuracy;
-
c)
limitations on critical parameters that will ensure the desired level of accuracy.
For validation of a specific test, limitations may include factors such as input DNA concentration or details on how DNA extraction needs to be performed. When a technology is being validated (as opposed to a specific test), there may also be limitations related to physical parameters such as PCR fragment length or G+C content. It should be stressed that a performance specification will only apply within particular limits of certain critical parameters; hence, care should be taken to ensure that the new test falls within these limits. For example, the performance specification for a hypothetical method for mutation scanning (>95% sensitivity for mutations in fragments <300 bp long and 25–60% G+C content) would not be applicable to a new test involving a 400-bp fragment or fragments with 70% G+C content.
Validation
Full validation is required when there is no suitable performance specification available, for example, with novel tests or technologies. This process involves assessing the performance of the test in comparison with a ‘gold standard’ or reference test that is capable of assigning the sample status without error (ie, a test that gives ‘true’ results). In simple terms, validation can be seen as a process to determine whether we are ‘performing the correct test’. In the field of medical genetics, with the almost complete absence of reference tests or certified reference materials, the reference should be the most reliable diagnostic method available. It is worth noting that the gold standard does not have to comprise results from a single methodology; different techniques could be used for different samples and in some cases the true result may represent a combination of results from a portfolio of different tests. To avoid introducing bias, the method under validation must not, of course, be included in this portfolio.
Validation data can be used to assess the accuracy of either the technology (eg, sequencing for mutation detection) or the specific test (eg, sequencing for mutation detection in the BRCA1 gene). Generally speaking, the generic validation of a novel technology should be performed on a larger scale, ideally in multiple laboratories (interlaboratory validation), and include a much more comprehensive investigation of the critical parameters relevant to the specific technology to provide the highest chance of detecting sources of variation and interference.
Verification
If a suitable performance specification is available, it is necessary to establish that the new test meets this specification within the laboratory; this process is called verification. In simple terms, verification can be seen as a process to determine that ‘the test is being performed correctly’.
Verification should usually be appropriate for CE-marked IVDD-compliant kits, but care should be taken to ensure that the performance specification is sufficient for the intended use of the kit, particularly with kits that are self-certified. Most diagnostic genetic tests are classified by the IVD directive as ‘low-risk’ and can be self-certified by the manufacturer without assessment by a third party. Such tests can be identified by the absence of a number following the CE mark (Article 9: IVDD Directive 98/79/EC).14, 15 If, at any stage, the test procedure associated with the performance specification is modified (eg, if reaction volumes of a CE-marked kit are reduced), verification is not appropriate and validation is required.16
Other applications of verification may include a new test being implemented using a technology that is already well established in a laboratory (eg, a sequencing assay for a new gene), or a test for which a suitable performance specification is available from another laboratory in which the test has already been validated. In all cases, it is essential that laboratories obtain as much information as possible with regard to the validation that has been performed.
Reporting validation and verification
The plan, experimental approach, results and conclusions of the validation or verification should all be recorded in a validation file, along with any other relevant details (see the section ‘Reporting the results’). In addition, the validation plan and outcome should be formally reviewed and approved. When reporting validations or verifications in peer-reviewed publications, it is strongly recommended that the STARD initiative (Standards for Reporting of Diagnostic Accuracy)17 be followed as far as possible.
Performance monitoring (ongoing validation)
Once a test validation has been accepted (ie, the use and accuracy have been judged to be fit for the intended diagnostic purpose), it is ready for diagnostic implementation. However, this is not the end of performance evaluation. The performance specification derived from the validation should be used to assess the ‘validity’ of each test run and this information should be added to the validation file at appropriate intervals. In many cases, the accumulation of data over time is an important additional component of the initial validation, which can be used to continually improve the assessment of test accuracy and quality. The ongoing validation should include results of internal quality control, external quality assessment and nonconformities related to the test or technique as appropriate.
Types of test
The core aim of validation is to show that the accuracy of a test meets the diagnostic requirements. Essentially, all tests are based on a quantitative signal, even if this measurement is not directly used for the analysis. Although measuring the proportion of a particular mitochondrial variant in a heteroplasmic sample is, for example, clearly quantitative, the presence of a band on a gel is commonly considered as a qualitative outcome. However, the visual appearance of the band is ultimately dependent on the number of DNA molecules that are present, even though a direct measurement of this quantity is rarely determined. These differences in the nature of a test affect how estimates of accuracy can be calculated and expressed.
For the purpose of this paper, we are concerned with two types of accuracy. Determining how close the fundamental quantitative measurement is to the true value is generally termed ‘analytical accuracy’. However, it is often necessary to make an inference about the sample or the patient on the basis of the quantitative result. For example, if the presence of a band on a gel signifies the presence of a particular mutation, test results are categorized as either ‘positive’ or ‘negative’ for that mutation, on the basis of the visible presence of the band. Such results are inferred from the quantitative result, but are not in themselves quantitative. Determination of how often such a test gives the correct result is termed ‘diagnostic accuracy’. The term diagnostic accuracy is generally used to describe how good a test is at correctly determining a patient's disease status. However, genotype does not necessarily equate directly to disease status (phenotype) for various reasons, including incomplete penetrance/modifying factors or simply because the patient is presymptomatic. The purpose of these guidelines is to enable laboratories to establish how good their tests are at correctly determining genotype; clinical interpretation of the genotype is not considered in this context. Therefore, for the purpose of this paper, the term diagnostic accuracy will be taken to relate exclusively to the ability of a test to correctly assign genotype irrespective of any clinical implication.
We distinguish three broad test types (quantitative, categorical and qualitative) that can be subdivided into five groups according to the method for interpreting the raw quantitative value to yield a meaningful result.
The following sections discuss each of these test types in more detail and provide guidance on appropriate measurement parameters in each case. A summary of the characteristics of the different test types and examples is given in Table 2, together with recommendations for appropriate measurement parameters and timing of validation.
Type A quantitative tests
For a quantitative test, the result is a number that represents the amount of a particular analyte in a sample. This can be either a relative quantity, for example, determining the level of heteroplasmy for a particular mitochondrial allele, or an absolute quantity, for example, measuring gene expression. In either case, the result of a quantitative test can be described as continuous as it can be any number (between two limits), including decimal numbers.
Two components of analytical accuracy are required to characterize a quantitative test: trueness and precision.18, 19 Trueness expresses how close the test result is to the reference value. Typically, multiple measurements are made for each point and the test result is taken to be the mean of the replicate results (excluding outliers if necessary). As quantitative assays measure a continuous variable, mean results are often represented by a regression of data (a regression line is a linear average). Any deviation of this regression from the reference (ie, the line where reference result equals test result) indicates a systematic error, which is expressed as a bias (ie, a number indicating the size and direction of the deviation from the true result).
There are two general forms of bias. With constant bias, test results deviate from the reference value by the same amount, regardless of that value. With proportional bias, the deviation is proportional to the reference value. Both forms of bias can exist simultaneously (Figure 2).
Although measurement of bias is useful (Figure 3), it is only one component of the measurement uncertainty and gives no indication of how dispersed the replicate results are (ie, the degree to which separate measurements differ). This dispersal is called precision and provides an indication of how well a single test result is representative of a number of repeats. Precision is commonly expressed as the standard deviation of the replicate results, but it is often more informative to describe a confidence interval (CI) around the mean result. For example, a result for a test investigating mutation load in a tumour sample might be described as 7% (95% CI: 5–10%).
Performance characteristics, error types and measurement metrics used for quantitative tests (adapted from Menditto et al)19.
Precision is subdivided according to how replicate analyses are handled and evaluated. Here, there is some variability in the use of terminology; however, for practical purposes, we recommend the following scheme based on ISO 3534-120 and the International Vocabulary of Metrology:21
Repeatability refers to the closeness of agreement between results of tests performed on the same test items, by the same analyst, on the same instrument, under the same conditions in the same location and repeated over a short period of time. Repeatability therefore represents ‘within-run precision’.
Intermediate precision refers to closeness of agreement between results of tests performed on the same test items in a single laboratory but over an extended period of time, taking account of normal variation in laboratory conditions such as different operators, different equipment and different days. Intermediate precision therefore represents ‘within-laboratory, between-run precision’ and is therefore a useful measure for inclusion in ongoing validation.
Reproducibility refers to closeness of agreement between results of tests carried out on the same test items, taking into account the broadest range of variables encountered in real laboratory conditions, including different laboratories. Reproducibility therefore represents ‘inter-laboratory precision’.22
In practical terms, internal laboratory validation will only be concerned with repeatability and intermediate precision and in many cases both can be investigated in a single series of well-designed experiments. Reduced precision indicates the presence of random error. The relationship between the components of analytical accuracy, types of error and the metrics used to describe them is illustrated in Figure 3.
Any validation should also consider robustness, which, in the context of a quantitative test, could be considered as a measure of precision. However, robustness expresses how well a test maintains precision when faced by a specific designed ‘challenge’, in the form of changes in preanalytic and analytic variables. Therefore, reduced precision does not represent random error. Typical variables in the laboratory include sample type (eg, EDTA blood, LiHep blood), sample handling (eg, transit time or conditions), sample quality, DNA concentration, instrument make and model, reagent lots and environmental conditions (eg, humidity, temperature). Appropriate variables should be considered and tested for each specific test. The principle of purposefully challenging tests is also applicable to both categorical and qualitative tests and should be considered in these validations as well. Robustness can be considered as a useful prediction of expected intermediate precision.
As trueness and precision represent two different forms of error, they need to be treated in different ways. In practice, systematic error or bias can often be resolved by using a correction factor; constant bias requires an additive correction factor, whereas proportional bias requires a multiplicative correction factor. For example, results from a test that has +5% bias can be multiplied by 100/105. Random error, in contrast, cannot be removed, but its effects can generally be reduced to acceptable levels by performing an appropriate number of replicate tests.
For the purpose of this paper, a basic understanding of the concepts described above is the main objective. However, it is worth outlining some of the complexities that can arise in estimating the analytical accuracy of quantitative tests. In molecular genetics, quantitative measurements are most often relative, that is, two measurements are taken and the result is expressed as a proportion (eg, the percentage of heteroplasmy of a mitochondrial mutation). In such cases, it is preferable to perform both measurements in a single assay to minimize the effects of proportional bias, as the assay conditions are likely to affect both the measurements in a similar way.
If the measurements must be taken in separate assays, each measurement is effectively an absolute measurement and must be quantified in comparison with a set of calibration standards run with each test batch. In this scenario, it is important to assess the variation in each test/standard pair, as even minor variation can dramatically affect the overall analytical accuracy. This is most effectively achieved by monitoring the efficiencies of the two reactions over time.23
For quantitative tests, particularly those requiring absolute quantification, it is most effective to estimate analytical accuracy on an ongoing basis by running a set of calibration standards (standard curve) with each batch or run. In this case, it is important that linearity be evaluated24 and that the lower and upper standards are respectively below and above the expected range of the results as precision cannot be assessed on extrapolated results. Where possible, calibration standards should be traceable to absolute numbers or to recognized international units.
Other factors that may need to be evaluated include the limit of detection defined as the lowest quantity of analyte that can be reliably detected above background noise levels and the limits of quantification that define the extremities at which the measurement response to changes in the analyte remains linear.
A detailed description of the determination of these limits is given in CLSI document EP17-A.25 In situations in which test results are likely to fall close to these extremities or there are significant clinically relevant boundaries within the linear range (eg, the intermediate expansion/mutation boundary in Huntington's disease), it is useful for both implementation and ongoing validation to use controls on or close to the boundary.
It should be noted that limit of detection is sometimes referred to as ‘sensitivity’; that is, how sensitive a methodology is to detecting low levels on a particular analyte in a large background. Use of the term ‘sensitivity’ in this context should be avoided, as it may be confused with sensitivity described in the section ‘Qualitative tests’ (ie, the proportion of positive results correctly identified by a test).
It can be seen that the analysis of all but the simplest quantitative assays can be complex and it is recommended that statistical advice be sought to determine those factors that need to be measured and the best way to achieve it.
Categorical tests
Categorical tests (sometimes referred to as semiquantitative26) are used in situations in which quantitative raw data, which could have any value including decimals, are grouped into categories to yield meaningful results. For example, fluorescent capillary analysis might be used to determine the size of PCR products (in base pairs) by analysing the position of the peaks relative to an internal size standard. The quantitative results from this analysis will include numbers with decimal fractions, but the length of the product must be a whole number of base pairs; a fragment cannot be 154.3 bp long. Therefore cutoffs must be used to assign quantitative results to meaningful categories. The parameters used to describe the estimates of analytical accuracy for a quantitative test (Figure 3) can be used to describe the performance of the categorical test in much the same way. However, there is an added level of complexity here, as the primary (quantitative) result is manipulated (ie, placed into a category). The categorized results for these tests retain a quantitative nature (although this is distinct from the quantitative primary data) and, in practice, trueness and precision can be determined at the category level, as well as at the level of the primary result. We divide categorical tests into two subgroups, depending on the number and type of categories and the degree of importance placed on knowing how accurate a result is (Figure 4).
(a) A type-B categorical test to size PCR fragments. Each category (indicated by alternating shading) has an upper cutoff that is also the lower cutoff of the next category. Results marked with arrows are not precise but fall within the given accuracy for the test of ±1 bp. (b) A type-C categorical test for allele quantification. Each category (shaded) has unique upper and lower cutoffs. Results falling between categories are classed as unreportable (marked with an arrow). A dosage quotient (DQ) of 0.5 represents a sample with a deleted allele, 1.0 represents normal and 1.5 represents a sample with a duplicated allele.
Type B categorical tests
This group includes tests in which there are (essentially) unlimited categories, such as the sizing example cited above. In this case, each cutoff forms the upper boundary of one category and the lower boundary of the next, so that all results can be categorized (except for those that have failed). Generally, less-stringent levels of accuracy are acceptable with this type of test. In this case, estimation of precision can be performed before implementation (eg, ±1 bp), whereas trueness is dealt with by running a standard curve with each experiment (ie, a size standard).
Type C categorical tests
When the number of predefined categories is limited, for example, with allele copy number determination, accuracy tends to be critical and a more definitive approach is often required. The most informative way to express accuracy for this type of test is the probability that a particular (quantitative) result falls into a particular category. Here, cutoffs are defined at particular level(s) of probability, typically 95% CI, which means that each category has its own unique upper and lower boundaries with regions in between, where results would be classified as unreportable.
Results can be assigned to the appropriate categories by a process of competitive hypotheses testing. For example, a test to determine constitutional allele copy number has three expected results: normal (2n), deleted (n) and duplicated (3n). The odds ratios p(2n):p(n) and p(2n):p(3n) can be used to assign results (Figure 5). It should be noted that mosaic variants may give rise to intermediate values; detection of mosaics should be considered under quantitative tests. A good example of this methodology is described in the MLPA spreadsheet analysis instructions that are freely available from NGRL (Manchester).27 In this case, the validation of accuracy is predominantly carried out on an ongoing basis by running replicate control assays during the actual test run to determine the extent of the random error observed within that particular run.
Multiplex ligation-dependent probe amplification to detect exon copy number (Categorical test type C). Dosage quotient (DQ)=relative height of test peak compared with control peaks. DQ=0.5 represents exon deletion, DQ=1.0 represents wild type and DQ=1.5 represents exon duplication. Population distributions of DQs are shown with 95% confidence intervals shaded. Results falling between categories are unreportable.
Qualitative tests
This is the extreme form of a categorical test, in which there are only two result categories, positive and negative. This binary categorization can be based either on a cutoff applied to a quantitative result, for example, peak height or a mathematical measure representing peak shape, or on direct qualitative observation by the analyst, for example, the presence or absence of a peak (in the latter case, as discussed in the section ‘Types of test’, the underlying data will generally be quantitative in nature, even though no formal quantification is performed). In terms of accuracy, categorization can be either correct or incorrect with respect to the ‘true’ (reference) result. A simple contingency table can be used to describe the four possible outcomes (Table 3).
The diagnostic accuracy of a qualitative test can be characterized by two components, both of which can be calculated from the figures in the contingency table:
-
i)
Sensitivity – the proportion of positive results correctly identified by the test=TP/(TP+FN);
-
ii)
Specificity – the proportion of negative results correctly identified by the test=TN/(TN+FP).
In addition, the overall accuracy can be characterized by the total number of true results as a proportion of the total results ((TP+TN)/(TP+TN+FP+FN)), although, in practice, this parameter is rarely used. For comparison with quantitative tests (Figure 3), the relationship between the components of accuracy is depicted in Figure 6.
The relationship between performance characteristics, error and measurement uncertainty used for qualitative tests (adapted from Menditto et al)19.
There is an inverse relationship between sensitivity and specificity (Figure 7). As more stringent cutoffs are used to reduce the number of false positives (ie, increase specificity), the likelihood of false negatives increases. Therefore, the desirable characteristics of a test must be considered in the context of the required outcome and the diagnostic consequences. For example, laboratory procedures for mutation scanning tests often involve a primary screen to determine which fragments carry mutations, followed by a second confirmatory test by sequencing to characterize the mutations present. In the primary screen, sensitivity is much more critical than specificity, to avoid missing mutations that are present; the only consequence of poor specificity is increase in the workload for confirmatory sequencing. Obviously, there is a limit to the lack of specificity that can be tolerated, even if only on the grounds of cost and efficiency.
(a) The relationship between sensitivity and specificity. The figure shows frequency distributions of the primary quantitative results for a qualitative (binary) test. Solid line represents gold standard negatives (wild type), broken line represents gold standard positives (mutant). Using a single cutoff to categorize the results as either positive or negative gives rise to both false negatives and false positives. (b) Cutoff location. Positioning the cutoff to the right encompasses more of the negative distribution, giving a low false-positive rate but a high false-negative rate (shaded). As the cutoff is moved to the left, the false-negative rate is reduced but the false-positive rate increases. (c) Use of two cutoffs. It is possible to minimize both false-positive and false-negative rates by using two cutoffs. In this case, results falling between the two cutoffs can either be classified as test failures or be passed for further analysis.
In situations in which sensitivity and specificity are both critical, it is desirable to use two cutoffs to minimize both false-positive and false-negative rates. In this case, results falling between the two cutoffs can either be classified as test failures or be passed for further analysis.
Type D qualitative tests
In many cases, particularly for mutation scanning methods, it is necessary to use a qualitative description to distinguish between a single normal state (negative result) and any number of mutated states (positive result). Although quantitative results for the normal state would be expected to be normally distributed, positive results would not, as they combine many (known or potential) different mutations, each with its own distribution (Figure 8b). Although it is still theoretically possible to use basic hypothesis testing to assign a probability that a result is not normal, competitive hypotheses cannot be used, as it is not possible to know the mean quantitative result for all possible mutations (unless they have all been tested). In this scenario, assessment of accuracy is therefore best performed in a preimplementation validation using a suitable number of positive (ie, known mutant) and negative (known normal) samples (see the section ‘Study design’).
(a) Truly binary test. Only two states of the analyte can be measured: one wild type (solid line) and one mutant (broken line). Competitive hypothesis testing could be used to determine the relative probability (odds ratio) that a result is either wild type or mutant. (b) Arbitrary binary test. There are many different possible states of the analyte; one wild type (solid line) and five different mutants (broken lines) are shown. The mutant state indicated is essentially indistinguishable from the wild type. Hypothesis testing could be used to estimate the probability that a result is not negative, but not that it is not positive.
Type E qualitative (binary) tests
In cases in which the test is designed to measure only two states of the analyte (eg, a specific SNP genotyping assay), the quantitative results for each state can be expected to be normally distributed (Figure 8a). In this case, results can be assigned to appropriate categories by competitive hypothesis testing, as described for type C categorical tests (see the section ‘Type C categorical tests’). Again, this model can be used as an ongoing validation method, minimizing the need for implementation validation. Test accuracy can also be described in terms of sensitivity and specificity, given particular cutoffs. This method would require much more stringent validation before implementation (see the section ‘Study design’).
Sequencing
Direct sequencing (currently, fluorescent dideoxy-terminator sequencing by capillary electrophoresis) is the method of choice for a wide range of clinical genetic tests and is widely considered to be the ‘gold standard’ (reference) method for identifying and characterizing DNA variations. As such, it is often not possible to develop a suitable reference for comparative validation of new sequencing-based tests. In this situation, it is recommended that validation be treated as a verification that sequencing is being performed to the required standard, in the context of the new test. Factors to be considered should include confirmation that the new test specifically targets the region of interest (ie, BLAST primers, and check sequence), that both alleles are reliably amplified (ie, ensure that no SNPs are located in primer binding sites) and that the sequencing data generated are consistently of suitable quality across the whole region of interest (eg, monitoring PHRED scores across the region of interest). It is important to note that, as sequencing methodologies can vary, for example, by the cleanup method, thermal cycling regime, or whether single or bidirectional sequencing is used for analysis, the validation scheme should be carefully tailored to the application. This is of particular importance when a new sequencing test is being ‘imported’ from another laboratory, as most laboratories will have their own particular sequencing methodology and this is unlikely to be identical to the local method.
As with other tests, it is important to participate regularly in an external quality assurance (EQA) scheme where possible. In the case of sequencing, this may be dealt with at the technology level in addition to disease-specific schemes; for example, the MSCAN and SEQ DNA schemes run by the European Molecular Genetics Quality Network (EMQN).28
Considerations for experimental design
Extrapolation of results (validation constraints)
The results of a validation can be applied beyond its immediate coverage; however, some rationale needs to be applied to such extrapolation. Let us consider the validation of a mutation-scanning technology that tested 100 different mutations in a particular gene (5000 bp) resulting in a sensitivity of ‘≥97% (95% CI)’ (see the section ‘Qualitative tests’ for calculating and reporting sensitivities). What does this actually mean in practice?
Only a very small number of the possible mutations in the region of interest were actually covered; there are 15 000 possible single-base substitutions in 5000 bp and virtually limitless insertion/deletion mutations. If only substitutions were tested in the validation, the estimated sensitivity could only reasonably be considered to apply to these types of mutations. However, assuming that all different types of mutations were broadly covered by the validation (eg, all possible nucleotide substitutions, different lengths of insertion and deletion and so on), it would be reasonable to say that sensitivity of mutation scanning in this gene using this method had been shown to be ≥97% (95% CI).
It is often appropriate to examine particular categories separately on the basis of specific knowledge of a test system. For example, it is known that certain single-base insertions or deletions in homopolymer stretches can be refractory to detection by high-resolution melting. To gain a realistic understanding of how relevant this might be, particular attention might be paid to this group of variations by including a disproportionate number in the validation. The specific gene or disease should also be considered: if only amino-acid substitutions are expected, a reduced sensitivity to single-base insertions would be irrelevant.
Broadly speaking, the limits of extrapolation can be defined by coverage of the parameters considered to be critical to the successful outcome of the test. That is, if mutation type is considered as a critical factor in achieving a correct result with the given test, then as many different types of mutations need to be included in the validation as possible. Equally, if the G+C content of the template is considered a critical factor, validation is only applicable to fragments within the G+C content range covered by the validation. This means that a validation of a technology could be applicable to a new gene even if the validation was carried out exclusively on another gene or genes, provided the test carried out on the new gene falls within the critical parameters of the validation (obviously, in this case, it is critical to ensure that the correct fragments are being amplified). In this case, the gene itself is not a critical factor.
Potentially critical factors should be identified and evaluated at the development stage, on the basis of previous experience and expertise with the technology being validated. However, with primary validation of new technology, attempts should be made to identify the key parameters by performing an evaluation covering as many different potential factors as possible (full or partial factorial). It is also recommended that interlaboratory reproducibility be evaluated (see the section ‘Type A quantitative tests’).
Sample selection
The limits of extrapolation of the validation results is ultimately defined by the choice of samples, which itself is generally limited by the availability of positive controls. For this reason, it is essential that the sample profile be clearly detailed in the validation report, together with an analysis of how this relates to the factors considered critical to the performance of the test.
Positive (mutant) samples should be chosen to represent as broad a range of results as possible, bearing in mind the desire or requirement for extrapolation of the results. This will depend on the purpose of the test under validation. For example, for validation of a method for mutation scanning, samples containing as many different mutations as possible should be included in the validation. In this context, it is not normally important that the mutations/variations chosen for the validation are actually pathogenic, as this is not normally relevant to whether they will be detectable. It should be noted that including multiple examples of the same mutation in the same amplicon will not increase the power of the study to determine sensitivity, as each repeat cannot be considered different with respect to sensitivity. It is also valuable to include examples in which potentially confounding variations exist (ie, is it possible to detect a mutation in a fragment containing a certain common polymorphism?).
In general, it is desirable to include samples containing mutations that represent the range of possible variation in parameters that are important to the technique under test. For example, key parameters for a technique that relies on heteroduplexing or melting would include the G+C content of the fragment, the position of the mutation in the fragment and the actual nucleotide change.
In some cases, particularly when validating a new technology, local limitations of sample availability may necessitate an interlaboratory collaboration to collect a suitable number of samples to attain the required power for diagnostic validation.
Sample size (numbers)
The number of samples used in a validation determines its statistical power, which is a measure of how much confidence can be placed on the results of the validation. Therefore, validation sample size is ultimately one of the most important factors in determining the analytical use of the test. Unfortunately, definitive guidelines defining specific sample sizes cannot be realistically given, as the requirement is so dependent on a wide range of factors, including the nature and performance of the test, the critical parameters, the way in which the test will be used in practice and the confidence level required for clinical use. A large number of tools for determining sample size, given certain input criteria (eg, confidence interval), are freely available on the internet (eg http://www.statpages.org/#Power, accessed May 2010).
The Clinical and Laboratory Standards Institute provides a number of evaluation protocols (prefixed EP) making reference to sample size requirements for a variety of situations.29, 30, 31, 32, 33 Although these tools will give useful estimates of the numbers of samples required, the limiting factor is often the availability of suitable control samples,34 even in the case of verification, which requires less-stringent analysis and therefore fewer samples. In this case, it is critical to understand the statistical relevance of using the given sample size and how the confidence level achievable with this sample size affects the utility of the test. It is recommended that statistical advice be sought and this is carefully reviewed in the context of clinical utility. As Jennings et al,5 state: ‘Although supporting evidence is essential to scientific understanding, it must be recognised that statistically significant data may not be available for every aspect of every validation study. There is simply not enough patient material, financial support, or scientist/technologist time to support such rigorous data collection. Therefore, medical judgement in the context of in-laboratory data and the larger health-care system is essential to deciding when a test is ready to be introduced for patient care.’
Whatever the availability of samples or the outcome of the validation, it is important to accurately record all details in the validation file, including confidence levels and the basis of any decisions made.
Qualitative tests
Estimating power
In the case of qualitative tests, there is a useful rule of thumb that can be used to estimate the power of a study given a particular number of samples. This can be illustrated by the following two qualitative validations of a methodology for mutation scanning:
-
a)
Validation using 30 different mutations.
-
b)
Validation using 300 different mutations.
If all mutations were correctly identified in both validations, the measured sensitivity would be 100% in both cases. However, we are likely to be much more confident in the results of validation (b) because a wider range of different mutations has been tested. This difference relates to the confidence that certain mutations, which cannot be detected by the technique, have not been excluded from the validation by the random selection of samples. This confidence increases as more different mutations are tested. This problem is referred to in statistics as sampling error. For a qualitative test, the goal is to determine a sample size that will provide sufficient power to determine sensitivity and specificity to the desired level of confidence for the particular application.
Precise calculations can be complex, but for practical purposes the ‘rule of 3’ provides a sufficiently accurate estimate of power according to sample size.35, 36, 37 This states that, at 95% confidence, the probability of an event that is not seen in validation of sample size n is 3/n. An illustration of the use of the ‘rule of 3’ using the examples above is given in Table 4.
With molecular genetic tests, technologies are often highly sensitive and the target of validation is often a sensitivity approaching 100%; a test that does not achieve a measured sensitivity of 100% is often not considered suitable for diagnostic purposes. Although it is likely that a false negative would be found given a big enough sample size, this expectation does mean that sample numbers calculated using the ‘rule of 3’ generally yield the required results.
In practical terms, the ‘rule of 3’ will give very accurate estimates for studies in which n>60; below this, the estimates become overcautious, which is not a bad thing in diagnostics. This rule is valid for any proportion; therefore, it can be used for either sensitivity or specificity.
Study design
As we have seen, the power of validation data is related to sample size. The number of positive samples (mutant) will prescribe the power to estimate sensitivity, and the number of negative samples (normal) that of specificity. For most applications it is sufficient to include equal numbers of mutant and wild-type samples; this will yield equal power to estimate both sensitivity and specificity.
This has a practical implication: for the validation of a mutation scan of over 50 amplicons using 100 mutant samples, it is not useful to screen all samples for all amplicons (ie, total of 50 × 100=5000 tests). This equates to 100 analyses of mutant samples (power to estimate sensitivity=97% (1−3/100 by ‘rule of 3’)) but to 4900 analyses of normal samples (power to estimate specificity=99.94% (1−3/4900)). There is clearly a disproportionate power to estimate specificity, which in this case is likely to be the less important measure. It would be sufficient to perform 100 analyses of normal samples (total analyses 200), although it would be sensible to evenly distribute these analyses among the amplicons. In situations in which sensitivity or specificity is considered to be particularly important, it may be appropriate to weight the number of mutant and normal samples appropriately.
It is critical that validation be performed without any knowledge of the actual status of each sample (ie, blinded analysis), especially in the case of categorical and qualitative tests. To eliminate systematic errors or bias, consideration should also be given to sample order, which should be randomized as much as is practically possible. It may also be beneficial to introduce redundancy into the experiment (eg, by duplication) to ensure coverage of all the required results. Although this is not critical to the validation results per se, it can save time repeating failed analyses. In addition, these data can be used in the determination of precision (repeatability and/or reproducibility).
Reporting the results
Comprehensive and clear documentation of validation is extremely important, both during the preimplementation phase and during ongoing validation. When reporting the results of a validation experiment, it is important to include the derived estimates of diagnostic accuracy, including confidence intervals and all details that may affect the interpretation of these estimates, including the following:
-
Sample inclusion criteria
-
Nature of the samples
-
Details of reference method
-
Technical details
-
Handling of failures
-
Critical parameters tested
-
Equipment details.
Reporting estimates of accuracy
Quantitative and categorical tests
In all cases in which estimates of accuracy are reported, some measure of the confidence that is applicable to the estimate should also be given. The confidence applied to quantitative measures is essentially the precision (with the exception of measures of probability, which are measures of confidence in themselves). This can most usefully be expressed as a confidence interval around the mean of the replicate results. The following is a simple guide to calculating confidence intervals:
-
1
Calculate the mean of the replicates (M)
-
2
Calculate the standard deviation of the replicates (SD)
-
3
Calculate the standard error sM=s/√N (where N=number of replicates)
-
4
Calculate degrees of freedom, d.f.=N−1
-
5
Find t for this d.f. using a Student's t table
-
6
Lower confidence limit=M−(t × sM)
-
7
Upper confidence limit=M+(t × sM).
For a comprehensive discussion on expression of uncertainty in relation to quantitative tests, refer to the European co-operation for Accreditation document EA-4/14.38
Qualitative tests
When reporting estimates of accuracy for a qualitative test, the measured sensitivity and specificity are not useful figures on their own, as they only relate to the specific samples tested in the validation (eg, the proportion of gold standard positives correctly identified). To apply the estimates to a wider population and to allow the validation results to be realistically compared with others, a confidence interval must be given. This is a function of the measured results and the sample size. Table 5 gives an example of the results of three experiments with different sample sizes but for which the measured sensitivities were identical. It is clear that the larger sample size of experiment C gives a much smaller confidence interval.
Such ambiguities are very common in the reporting of diagnostic accuracy.39 At best, they can preclude any realistic comparison of different validation experiments; at worst, they can provide misleading diagnostic information with potentially serious consequences for patient care.
To improve this situation, estimates of accuracy should always be based on valid calculations and be given with appropriate confidence intervals; for example, the lower and upper limits between which there is 95% confidence that the sensitivity/specificity for the wider population falls.
In cases in which the measured sensitivity and/or specificity is 100% and the sample size is ≥60, the ‘rule of 3’ (as described in the section ‘Estimating power’) reference is sufficiently accurate to determine the confidence interval. Only the lower confidence limit need be stated, as the upper limit is 100%.
It is important to note that using the ‘rule of 3’ in this context is only valid if all tested mutations are detected. In situations in which the measured diagnostic accuracy is less than 100%, more complex statistics are required to calculate the confidence interval. It is recommended that the exact method based on the binomial distribution be used, as confidence intervals near 100% need to be skewed (ie, the interval above and below the measured result is not equal) to avoid upper confidence limits ‘>100%’. A detailed description of the method, together with instructions on performing the calculations in Microsoft Excel, is available on the NIST engineering statistics handbook website.40 In all cases in which measured diagnostic accuracy is less than 100%, it is recommended to consult a competent statistician.
Conclusion
This paper has outlined the basic principles for including validation and verification in an implementation process for molecular genetic testing. We have described the different types of tests and the key components for validation, and suggested some relevant statistical approaches. The standardized validation pro forma provided in the Supplementary data can be used to guide and record validations and verifications for the purposes of quality management and accreditation. Any suggestions for additions or alterations should be addressed to the corresponding author.
References
Haddow JE, Palomaki GE : ACCE: a model process for evaluating data on emerging genetic tests; in Khoury M, Little J, Burke W (eds): Human Genome Epidemiology: A Scientific Foundation for Using Genetic Information to Improve Health and Prevent Disease. Oxford University Press: New York, 2003, pp 217–233.
International Organization for Standardization: Medical laboratories – Particular requirements for quality and competence. ISO 15189: 2007.
International Organization for Standardization: General requirements for the competence of testing and calibration laboratories. ISO/IEC 17025: 2005.
EuroGentest, EU Contract No.: FP6-512148, http://www.eurogentest.org.
Jennings L, Van Deerlin VM, Gulley ML : Recommended principles and practices for validating clinical molecular pathology tests. Arch Pathol Lab Med 2009; 133: 743–755.
Prence EM : A practical guide for the validation of genetic tests. Genet Test 1999; 3: 201–205.
Standards Unit, Evaluations and Standards Laboratory. QSOP23 – Commercial and in-house diagnostic tests: evaluations and validation. http://www.hpa-standardmethods.org.uk/documents/qsop/pdf/qsop23.pdf (accessed May 2010).
Eurachem: The fitness for purpose of analytical methods a laboratory guide to method validation and related topics http://www.eurachem.org/guides/valid.pdf (accessed May 2010).
Zhang Z, Schwartz S, Wagner L, Miller W : A greedy algorithm for aligning DNA sequences. J Comput Biol 2000; 7: 203–214.
Amos Wilson J, Zoccoli MA, Jacobson JW et al: Validation and Verification of Qualitative Multiplex Nucleic Acid Assays, Approved Guideline. Wayne, PA: Clinical Laboratory and Standards Institute, 2008 (CLSI document MM17).
Longo MC, Berninger MS, Hartley JL : Use of uracil DNA glycosylase to control carryover contamination in the polymerase chain reaction. Gene 1990; 93: 125–128.
Pruvost M, Grange T, Geigl EM : Minimizing DNA contamination by using UNG-coupled quantitative real-time PCR on degraded DNA samples: application to ancient DNA studies. Biotechniques 2005; 38: 569–575.
Hartley JL, Rashtchian A : Dealing with contamination: enzymatic control of carryover contamination in PCR. Genome Res 1993; 3: S10–S14.
In-Vitro Directives Division Directive 98/79/EC, http://www.mdss.com/IVDD/IVDD_Directive.htm (accessed May 2010).
MHRA Bulletin 20: Conformity Assessment Procedures under the In Vitro Diagnostic Medical Devices Directive 98/79/EC, http://www.mhra.gov.uk/Howweregulate/Devices/InVitroDiagnosticnMedicalDevicesDirective/Conformityassessment/index.htm (accessed May 2010).
Camajova J, Berwouts S, Matthijs G, Macek Jr M, Dequeker E : Variability in the use of CE-marked assays for in vitro diagnostics of CFTR gene mutations in European genetic testing laboratories. Eur J Hum Genet 2009; 17: 537–540.
Bossuyt PM, Reitsma JB, Bruns DE et al: Towards complete and accurate reporting of studies of diagnostic accuracy: the STARD initiative. Standards for Reporting of Diagnostic Accuracy. Clin Chem 2003; 49: 1–6.
Hauck WW, Kock W, Abernethy D, Williams RL : Making sense of trueness, precision, accuracy and uncertainty. Pharmacopeial Forum 2008; 34: 838–842.
Menditto A, Patriarca M, Magnusson B : Understanding the meaning of accuracy, trueness and precision. Accred Qual Assur 2007; 12: 45–47.
International Organization for Standardization: Statistics – Vocabulary and Symbols – Part 1: General Statistical Terms and Terms Used in Probability. ISO 3534-1:2006.
International Vocabulary of Metrology: Basic and General Concepts and Associated Terms VIM, 3rd edn. ISO/IEC Guide 99:2007.
International Organization for Standardization: Laboratory medicine-requirements for reference measurement laboratories. ISO 15195: 2003.
Sivaganesan M, Seifring S, Varma M, Haugland RA, Shanks OC : A Bayesian method for calculating real-time quantitative PCR calibration curves using absolute plasmid DNA standards. BMC Bioinformatics 2008; 9: 120–131.
Tholen DW, Kroll M, Astles JR et al: Evaluation of the Linearity of Quantitative Measurement Procedures: A Statistical Approach; Approved Guideline, 2003 (NCCLS Document EP6-A).
Tholen DW, Kondratovich M, Armbruster DA et al: Protocols for Determination of Limits of Detection and Limits of Quantitation. Approved Guideline, 2004 (NCCLS document EP17-A).
COFRAC. Guide de validation de méthodes en biologie médicale, 2004 (LAB GTA 04).
Wallace A : MLPA Analysis Spreadsheets – User Guide, Manchester: National Genetics Reference Laboratory, 2006 http://www.ngrl.org.uk/Manchester/mlpapubs.html (accessed May 2010).
The European Molecular Genetics Quality Network, http://www.emqn.org/emqn/Schemes.html (accessed May 2010).
Tholen DW, Kallner A, Kennedy JW, Krouwer JS, Meier K : Evaluation of Precision Performance of Quantitative Measurement Methods; Approved Guideline, 2nd edn., 2004.
Krouwer JS, Tholen DW, Garber CC et al: Method Comparison and Bias Estimation Using Patient Samples; Approved Guideline, 2nd edn., 2002 (NCCLS document EP9-A2).
Krouwer JS, Cembrowski GS, Tholen DW : Preliminary Evaluation of Quantitative Clinical Laboratory Methods; Approved Guidelines, 3rd edn., 2006 (NCCLS document EP10-A3).
Garrett PE, Lasky FD, Meier KL User Protocol for Evaluation of Qualitative Test Performance; Approved Guideline, 2nd edn., 2008 (NCCLS document EP12-A2).
Carey RN, Anderson FP, George H et al: User Verification of Performance for Precision and Trueness; Approved Guideline, 2nd edn., 2005 (NCCLS document EP15-A2).
Maddalena A, Bale S, Das S, Grody W, Richards S : Technical standards and guidelines: molecular genetic testing for ultra-rare disorders. Genet Med 2005; 7: 571–583.
Jones SR, Carley S, Harrison M : An introduction to power and sample size estimation. Emerg Med J 2003; 20: 453–458.
Hanley JA, Lippman-Hand A : If nothing goes wrong, is everything all right? Interpreting zero numerators. JAMA 1983; 249: 1743–1745.
Rümke CL : Uncertainty as to the acceptance or rejection of the presence of an effect in relation to the number of observations in an experiment. Triangle 1968; 8: 284–289.
EA guidelines on the expression of uncertainty in quantitative testing, 2003 (document reference EA 4/16).
Harper R, Reeves B : Reporting of precision of estimates for diagnostic accuracy: a review. BMJ 1999; 318: 1322–1323.
NIST/SEMATECH e-Handbook of Statistical Methods http://www.itl.nist.gov/div898/handbook/prc/section2/prc241.htm (accessed May 2010).
Acknowledgements
We thank Patrick M Bossuyt (Department of Clinical Epidemiology, Biostatistics & Bioinformatics, Amsterdam, The Netherlands), Ian Mann (Swiss Accreditation Service) and Paul Strike (Salisbury Foundation Trust, UK) for their help and advice, and Milan Macek Jr (Charles University, Prague) for hosting the meetings. This study was funded by Eurogentest and by the UK Department of Health.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Useful web sites and documents
Statistics: http://www.fao.org/docrep/W7295E/w7295e08.htm (accessed May 2010) – Basic statistics http://davidmlane.com/hyperstat/index.html (accessed May 2010) – Comprehensive handbook of statistics http://www.itl.nist.gov/div898/handbook/index.htm (accessed May 2010) – Comprehensive handbook of statistics http://en.wikipedia.org/wiki/Statistics (accessed May 2010) – Useful descriptions of statistical tests http://faculty.vassar.edu/lowry/vsmap2.html (accessed May 2010) – A useful resource for statistical tests (with calculators). Validation procedures: Clinical laboratory standards institute (CLSI) publish a range of protocols and standards that may be useful for diagnostic genetic applications. http://www.clsi.org/Source/Custom/Currentdocs.cfm?Section=Current_Versions_of_CLSI_Documents (accessed May 2010).
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
APPENDIX
APPENDIX
EuroGentest (Unit 1) working group: Mike Zoccoli (Celera, Alameda, USA), Jana Camajova, Petra Křenková, Patricia Norambuena, Alexandra Stambergova and Milan Macek (Charles University Prague, 2nd Faculty of Medicine and University Hospital Motol, Department of Biology and Medical Genetics; Prague, Czech Republic), Isabelle Moix (Molecular Diagnostic Laboratory, Service of Genetic Medicine, Geneva University Hospitals, Switzerland), Patrick M. Bossuyt (Department of Clinical Epidemiology, Biostatistics & Bioinformatics, Amsterdam, The Netherlands), Els Voorhoeve and Bert Bakker (Department of Human and Clinical Genetics, LUMC St Radboud, Leiden, The Netherlands), Sarah Berwouts, Tom Janssens and Ivo Salden (KU Leuven, Centre for Human Genetics, Leuven, Belgium), Trudi McDevitt and David Barton (National Centre for Medical Genetics, Dublin, Ireland), Jean Amos-Wilson (Sequenom, San Diego, USA), Ian Mann (Swiss Accreditation Service SAS, Lausanne, Switzerland), Hans Scheffer (University Medical Centre Nijmegen, Nijmegen, The Netherlands).
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Mattocks, C., Morris, M., Matthijs, G. et al. A standardized framework for the validation and verification of clinical molecular genetic tests. Eur J Hum Genet 18, 1276–1288 (2010). https://doi.org/10.1038/ejhg.2010.101
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.101
Keywords
This article is cited by
-
EMQN best practice guidelines for genetic testing in hereditary breast and ovarian cancer
European Journal of Human Genetics (2024)
-
Analytical validation and initial clinical testing of quantitative microscopic evaluation for PD-L1 and HLA I expression on circulating tumor cells from patients with non-small cell lung cancer
Biomarker Research (2022)
-
Evaluation of the Suitability of RNAscope as a Technique to Measure Gene Expression in Clinical Diagnostics: A Systematic Review
Molecular Diagnosis & Therapy (2022)
-
EMQN best practice guidelines for molecular genetic testing and reporting of 21-hydroxylase deficiency
European Journal of Human Genetics (2020)
-
Comparison of Mendeliome exome capture kits for use in clinical diagnostics
Scientific Reports (2020)