Abstract
Hereditary hearing loss (HL) is a very heterogeneous trait, with 46 gene identifications for non-syndromic HL. Mutations in GJB2 cause up to half of all cases of severe-to-profound congenital autosomal recessive non-syndromic HL, with 35delG being the most frequent mutation in Caucasians. Although a genotype–phenotype correlation has been established for most GJB2 genotypes, the HL of 35delG homozygous patients is mild to profound. We hypothesise that this phenotypic variability is at least partly caused by the influence of modifier genes. By performing a whole-genome association (WGA) study on 35delG homozygotes, we sought to identify modifier genes. The association study was performed by comparing the genotypes of mild/moderate cases and profound cases. The first analysis included a pooling-based WGA study of a first set of 255 samples by using both the Illumina 550K and Affymetrix 500K chips. This analysis resulted in a ranking of all analysed single-nucleotide polymorphisms (SNPs) according to their P-values. The top 250 most significantly associated SNPs were genotyped individually in the same sample set. All 192 SNPs that still had significant P-values were genotyped in a second independent set of 297 samples for replication. The significant P-values were replicated in nine SNPs, with combined P-values between 3 × 10−3 and 1 × 10−4. This study suggests that the phenotypic variability in 35delG homozygous patients cannot be explained by the effect of one major modifier gene. Significantly associated SNPs may reflect a small modifying effect on the phenotype. Increasing the power of the study will be of greatest importance to confirm these results.
Similar content being viewed by others
Introduction
Hearing loss (HL) is the most common birth defect in industrialised countries and also the most frequent sensorineural disorder, affecting about 1 in 500 newborns.1 The causes of HL are diverse, but a genetic defect can be found in at least 50% of the cases. The identification of deafness genes for hereditary HL has evolved very quickly during the past 10 years, with 46 different genes known to cause non-syndromic hearing impairment (Hereditary Hearing Loss Homepage: http://webh01.ua.ac.be/hhh/). This large number of deafness genes makes HL a very heterogeneous trait, with most genes being causative in only a small percentage of patients. However, the GJB2 gene (gap junction protein β-2 encoding for connexin 26) is responsible for up to 50% of the cases with autosomal recessive non-syndromic sensorineural HL (ARNSHL).2
Mutations in GJB2 are the most frequent cause of ARNSHL, and according to the connexin deafness homepage, more than 100 different mutations have been identified (http://davinci.crg.es/deafness/index.php). GJB2 mutations also cause autosomal dominant HL, often in combination with skin disorders. Although most mutations are not very frequent, 35delG has been found in more than 50% of Caucasians with GJB2 mutations. Carrier frequencies in Europe range from 0.5 to 3.5%, with the highest frequencies found in the Mediterranean region.3 A large multicentre study has established genotype–phenotype correlations for connexin 26-related HL, based on audiometric data of 1531 patients from 26 laboratories.4 All patients suffered from non-syndromic mild-to-profound HL because of biallelic GJB2 mutations. The study clearly demonstrated that inactivating mutations of GJB2 cause a more severe phenotype than non-inactivating mutations. In addition, mutations M34T and V37I were shown to cause a mild HL when in compound heterozygosity with an inactivating mutation. Although specific genotype–phenotype correlations were established for most genotypes, the correlation could explain only a part of the phenotypic variability. For most genotypes, a certain degree of variation was still present that was mostly pronounced within the group of patients with a homozygous 35delG genotype. Both inter- and intrafamilial variations were observed, with the HL ranging from mild or moderate (least often) to severe and profound (most frequent). This phenotypic variation may be explained by the influence of environmental factors and/or by the influence of modifier genes.
Several strategies can be used to identify the modifier genes for human disorders. In mice, the identification can be performed by crossing parental inbred strains that carry the disease-causing mutation and exhibit a difference in phenotype. Two examples are the modifier gene Mdfw in deafwaddler mice with an Atp2b2 mutation5 and the modifier gene Moth1 in the Tubby mouse.6 There is, however, a limitation in available inbred strains, and some mouse mutants are not an accurate model for the human phenotype. In addition, Gjb2 knockout mice are embryonically lethal.7 Using human patients, two strategies can be adopted. The first approach uses families and linkage analysis. A whole-genome search is performed to investigate the phenotypic difference among patients from multiplex families with sufficient intrafamilial variation. In this way, the DFNM1 locus was identified as a modifier of DFNB26-linked HL and the 1555A>G mutation in MTRNR1 was shown to be modified by three different genes: MTO1, TFB1M and GTPBP3.8 The second strategy uses association studies in unrelated patients. Here, the correlation between the phenotypic variation and genetic markers is analysed statistically. The development of single-nucleotide polymorphism (SNP) chips, by Illumina and Affymetrix, containing hundreds of thousands of SNPs covering the complete genome makes these whole-genome association (WGA) studies possible. Associated genetic factors involved in complex diseases are identified in a hypothesis-free way. However, WGA studies for individual samples are costly. An alternative strategy is to use pooled genomic DNA. The research institute TGen (Translational Genomics Research Institute, Phoenix, AZ, USA) has developed a pooling-based WGA approach that is of high quality and has been proven successful in the identification of important associations for complex traits.9, 10, 11, 12, 13 An important note is that pooling-based WGA studies are effective in identifying only common variations with a large effect on the disease.
In this report, we sought to identify modifier genes for connexin 26-related hearing impairment by performing a WGA with pooled DNA samples in a set of 35delG homozygous patients. The population is suitable for identifying modifier genes, because it is a genetically homogeneous sample set, eliminating the effect of the mutation on the phenotype as a confounding factor. The 35delG mutation is very common in Caucasians, making it possible to collect a relatively large sample set. Characterising modifier genes for GJB2 is important for several reasons. First, it could lead to improved diagnosis and genetic counselling by facilitating a more accurate prediction of the phenotype based on the genotype of both GJB2 and the modifier gene. Second, it could provide a better definition of the auditory pathways in which the primary mutation functions and gain potentially novel insights into the molecular pathology of HL. Finally, knowledge of modifiers may allow the development of new therapies, making GJB2-related deafness more amenable to treatment.
Materials and methods
Sample and data collection
The study was approved by the ethical committee of the University of Antwerp and by the ethical committees of all research institutes collaborating in this study. Informed consent was obtained by each research centre from every participant or from the parents of minors. For every patient, a series of data were collected, including general data (eg, date of birth, gender, ethnicity and the presence of additional clinical features apart from the HL) and audiometric data (age of onset, date of audiometry, the audiometric technique used and hearing thresholds). In addition, a DNA sample of good quality of the 35delG homozygous patient was collected. These data were used to select the appropriate samples. In total, 25 centres from 14 different countries across Europe and North America collaborated in this study. This strategy resulted in the collection of 1277 unrelated samples with a clear excess of patients with profound HL (Figure 1). The sample set consisted solely of unrelated samples to avoid bias. Names were not used to identify any of the samples to guarantee anonymity of the patients.
Audiometric data were obtained by the use of different techniques. In most cases, hearing levels were obtained by pure tone audiometry (PTA) by making measurements at frequencies ranging from 0.125 to 8 kHz. A few centres also used other techniques apart from PTA, in most cases to obtain hearing thresholds in very young children. On the one hand, electrical responses to sound were measured by the use of automated brainstem response, steady state-evoked potentials, auditory steady-state response and electrocochleogram. On the other hand and in a minority of cases, two behavioural tests, visual reinforcement audiometry and conditioned orientation reflex were used for measuring the hearing thresholds.
To verify the 35delG genotype in all samples obtained, the Snapshot (Applied Biosystems) detection method was used according to the manufacturer's instructions. The samples were analysed with an ABI PRISM 3100 Genetic analyser (Applied Biosystems). Samples with an unclear Snapshot result were analysed by DNA sequencing to determine the correct genotype.
Study design
The identification of modifier genes was performed using a pooling-based WGA study in a 35delG homozygous patient cohort.10 The PTA0.5,1,2 kHz was the variable under investigation and was used to create a ‘case’ and ‘control’ group of patients, using only the two extremes of the PTA spectrum. As the sample set consisted of patients with 14 different nationalities, we chose to match the samples for ethnicity, based on the centre where the samples originated. Within each ethnicity, patients were divided into a mild/moderate group (the ‘case’ group) and a profound group (the ‘control’ group), based upon their PTA0.5,1,2 kHz. To create the mild/moderate group, all mild and moderate cases with a PTA0.5,1,2 kHz ≤70 dB within each ethnicity were selected. To create the profound group, each sample of the mild/moderate group was matched to two samples from the profound group with a PTA0.5,1,2 kHz ≥100 dB from the same ethnicity. Using this approach, every sample from the mild/moderate group had two ethnically matched samples from the profound group, making the profound group twice as large as the mild/moderate group.
The WGA was designed in three stages: (1) pooling-based WGA on mild/moderate and profound pools from the first sample set, analysed on two SNP chips; (2) individual genotyping of the top 250 most significantly associated SNPs in the same sample set; and (3) individual genotyping of the significant SNPs in an independent replication population.
Creating DNA pools for WGA
The WGA study on pooled DNA was performed in an initial set of 255 samples. Before quantification, all DNA samples were loaded on a 1.5% agarose gel to exclude degraded samples from the pooling analysis. Genomic DNA concentrations of all samples were measured in triplicate with the QuantiT PicoGreen dsDNA Assay Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. The median concentration was calculated for each sample and was used to determine the volume of DNA needed in the DNA pool to obtain equivalent molar amounts. Eighty-five samples from the mild/moderate group were pooled, as well as 170 samples from the profound group. To control for pipetting errors, the pools were created in quadruplicate to have microarray technical replicates, providing eight individual ethnically matched pools. Through these replicates, variance arising from pooled allelotyping can be measured and a quality control can be provided to eliminate poorly performing SNPs and failed assays.
Genotyping with Illumina and Affymetrix chips
The WGA analysis of the first sample set of 35delG homozygous samples was performed using two different SNP chips: the Sentrix®humanhap550 genotyping beadchip (Illumina, San Diego, CA, USA), containing 555 000 tag SNPs and the Affymetrix 500K chip (Affymetrix, Santa Clara, CA, USA) containing 500 000 SNPs with an average distance of 5.8 kb. For both SNP chips, the pools were assayed in four technical replicates following the protocols for individual genotyping for both platforms. Combining the two genotyping platforms leaves only a few gaps in the overall genome-wide SNP coverage, reducing the number of genomic regions that are not analysed.10
The first step in the data analysis was scoring and ranking the SNPs from both Illumina and Affymetrix platforms separately, based on GenePool silhouette scores.10 The top 5000 SNPs of both platforms were taken along for calculation of allele frequencies. The allele frequencies were inferred from the pool data using k-correction factors and, subsequently, t-test P-values were calculated. The SNPs were then sorted according to the allele frequency-based P-values. The k-correction method was used to estimate the pooling accuracy as described by Craig et al.9 SNPs with a k-correction factor below 0.5, above 2 and exactly 1 were not analysed further because their estimation of allele frequencies was deemed not reliable enough. In addition, SNPs with minor allele frequencies less than 3% according to Hapmap were excluded, as well as the top 1% most variable SNPs. A final selection of 250 SNPs from both platforms was taken forward for individual genotyping. No multiple testing correction was performed to make the SNP selection.
Individual genotyping
The 250 selected SNPs from both platforms were individually genotyped in the same sample set by Kbioscience (Hoddesdon, UK), using the KASPar SNP genotyping chemistry. The data analysis and quality control were performed in SAS (SAS 9.1.2; SAS Institute Inc., Cary, NC, USA). First, several quality-checking steps were performed. Samples with more than 15% missing genotyping data were excluded. Next, the percentage of missing data per SNP was calculated and SNPs with more than 3% missing genotypes were excluded. Hidden relatives were removed from further analysis with Graphical Relationship Representation.14 Checkhet was used to look for genetically abnormal samples per ethnicity, based on genotypes of multiple SNPs.15 Finally, the Hardy–Weinberg equilibrium was checked per ethnicity, and SNPs with a P-value below 0.001 were also excluded. For the statistical analysis, P-values and allelic odds ratios were calculated for each SNP with the Armitage's Trend Test by the use of the statistical software program R (www.r-project.org). This trend test uses an additive model that has been proven successful for the detection of additive and dominant disease susceptibility loci.16
In a next step, individual genotyping of the significant SNPs from the previous genotyping was performed in an independent replication population to confirm their significance. A replication set of 297 samples was collected, containing 99 mild/moderate samples and 198 profound samples. The same quality control and statistical analysis was performed as for the individual genotyping in the first sample set. A combined P-value for SNPs that were significant in both sample sets was obtained with the Mantel–Haenszel test. This test measures the strength of association by estimating the common odds ratio. A significant interaction P-value indicates a different effect of the genotype on the phenotype in the two sample sets. If the interaction term is not significant, a common odds ratio and a combined P-value can be calculated using Mantel–Haenszel statistics.
For genes that were in linkage disequilibrium (LD) with a significantly associated SNP, their expression in the cochlea was checked on the basis of the Morton human foetal cochlear EST database (http://www.brighamandwomens.org/bwh_hearing/human-cochlear-ests.aspx) and the Unigene database (http://www.ncbi.nlm.nih.gov/).
Results
Pooling-based WGA
The pooling-based WGA study was performed on a first set of 255 samples (85 mild/moderate samples and 170 profound samples) on both the Illumina and Affymetrix platforms. The analysis of the genotyping data resulted in a ranking of SNPs for both platforms separately, according to the allele frequency-based P-values. For each SNP in the top 250, its position was evaluated in relation to the neighbouring genes. Only SNPs in proximity to a gene were selected. SNPs that were not in LD with any of the genes in a region of 500 kb upstream and downstream of the gene were excluded. If two SNPs were in complete LD with each other, only one of them was included in the selection. As the Illumina chip provides higher quality data than the Affymetrix chip with regard to variance measurements, more Illumina SNPs were taken along.17 In this way, 155 Illumina SNPs and 95 Affymetrix SNPs were selected for further analysis.
The 250 selected SNPs were individually genotyped in the same sample set. Quality control resulted in the exclusion of 21 samples and 14 SNPs. The Armitage's Trend Test was performed for 236 SNPs in 234 35delG homozygous samples. P-values ranged between 1.48.10−6 and 0.68. One hundred and ninety-eight SNPs had P-values below 0.05 and were subsequently genotyped in the replication set.
The replication population was collected in the same way as the first sample set. Genotyping analysis was performed for those 198 SNPs that were significant in the first set, in 297 samples (99 mild/moderate samples and 198 profound samples). The quality check resulted in the exclusion of 27 samples and six SNPs. Subsequently, 192 SNPs were statistically analysed by the Armitage's Trend Test in 270 samples. Twelve SNPs out of 192 still showed significant P-values in the replication population and are listed in Table 1. The Mantel–Haenszel test was used to calculate the interaction term and, if allowed, the combined P-values and odds ratios for the complete sample set. Three SNPs had a significant interaction P-value and the odds ratios of both independent sample sets indicated an opposite effect. The remaining nine SNPs all had the same effect on the phenotype and had combined P-values between 1 × 10−4 and 3 × 10−3 and combined odds ratios between 0.38 and 1.88 (Table 1).
Candidate genes
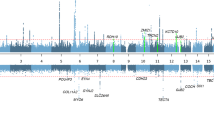
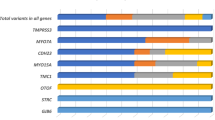
Eleven genes were found to be in LD with the significantly associated SNPs in both populations. SNP rs2215128 was in LD with five different genes and two of them, RPS23 and ATG10, were expressed in the cochlea according to the Morton human foetal cochlear EST database and/or Unigene. All other genes in LD with one of the nine SNPs were not expressed in the cochlea (Table 2).
Discussion
In this study, we sought to identify modifier genes for connexin 26-related hearing impairment in a set of 1277 35delG homozygous patients from 14 countries. As they share the same homozygous mutation, phenotypic variation on the basis of the mutation is already excluded. Figure 1 shows the excess of profound cases compared with mild and moderate cases. As the distribution of samples in the different HL categories is not normally distributed, we chose not to study the PTA0.5,1,2 kHz as a quantitative trait. Instead, we chose to work under a case–control paradigm by selecting the two extreme spectra of the curve. However, as a consequence, we were restricted in the total number of cases for analysis, given the limited number of mild/moderate cases. We had to compromise between, on the one hand, having a large enough power by analysing enough samples and, on the other hand, selecting mild/moderate cases and profound cases from the two spectra for which the PTA0.5,1,2 kHz values showed a difference that was large enough. Therefore, we chose cutoff values of 70 and 100 dB for the mild/moderate and profound groups, respectively.
To pick up modifier genes for connexin 26, we chose to perform a WGA study as the genetic pathways in which connexin 26 functions are only partly understood, making it difficult to select candidate genes. Pooled WGA studies are a very good and cost-effective alternative to detect common variants with a large effect on the phenotype. Finding true associations can only be achieved when a number of requirements are fulfilled. A number of parameters need to be of sufficient size, including the allele frequency of the causal variant, the LD between the causal variant and the typed SNPs, the odds ratio and the sample size. In addition, several extra factors need to be taken into account when pooling is used. We tried to solve most of these issues as discussed by Pearson et al.10 For example, the DNA concentrations were measured in triplicate, degraded DNA samples were excluded and the pools were constructed in quadruplicate. Population stratification and admixture have largely been excluded by matching all mild/moderate and profound samples per ethnicity. As pooling-based WGA studies are much more affordable, we were able to use both the Illumina 550K and the Affymetrix 500K platforms, typing about 800 000 different SNPs without leaving large gaps in the genome. A quality control for the accuracy of the pooling phase in pooling-based WGA studies is the number of significant associations that still remain after individual genotyping of the top 250 SNPs of the pooled analysis. In our study, 198 out of 236 successfully genotyped SNPs had a P-value below 0.05 that corresponds to 84%. This is a high percentage, indicating that the allele frequencies on the basis of the pooled data are reasonably accurate.
A few issues, however, remain. The major restriction of pooling-based WGAs is the ability to detect only large effects. The first reason for this is the problem to reach a very high accuracy of allele frequency measurements. As the measurement error is usually around 2%, it becomes more difficult to accurately rank the SNPs after pooling. Second, you lose the ability to compare subphenotypes of pools to directly measure genotypes and detect gene–gene interactions. The number of individuals pooled in this study was not very high, but as discussed before, our study design was chosen very carefully and major efforts were performed to collect a large amount of samples.
The results from the genome-wide association study do not show a very strong association in the population we studied. This observation may suggest that the phenotypic variability in 35delG homozygous patients cannot be explained by the effect of one major modifier gene. If this would have been the case, we should have picked up this gene in this study, despite the pooling strategy and the restricted number of samples we have. The SNPs that were significantly associated in both populations tested may have a small modifying effect. For two cochlear expressed genes, RPS23 and ATG10, an associated SNP was found in LD.
RPS23 encodes the ribosomal protein S23, which is related to Saccharomyces cerevisiae ribosomal protein S28. RPS23 belongs to the small 40S subunit of the ribosomes. Although the gene is ubiquitously expressed, the highest number of ESTs is found in the ear, and 26 ESTs of RPS23 were present in the Morton database. In addition, the gene was found in a human vestibular cDNA library.18 The expression in both the cochlea and the vestibular system might indicate a special role for RPS23 in the auditory and vestibular function. No direct link between RPS23 and GJB2 could be found in the literature.
ATG10 encodes the autophagy-related 10 homologue of the yeast S. cerevisiae. Its protein ATG10 is an E2-like enzyme involved in autophagy, a cellular mechanism for bulk degradation of cellular proteins and organelles in lysosomes.19 Although many E2 enzymes have been identified in eukaryotes, including the ubiquitin-conjugating enzyme family (Ubc), ATG10 does not show any homology to other known E2 enzymes.20 The gene is not present in the Morton database, but according to the Unigene expression profile, it is expressed in the ear. No known function in the hearing process has been reported for ATG10.
The other candidate genes in LD with significantly associated SNPs can also not be ruled out as modifier genes. Their lack of expression in the cochlea according to the Morton database and Unigene does not completely rule out their presence in the ear. SNPs that are not in immediate LD with a gene may also have a modifying effect, for example, by being located in cis-acting elements, which control the expression of more distant genes. Libioulle et al21 found a possible association between Crohn disease and a SNP in a gene desert. This SNP is located 270 kb from the closest gene PTGER4 and was found to regulate the expression level of this gene, suggesting its involvement in Crohn disease.
In the future, there are several options to extend this study. On the one hand, RPS23 and ATG10 could be good candidate modifiers with a smaller effect on the phenotype as they are expressed in the human cochlea. Finemapping of the region around rs2215128 should be the first step in confirming the association. Further on, functional research may be performed to elucidate the specific role of RPS23 and ATG10 in the hearing process and link them to connexin 26. It may also be interesting to look for cis-acting elements in the regions around the significantly associated SNPs to identify effects on genes at a larger distance. In this regard, it may also be of interest to individually genotype SNPs from the top 250 of the pooling experiment, which are not in LD with a nearby gene. One of the most important aims should be to increase the power of the study. This could mainly be performed by collecting additional samples. The current sample set was collected by 25 centres across Europe and North America. We are convinced that it should be possible to collect additional samples from more centres, as diagnostic testing of GJB2 is widespread and 35delG homozygous patients are very frequently picked up. We hope to find additional contributors in the future to extend this study. Another way to increase the power is by genotyping all collected samples on individual SNP chips. This strategy has the major advantage of having separate and accurate genotyping data for all samples and gives the opportunity to look for gene–gene interactions. These interactions might be of great importance for this study. As the results of this study suggest that no major modifier gene is present, we assume that the phenotypic variation will be caused by a smaller effect of different interacting genes.
References
Morton CC, Nance WE : Newborn hearing screening – a silent revolution. N Engl J Med 2006; 354: 2151–2164.
Kenneson A, Van Naarden Braun K, Boyle C : GJB2 (connexin 26) variants and nonsyndromic sensorineural hearing loss: a HuGE review. Genet Med 2002; 4: 258–274.
Gasparini P, Rabionet R, Barbujani G et al: High carrier frequency of the 35delG deafness mutation in European populations. Genetic analysis consortium of GJB2 35delG. Eur J Hum Genet 2000; 8: 19–23.
Snoeckx RL, Huygen PL, Feldmann D et al: GJB2 mutations and degree of hearing loss: a multicenter study. Am J Hum Genet 2005; 77: 945–957.
Bryda EC, Kim HJ, Legare ME, Frankel WN, Noben-Trauth K : High-resolution genetic and physical mapping of modifier-of-deafwaddler (mdfw) and Waltzer (Cdh23v). Genomics 2001; 73: 338–342.
Ikeda A, Zheng QY, Zuberi AR, Johnson KR, Naggert JK, Nishina PM : Microtubule-associated protein 1A is a modifier of tubby hearing (moth1). Nat Genet 2002; 30: 401–405.
Gabriel HD, Jung D, Butzler C et al: Transplacental uptake of glucose is decreased in embryonic lethal connexin26-deficient mice. J Cell Biol 1998; 140: 1453–1461.
Bykhovskaya Y, Mengesha E, Wang D et al: Phenotype of non-syndromic deafness associated with the mitochondrial A1555G mutation is modulated by mitochondrial RNA modifying enzymes MTO1 and GTPBP3. Mol Genet Metab 2004; 83: 199–206.
Craig DW, Huentelman MJ, Hu-Lince D et al: Identification of disease causing loci using an array-based genotyping approach on pooled DNA. BMC Genomics 2005; 6: 138.
Pearson JV, Huentelman MJ, Halperin RF et al: Identification of the genetic basis for complex disorders by use of pooling-based genomewide single-nucleotide-polymorphism association studies. Am J Hum Genet 2007; 80: 126–139.
Steer S, Abkevich V, Gutin A et al: Genomic DNA pooling for whole-genome association scans in complex disease: empirical demonstration of efficacy in rheumatoid arthritis. Genes Immun 2007; 8: 57–68.
Shifman S, Bhomra A, Smiley S et al: A whole genome association study of neuroticism using DNA pooling. Mol Psychiatry 2007; 13: 302–312.
Dunckley T, Huentelman MJ, Craig DW et al: Whole-genome analysis of sporadic amyotrophic lateral sclerosis. N Engl J Med 2007; 357: 775–788.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR : GRR: graphical representation of relationship errors. Bioinformatics 2001; 17: 742–743.
Curtis D, North BV, Gurling HM, Blaveri E, Sham PC : A quick and simple method for detecting subjects with abnormal genetic background in case–control samples. Ann Hum Genet 2002; 66: 235–244.
Lettre G, Lange C, Hirschhorn JN : Genetic model testing and statistical power in population-based association studies of quantitative traits. Genet Epidemiol 2007; 31: 358–362.
Macgregor S, Zhao ZZ, Henders A, Nicholas MG, Montgomery GW, Visscher PM : Highly cost-efficient genome-wide association studies using DNA pools and dense SNP arrays. Nucleic Acids Res 2008; 36: e35.
Abe S, Koyama K, Usami S, Nakamura Y : Construction and characterization of a vestibular-specific cDNA library using T7-based RNA amplification. J Hum Genet 2003; 48: 142–149.
Nemoto T, Tanida I, Tanida-Miyake E et al: The mouse APG10 homologue, an E2-like enzyme for Apg12p conjugation, facilitates MAP-LC3 modification. J Biol Chem 2003; 278: 39517–39526.
Ohsumi Y : Molecular dissection of autophagy: two ubiquitin-like systems. Nat Rev Mol Cell Biol 2001; 2: 211–216.
Libioulle C, Louis E, Hansoul S et al: Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS Genet 2007; 3: e58.
Acknowledgements
We are very grateful to all patients who cooperated in this study. This study was supported in part by grants from EUROHEAR (LSHG-CT-2004-512063), the Fund for Scientific Research Flanders (FWO-F, Grant G.0138.07), the MNiSW Grant 0711/P01/2006/31, the Oticon Foundation (Denmark, MBP) and the National Institutes of Health (NIDCD DC02842 to RJHS). Nele Hilgert is a fellow of the Fund for Scientific Research Flanders (FWO-F).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hilgert, N., Huentelman, M., Thorburn, A. et al. Phenotypic variability of patients homozygous for the GJB2 mutation 35delG cannot be explained by the influence of one major modifier gene. Eur J Hum Genet 17, 517–524 (2009). https://doi.org/10.1038/ejhg.2008.201
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.201
Keywords
This article is cited by
-
Beyond Mendelian Inheritance: Genetic Buffering and Phenotype Variability
Phenomics (2022)
-
Characterization of GJB2 cis-regulatory elements in the DFNB1 locus
Human Genetics (2019)
-
Connexin 26 (GJB2) mutation in an Argentinean patient with keratitis-ichthyosis-deafness (KID) syndrome: a case report
BMC Medical Genetics (2016)
-
Mutations of GJB2 encoding connexin 26 contribute to non-syndromic moderate and severe hearing loss in Pakistan
European Archives of Oto-Rhino-Laryngology (2015)
-
The promoter mutation c.−259C>T (−3438C>T) is not a common cause of non-syndromic hearing impairment in Austria
European Archives of Oto-Rhino-Laryngology (2015)