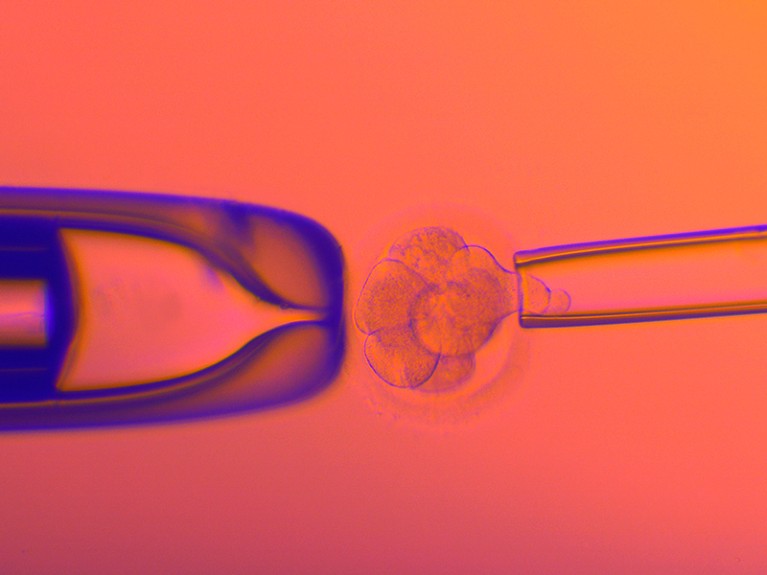
A cell is plucked from a human embryo created using in vitro fertilization so that it can be screened for genetic disorders.Credit: Pascal Goetgheluck/SPL
The emergence of companies that offer prospective parents complex genetic tests on embryos ahead of in vitro fertilization (IVF) has alarmed geneticists and bioethicists alike. The companies claim to be able to predict the risk of many common diseases — including those influenced by dozens or even hundreds of genes. People undergoing IVF are then offered the chance to select an embryo with a perceived low relative risk of developing such diseases.
Read the paper: Whole-genome risk prediction of common diseases in human preimplantation embryos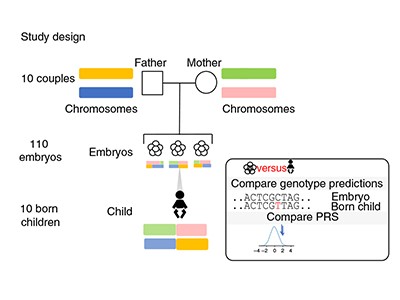
Researchers are right to be concerned. The selection of embryos on the basis of these predictions is not yet supported by science. Moreover, the societal implications of using complex genetic tests to choose embryos has not yet been fully considered. Some scientists are completely opposed to the practice, whereas others recognize that, as more data accrue, there might be benefits, but realize that it must be carefully regulated. A study published in Nature Medicine1 on 21 March that explains some of the methodology behind the determination of what are called polygenic risk scores draws attention to the practice — but does not allay scientists’ fears.
Some health authorities around the world do regulate the use of simple genetic testing alongside IVF, but many do not. The aim of these tests is to reduce the chances of a parent transmitting an inherited disease to their unborn child. This is typically done in the case of rare, devastating conditions caused by mutations in a single gene. In the United Kingdom, for example, tests have been approved by the Human Fertilisation and Embryology Authority for more than 600 inherited disorders, including Tay–Sachs disease and breast cancers caused by mutations in the genes BRCA1 and BRCA2.
But the most common diseases, such as type 2 diabetes, are associated with mutations not in a single gene, but in many — potentially even thousands. To understand genetic contributions to such conditions, researchers have been analysing DNA sequences from many thousands of people with the disease, and comparing them with the DNA of people who do not have it, looking for genetic variants that are associated with a higher risk of developing the condition. This information is then converted into an overall risk score that estimates a person’s relative risk of developing a given disorder.
The ambition is that, as genetic studies continue to sample populations more broadly and more deeply, polygenic risk scores will become more refined, and could eventually be used to guide treatment and prevention strategies. But there’s a consensus that the scores are not yet ready to be used beyond research studies.
Feasibility study
In the latest study, the authors — most of whom work for companies involved in IVF or genetic testing — address the technical challenge of predicting accurate genome sequences from the small amounts of DNA available from one or two cells biopsied from an embryo. The researchers constructed sequences for more than 100 embryos by analysing hundreds of thousands of sites in the embryos’ genomes using a technique called genotyping, which requires less DNA than whole-genome sequencing does. They then combined these data with whole-genome sequences from the parents to fill in the rest of the DNA sequences and, in ten cases, compared the reconstructed genome from an embryo with the full genome sequence of the resulting born child. They found that they were able to infer the correct genome sequence at sites used to calculate polygenic risk scores for 12 conditions — including diabetes, certain types of heart disease and several forms of cancer and autoimmune disorder — with 97–99% accuracy.
Polygenic embryo testing: understated ethics, unclear utility
The authors say that this technique — which has been peer reviewed — establishes the feasibility of assessing genomic regions necessary to calculate a polygenic risk score for an embryo. But this technical capability is not the main reason for concern and debate over the use of polygenic risk scores in embryo selection for IVF.
There are many other concerns surrounding the practice. One is that the scores have been developed on the basis of genome-wide association studies that have heavily sampled DNA from people of European descent. Although efforts are under way to diversify such databases, the scores currently available are not based on an appropriately diverse subset of people. Even among white European people, polygenic risk scores are sometimes predictive only within narrow subsets of that population — potentially, in part, because of poorly understood interactions between the genetic and environmental contributions to a condition.
Furthermore, scientists do not yet fully understand how the selection of embryos with a lower relative risk for one disease might influence susceptibility to other conditions. Genetic variation can have a number of effects — a phenomenon known as pleiotropy — and a DNA sequence associated with one beneficial characteristic could also increase the risk of a detrimental one.
Many of these polygenic scores are being used to predict the risk of disorders that occur later in life, without any way of incorporating potential changes in environment that could occur over that time. A child born today will probably not experience heart disease or diabetes for decades, and there is no way of knowing what treatments or preventive steps will be available by then, or what changes in the environment might have taken place.
Potential harms
Concerns about under-represented populations and pleiotropy might be addressed with further research. But polygenic risk assessments are already being marketed directly to consumers (and not only for IVF) in some countries, including the United States and Japan. It is not clear to what extent individuals are counselled on the technique’s uncertainties and risks. Meanwhile, the scores of such assessments could be harmful. They could trigger the unnecessary destruction of viable embryos or induce women to undergo extra cycles of ovarian stimulation to collect more oocytes.
The uncertain science of preimplantation and prenatal genetic testing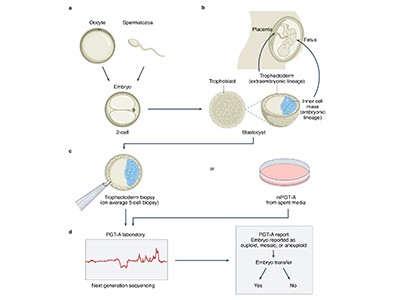
In some regions, there is little regulation of IVF and a long history of IVF technologies sweeping the market with little evidence that they improve the chances of conception or offer health benefits to children born as a result of the technology. Meanwhile, IVF is big business: the global market was about US$14 billion in 2020 and is predicted to more than double, to $34 billion, by 2028, according to Grand View Research, a market-research company based in San Francisco, California.
For now, prospective parents seeking IVF should not be offered polygenic risk scores for diseases unless they are part of rigorous clinical trials. Professional societies should make this clear to their members — as some have already done — and should publish guidelines on how to counsel participants in such trials to avoid giving them false hopes or fears about their children’s health. Genetic counsellors must be trained to do the same.
These tests demand a broader societal discussion. By nature of their complexity, polygenic risk scores also open the door to evaluating not only disease risk, but traits such as height or intelligence. At present, not enough is known about the genetic contributors to such traits to develop meaningful tests that would allow prospective parents to select embryos. But those data are on the way and the technology is going to move quickly — it is well past time to discuss how far it should go.

 Read the paper: Whole-genome risk prediction of common diseases in human preimplantation embryos
Read the paper: Whole-genome risk prediction of common diseases in human preimplantation embryos
 Polygenic embryo testing: understated ethics, unclear utility
Polygenic embryo testing: understated ethics, unclear utility
 The uncertain science of preimplantation and prenatal genetic testing
The uncertain science of preimplantation and prenatal genetic testing
 Personalized profiles for disease risk must capture all facets of health
Personalized profiles for disease risk must capture all facets of health
 Improving reporting standards for polygenic scores in risk prediction studies
Improving reporting standards for polygenic scores in risk prediction studies
 Don’t ignore data from minority populations
Don’t ignore data from minority populations