- NEWS AND VIEWS
A base-pair view of interactions between genes and their enhancers
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 595, 36-37 (2021)
doi: https://doi.org/10.1038/d41586-021-01494-x
References
Hua, P. et al. Nature 595, 125–129 (2021).
The ENCODE Project Consortium et al. Nature 583, 699–710 (2020).
Lambert, S. A. et al. Cell 172, 650–665 (2018).
de Laat, W. & Duboule, D. Nature 502, 499–506 (2013).
Stadhouders, R., Filion, G. J. & Graf, T. Nature 569, 345–354 (2019).
Furlong, E. E. M. & Levine, M. Science 361, 1341–1345 (2018).
Schoenfelder, S. & Fraser, P. Nature Rev. Genet. 20, 437–455 (2019).
de Wit, E. & de Laat, W. Genes Dev. 26, 11–24 (2012).
Bonev, B. & Cavalli, G. Nature Rev. Genet. 17, 661–678 (2016).
Fudenberg, G., Abdennur, N., Imakaev, M., Goloborodko, A. & Mirny, L. A. Cold Spring Harb. Symp. Quant. Biol. 82, 45–55 (2017).
Davies, J. O. J. et al. Nature Methods 13, 74–80 (2016).
Krietenstein, N. et al. Mol. Cell 78, 554–565 (2020).
Dowen, J. M. et al. Cell 159, 374–387 (2014).
Kagey, M. H. et al. Nature 467, 430–435 (2010).
Chien, R. et al. J. Biol. Chem. 286, 17870–17878 (2011).
Liu, N. Q. et al. Nature Genet. 53, 100–109 (2021).
Competing Interests
The authors declare no competing interests.

 Read the paper: Defining genome architecture at base-pair resolution
Read the paper: Defining genome architecture at base-pair resolution
 One ring to rule them all
One ring to rule them all
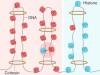 How to build a cohesive genome in 3D
How to build a cohesive genome in 3D