- NEWS AND VIEWS
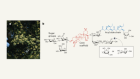
Previously unknown type of protein crosslink discovered
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 593, 343-344 (2021)
doi: https://doi.org/10.1038/d41586-021-01135-3
References
Wensien, M. et al. Nature 593, 460–464 (2021).
Mor-Cohen, R. Antioxid. Redox. Signal. 24, 16–31 (2016).
Hassan, A., Ibrahim, Y. R. & Shawky, A. M. J. Sulfur Chem. 28, 211–222 (2007).
Challis, B. C. & Butler, A. R. in The Chemistry of the Amino Group (ed. Patai, S.) Ch. 6 (Interscience, 1968).
Sawyer, D. T. Oxygen Chemistry Ch. 5–6 (Oxford Univ. Press, 1991).
Lang, P. T. et al. Protein Sci. 19, 1420–1431 (2010).

 Read the paper: A lysine–cysteine redox switch with an NOS bridge regulates enzyme function
Read the paper: A lysine–cysteine redox switch with an NOS bridge regulates enzyme function
 Enzymes can adapt to cold by wiggling regions far from their active site
Enzymes can adapt to cold by wiggling regions far from their active site
 Molecular architecture of the key precursor of thyroid hormones revealed
Molecular architecture of the key precursor of thyroid hormones revealed