- NEWS AND VIEWS
A catalogue of cancer-driving mutations in healthy tissue
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 580, 595-596 (2020)
doi: https://doi.org/10.1038/d41586-020-01081-6
References
Moore, L. et al. Nature 580, 640–646 (2020).
Bray, F. et al. CA Cancer J. Clin. 68, 394–424 (2018).
Siegel, R. L., Miller, K. D. & Jemal, A. CA Cancer J. Clin. 70, 7–30 (2020).
Joshi, A., Miller, C. Jr., Baker, S. J. & Ellenson, L. H. Am. J. Pathol. 185, 1104–1113 (2015).
Yang, H. P. et al. Br. J. Cancer 112, 925–933 (2015).
Levine, D. A. & The Cancer Genome Atlas Research Network. Nature 497, 67–73 (2013).
Sangtani, A. et al. Gynecol. Oncol. 156, 387–392 (2020).
Wang, Y. et al. Sci. Transl. Med. 10, eaap8793 (2018).
Wan, J. C. M. et al. Nature Rev. Cancer 17, 223–238 (2017).

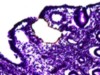 Read the paper: The mutational landscape of normal human endometrial epithelium
Read the paper: The mutational landscape of normal human endometrial epithelium
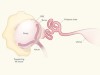 Anatomy of an ovarian cancer
Anatomy of an ovarian cancer
 Global genomics project unravels cancer’s complexity at unprecedented scale
Global genomics project unravels cancer’s complexity at unprecedented scale