Host: Shamini Bundell
Welcome back to the Nature Podcast. This week, working towards a quantum internet…
Host: Nick Howe
And the pigment posing a chemical puzzle. I’m Nick Howe.
Host: Shamini Bundell
And I’m Shamini Bundell.
[Jingle]
Interviewer: Nick Howe
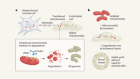
First up on the show, let’s talk about painting. Now, don’t worry, this is still the Nature Podcast. I’ve not started my spin-off art show… yet. But I do want to talk to you about a particular shade of deep blue – Prussian blue, in fact. It’s been used for centuries for all sorts of applications from art to medicine. For example, it can be used to treat people who have heavy metal poisoning by soaking up the toxins. And it’s not just Prussian blue itself that has proven useful. Prussian blue analogues, materials with similar but not identical structures, have even more uses, such as decontaminating radioactive materials, acting as catalysts and making next-generation batteries. The reason these chemicals are in such demand is because they have little gaps in their structures known as vacancies which allow ions to pass through them, like holes in a sponge. This allows them to do things like capture radioactive waste and conduct electricity. But to take advantage of this, scientists need to know where those vacancies are. That, it turns out, is quite tricky.
Interviewee: Andrew Goodwin
Our main way of determining the structure of a solid is using crystallography, but the whole premise of that technique is that the atoms are arranged in a repeating pattern, and so I’d say one of the big challenges of twenty-first century structural science is working out what to do when you don’t have that periodicity or that pattern, and the Prussian blue analogues are an extremely good example of exactly that type of material.
Interviewer: Nick Howe
This is Andrew Goodwin, a chemist from the University of Oxford. Crystallography works by crystallising materials and then pummelling them with radiation, often X-rays. This creates a diffraction pattern – a sort of shape made from light – which can then be used to work out where the atoms, and in this case the vacancies, are in materials. But in disordered materials, like the Prussian blue analogues, this diffraction pattern is hard to interpret.
Interviewee: Andrew Goodwin
When you take a crystal where things do repeat and you shine radiation on it, if there is that pattern, that periodicity, then you see a pattern that itself has a bunch of spots in it and those spots, their arrangement and their intensities, that’s the thing that we’ve learnt how to interpret. As soon as you have this disorder, in additional to all of these spots, you get this sort of streaking or beautiful patterns, and that’s much harder to interpret.
Interviewer: Nick Howe
But Andrew may have come up with a solution. This week in Nature, he’s got a new paper describing the structure of Prussian blue analogues and the positions of their vacancies. So, how did he interpret the uninterpretable?
Interviewee: Andrew Goodwin
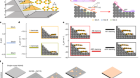
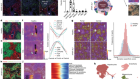
We took a bit of a guess at what chemical forces or interactions might be controlling the arrangement of the vacancies in this Prussian blue, so we just used our knowledge of the chemistry of the systems to say if I were a vacancy, how would I try and arrange myself, and we came up with two basic rules.
Interviewer: Nick Howe
The first of those two rules is that the vacancies try and distribute themselves as evenly as possible. The second rule is that depending on the chemistry of the particular material, any two vacancies will arrange themselves either next to or opposite each other around a central point.
Interviewee: Andrew Goodwin
And on the basis of those, what we can do is we can create a computer model that tries to arrange vacancies within the model, subject to our little rules, our guesses, and once we have a model in our computer of how we think the vacancies might arrange, we can calculate what this crazy scattering pattern I was talking about with all its streaks and so on, we can calculate what we think that would look like.
Interviewer: Nick Howe
Once Andrew had this model, he could grow the Prussian blue analogues as crystals and perform crystallography. He could then compare the diffraction patterns he observed with his computer models. If they matched, then he had found the structure and, importantly, where the vacancies were. So, how were the vacancies arranged?
Interviewee: Andrew Goodwin
The first thing that we can see even just for the experimental data was that they’re not random, and we can see that irrespective of whatever chemistry we used, whatever elements were there. The vacancies are not random. In a sense, that’s kind of important for the field because, really, to date, for lots of good reasons, people have been forced to kind of assume that they’re arranged randomly. If we change the chemistry – if I use different metals when I make one of these analogues – then the scattering patterns themselves changed in a way that told us that these disordered arrangements, even though they’re not random, they’re different for different Prussian blue analogues, and that’s telling us that the chemistry is going to allow us to tune the arrangement of these holes in our sponge, effectively.
Interviewer: Nick Howe
As the vacancies are what give Prussian blue analogues their interesting properties, being able to predict how they will arrange themselves depending on the chemistry of the material could allow researchers to make the best Prussian blue analogue for their particular purpose.
Interviewee: Andrew Goodwin
So, to take an example from this Prussian blue analogue paper, we can work out, from our models of the arrangements of these vacancies, which ones are going to have the best network of vacancies for taking up materials, and it turns out that not only is it that you need disorder to maximise that property, but a very specific type of this will give you the best results.
Interviewer: Nick Howe
To put this into practice may be a little more complicated though. To perform the crystallography, the analogues were grown as crystals. But when they’re used in batteries and for other applications, they’re powders. So, it’s possible that the vacancies would be positioned differently. Andrew has some evidence that the powders have similar arrangements of vacancies, but he can’t be sure. Another concern is that the model may not be representing the structure completely accurately, but Andrew thinks this is part of the nature of disordered materials.
Interviewee: Andrew Goodwin
So, we’re used to seeing papers like the structure of DNA, for example. There is a solution to that. When something is disordered, there’s no individual solution to the structure. All we can do is try and understand the properties of a representative model.
Interviewer: Nick Howe
For these disordered materials, the vacancies will be in different positions each time you look at the structure, but the characteristic pattern will remain the same. So probabilistically, the vacancies will be arranged in the same way. Andrew’s next steps are to try and see if he can indeed predict the properties of Prussian blue analogues based on their chemistry. He’s also hopeful that a similar kind of modelling approach that he used here will work on other kinds of disordered materials.
Interviewee: Andrew Goodwin
We’re working on lots of other systems where just completely different chemistries but again, coming back to this idea of can we exploit disorder of particular types to make materials that can do things you couldn’t otherwise do?
Interviewer: Nick Howe
That was Andrew Goodwin from the University of Oxford. You can find his paper, along with an accompanying News and Views article, over at nature.com.
Host: Shamini Bundell
Later on, we’ll have an update on the continuing coronavirus outbreak and hear about a technique to boost the accuracy of gene editing. Right now, though, it’s time for the Research Highlights, read this week by Dan Fox.
[Jingle]
Dan Fox
As I approach my mid-thirties, I’m happy to say that I am a morning person. I spring out of bed and get some of my best work done before lunch. But when I was a teenager, it was a different story. I was lucky to wake up before midday. In other words, I had what is known as a late chronotype. This may have explained my lacklustre performance in morning lessons, as now, new research shows that teenagers chronotypes can influence academic performance. By selecting 753 Argentinian students to start school either in the morning, afternoon or evening, researchers found that in the morning classes, early chronotype students performed better than their late chronotype peers in all subjects. This advantage disappeared in the evening lessons, though, as then, late chronotype students were the best performers. The researchers suggest that arranging school subjects around students’ chronotype might be a simple way to improve academic performance. Read the rest of that research, at a time that suits you, at Nature Human Behaviour.
[Jingle]
Dan Fox
Prehistoric creatures are often mysterious, fearsome, and in the case of one enormous rodent that lived in Brazil 10 million years ago, particularly tiny-brained. Researchers looked at two skulls of the large dog-sized rodent Neoepiblema acreensis, using CT scanners. By measuring the space inside the skulls, they estimated the volume of the brain before calculating that the 80-kilogram creature had a brain that weighed only 47 grams. This puts their encephalization quotient – a measure of the difference between expected and actual brain size – at around five times smaller than that of modern South American rodents. The researchers suggest that since South America was isolated while the hefty rodent was alive, they had few predators to outsmart, and so a large brain simply wasn’t worth the energy of maintaining it. Put your much larger brains to work reading that research at Biology Letters.
[Jingle]
Host: Shamini Bundell
Next up on the podcast, Benjamin Thompson is taking a step into the world of quantum communication.
Interviewer: Benjamin Thompson
The internet is great, right? It connects people and lets us send data around the world in a flash. It even allows us to visit amazing websites like nature.com/podcast. But the internet doesn’t just include computers. You’ve got fridges, printers, modems – you name it. Each of these are called nodes, but this is the classic internet. Now, researchers around the world are looking even further. They’re trying to build the quantum internet by connecting quantum nodes together. It’s hoped that a quantum internet would offer new computational opportunities that aren’t possible using a regular network. Here’s Tracy Northup, a quantum physicist from the University of Innsbruck in Austria.
Interviewee: Tracy Northup
One aspect is the idea that small-scale quantum computers might not be powerful enough by themselves to do meaningful computations but that by linking them together we can gain this new computational power. And also, there have been proposals to link together things like telescopes or atomic clocks, and then if we can harness quantum mechanics for this link, we can gain new sensitivity for these kinds of measurements.
Interviewer: Benjamin Thompson
Connecting distant nodes relies on a quantum effect called entanglement. Once two nodes are entangled, the state of one node affects the other, and so information can be shared between the two. One of the ways that two nodes can be entangled involves sending photons down optical fibres, but this method does have its drawbacks, explains Jian-Wei Pan from the University of Science and Technology of China.
Interviewee: Jian-Wei Pan
The quantum signal has its drawbacks of the photon loss caused by unavoidable absorption of the fibre, so then the photo signal will be weak, so then the distance of quantum communication is very much limited. So, we have to find a way how to send the photon signal over larger distances.
Interviewer: Benjamin Thompson
Because photons produced by nodes can be readily absorbed by the optical fibres that carry them, it’s hard to get entanglement over longer distances – something that will be needed for the quantum internet to become a reality. So far, the longest entanglement between two nodes using this method is 1.3 kilometres, but that might be about to change. This week, Jian-Wei and his colleagues have published a paper in Nature demonstrating entanglement of two nodes over dozens of kilometres, by combining a number of techniques. Now, these two nodes were actually in the same lab, but they were connected by fibre-optic cables to an intermediate station across town. At the heart of these nodes was a cloud of laser-cooled atoms. To entangle the nodes, each cloud was coaxed to produce a photon, which were fired along the fibre-optic cables. If these photons met in the intermediate station at precisely the right time, the nodes would become entangled together. This might sound simple, but there is a catch if you want to entangle nodes over longer distances using optical fibres. Owing to the wonders of quantum mechanics, photons don’t only exist as particles – they are also waves. And the wavelength of a photon influences how much it is absorbed by the fibres its travelling down. In this case, Jian-Wei’s cold atom clouds produced photons at an unhelpful wavelength.
Interviewee: Jian-Wei Pan
Normally, our cold atomic clouds only emit photons with a wavelength of about 800 nanometres. Then if you send a special photon to the fibre, then the photon loss is huge.
Interviewer: Benjamin Thompson
At 800 nanometres, many of the photons would be absorbed by the fibres, making entanglement difficult. One of the ways that Jian-Wei and his colleagues used to get around this involved converting the wavelengths of the photons produced by the nodes.
Interviewee: Jian-Wei Pan
We managed to convert our frequency from 795 nanometres to 1,342 nanometres. That’s one of the key technologies we have been pursuing for the last ten years.
Interviewer: Benjamin Thompson
This change in frequency put the photons firmly inside a range used for telecommunications around the world. Photons at this frequency are much less likely to be absorbed by the fibres, making it a much more efficient way of establishing entanglement. Overall, Jian-Wei and colleagues demonstrated they were able to get entanglement between two nodes to occur at over 50 kilometres, and he says they’re looking to go even further. Tracy Northup, who you heard from earlier and who wasn’t part of the research, was impressed with the work.
Interviewee: Tracy Northup
I am impressed. I think there’s a handful of technological advancements, and combining those in a single experiment is really challenging, so they’ve just done a lot of things to try to scale up the system and it’s challenging to do those all at once.
Interviewer: Benjamin Thompson
There is, of course still a lot of work to do, before a system like this could become part of a larger network. For example, Tracy explained that entangled states are fragile and can easily be lost, and right now, in Jian-Wei’s system, they are lost faster than new ones can be generated, but she thinks that this is something that can be overcome. While some of the principals of quantum mechanics have been used in a limited form to encrypt data along regular communication networks for a while, a network of multiple interconnected quantum nodes around the globe remains a way off just yet. This work appears to be another step in the right direction, but it could be a long road before we reach a quantum internet and its promise of radically different applications in science and computing.
Interviewee: Tracy Northup
I think that for all of these different applications, we’ll just start to see kind of the first test beds emerging, and then I think we’ll start to see, within the next ten years, that we can link together quantum computers and do computations over these distributed networks or distributed sensors. That’s something I’m certainly looking forward to.
Host: Shamini Bundell
That was Tracy Northup. You also heard from Jian-Wei Pan, and you can read his paper over at nature.com.
Interviewer: Nick Howe
Finally, on the show, it’s time for the News Chat, and I’m joined in the studio by Nisha Gaind, Nature’s European Bureau Chief. Nisha, hi.
Interviewee: Nisha Gaind
Hi, Nick.
Interviewer: Nick Howe
So, Nisha, we’re actually sat in the studio recording this on Wednesday morning, and this is a very fast-moving story. By the time we go out, things may have changed, but it’s of course time to talk about the coronavirus, which we’ve been keeping track of each week. So, Nisha, what more do we know since last week?
Interviewee: Nisha Gaind
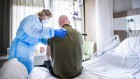
Since last week, of course, cases have continued to rise. As of this morning, there are about 45,000 reported cases of this outbreak in China alone, and that’s obviously where the cases are concentrated. We, in the past week, have passed a couple of milestones in terms of case numbers. The number of deaths that have occurred because of this virus have passed those that were caused by SARS (Severe Acute Respiratory Syndrome) – that’s the disease outbreak that happened in 2002-2003 that also emerged from China – and now, the deaths caused by this current coronavirus have passed 1,000, which is another sort of symbolic milestone. So, cases have continued to rise, and there is still a fervent response to trying to control this outbreak.
Interviewer: Nick Howe
Right, and as you said, there’s about 45,000 cases there, but is it starting to level off slightly?
Interviewee: Nisha Gaind
So, that’s something that is very difficult to tell, and that’s the question that epidemiologists will be trying to answer as best they can. They do that in a number of ways. They use models where they plug in the data and try to predict how the outbreak might continue, but some of the key information that is needed for those models are things like the incubation rate – that’s the time between somebody being infected and showing symptoms – and it’s actually not clear what that time is at the moment. There have been estimates from 3-14 days.
Interviewer: Nick Howe
So, we may not have reached the peak of the outbreak yet – we just don’t know. And I also understand that the WHO has named the virus.
Interviewee: Nisha Gaind
Yes, that’s right, and that was quite a big piece of news. That happened this week. The WHO convened a meeting where they are primarily talking about research priorities in this outbreak, and one of the first things that they announced from this meeting is a name for the disease that this virus has caused. Now, that’s quite an important distinction. There are different names for the virus and the disease it causes. The WHO chose to name the disease COVID-19 – that’s an abbreviation, essentially, for coronavirus disease 19, the year in which it initially emerged. And there had been a lot of discussion about the name of this virus. It had been going by several different monikers, things that journalists had been using, different health agencies had been using, things like the Wuhan virus and the coronavirus. But the WHO made clear that it was really important that the name for the disease did not refer to a geographical location or an animal or an individual or a group of people because these are all things that can be stigmatising or inaccurate.
Interviewer: Nick Howe
And scientists have been desperately trying to find this for a while and we talked about this a couple of times, but do we have any more idea of where this virus has come from?
Interviewee: Nisha Gaind
Yes, so this is another key question that researchers have been really desperate to find out about this virus – where did it emanate from – because we know from previous viruses like SARS and MERS (Middle East Respiratory Syndrome) that they came originally from bats and then it leapt to humans through an intermediate animal. Now, there’s been quite a lot of discussion about what this intermediate animal might be for this current virus, and that’s because it originally seemed to emerge in this animal and seafood market in Wuhan, the epicentre of the outbreak. There have been several suggestions – one, a few weeks ago, was snakes and that was roundly dismissed by scientists – but the latest suggestion is that pangolins, which are these scaly mammals that are quite a protected species, may have been this intermediate animal from which the coronavirus leapt to humans.
Interviewer: Nick Howe
And what’s the consensus at the moment for pangolins being an intermediary?
Interviewee: Nisha Gaind
Well, at this point, it’s really important to stress that this research and a lot of the research being done in response to this outbreak is preliminary research. This particular piece of work was done by scientists in China, and it was a genetic analysis that compared a coronavirus taken from pangolins to the coronavirus that has been taken from humans in the outbreak. This research hasn’t been published yet and it hasn’t been peer reviewed. It was announced by researchers at a press conference and announced on their university website, so there is caution to be had there. But the response from other researchers who have looked at the work say actually, this seems plausible. The genetic sequences of these viruses are 99% similar, according to the researchers, and they say that because pangolins are prized in China – they’re often used in traditional Chinese medicine – it’s plausible that they could have been in this animal market in Wuhan and where they think the virus originated.
Interviewer: Nick Howe
And other than trying to find the source of this virus, what else are scientists doing in order to try and combat this outbreak?
Interviewee: Nisha Gaind
Well, of course, there is the question of drugs and vaccines, and these are things that take a lot of time to develop. They are difficult to create in a rapid way in response to an escalating outbreak that is causing worldwide concern, but that is exactly what the WHO is meeting about this week. They are having a meeting about diagnostics and research priorities and drugs and vaccines and we should be hearing the outcome of that in the following days. But we know already that there are some drug trials co-opting drugs that are already used to treat viruses such as HIV that are being tried against this coronavirus, and we will probably hear some updates on vaccine development efforts in the next day or two.
Interviewer: Nick Howe
So, we’ll keep an eye out for those updates, but to move on to our second story, researchers have been trying to improve upon gene editing, specifically a tool called base editing. Before we get too much into that, for people who might not be familiar, what is base editing?
Interviewee: Nisha Gaind
Yeah, so base editing is another type of gene editing, and I’m sure all listeners will be familiar with CRISPR-Cas9, which is this revolutionary gene-editing technique. The difference is that CRISPR-Cas9 cuts both strands of DNA and the DNA repair process makes changes to the genome. Base editing is an even more precise version of this, and the way in which it is more precise is that instead of cutting the double strand of DNA and having the DNA repair mechanism make changes, it can actually make single-letter changes to the DNA.
Interviewer: Nick Howe
This sounds like it could be a really useful tool for scientists, but there are some problems in actually applying it.
Interviewee: Nisha Gaind
That’s right. None of these tools is perfect. They all have some pitfalls. So, one of the main concerns in gene editing is that the techniques make these things called off-target mutations. That means that if you’re a researcher and you want to target a particular gene, the technique that you’re using might also make changes in places that you don’t want it to make, and those can be potentially dangerous.
Interviewer: Nick Howe
But scientists have been working to sort of improve this and get rid of these off-target edits. How have they done this?
Interviewee: Nisha Gaind
Yeah, that’s right. So, this week, in Nature Biotechnology, there are a couple of papers that demonstrate a way to improve base editing, and in particular, this issue of off-target mutations. So, the researchers developed a method in which they inserted base editors into bacteria and ultimately, they were able to screen various different enzymes, both naturally occurring and engineered ones, in search of these base-editing enzymes that were much better at making these super-precise single-letter changes i.e. those that could convert C to T without causing as many off-target mutations.
Interviewer: Nick Howe
So, why might this be a useful tool for scientists?
Interviewee: Nisha Gaind
So, ultimately, the aim of all of these gene-editing technologies is to create therapies that can be used in medicine and treat diseases, and in this case, one example of a disease that could really benefit from base editing is sickle-cell anaemia, which is a genetic human disease and that can often be caused by single-letter changes to DNA, so you can see there how base editing might be a really attractive approach for treating diseases like that.
Interviewer: Nick Howe
Well, we’ll keep an eye on these developments as they come. Listeners, for more on those stories, head over to nature.com/news. There you’ll also find a live blog where we’re keeping up to date with all the coronavirus news, and if you’re a researcher that’s been affected by coronavirus or it’s affected your research then please let us know. We have a little survey in that live blog where you can tell us all about it. All that remains to be said then is, Nisha, thanks for joining me.
Interviewee: Nisha Gaind
Thanks.
Host: Shamini Bundell
That’s all we’ve got time for this week, but if you still want some more science, we’ve been busy making a lot of videos for you. We have a three-minute guide on how scientists are fighting the coronavirus, a three-part video series about Japan’s big physics, and a study of bird flight using bubbles. Find all of that over at youtube.com/NatureVideoChannel. I’m Shamini Bundell.
Host: Nick Howe
And I’m Nick Howe. Thanks for listening.

 Read the paper: Hidden diversity of vacancy networks in Prussian blue analogues
Read the paper: Hidden diversity of vacancy networks in Prussian blue analogues
 Latest news on the coronavirus
Latest news on the coronavirus
 Super-precise CRISPR tool enhanced by enzyme engineering
Super-precise CRISPR tool enhanced by enzyme engineering