- NEWS AND VIEWS
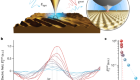
Getting the measure of living biomaterials
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 572, 38-39 (2019)
doi: https://doi.org/10.1038/d41586-019-02263-7
References
Hughes, A. J. et al. Dev. Cell 44,165–178 (2018).
Nerger, B. A., Brun, P.-T. & Nelson, C. M. Soft Matter 15, 5539–5780 (2019).
Morley, C. D. et al. Nature Commun. 10, 3029 (2019).
Murphy, S. V. & Atala, A. Nature Biotechnol. 32, 773–785 (2014).
Kolesky, D. B., Homan, K. A., Skylar-Scott, M. A. & Lewis, J. A. Proc. Natl Acad. Sci. USA 113, 3179–3184 (2016).
Bhattacharjee, T. et al. Sci. Adv. 1, e1500655 (2015).
Hinton, T. J. et al. Sci. Adv. 1, e1500758 (2015).
Gladman, A. S., Matsumoto, E. A., Nuzzo, R. G., Mahadevan, L. & Lewis, J. A. Nature Mater. 15, 413–418 (2016).
Bade, N. D., Xu, T., Kamien, R. D., Assoian, R. K. & Stebe, K. J. Biophys. J. 114, 1467–1476 (2018).
Grimes, A. et al. Lab Chip 8, 170–172 (2008).
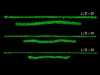
 Read the paper: Quantitative characterization of 3D bioprinted structural elements under cell generated forces
Read the paper: Quantitative characterization of 3D bioprinted structural elements under cell generated forces
 Versatile gel assembly on a chip
Versatile gel assembly on a chip
 Radicals promote magnetic gel assembly
Radicals promote magnetic gel assembly