- NEWS AND VIEWS
Detailed view of human telomerase enzyme invites rethink of its structure
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 557, 174-175 (2018)
doi: https://doi.org/10.1038/d41586-018-04756-3
References
Palm, W. & de Lange, T. Ann. Rev. Genet. 42, 301–334 (2008).
Vulliamy, T. J. & Dokal, I. Biochimie 90, 122–130 (2008).
Kim, N. W. et al. Science 266, 2011–2015 (1994).
Nguyen, T. H. D. et al. Nature 557, 190–195 (2018).
Egan, E. D. & Collins, K. RNA 18, 1747–1759 (2012).
Kiss, T., Fayet-Lebaron, E. & Jády, B. E. Mol. Cell 37, 597–606 (2010).
Cristofari, G. & Lingner, J. EMBO J. 25, 565–574 (2006).
Sauerwald, A. et al. Nature Struct. Mol. Biol. 20, 454–460 (2013).
Chan, H., Wang, Y. & Feigon, J. Ann. Rev. Biophys. 46, 199–225 (2017).
Li, L. & Ye, K. Nature 443, 302–307 (2006).
Venteicher, A. S. et al. Science 323, 644–648 (2009).
Zappulla, D. C. & Cech, T. R. Proc. Natl Acad. Sci. USA 101, 10024–10029 (2004).
Jiang, J. et al. Nature 496, 187–192 (2013).
Alves, D. et al. Nature Chem. Biol. 4, 287–289 (2008).
Egan, E. D. & Collins, K. Mol. Cell. Biol. 30, 2775–2786 (2010).
Egan, E. D. & Collins, K. Mol. Cell. Biol. 32, 2428–2439 (2012).
Stone, M. D. et al. Nature 446, 458–461 (2007).

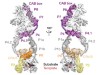 Read the paper: Cryo-EM structure of substrate-bound human telomerase holoenzyme
Read the paper: Cryo-EM structure of substrate-bound human telomerase holoenzyme
 The telomerase enzyme and liver renewal
The telomerase enzyme and liver renewal
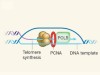 Telomere-lengthening mechanism revealed
Telomere-lengthening mechanism revealed