- NEWS AND VIEWS
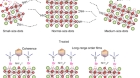
DNA self-assembly scaled up
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 552, 34-35 (2017)
doi: https://doi.org/10.1038/d41586-017-07690-y
References
Whitesides, G. M. & Grzybowski, B. Science 295, 2418–2421 (2002).
Tikhomirov, G. et al. Nature 552, 67–71 (2017).
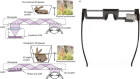
Wagenbauer, K. F. et al. Nature 552, 78–83 (2017).
Ong, L. L. et al. Nature 552, 72–77 (2017).
Praetorius, F. et al. Nature 552, 84–87 (2017).
Chen, Y.-J., Groves, B., Muscat, R. A. & Seelig, G. Nature Nanotechnol. 10, 748–760 (2015).
Li, J., Green, A. A., Yan, H. & Fan, C. Nature Chem. 9, 1056–1067 (2017).
Zheng, J. et al. Nature 461, 74–77 (2009).
Rothemund, P. W. K. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Nature 459, 414–418 (2009).
Wei, B., Dai, M. & Yin, P. Nature 485, 623–626 (2012).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Science 338, 1177–1183 (2012).
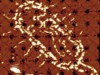
 Read the paper: Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns
Read the paper: Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns
 Read the paper: Gigadalton-scale shape-programmable DNA assemblies
Read the paper: Gigadalton-scale shape-programmable DNA assemblies
 Read the paper: Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components
Read the paper: Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components
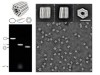 Read the paper: Biotechnological mass production of DNA origami
Read the paper: Biotechnological mass production of DNA origami
 Pathfinder for DNA constructs
Pathfinder for DNA constructs
 Molecular robots on the move
Molecular robots on the move