Abstract
The Small ubiquitin-related modifier (SUMO) conjugation to a variety of proteins regulates diverse cellular processes, including transcription, cell cycle regulation and maintenance of genome integrity. To investigate in vivo biological function of SUMO paralogs, we inactivated them in the early development of zebrafish. While zebrafish embryos deficient for all three SUMO paralogs, as Ubc9-deficient ones, displayed severe defects, loss of individual SUMO paralog was compatible with a normal development. SUMO-deficient embryos can be rescued by a single human or zebrafish SUMO. While key structural basic lysine residues and N-terminal unstructured stretch of SUMO are critical for in vivo rescue, the consensus K11 sumoylation site of SUMO2 is dispensable, implying that chain formation on this potential site is unessential for normal development. Inactivation of all three SUMOs triggered p53-dependent apoptosis and further inactivation of p53 restored normal zebrafish development. Interestingly, we also demonstrate that the dominant negative truncated form of p53, Δ113p53, significantly blunts SUMO depletion-induced p53 activity in vivo. Taken together, our results suggest that SUMO paralogs are indispensable, but redundant, in the early development of zebrafish.
Similar content being viewed by others
Introduction
The small ubiquitin-related modifier (SUMO) is highly conserved through evolution, and belongs to the growing family of ubiquitin-like proteins (UBLs) involved in post-translational protein modification. SUMO conjugation is carried out by a multistep enzymatic pathway consisting of a unique heterodimeric SUMO-activating enzyme (E1), Aos1/Uba2, a unique SUMO-conjugating enzyme (E2), Ubc9, and three families of SUMO ligase (E3), which eventually attach SUMO to the substrates 1, 2, 3, 4. Protein sumoylation plays important roles in a wide variety of biological processes such as transcriptional regulation, maintenance of genome integrity and subcellular localization 5, 6, 7, 8, 9.
The sumoylation pathway is conserved from yeast to human. Four SUMO paralogs, SUMO1, SUMO2, SUMO3 and SUMO4, exist in mammals. While SUMO2, SUMO3 and SUMO4 share high homology with each other 10, SUMO1 consists of 101 amino acids and shares about 50% sequence identity with SUMO2, SUMO3, SUMO4 and 18% with ubiquitin. In mammalian cells, different SUMO paralogs appear to share common conjugation properties, but also have some specificities, such as subcellular distribution and substrate preferences. For instance, it is reported that, within cells, there is a larger pool of free, non-conjugated SUMO2/3 than that of SUMO1 11. The distribution of the SUMO paralogs within cells also seems to be different. SUMO1 is found within the nucleoli, the nuclear envelope and cytoplasmic foci, whereas SUMO2/3 are dominant on chromosomes. Moreover, while the expression of SUMO1 and SUMO2/3 is ubiquitous, SUMO4 is found predominantly in kidney and immune system 10, 12. The conjugation of SUMO2/3 to substrates is usually induced when the cells are subjected to protein-damaging stimuli, such as acute temperature fluctuation, suggesting a role for SUMO2/3 in the cellular responses to environmental stress 11. Covalent conjugation of SUMO4 to substrates is still under debate 13, 14. Finally, for some substrates, strong paralog preferences have been reported. For instance, promyelocytic leukemia protein (PML) could be conjugated to all three SUMO paralogs 15, 16. RanGAP1 is preferentially modified by SUMO1 11, and topoisomerase II is sumoylated by SUMO2/3 during mitosis 17.
In most model organisms, loss of Ubc9 leads to severe defects and lethality 18, 19, 20, 21, 22, 23. However, the respective biological impact of each SUMO paralog is poorly explored in vivo. It is known that SUMO is required for normal embryonic development in the organisms with one single SUMO gene 20, 24. More recently, it has been reported that SUMO1 is dispensable in mouse development 25, but the importance of SUMO2/3 in vivo is unknown. Here, we have investigated the functional role of SUMO paralogs during the early embryonic development of zebrafish. We found that all three SUMOs were maternally provided and ubiquitously expressed at early developmental stages. While the maternally supplied SUMO proteins may still be sufficient to carry out essential functions for cell viability during early hours of embryonic development, loss of zygotic SUMO paralogs by antisense morpholino oligonucleotides (MOs) causes severe embryonic developmental defects. Further analysis reveals that cells undergoing p53-dependent apoptosis may account for the SUMO deficiency-triggered developmental defects.
Results
Spatial and temporal expression patterns of SUMO paralogs during zebrafish early embryonic development
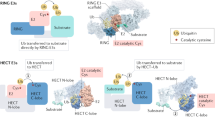
Three SUMO paralogs of zebrafish were identified via bioinformatic searching. Zebrafish SUMO paralogs and Ubc9 share high homology with their human counterparts (Figure 1A). Their embryonic expression was examined by whole-mount mRNA in situ hybridization. Similar spatial and temporal expression patterns were found for all three SUMO paralogs and Ubc9 (Figure 1B and Supplementary information, Figure S1). All three SUMO transcripts could be detected throughout all investigated stages, including one-cell stage (Figure 1B, a1-3), indicating that they were both maternally and zygotically supplied. During cleavage (Figure 1B, b1-3), blastula (Figure 1B, c1-3), gastrula (Figure 1B, d1-3 and e1-3) and early segmentation stages (data not shown), the three SUMO transcripts were ubiquitously expressed throughout the entire embryo. At 24 hours post-fertilization (hpf), expression was dominant in the head region, including brain, eyes, cranial neural crest cells (Figure 1B, f1-3 and g1-3), which are known as late proliferative zones. Double staining with neural crest marker dlx2a 26 confirmed that the three SUMOs were specifically expressed within this region (Figure 1B, h1-3). At 48-hpf, expression became further restricted to late proliferative zones. While expression was found in pectoral fin buds, it was much weaker within the regions of trunk and tail (Figure 1B, i1-3). At 72-hpf, expression had decreased in most tissues except in the branchial arches and the digestive organs (Figure 1B, j1-3).
Zebrafish SUMO paralog expression patterns during embryonic development. (A, upper panel) Zebrafish SUMO paralogs and Ubc9 share high homology with human counterparts. (A, lower panel) Amino acid sequence alignment of zebrafish SUMO paralogs. Asterisk indicates conserved amino acid. D.r, Danio rerio; H.s, Homo sapiens. (B) Whole-mount mRNA in situ hybridization of SUMO1, SUMO2 and SUMO3 was carried out at indicated stages of development. (B, h1-3) Double staining of SUMO paralog (red) and dlx2a (dark blue), a marker for cranial neural crest cells. dlx2a-positive crest streams are numbered. cnc, cranial neural crest; e, eye; pfb, pectoral fin bud; gi, gastrointestinal system; ga, gill arches.
Loss of SUMO expression leads to developmental defects
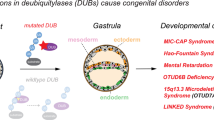
To study loss of functions of zebrafish SUMO paralogs during early embryonic development, SUMO paralog-specific MOs were injected individually or in different combinations at one-cell stage, and specific rescue experiments were performed in parallel. Injected embryos were followed until 7 days post-fertilization (dpf). The efficacy of each SUMO paralog-specific MO was first tested with a SUMO-EGFP reporter (Supplementary information, Figure S2A) and further confirmed by the diminution of endogenous SUMOs and Ubc9 proteins (Supplementary information, Figure S2B). Individual loss of SUMO1, SUMO2 and SUMO3, or pairwise loss of SUMO1 and SUMO2 had no overt phenotype (Supplementary information, Table S1). Severe developmental defects, similar to those observed in Ubc9-deficient embryos, were found only in the knockdown group for all three SUMO paralogs as soon as 3 dpf (Figure 2A and Supplementary information, Table S1). At around 7 dpf, the injected embryos died. A reduction in the sizes of brain and eyes, and malformations in the jaw were observed (Figure 2A, a-d). Surprisingly, the structures of the trunk, tail and otic placode remained unaffected. Hematoxylin-eosin staining showed a relative normal gross structure of eyes, but with reduced cell number within each structural cell layer (Figure 2A, i and j). Alcian blue staining demonstrated that the first to fourth pair of gill arches were dramatically reduced or absent. Interestingly, the fifth pair was still present (Figure 2A, e-h). Chondrocytes of gill arches of SUMO-deficient embryos were much larger compared to the control ones (Figure 2A, e'-h'), suggesting that the cell cycle might be affected. Pairwise loss of SUMO1 and SUMO3, or SUMO2 and SUMO3 paralogs led to a much lower incidence of developmental defects (Supplementary information, Table S1). Lower incidence and less severe defects of double SUMO knockdown morphants were probably due to the fact that there were less apoptosis events compared to that of triple SUMO knockdown, as revealed by the terminal deoxynucleotidyl transferase-mediated deoxyuridin triphosphate nick-end labeling (TUNEL) assay (Supplementary information, Figure S3).
Phenotype and mechanism analysis of zebrafish SUMO-deficient embryos. (A) Morphological defects of SUMO and Ubc9-deficient morphants. (a, e) Embryos were injected with mismatch MO. (b, f) SUMO1-SUMO2-SUMO3 MOs. (c, g) SUMO1-SUMO2-SUMO3 MOs plus zebrafish SUMO1-SUMO2-SUMO3 mRNAs and (d, h) Ubc9 MO. SUMO paralogs or Ubc9-deficient morphants displayed similar developmental defects, such as reduction in the size of head (red arrowhead) and eyes (red sharp arrow), and malformations in the jaw (red arrow). Note that the trunk and otic placode (blue arrow) are normal. Embryo co-injected with zebrafish SUMO1-SUMO2-SUMO3 mRNAs displayed relatively normal structure. (e-h) Embryos were stained with alcian blue to visualize chondrocytes of gill arches at 120 hpf. (e'-h') Highly magnified images of corresponding boxed chondrocytes from e-h. (i, j) Transverse sections through the eye at 72 hpf were stained with hematoxylin-eosin. The mismatch control larvae shows normal characteristic stratification, including ganglion cell layer (1, GCL), inner plexiform layer (2, IPL), inner nuclear layer (3, INL), outer nuclear layer (4, ONL) and retinal pigmented epithelium (5, RPE). Note that there was a severe reduction of cells in each layer, though the stratification of retinal layers still remained in SUMO paralog-deficient morphant. (k) Diameters of eye vesicles of indicated embryos were measured as following: lane 1, 312.53 μm ± 8.4 of mismatch group, (n = 10); lane 2, 205.75 μm ±14.77 of SUMO1-SUMO2-SUMO3 MOs group, (n = 10); lane 3, 209.46 μm ± 13.7 of Ubc9 MO group (n = 5); lane 4, 301.0 μm ±8.7 of SUMO1-SUMO2-SUMO3 MOs plus zebrafish SUMO1-SUMO2-SUMO3 mRNAs group (n = 10). * P < 0.01. cb, ceratobranchials of gill arches 1–5. (B) TUNEL, pH3 and FACS analysis. (a, b) TUNEL staining showed more apoptotic cells within head region of SUMO paralog-deficient morphant compared to that of control embryo. (c, d) SUMO paralog-deficient morphant displayed more phosphorylated histone H3 (pH3)-positive cells within the head region. (c′, d′) Highly magnified images of corresponding boxed region from c and d. (e, f) FACS analysis of propidium-iodide (PI) labeled cells pooled from 30 embryos showed that there was a significant higher fraction of G2/M-phase cells in the population from head region of SUMO paralog-deficient embryos at 72 hpf. (g, h) No obvious difference was detected in cell populations from trunk and tail regions. The data are representative of three independent experiments.
To understand whether the size reduction of brain and eyes in SUMO-deficient morphants was due to reduced cell proliferation and/or increased cell death, TUNEL assay and anti-phosphorylated histone H3 (pH3) immunostaining, a G2/M-phase marker, were performed. The TUNEL assay revealed that SUMO paralog-deficient embryos displayed more apoptotic cells in the brain and eye vesicles (Figure 2B, b and Supplementary information, Figure S3J), consistent with the SUMO expression pattern. There was also an increase of pH3-positive cells within head region (Figure 2B, d). A more detailed analysis by flow cytometric assay revealed that cell populations from the head region of SUMO paralog-deficient embryos had a significant higher fraction of cells in the G2/M-phase, which was not the case for the cells from trunk and tail (Figure 2B, e-h). Furthermore, using dominant negative non-conjugation SUMO mutants (SUMOΔGG-GFP), we found that cells expressing SUMO2ΔGG-GFP and SUMO3ΔGG-GFP also accumulated in the G2/M-phase (data not shown).
Rescue experiments suggest that developmental defects are specifically caused by the loss of SUMO paralogs
To confirm that the developmental defects of the SUMO paralog-deficient morphants were specifically due to the loss of SUMO expression, a series of rescue experiments were carried out. Full-length zebrafish and human SUMO paralog mRNAs were co-injected with SUMO MOs in different combinations. While co-injection of all three zebrafish SUMO mRNAs completely rescued developmental defects (Figure 2A, c, g and k), more interestingly, each individual SUMO paralog, either of zebrafish or human origin, was also able to restore a normal phenotype, with SUMO2 and SUMO3 being the most efficient (Table 1). A sumoylation consensus Ψ-K-X-D/E motif also exists in SUMO2 and SUMO3, but not in SUMO1, suggesting that SUMO2 and SUMO3 may form poly-SUMO chains 27. To test if this potential poly-SUMO formation lysine is required for SUMO rescue function in vivo, we constructed a zebrafish SUMO2 mutant where the conserved lysine 11 was changed to glutamine (K11E). Interestingly, this mutant had the same rescue efficiency as the wild-type (wt) one, suggesting that the potential poly-sumoylation of SUMO2 itself is not required in this setting (Table 1). The unstructured stretch of first 21 N-terminal amino acids of SUMO is not conserved in ubiquitin. A delta N21 SUMO2 mutant was constructed and used for rescue assay. Interestingly, the mutant was unable to restore the normal phenotype. It was also reported that charge-reversal mutations of K33/35/42 within a highly conserved surface of SUMO dramatically abrogated its repressive effect 28, 29. Consistent with this, our zebrafish SUMO2 K33/35/42E mutant had completely lost its rescue function, despite that the mutant could still efficiently conjugate to substrates (Table 1 and data not shown).
cDNA microarray and real-time PCR analysis revealed specific activation of the p53 pathway in SUMO paralog-deficient morphants
To further investigate the possible molecular mechanisms underlying the developmental defects caused by loss of all three SUMO paralogs, cDNA microarray analysis was performed on Affymetrix GeneChip Zebrafish Genome Array. About 100 genes were found upregulated or downregulated by at least twofold. Genes involved in the development of neurons and eyes, for instance, crx 30, rcv1 31, neurod 32, vsx1 33, rho 34, atoh2a 35 and rorab 36 were found downregulated (data not shown). Interestingly, the tumor suppressor p53 signaling pathway was found to be activated. Quantitative real-time PCR analysis further demonstrated that most p53 target genes, including p21, mdm2 and cyclin G1, were upregulated in a time-dependent manner with peaks appearing at 24 to 48 hpf (Figure 3A). Moreover, using isoform-specific primers, real-time PCR analysis showed that it was the truncated form of p53 (Δ113p53), lacking the transactivation domain and part of the DNA-binding domain, but not the full-length p53 (FL p53) that was drastically upregulated (Figure 3A and 3B). While bax was marginally activated, significant activation of other target genes were further confirmed using mRNAs isolated from morphants head region (Figure 3B). Importantly, Δ113p53 was not induced by extinction of individual SUMO paralog (Figure 3B).
Loss of SUMO paralogs activates the p53 pathway. (A) Time course of quantitative RT-PCR analysis of p53 target genes′ activation in SUMO-deficient embryos. (B) Quantitative RT-PCR analysis of p53 target genes at 48 hpf of the head region origin mRNA of embryos injected with mismatch MO (black), SUMO1-SUMO2-SUMO3 MOs (red), SUMO1-SUMO2-SUMO3 MOs plus FL p53 MO (green), SUMO1-SUMO2-SUMO3 MOs plus Δ113p53 MO (yellow), each individual SUMO MO (blue, purple and clear blue). Note that Δ113p53 was upregulated only in three SUMO paralog-deficient embryos, but not in any single SUMO paralog-deficient ones.
Given the fact that the p53 pathway was activated and Δ113p53 was strongly upregulated in embryos deficient for all three SUMO paralogs, MO against FL p53 or Δ113p53 was co-injected with the three SUMO paralog MOs. Interestingly, FL p53 MO eradicated apoptotic cells in the head region and alleviated morphological developmental defects (Figure 4A and 4B), and diminished the activation of p53 target gene transcription (Figure 3B). As a result, the embryos survived and continued to develop normally. On the contrary, Δ113p53 MO caused enhanced activation of p53 target genes and resulted in more severe and earlier onset of embryonic developmental defects due to cell apoptosis (Figure 3B, 4A and 4B). Consistent with a previous report 37, we also demonstrated that Δ133p53, the human counterpart of zebrafish Δ113p53, was a possible p53 direct target gene driven by an alternative internal promoter within intron 4 of human p53 (data not shown). Luciferase assays were performed on a series of p53 target gene promoters, including mdm2, fas and PIG3. While Δ133p53 by itself could not activate these promoters, it attenuated the transactivation effect of FL p53 in a dose-dependent manner (Figure 4C-E). Then we questioned if Δ133p53 may regulate FL p53 activity through its oligomerization-dependent protein stabilization effect, as the oligomerization domain was still retained in Δ133p53. Pulse chase result showed that the turnover rate of p53 was not affected by Δ133p53 (Figure 4F). It has been reported that p53 itself could be sumoylated on K386, and the consequence of p53 sumoylation on transcription regulation was still under debate 38, 39, 40, 41. Our luciferase assay results showed no significant difference between wt p53 and p53 K386R on the tested promoters (Supplementary information, Figure S4). In addition, turnover rate of the two proteins was quite similar (data not shown). These data suggest that hypo-sumoylation of p53 may not have a direct impact on its transactivation.
FL p53 MO specifically rescues the developmental defects resulted from SUMOs deficiency. (A) SUMO paralog-deficient morphant displayed significantly more apoptotic cells within brain and eye vesicle regions compared to that of mismatch controls, as revealed by TUNEL assay. While co-knockdown of FL p53 diminished apoptotic cells, co-knockdown of Δ113p53 augmented cell apoptosis in SUMO paralog-deficient morphant. (B) Morphology of embryos injected with mismatch MO plus FL p53 MO, SUMO MOs, SUMO MOs plus FL p53 MO and SUMO MOs plus Δ113p53 MO at 72 hpf. Note that knockdown of FL p53 alone had no effect on embryonic development. (C-E) Luciferase activity assay of human mdm2, fas and PIG3 promoter was carried out in the presence of empty vector (pSG5), FL p53, Δ133p53 or FL p53 plus Δ133p53 with indicated ratio in H1299 cells. Renilla luciferase plasmid was used as internal control. (F) H1299 cells were transfected with FL p53 in the absence or presence of Δ133p53. Cells were treated with cycloheximide (CHX) 24 h after transfection and harvested at indicated time point after CHX treatment. Cell extracts were blotted with anti-p53 antibody.
Discussion
Sumoylation and desumoylation have essential effects on a variety of biological events and involve a multienzyme process. It has been reported that loss of the unique E2 conjugating enzyme Ubc9 is embryonic lethal in zebrafish 23 and mice 22. In contrast, disruption of the E3 ligase PIAS family genes leads to modest defects in mice. PIASx-mutant mice have reduced testis weight and an increased rate of apoptosis 42, PIASy-deficient mice display modest defects in IFN and Wnt signaling pathways 43, 44 and PIAS1 knockout mice do not have histological defects 45. These results suggest that there is functional redundancy among SUMO E3 ligase PIAS family members during mouse development. Inactivation of the desumoylating enzyme, SENP1, causes an increase in the level of SUMO1 conjugation but not in that of SUMO2 or SUMO3, and results in prenatal lethality in mice 46. SENP1 knockout mice display severe fetal anemia in midgestation 47. But up to date, the respective biological function of each SUMO paralog in the early embryonic development remains unknown. Here, we report that similar to the loss of Ubc9, simultaneous knockdown of all three SUMO paralogs was lethal in most morphants. It is worth noting that the loss of Ubc9 leads to a higher percentage of defective embryos than does the loss of three SUMO paralogs (97.5% vs 84.9%). This observation may be explained, in part, by the fact that reduction of Ubc9 by MO is more efficient than that of SUMO paralogs (Supplementary information, Figure S2B). Interestingly, we find that loss of single paralog or pairwise loss of SUMO1 and SUMO2 leads to no overt defect. SUMO1 and SUMO3, or SUMO2 and SUMO3 double-deficient embryos displayed a lower incidence and less severity of developmental defects, suggesting that a partial functional redundancy may exist between SUMO paralogs and that SUMO3 might have a more critical or broader spectrum of substrates during the early embryonic development. In accordance with these findings, we demonstrate that the developmental defects associated with loss of the three SUMO paralogs could be rescued by any single SUMO paralog. The rescue effect of SUMO2 and SUMO3 is much better than that of SUMO1, again arguing for the importance of SUMO2 and SUMO3.
Although we do not have the direct in vivo evidence that different paralogs can be used interchangeably, our findings suggest that there is a significant functional redundancy of SUMO paralogs and that the functions of one specific SUMO can be compensated by other paralogs. Our observation is further supported by the recent findings that SUMO1 is dispensable in mouse development 25 and that SUMO1 and SUMO2 are functionally redundant genes in Arabidopsis as only double-homozygous mutants in embryo are lethal 48.
Sumoylation of transcription factors normally leads to transcriptional repression through the interaction with co-repressors such as Daxx or HDACs. A basic residue-rich region of SUMO (K33, K35, K42, R50) defines the interaction surface for the co-repressors and accounts for the inhibitory properties of SUMO 28, 29. These sites are highly conserved in all zebrafish and human SUMO paralogs. In our rescue experiments, the zebrafish SUMO2 K33/35/42E charge-reversed mutant failed to function, further emphasizing the critical role of these residues for SUMO function in vivo. In SUMO2 and SUMO3, the presence of a potential sumoylation consensus sequence could allow SUMO2 and SUMO3 to form poly-SUMO chain 27. Though we could not rule out the possibility that other lysine residues may also be used for chain formation, the SUMO2 K11E consensus sequence mutant has no impact on the rescue effect, suggesting that the possible poly-SUMO chain formation on this potential site might not be required for its in vivo function in this setting. Incapability of the SUMO2 delta N21 mutant to restore the normal phenotype suggests that the N-terminal unstructured region of SUMO, which is not conserved in ubiquitin, may possess yet unidentified properties required for its biological function.
In our experiments, we show that the p53 pathway was activated in morphants deficient for all three SUMO paralogs. It has been reported that the MO knockdown technology may elicit undesirable off-target effects and that the major off-target effect is mediated through p53 activation accompanied by diagnostic transcription of Δ113p53 49. Our results do not support this scenario. First, as mentioned above, knockdown of single SUMO paralog did not lead to any developmental defects and p53 activation. Second, single SUMO paralog, of either zebrafish or human origin, could specifically reverse the phenotype caused by SUMO deficiency, suggesting that the developmental defects are strictly SUMO-related. Finally, MO targeting FL p53 could alleviate cell apoptosis and rescue morphological defects associated with the loss of SUMO paralogs.
We also found that the dominant negative isoform Δ113p53 was drastically upregulated, and Δ113p53 MO caused further activation of p53 target genes and earlier onset of more severe embryonic developmental defects. Consistent with this, it has been very recently reported that zebrafish Δ113p53 is a p53 target gene and antagonizes p53 apoptotic activity in vivo through Bclxl activation 50.
It is still unclear how loss of SUMO paralogs activates the p53 pathway. This is unlikely due to the direct consequence of hypo-sumoylation of p53 protein caused by SUMO knockdown, as the p53 sumoylation-defective mutant (p53 K386R) shares similar transactivation and protein stability properties with the wt one. This is not surprising since only a limited fraction of p53 is sumoylated in vivo. We favor the hypothesis that p53 activation could be a secondary effect resulting from the loss of SUMOs. A plausible mechanism is suggested by the recent findings that SUMO paralogs are involved in the mitotic process of mammalian cells, and sumoylation is a key regulator of cell cycle 51. Inhibition of sumoylation may block sister chromatid segregation and cause cell cycle arrest. Cells may eventually succumb to a secondary, p53-dependent apoptotic cell death.
Taken together, our in vivo probing of the SUMO system in zebrafish embryo demonstrates that SUMO extinction triggers p53 activation and cell death, but that the three SUMOs are functionally redundant. SUMO K11 chain formation appears to be dispensable for early developmental processes.
Materials and Methods
Fish maintenance
Zebrafish maintenance and staging were performed as described previously 52.
Generation of constructs
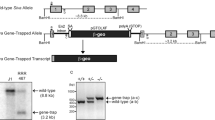
SUMOs and Ubc9 were amplified by RT-PCR with indicated primes for either WISH assay or rescue assay (Table S2), and cloned into pCS2+ vector. For MO specificity assay, a 5′ fragment of each SUMO paralog covering the ATG site was amplified with indicated primers (Table S2) and cloned in frame with egfp into pCS2+ vector. The zebrafish SUMO2 K11E and K33/35/42E mutants were generated by QuickChange Site-Directed Mutagenesis Kit (Stratagene) with indicated primers (Table S2). The zebrafish SUMO2ΔN21 was generated by PCR with indicated primers (Table S2). The human p53 intron 4 sequences were amplified from human genomic DNA with indicated primers (Table S2). Human Δ133p53 was cloned into pSG5 expression vector with indicated primers (Table S2).
Whole-mount in situ hybridization
Digoxigenin-labeled or fluorescein-labeled antisense RNA probes were transcribed from linearized cDNA constructs using T3 or T7 polymerase (Roche). Whole-mount in situ hybridization was performed as described previously 53. The probes were detected using alkaline phosphatase-coupled anti-digoxigenin Fab fragment antibody (Roche) or anti-fluorescein Fab fragment antibody (Roche) with BCIP/NBT staining (Vector Laboratories) or Fast red staining (Roche).
MOs and mRNAs microinjection
MOs were purchased from Gene Tools. SUMO1 MO 5′-GTC TCC GTG TCT GAC ATG ATA TTC C-3′; SUMO2 MO 5′-CAT GGT TAT TGT ATT TGC GCT TCT C-3′; SUMO3 MO 5′-TAG GCT TGT CTT CGG ACA TTT TTG C-3′; Ubc9 MO 5′-TCA GAG CAA TGC CAG ACA TGA CCA C-3′; p53 MO 5′-TCT TGG CTG TCG TTT TGC GCC ATT G-3′; Δ113p53 MO 5′-GCA ACG TCC ACC ACC ATT TGA ACG G-3′; mismatch MO 5′-GTG TCC CTG TCT CAC ATC ATA TAC C-3′. The doses injected per embryo were: mismatch, 12.46 ng; MO SUMO1, 12.46 ng; MO SUMO2, 12.46 ng; MO SUMO3, 12.46 ng; MO Ubc9, 12.46 ng; MO SUMO1-SUMO2 combination, 4.15 ng each; MO SUMO1-SUMO3 combination, 4.15 ng each; MO SUMO2-SUMO3 combination, 4.15 ng each; MO SUMO1-SUMO2-SUMO3 combination, 4.15 ng each; MO FL p53, 4.15 ng; MO Δ113p53, 4.15 ng. 200 pg capped mRNAs were delivered per embryo with MOs.
Immunoblotting
Embryos were harvested at indicated time points and de-yolked as described previously 54. Western blotting was performed with rabbit anti-SUMO1 (Zymed), rabbit anti-SUMO2/3 (Zymed), rabbit anti-Ubc9 (Zymed), goat anti-p53 (Santa Cruz) and rabbit anti-actin (Sigma) antibodies. The blots were detected by SuperSignal West Pico Substrate (Pierce). P53 pulse chase assay was carried out in H1299 cells. Cells were transfected with different p53 constructs and treated with 50μg/ml cycloheximide (CHX) 24 h after transfection. Cells were harvested at indicated time points and subjected to immunoblotting analysis.
Histology and alcian blue staining
Embryos were fixed in 4% paraformaldehyde/PBS, dehydrated and embedded in paraffin. Specimens were cut into 4 μm sections that were stained with hematoxylin and eosin solution. Alcian blue staining was performed as described previously 49.
TUNEL assay, phosphorylated histone H3 labeling and fluorescence-activated cell sorting analysis (FACS)
TUNEL was performed using the in situ Cell Death Detection Kit, TMR red (Roche) according to the manufacturer's recommendations. Briefly, embryos were dechorionated and fixed in 4% paraformaldehyde/PBS overnight, dehydrated and stored in methanol at −20 °C. After rehydration in PBS, embryos were permeabilized with 0.1% sodium citrate and 0.1% Triton X-100 for 15 min at room temperature and subjected to the TUNEL assay. Phosphorylated histone H3 labeling of fixed embryos was performed with the rabbit anti-phosphohistone H3 antibody (Santa Cruz) at 4 °C overnight and revealed with Alexa Fluor 488 goat anti-rabbit secondary antibody (Invitrogen). FACS was performed as described previously 23.
cDNA microarray and quantitative real-time PCR
RNA of injected embryos were extracted with RNeasy Mini Kit (Qiagen) according to the manufacturer's recommendations. Microarray assay was performed by Shanghai Biochip Co. Ltd. The quantitative real-time PCR was performed by a LightCycler 1.5 (Roche) following manufacturer's protocols, and the amount of transcribed cDNA was normalized to β-actin. The quantitative real-time PCR were performed with indicated primers (Table S3).
Luciferase reporter assay
H1299 cells were transfected with indicated plasmid using FuGENE HD Transfection Reagent (Roche). Cells were harvested 48 h after transfection and luciferase activities were analyzed using the Dual Luciferase Reporter Assay Kit (Promega) according to the manufacturer's protocols. Luciferase activity was normalized to Renilla activity.
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Melchior F . SUMO--nonclassical ubiquitin. Annu Rev Cell Dev Biol 2000; 16:591–626.
Kim KI, Baek SH, Chung CH . Versatile protein tag, SUMO: its enzymology and biological function. J Cell Physiol 2002; 191:257–268.
Muller S, Hoege C, Pyrowolakis G, Jentsch S . SUMO, ubiquitin's mysterious cousin. Nat Rev Mol Cell Biol 2001; 2:202–210.
Hay RT . SUMO: a history of modification. Mol Cell 2005; 18:1–12.
Matunis MJ, Coutavas E, Blobel G . A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J Cell Biol 1996; 135:1457–1470.
Muller S, Matunis MJ, Dejean A . Conjugation with the ubiquitin-related modifier SUMO-1 regulates the partitioning of PML within the nucleus. EMBO J 1998; 17:61–70.
Lallemand-Breitenbach V, Jeanne M, Benhenda S, et al. Arsenic degrades PML or PML-RARalpha through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol 2008; 10:547–555.
Desterro JM, Rodriguez MS, Hay RT . SUMO-1 modification of I kappa B alpha inhibits NF-kappaB activation. Mol Cell 1998; 2:233–239.
Gill G . Something about SUMO inhibits transcription. Curr Opin Genet Dev 2005; 15:536–541.
Guo D, Li M, Zhang Y, et al. A functional variant of SUMO4, a new I kappa B alpha modifier, is associated with type 1 diabetes. Nat Genet 2004; 36:837–841.
Saitoh H, Hinchey J . Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J Biol Chem 2000; 275:6252–6258.
Bohren KM, Nadkarni V, Song JH, Gabbay KH, Owerbach D . A M55V polymorphism in a novel SUMO gene (SUMO-4) differentially activates heat shock transcription factors and is associated with susceptibility to type I diabetes mellitus. J Biol Chem 2004; 279:27233–27238.
Owerbach D, McKay EM, Yeh ET, Gabbay KH, Bohren KM . A proline-90 residue unique to SUMO-4 prevents maturation and sumoylation. Biochem Biophys Res Commun 2005; 337:517–520.
Wei W, Yang P, Pang J, et al. A stress-dependent SUMO4 sumoylation of its substrate proteins. Biochem Biophys Res Commun 2008; 375:454–459.
Kamitani T, Nguyen HP, Kito K, Fukuda-Kamitani T, Yeh ET . Covalent modification of PML by the sentrin family of ubiquitin-like proteins. J Biol Chem 1998; 273:3117–3120.
Sternsdorf T, Puccetti E, Jensen K, et al. PIC-1/SUMO-1-modified PML-retinoic acid receptor alpha mediates arsenic trioxide-induced apoptosis in acute promyelocytic leukemia. Mol Cell Biol 1999; 19:5170–5178.
Azuma Y, Arnaoutov A, Dasso M . SUMO-2/3 regulates topoisomerase II in mitosis. J Cell Biol 2003; 163:477–487.
Seufert W, Futcher B, Jentsch S . Role of a ubiquitin-conjugating enzyme in degradation of S- and M-phase cyclins. Nature 1995; 373:78–81.
al-Khodairy F, Enoch T, Hagan IM, Carr AM . The Schizosaccharomyces pombe hus5 gene encodes a ubiquitin conjugating enzyme required for normal mitosis. J Cell Sci 1995; 108 (Pt 2):475–486.
Jones D, Crowe E, Stevens TA, Candido EP . Functional and phylogenetic analysis of the ubiquitylation system in Caenorhabditis elegans: ubiquitin-conjugating enzymes, ubiquitin-activating enzymes, and ubiquitin-like proteins. Genome Biol 2002; 3: research0002.1–research0002.15.
Epps JL, Tanda S . The Drosophila semushi mutation blocks nuclear import of bicoid during embryogenesis. Curr Biol 1998; 8:1277–1280.
Nacerddine K, Lehembre F, Bhaumik M, et al. The SUMO pathway is essential for nuclear integrity and chromosome segregation in mice. Dev Cell 2005; 9:769–779.
Nowak M, Hammerschmidt M . Ubc9 regulates mitosis and cell survival during zebrafish development. Mol Biol Cell 2006; 17:5324–5336.
Tanaka K, Nishide J, Okazaki K, et al. Characterization of a fission yeast SUMO-1 homologue, pmt3p, required for multiple nuclear events, including the control of telomere length and chromosome segregation. Mol Cell Biol 1999; 19:8660–8672.
Zhang FP, Mikkonen L, Toppari J, et al. Sumo-1 function is dispensable in normal mouse development. Mol Cell Biol 2008; 28:5381–5390.
Elsen GE, Choi LY, Millen KJ, Grinblat Y, Prince VE . Zic1 and Zic4 regulate zebrafish roof plate specification and hindbrain ventricle morphogenesis. Dev Biol 2008; 314:376–392.
Tatham MH, Geoffroy MC, Shen L, et al. RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol 2008; 10:538–546.
Chupreta S, Holmstrom S, Subramanian L, Iniguez-Lluhi JA . A small conserved surface in SUMO is the critical structural determinant of its transcriptional inhibitory properties. Mol Cell Biol 2005; 25:4272–4282.
Rosendorff A, Sakakibara S, Lu S, et al. NXP-2 association with SUMO-2 depends on lysines required for transcriptional repression. Proc Natl Acad Sci USA 2006; 103:5308–5313.
Shen YC, Raymond PA . Zebrafish cone-rod (crx) homeobox gene promotes retinogenesis. Dev Biol 2004; 269:237–251.
Dekomien G, Epplen JT . The canine Recoverin (RCV1) gene: a candidate gene for generalized progressive retinal atrophy. Mol Vis 2002; 8:436–441.
Korzh V, Sleptsova I, Liao J, He J, Gong Z . Expression of zebrafish bHLH genes ngn1 and nrd defines distinct stages of neural differentiation. Dev Dyn 1998; 213:92–104.
Passini MA, Kurtzman AL, Canger AK, et al. Cloning of zebrafish vsx1: expression of a paired-like homeobox gene during CNS development. Dev Genet 1998; 23:128–141.
Vihtelic TS, Doro CJ, Hyde DR . Cloning and characterization of six zebrafish photoreceptor opsin cDNAs and immunolocalization of their corresponding proteins. Vis Neurosci 1999; 16:571–585.
Liao J, He J, Yan T, Korzh V, Gong Z . A class of neuroD-related basic helix-loop-helix transcription factors expressed in developing central nervous system in zebrafish. DNA Cell Biol 1999; 18:333–344.
Katsuyama Y, Oomiya Y, Dekimoto H, et al. Expression of zebrafish ROR alpha gene in cerebellar-like structures. Dev Dyn 2007; 236:2694–2701.
Bourdon JC, Fernandes K, Murray-Zmijewski F, et al. p53 isoforms can regulate p53 transcriptional activity. Genes Dev 2005; 19:2122–2137.
Gostissa M, Hengstermann A, Fogal V, et al. Activation of p53 by conjugation to the ubiquitin-like protein SUMO-1. EMBO J 1999; 18:6462–6471.
Rodriguez MS, Desterro JM, Lain S, et al. SUMO-1 modification activates the transcriptional response of p53. EMBO J 1999; 18:6455–6461.
Schmidt D, Muller S . Members of the PIAS family act as SUMO ligases for c-Jun and p53 and repress p53 activity. Proc Natl Acad Sci USA 2002; 99:2872–2877.
Wu SY, Chiang CM . Crosstalk between sumoylation and acetylation regulates p53-dependent chromatin transcription and DNA binding. EMBO J 2009; 28:1246–1259.
Santti H, Mikkonen L, Anand A, et al. Disruption of the murine PIASx gene results in reduced testis weight. J Mol Endocrinol 2005; 34:645–654.
Roth W, Sustmann C, Kieslinger M, et al. PIASy-deficient mice display modest defects in IFN and Wnt signaling. J Immunol 2004; 173:6189–6199.
Wong KA, Kim R, Christofk H, et al. Protein inhibitor of activated STAT Y (PIASy) and a splice variant lacking exon 6 enhance sumoylation but are not essential for embryogenesis and adult life. Mol Cell Biol 2004; 24:5577–5586.
Liu B, Mink S, Wong KA, et al. PIAS1 selectively inhibits interferon-inducible genes and is important in innate immunity. Nat Immunol 2004; 5:891–898.
Yamaguchi T, Sharma P, Athanasiou M, et al. Mutation of SENP1/SuPr-2 reveals an essential role for desumoylation in mouse development. Mol Cell Biol 2005; 25:5171–5182.
Cheng J, Kang X, Zhang S, Yeh ET . SUMO-specific protease 1 is essential for stabilization of HIF1alpha during hypoxia. Cell 2007; 131:584–595.
Saracco SA, Miller MJ, Kurepa J, Vierstra RD . Genetic analysis of SUMOylation in Arabidopsis: conjugation of SUMO1 and SUMO2 to nuclear proteins is essential. Plant Physiol 2007; 145:119–134.
Robu ME, Larson JD, Nasevicius A, et al. p53 activation by knockdown technologies. PLoS Genet 2007; 3:e78.
Chen J, Ng SM, Chang C, et al. p53 Isoform delta113p53 is a p53 target gene that antagonizes p53 apoptotic activity via BclxL activation in zebrafish. Genes Dev 2009; 23:278–290.
Zhang XD, Goeres J, Zhang H, et al. SUMO-2/3 modification and binding regulate the association of CENP-E with kinetochores and progression through mitosis. Mol Cell 2008; 29:729–741.
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF . Stages of embryonic development of the zebrafish. Dev Dyn 1995; 203:253–310.
Bennett CM, Kanki JP, Rhodes J, et al. Myelopoiesis in the zebrafish, Danio rerio. Blood 2001; 98:643–651.
Link V, Shevchenko A, Heisenberg CP . Proteomics of early zebrafish embryos. BMC Dev Biol 2006; 6:1.
Acknowledgements
We thank Dr. Jiang Zhu (Shanghai institute of hematology, Rui Jin hospital) and Dr. Nelly Kieffer (CNRS LIA, Rui Jin hospital) for their comments. This work was supported by grants from the National High Tech Program of China (863, 2006AA02Z150), the National Science Foundation of China (30525006), the Science and Technology Commission of Shanghai Municipality (07XD14022, 06PJ14068), ATIP program and BNP PARIBAS.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information, Figure S1
Expression patterns of zebrafish Ubc9. (PDF 41 kb)
Supplementary information, Figure S2
Morpholino efficacy assay. (PDF 205 kb)
Supplementary information, Table S1
Percentage of defective embryos of SUMO deficient zebrafish (PDF 12 kb)
Supplementary information, Figure S3
Comparison of the phenotypes induced by different SUMO MOs combinations. (PDF 454 kb)
Supplementary information, Figure S4
Characterization of transactivation property of p53 SUMO defective mutant p53 K386R. (PDF 34 kb)
Supplementary information, Table S2
Primer sequences for cloning (PDF 32 kb)
Supplementary information, Table S3
Primer sequences of quantitative real-time PCR (PDF 59 kb)
Rights and permissions
About this article
Cite this article
Yuan, H., Zhou, J., Deng, M. et al. Small ubiquitin-related modifier paralogs are indispensable but functionally redundant during early development of zebrafish. Cell Res 20, 185–196 (2010). https://doi.org/10.1038/cr.2009.101
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.101
Keywords
This article is cited by
-
SUMOylation controls stem cell proliferation and regional cell death through Hedgehog signaling in planarians
Cellular and Molecular Life Sciences (2018)
-
Sumoylation of CCAAT/enhancer-binding protein α is implicated in hematopoietic stem/progenitor cell development through regulating runx1 in zebrafish
Scientific Reports (2015)
-
CCAAT/enhancer-binding protein α is required for hepatic outgrowth via the p53 pathway in zebrafish
Scientific Reports (2015)
-
Mutation of kri1l causes definitive hematopoiesis failure via PERK-dependent excessive autophagy induction
Cell Research (2015)
-
Overexpression of SUMO perturbs the growth and development of Caenorhabditis elegans
Cellular and Molecular Life Sciences (2011)