Abstract
MiR-133 was found to be specifically expressed in cardiac and skeletal muscle in previous studies. There are two members in the miR-133 family: miR-133a and miR-133b. Although previous studies indicated that miR-133a was related to myogenesis, the signaling pathways regulated by miR-133 were still not very clear. In this study, we showed that both miR-133a and miR-133b were upregulated during myogenesis through Solexa sequencing. We confirmed that miR-133 could promote myoblast differentiation and inhibit cell proliferation through the regulation of the extracellular signal-regulated kinase (ERK) signaling pathway in C2C12 cells. FGFR1 and PP2AC, which both participate in signal transduction of the ERK1/2 pathway, were found to be negatively regulated by miR-133a and miR-133b at the post-transcriptional level. Also, downregulation of ERK1/2 phosphorylation by miR-133 was detected. FGFR1 and PP2AC were also found to repress C2C12 differentiation by specific siRNAs. In addition, we found that inhibition of ERK1/2 pathway activity can inhibit C2C12 cell proliferation and promote the initiation of differentiation but form short and small myotubes. Furthermore, we found that the expression of miR-133 was negatively regulated by ERK1/2 signaling pathway. In summary, we demonstrated the role of miR-133 in myoblast and further revealed a new feedback loop between miR-133 and the ERK1/2 signaling pathway involving an exquisite mechanism for regulating myogenesis.
Similar content being viewed by others
Main
MicroRNAs (miRNAs), a class of small, non-coding RNAs (∼22 nt in length), can repress gene expression at the post-transcriptional level through interaction with the 3′-untranslated region (3′-UTR) of target mRNA.1 Previous studies found miRNA involved in many biological processes (such as tissue development, metabolism and cancer).2 The first identified miRNA, lin-4, was found to regulate positively larval developmental timing in Caenorhabditis elegans.3, 4 Subsequent studies showed that miRNAs participated in many developmental processes, such as early embryonic development, neuronal development, muscle development and lymphocyte development.5 Several miRNAs were identified as important regulators of skeletal muscle development in recent years, including miR-1, miR-133, miR-206, miR-181, miR-24, miR-26 and miR-29.6, 7, 8, 9, 10, 11, 12, 13
The miR-133 family has two members, namely, miR-133a and miR-133b. The sequences of these miRNAs are different only at the last nucleotide of the 3′ terminus, but they are from different chromosomes in mouse. Mmu-miR-133a is encoded by mmu-mir-133a-1 and mmu-mir-133a-2, which are located on chromosome 18 and chromosome 2, respectively. Mmu-miR-133b is encoded by mmu-mir-133b, which is located on chromosome 1. The expression of miR-133a and miR-133b was first observed in microarrays to be significantly upregulated during myoblast differentiation. Northern blot analysis using completely complementary probes to miR-133a demonstrated that miR-133 was specifically expressed in cardiac and skeletal muscle.7 A functional study in zebrafish found actin organization in the sarcomere was disrupted after injecting an miR-133 inhibitor (completely complementary to miR-133a and miR-133b) into zebrafish embryos.14 While, the results of a study that focused on miR-133a function showed that miR-133a inhibited the formation of myotubes (differentiation) and promoted myoblast proliferation by targeting serum reaction factor.7 A recent study reported miR-133 suppressed myoblast proliferation by targeting SP1.15 The function of miR-133 in myoblast seemed controversial. The regulatory roles between miR-133 and the important pathways in myogenesis still need further investigation.
To further investigate the molecular mechanism of regulating myogenesis by miR-133, we identified target genes of miR-133 during myogenesis. Predicted targets of miR-133 were identified using the program TargetScan (http://www.targetscan.org/). Among hundreds of predicted target genes of miR-133, we found two genes involved in the regulation of extracellular signal-regulated kinase 1/2 (ERK1/2) activity that had a high aggregate PCT score. Those genes are fibroblast growth factor receptor 1 (FGFR1) and protein phosphatase 2 A catalytic subunit (PP2AC, including Ppp2ca and Ppp2cb). Both genes participated in the signal transduction of the ERK1/2 cascade.16, 17 Previous studies showed that the activated ERK1/2 pathway negatively regulates myoblast differentiation. The ERK1/2 pathway is a part of the mitogen-activated protein kinase (MAPK) pathway, which mainly participates in signal transduction by responding to extracellular stimuli that regulate many facets of cellular processes.18 Upstream signals activate phosphorylation of ERK1/2, giving ERK1/2 the ability to regulate cell proliferation and differentiation.18 In C2C12 cells, activity of ERK1/2 was significantly downregulated after initiation of differentiation. Inhibition of ERK1/2 activity by overexpression of MAPK phosphatase-1 promotes differentiation of C2C12 cells.19 Another study found that myostatin could repress differentiation of C2C12 cells by activating ERK1/2 phosphorylation.20
In this study, to explore the role of miR-133 in myogenesis, we analyzed how miR-133 influences myoblast proliferation and differentiation by gain- and loss-of-function studies in C2C12 cells. Our study is the first to investigate the regulation of ERK1/2 activity by miR-133 family members in myoblast.
Results
MiR-133 represses the myoblast proliferation and promotes its differentiation
We performed both overexpression and loss-of-function studies for miR-133 in C2C12 cells to confirm the role of miR-133 in myogenesis. Double-stranded RNA mimics with miR-133a or miR-133b sequence were used to mimic miR-133 function. The expression level of miR-133 in C2C12 cells transfected with miR-133a mimics or miR-133b mimics was significantly higher than that of the control (Figures 1a and b). And, a specific 2′-O-methyl antisense oligonucleotide inhibitor of miR-133b was used to knockdown the expression of miR-133 in myoblasts. MiR-133b inhibitor could interact with both miR-133a (Figure 1a) and miR-133b (Figure 1b), and significantly knocked down endogenous miR-133 expression in myoblasts (Figure 2a). Following transfection of miR-133a/b mimics or miR-133b inhibitor into C2C12 cells, myoblast cells were subjected to cell cycle analysis 24 h after transfection or were switched to differentiation medium for a predetermined time, after which the expression of myogenic marker genes (myogenin and MyHC) was analyzed to investigate the role of miR-133 in myoblast differentiation.
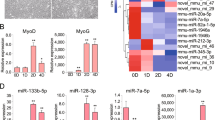
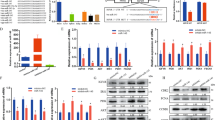
Overexpression of miR-133 repressed myoblast proliferation and promoted cell differentiation. (a and b) Expression of miR-133 in C2C12 cells after transfection with miR-133a mimics, miR-133b mimics or miR-133b inhibitor. MiR-133a mimics, miR-133b mimics, Duplex NC or miR-133b inhibitor were transfected into C2C12 cells, respectively. After transfection, cells were switched into the growth medium (GM) for 24 h. Then, total RNA of transfected cells was isolated, and the expression of miR-133 was analyzed by qPCR. (c) Overexpression of miR-133a/b repressed myoblast proliferation. C2C12 cells transfected with miR-133a mimics, miR-133b mimics or Duplex NC were collected for cell cycle analysis approximately 30 h after transfection. Cells were stained by propidium iodide and subjected to fluorescence-activated cell sorting analysis by flow cytometry. (d) Overexpression of miR-133a/b promoted C2C12 cell differentiation. Duplex NC, miR-133a mimics or miR-133b mimics were transfected into C2C12 cells, respectively. After transfection, cells were switched into the GM for 24 h. Then, cells were transferred to the differentiation medium (DM) for another 4 days before immunostaining for MyHC (green). Nuclei were stained with DAPI. (e) Area of myotubes stained in (d). Myotube area was calculated by Image Pro Plus software. Error bars represent S.D. from three independent experiments (*P<0.05; **P<0.01)
Knockdown of miR-133 repressed myoblast differentiation and promoted cell proliferation. (a) In myoblasts and myotubes, miR-133b inhibitor significantly repressed endogenous expression of miR-133. Inhibitor NC and miR-133b inhibitor were transfected into C2C12 cells cultured in six-well plates. Total RNA of transfected cells was isolated from cells cultured in the growth medium (GM) for 24 h or differentiation medium (DM) for 36 h after transfection. Expression of miR-133 was analyzed by qPCR. (b) Expression of myogenin and MyHC-2d was significantly downregulated by miR-133b inhibitor. Total RNA of C2C12 cells transfected with miR-133b inhibitor or inhibitor NC and cultured in the DM for 36 h after transfection was isolated, and the expression of myogenin and MyHC-2d was analyzed by qPCR. (c) Inhibition of miR-133 expression promoted myoblasts’ proliferation. C2C12 cells transfected with miR-133b inhibitor or inhibitor NC were collected for cell cycle analysis approximately 30 h after transfection. Cells were stained by propidium iodide and subjected to fluorescence-activated cell sorting analysis by flow cytometry. (d) Inhibition of miR-133 expression repressed myoblasts differentiation. Inhibitor NC or miR-133b inhibitor was transfected into C2C12 cells cultured in 24-well plates, respectively. After transfection, cells were switched into the GM for 24 h. Then, cells were transferred to the DM for another 4 days before immunostaining for MyHC (green). Nuclei were stained with DAPI. (e) Area of myotubes stained in (d). Myotube area was calculated by Image Pro Plus software. Error bars represent S.D. from three independent experiments (*P<0.05)
The results of cell cycle analysis showed that both miR-133a mimics and miR-133b mimcs could result in increasing the percentage of C2C12 cells in the G1 phase, and decreasing the percentage of cells in the S phase (Figure 1c), whereas the percentage of C2C12 cells in the G1 phase was significantly decreased, and the percentage of cells in the S phase was significantly increased, with miR-133 knockdown (Figure 2c). The expression of MyHC was induced (Figure 1d) with miR-133 overexpression in C2C12 cells, whereas the expression of MyHC was significantly repressed by miR-133b inhibitor when cells were cultured in the differentiation medium for 36 h after transfection (Figures 2c and d). The expression of myogenin was also significantly repressed by miR-133b inhibitor during myoblasts differentiation (Figure 2c). The myotube area in cells transfected with miR-133a mimics or miR-133b mimcs was significantly greater than that of control (Figure 1e), whereas myotube area in cells transfected with miR-133b inhibitor was significantly smaller than that of control (Figure 2e). Taken together, these results indicate that miR-133 functions in repressing proliferation of myoblasts and promoting cell differentiation.
Expression levels of FGFR1 and PP2AC were downregulated during C2C12 cells differentiation, whereas miR-133 was upregulated
To compare the expression of miR-133, FGFR1 and PP2AC during myogenesis, total RNA and protein of C2C12 cells cultured in the growth medium (0d) and differentiation medium for 1 day (1d), 2 days (2d) or 4 days (4d) was isolated, respectively, for gene expression analysis. Expression levels of miR-133 and mRNA of FGFR1 and PP2AC were analyzed by reverse transcriptase-polymerase chain reaction (RT-PCR) and real-time quantitative PCR (qPCR). Simultaneously, expression levels of miR-133a and miR-133b were analyzed by Solexa sequencing. Expression levels of myogenic marker genes (myogenin and MyHC-2d) mRNA were detected simultaneously as a positive control of myoblast differentiation. Protein expression levels of FGFR1, PP2AC and phospho-ERK1/2 (p-ERK1/2) were analyzed by western blotting, whereas the expression of tubulin and ERK1/2 protein was used as control.
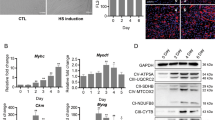
Exposure of C2C12 cells to the differentiation medium for increasing periods of time resulted in a concomitant increase in the formation of myotubes (Figure 3a) and an upregulation of myogenin and MyHC-2d expression (Figure 3c), which indicated normal differentiation of C2C12 cells. The result of Solexa sequencing analysis showed that miR-133a and miR-133b had similar expression patterns, and were both upregulated during myogenesis (Figure 3b). The miR-133 qPCR result was in concordance with that of the Solexa sequencing analysis (Figure 3c). The differences in expression levels of FGFR1 and PP2AC mRNA were not significant among all four samples (Figure 3c); however, the protein levels of FGFR1 and PP2AC were significantly downregulated during C2C12 cell differentiation (Figure 3d). These results suggested that the expression of FGFR1 and PP2AC were regulated at the post-transcriptional level during myogenesis. Furthermore, the expression trends of FGFR1, PP2AC and p-ERK1/2 (Figure 3d) were opposite to that of miR-133 during C2C12 cell differentiation, suggesting that miR-133 might be involved in regulating the ERK pathway by targeting FGFR1 and PP2AC.
Expression of miR-133, FGFR1, PP2AC and p-ERK1/2 during myoblast differentiation. (a) Morphologies of C2C12 cells cultured in the growth medium (0d), or differentiation medium for 1 day (1d), 2 days (2d) or 4 days (4d). (b) Expression of miR-133a and miR-133b during C2C12 differentiation. Total RNA of C2C12 cells collected at the four time points (0d, 1d, 2d and 4d) was isolated, and the expression of miR-133a and miR-133b was analyzed by Solexa sequencing. (c) Expression levels of FGFR1 and PP2AC mRNA during C2C12 differentiation did not significantly change, whereas the expression of miR-133 was significantly upregulated. Expression of miR-133 and other genes during C2C12 differentiation (using RNA samples from (b)) were analyzed by qPCR. (d) Protein of FGFR1 and PP2AC was downregulated during C2C12 differentiation, whereasile p-ERK1/2 was also downregulated. Total protein from C2C12 cells collected at the four time points (0d, 1d, 2d and 4d) was isolated, and protein expression of FGFR1, PP2AC and p-ERK1/2 was analyzed by western blotting, detecting tubulin and total ERK1/2 as controls. Error bars represent S.D. from three independent experiments (**P<0.01)
MiR-133 repressed ERK1/2 activity by targeting FGFR1 and PP2AC
The results of miR-133 target gene prediction by TargetScan showed that miR-133a and miR-133b had identical targets because of the same ‘seed region’. MiR-133a/b had two binding sites in the 3′-UTR of FGFR1, one binding site in the 3′-UTR of Ppp2ca and one binding site in the 3′-UTR of Ppp2cb (Figure 4a). To validate whether miR-133a and miR-133b directly targets these genes, we conducted a luciferase reporter assay.
MiR-133 targeted FGFR1 and PP2AC and downregulated p-ERK1/2 during myoblast differentiation. (a) Schematic diagram of five luciferase reporter constructs (psi-FGFR1, psi-F260, psi-F490, psi-Ppp2ca and psi-Ppp2cb) and four mutated reporter constructs (psi-FGFR1 Mut1, psi-FGFR1 Mut2, psi-Ppp2ca Mut and psi-Ppp2cb Mut). (b) MiR-133a/b, but not Duplex NC or miR-214, significantly inhibited the activities of the five luciferase reporters (as in (a)). Plasmids of the five reporters (as in (a)) were co-transfected into BHK-21 cells with miR-133a mimics, miR-133b mimics, miR-214 mimics or Duplex NC. Luciferase activity in treated cells was measured about 30 h after transfection. (c) MiR-133a/b did not inhibit the activities of the four mutated reporters (as in (a)). The experiments were performed as described in (b). (d) MiR-133a/b repressed the expression of FGFR1 and PP2AC, and also repressed the phosphorylation of ERK1/2 during myoblast differentiation. In addition, miR-133a mimics, miR-133b mimics and Duplex NC were transfected into C2C12 cells. Next, cells were switched into the differentiation medium approximately 6 h after transfection. The total protein from transfected cells was isolated about 24 h after culturing in the differentiation medium. Expression of the indicated proteins was analyzed by western blotting. Error bars represent S.D. from three independent experiments (**P<0.01)
A luciferase reporter construct, psi-FGFR1 (Figure 4a), was constructed by inserting a fragment of FGFR1 3′-UTR that included miR-133a/b and miR-214 binding sites into the luciferase reporter vector psi-CHECK-2. Next, the construct was co-transfected into BHK-21 cells with miR-133a mimics and miR-133b mimics, miR-214 mimics or Duplex NC. The luciferase assay showed that the luciferase activity was significantly repressed by miR-133a mimics and miR-133b mimics (P<0.01) but not by miR-214 mimics (Figure 4b). Luciferase reporter constructs psi-F260 (Figure 4a) containing a fragment of FGFR1 3′-UTR in which the miR-133a/b seed region binds from 260 to 266 bp or psi-F490 (Figure 4a) containing a fragment of FGFR1 3′-UTR in which the miR-133a/b seed region binds from 490 to 496 bp was co-transfected into BHK cells with miR-133b mimics or Duplex NC, respectively. The resulting luciferase activities from constructs psi-F260 and psi-F490 were significantly repressed by miR-133b mimics (P<0.01; Figure 4b). The luciferase activity from the luciferase reporter constructs psi-Ppp2ca (Figure 4a), containing a fragment of the Ppp2ca 3′-UTR that included the miR-133a/b binding site and psi-Ppp2cb (Figure 4a), containing a fragment of the Ppp2cb 3′-UTR that included miR-133a/b and miR-214 binding sites, were also significantly repressed by miR-133a mimics and miR-133b mimics (P<0.01; Figure 4b). However, miR-133a mimics and miR-133b mimics did not inhibit luciferase activity of mutated reporters (psi-FGFR1 Mut1, psi-FGFR1 Mut2, psi-Ppp2ca Mut and psi-Ppp2cb Mut) (Figure 4c), in which binding sites of miR-133a/b were changed (Figure 4a).
After the luciferase reporter assay, we performed western blotting to analyze whether miR-133a and miR-133b could repress protein expression of FGFR1 and PP2AC. C2C12 cells cultured in the growth medium were transfected individually with miR-133a mimics, miR-133b mimics, Duplex NC or both miR-133a mimics and miR-133b mimics. Approximately 6 h after transfection, cells were switched into the differentiation medium. Total protein from transfected cells was extracted approximately 24 h after culturing in the differentiation medium to detect the expression of FGFR1and PP2AC by western blotting. Phosphorylation of ERK1/2 was analyzed simultaneously. The expression levels of the FGFR1 and PP2AC proteins were repressed by both miR-133a mimics and miR-133b mimics compared with the Duplex NC group. ERK1/2 phosphorylation was also repressed with overexpression of miR-133 (Figure 4c).
Collectively, these results demonstrate that both miR-133a and miR-133b could repress expression of FGFR1 and PP2AC by binding to their 3′-UTRs during myoblast differentiation. Accordingly, phosphorylation of ERK1/2 was also repressed by miR-133a/b during myogenesis.
Knockdown of FGFR1 and PP2AC by siRNA promoted C2C12 differentiation
To evaluate the role of FGFR1 and PP2AC in C2C12 differentiation, specific siRNAs for FGFR1 or PP2AC were transfected into C2C12 cells. Firstly, we used four specific siRNAs for each of them to find the most effective one. The results showed that most of the siRNAs that we used could significantly knockdown the expression of FGFR1 (P<0.01; Figure 5a) or PP2AC (P<0.01; Figure 5b) at the mRNA level. We chose the si-F2 and si-P3 as the most effective siRNAs for FGFR1 and PP2AC, respectively. Results of western blotting indicated that the expression of FGFR1 was significantly reduced by si-F2 (Figure 5c), and the expression of PP2AC was significantly reduced by si-P3 (Figure 5d). Then, si-F2 and si-P3 were transfected into C2C12 cells, respectively, and cells were switched into the differentiation medium 24 h after transfection. Five days after transfection, cells were collected and expressions of myogenic genes were analyzed. The results showed that the expression level of MyHC, myogenin and MCK in cells transfected with si-P3 were all significantly higher than that of control (P<0.05), whereas only the expression level of MCK in cells transfected with si-F2 was significantly higher than that of the control (P<0.05), and the expression of MyHC and myogenin had no significant difference in cells transfected with si-F2 and NC (Figure 5e). These results suggested that FGFR1 and PP2AC may take part in repression of C2C12 differentiation.
Knockdown of FGFR1 and PP2AC by siRNA promoted C2C12 differentiation. (a) Three of four siRNAs significantly reduce mRNA expression of FGFR1 in C2C12 cells. The four siRNAs were transfected into C2C12 cells, respectively. One day after transfection, cells were collected and the expression of FGFR1 was analyzed by qPCR. (b) Three of four siRNAs significantly reduce mRNA expression of PP2AC in C2C12 cells. Experiments were conducted as described in (a). (c and d) Expression of FGFR1 and PP2AC were significantly reduced by si-F2 or si-P3, respectively. siRNA (si-F2 or si-P3) was transfected into C2C12 cells, and cells were collected 24 h after transfection. Expression of FGFR1 and PP2AC proteins was analyzed by western blotting. (e) Expression of MyHC, myogenin and MCK in cells transfected with si-F2, si-P3 or NC. Error bars represent S.D. from three independent experiments (*P<0.05; **P<0.01; ***P<0.001)
Inhibition of the ERK1/2 pathway enhanced the expression of miR-133 and induced myoblast differentiation; however, the activated ERK1/2 pathway was indispensable for early myogenic processes in myoblasts
To investigate how ERK1/2 pathway inhibition influences proliferation and differentiation of myoblasts and expression of miR-133, we used PD98059 to inhibit ERK1/2 pathway in C2C12 cells. DMSO was used to treat cells as a negative control. C2C12 cells cultured in the growth medium were treated with PD98059 or DMSO for 24 h, and cells were subsequently either subjected to cell cycle analysis or total RNA extraction for miR-133 and myogenic marker gene expression analysis. We found that cells treated with PD98059 were thinner and longer than control cells (Figure 6a), shown by morphologies of myoblasts initiating differentiation. After PD98059 treatment, the percentage of cells in the G1 phase significantly increased, and the percentage of cells in the S phase was significantly decreased, which showed that the proliferation of cells was repressed (Figure 6b). The expression levels of myogenin and miR-133 in cells treated with PD98059 were significantly higher than that of the control (P<0.01; Figure 6c). These results showed that inhibition of the ERK1/2 pathway in myoblast cells could result in a cell cycle arrest and induce differentiation simultaneously, whereas the expression of miR-133 was upregulated concomitantly.
Inhibition of the ERK1/2 pathway in C2C12 cells cultured in the growth medium significantly repressed cell proliferation and promoted the initiation of cell differentiation. (a) Cells treated with PD98059 were thinner and longer than control cells, showing morphologies of myoblasts initiating differentiation. C2C12 cells cultured in the growth medium (about 30% confluent) were treated with PD98059 (25 nM) or dimethyl sulfoxide (DMSO) (0.2%, control) for approximately 24 h. Morphologies of cells were observed by microscopy 24 h after treatment. (b) PD98059 inhibited myoblast proliferation. C2C12 cells treated as in (a) were subjected to cell cycle analysis by flow cytometry. (c) Expression of myogenin and miR-133 was upregulated in cells treated with PD98059. Total RNA from C2C12 cells was treated (as in (a)) and isolated, and the expression of myogenin and miR-133 was analyzed by qPCR. Error bars represent S.D. from three independent experiments (**P<0.01)
C2C12 cells were switched to the differentiation medium with either PD98059 or DMSO when cells were cultured to be approximately 70% confluent. The expression levels of myogenin (Figure 7a), MyHC-2d (Figure 7b) and miR-133 (Figure 7c) in cells treated with PD98059 were all significantly higher than those of the control (P<0.05) 20 or 36 h after treatment. However, 72 or 96 h after treatment, we found cells treated with PD98059 formed shorter and smaller myotubes than the control cells, shown by immunostaining of MyHC (Figure 7d). These results indicate that inhibition of the ERK1/2 pathway accelerated cell differentiation, which might result in the formation of shorter and smaller myotubes.
Inhibition of the ERK1/2 pathway in C2C12 cells accelerated cell differentiation during the early period of differentiation, which resulted in forming shorter and smaller myotubes. C2C12 cells cultured in the differentiation medium were treated with PD98059 (25 nM) or dimethyl sulfoxide (DMSO) (0.2%, control). Total RNA of cells treated for 20 or 36 h was isolated, respectively. Expression of myogenin (a), MyHC-2d (b) and miR-133 (c) was analyzed by qPCR. Then, differentiation of cells treated with PD98059 (25 nM) or DMSO (0.2%, control) for 72 or 96 h were monitored by immunostaining for MyHC (green). Nuclei were stained with DAPI (d). Error bars represent S.D. from three independent experiments (*P<0.05; **P<0.01)
Taken together, these results suggested that the expression of miR-133 was negatively regulated by ERK1/2 pathway, whereas ERK1/2 pathway was necessary for the early myogenic process such that enough myoblasts could be accumulated for subsequent cell fusion to form normal myotubes.
In summary, our results indicated a feedback loop between miR-133 and the ERK1/2 pathway involving an exquisite mechanism for regulating myogenesis (Figure 8). Activation of the ERK1/2 pathway maintains cell proliferation to accumulate enough myoblasts, whereas the expression of miR-133 is inhibited by activated ERK1/2. After initiation of differentiation, the expression of miR-133 is upregulated. Next, miR-133 represses myoblast proliferation and promotes cell differentiation by repressing the activity of the ERK1/2 pathway through targeting FGFR1 and PP2AC.
Discussion
In this study, we confirmed that both miR-133a and miR-133b have important roles in repressing myoblast proliferation and promoting cell differentiation during myogenesis in C2C12 cells, an in vitro model for skeletal muscle development.21 These roles are also in concordance with the expression pattern of miR-133 during C2C12 cell differentiation. In our previous study, we analyzed miRNA expression profiles in porcine fetal and adult longissimus muscle. We found that miR-133 had a high expression level in both fetal and adult longissimus muscle,22 suggesting that miR-133 might participate in more regulatory processes during skeletal muscle development.
We identified two new targets of miR-133 in myoblast cells, namely, FGFR1 and PP2AC. The differences in expression between mRNA and protein during C2C12 cell differentiation suggested that their expression might be regulated at the post-transcriptional level. Results from the luciferase reporter analysis and western blotting demonstrated that miR-133 directly targets FGFR1 and PP2AC by interaction with their 3′-UTRs. FGFR1 is one of the two FGFRs expressed in muscle cells.23, 24, 25, 26 Overexpression of FGFR1 in mouse myocytes promoted cell proliferation and delayed differentiation; conversely, the expression of mutated FGFR1 enhanced cell differentiation.27 The role of PP2AC in myoblast processes has yet to be investigated. In this study, we found knockdown of FGFR1 and PP2AC by specific siRNAs promoted C2C12 differentiation, which suggested that they may repress myoblast differentiation. Thus, it is possible that miR-133 influences myogenesis by repressing the expression of FGFR1 and PP2AC. When we prepared this manuscript, Belevych et al.28 reported that miR-133a could target and repress the expression of PP2AC in myocardial cell, which gave us more confidence of the role we presented here.
We observed a down-egulation of ERK1/2 phosphorylation along with a downregulation of protein levels for FGFR1 and PP2AC by miR-133 during C2C12 cell differentiation. FGF signal transduction and activation of ERK1/2 by FGFR1 was found to be notably important for proliferation of skeletal muscle cells. In fibroblasts, FGFs bind to FGFRs and activate the intracellular ERK1/2 cascade to maintain cell proliferation.16 In skeletal muscle cells, basic FGF could stimulate cell proliferation and repress myotube formation by activating ERK1/2.29, 30, 31, 32, 33 The expression of truncated FGFR1 (a mutant FGFR1 lacking the intracellular domain) in mouse muscle cells decreased cell responsiveness to exogenously supplied FGF-1 and repressed cell proliferation.29 Thus, we concluded that miR-133 functioned in myoblasts by repressing the activity of the ERK1/2 pathway through targeting FGFR1.
PP2AC is the catalytic subunit of PP2A holoenzyme that comprises a family of serine/threonine phosphatases and has a role in the dephosphorylation of protein.17 It is currently unknown how ERK1/2 activity is regulated by PP2A in muscle cells. Results of in vitro studies in other cell lines showed that PP2A could positively regulate the activity of the ERK1/2 pathway by activating Raf1, which is upstream of MEK1/2 in the ERK1/2 cascade. In COS cells, the A and C subunits of the PP2A holoenzyme were found to combine with Raf1 by immunoprecipitation.34 Raf1 was activated by dephosphorylation at serine 259 by PP2A.34, 35, 36, 37 A recent study in 293T cells found that PP2A positively regulated Raf1-MEK1/2-ERK1/2 signaling.38 We proposed that PP2A could also positively regulate the ERK1/2 signaling pathway in myoblasts such that ERK1/2 phosphorylation was downregulated, whereas the expression of PP2AC protein was repressed by miR-133b during C2C12 cell differentiation (Figure 4d).
Following functional studies and identification of target genes of miR-133, we analyzed whether miR-133 expression was regulated by the ERK1/2 pathway in myoblasts. The results showed that the expression of miR-133 was significantly upregulated by inactivation of the ERK1/2 signal during myoblast proliferation or differentiation. Simultaneously, we observed the effect on myoblast proliferation and differentiation after inhibition of ERK1/2 activity. We found that blocking the ERK1/2 pathway in C2C12 cells resulted in a cell cycle arrest and induction of cell differentiation and finally induced the formation of shorter, smaller myotubes. These results were concordant with results of a study interrupting FGF signaling in chicken embryos. In that study, chicken embryos ectopically expressed a truncated FGFR1-formed skeletal muscles with a lower myofiber density and weight. The primary muscle cells expressed a truncated FGFR1-formed myotubes with fewer myonuclei than the controls.39 Another study on ERK1/2 also found that knockdown of ERK2 significantly repressed the formation of multinucleated myotubes.40 Results of a study on analyzing skeletal muscle cell differentiation in vitro showed that fusion of muscle cells occurred 24 h after being cultured in the differentiation medium, and muscle cells mainly completed proliferation before the initiation of differentiation. Cells would continue to proliferate if they did not fuse into myotubes.41 Thus, activation of the ERK1/2 pathway is necessary in early myogenesis for proliferation of a sufficient number of myoblasts to form myotubes.
In conclusion, we confirmed the role of miR-133 in myoblasts and further revealed a new feedback loop between miR-133 and the ERK1/2 signaling pathway involving an exquisite mechanism for regulating myogenesis.
Materials and Methods
Cell culture
The C2C12 myoblast cell line was used to analyze the function of miR-133b during myogenesis. BHK-21 was used for luciferase reporter analysis. Cells stored in liquid nitrogen were thawed at 37 °C and cultured in the growth medium (DMEM; Hyclone, Logan, UT, USA) supplemented with fetal bovine serum (10%) at 37 °C in a humidified 5% CO2 atmosphere. To induce differentiation, C2C12 cells were switched to the differentiation medium (DMEM, 2% horse serum). The MEK1 inhibitor PD98059 (Beyotime, Haimen, China) was used to inhibit the activity of the ERK1/2 pathway in C2C12 cells. Approximately 24 h after aliquoting cells into six-well plates, either PD98059 (25 nM) or DMSO (control, 0.2%) was added to the cell medium.
RNA oligonucleotides and transfection
All RNA oligonucleotides used in this study (miRNA mimics, miRNA inhibitor, siRNAs and control oligonucleotides) were synthesized from GenePharma (Shanghai, China). The oligonucleotides’ sequences are listed in Supplementary Table S1. Cells were aliquoted into plates 24 h before transfection. Corresponding RNA oligonucleotides (50 nM for miRNA mimics or miRNA inhibitor, 100 nM for siRNAs) were transfected into cells with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. For cell proliferation analysis, transfection was conducted when the cells were approximately 40% confluent. For cell differentiation analysis after overexpression miR-133, transfection was conducted when the cells were approximately 40% confluent. For cell differentiation analysis after inhibition of miR-133 expression, transfection was conducted when the cells were approximately 70% confluent.
Expression detection of miR-133a and miR-133b by Solexa sequencing
To distinguish the expression levels of miR-133a and miR-133b during myoblast differentiation, the Solexa sequencing method was performed. Generally, the total RNA was isolated from C2C12 cells cultured in the growth medium and the C2C12 cells cultured in the differentiation medium for 1 day, 2 days and 4 days, respectively. The Solexa sequencing was carried out at Beijing Genomics institute (BGI, Beijing, China) and the miRNA annotation was performed according to our previous study.42 The expression levels of miR-133a and miR-133b were represented using their relative read numbers of one million reads.
cDNA synthesis, RT-PCR and qPCR
Total cell RNA was extracted with Trizol (Invitrogen) using the manufacturer’s instructions. RNA quality was assessed by 0.8% agarose gel electrophoresis. Reverse transcription was performed with a ReverTra Ace kit (Toyobo, Tokyo, Japan) according to the manufacturer’s manual. A specific stem-loop primer was used as the reverse transcription primer to synthesize cDNA for miR-133 amplification (RT-miR-133: 5′-CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTAGCTGGT-3′). The stem-loop primer was designed as described previously.43 PCR conditions for RT-PCR for miR-133 and other gene expression analysis were 94 °C for 40 s, 62 °C for 30 s and 72 °C for 15 s for 28 cycles. Differential expression of miR-133 and other genes were confirmed by quantitative analysis.44 SYBR Green Realtime PCR Master Mix (Toyobo) was used for qPCR. Internal controls for expression analysis were U6 for miR-133 and GAPDH for other genes. Primers used for miRNA and gene expression analysis are listed in Supplementary Table S2. All real-time quantitative PCR experiments used a LightCycler 480 (Roche, Basel, Switzerland) with 2 min at 95 °C, followed by 40 cycles of amplification (20 s at 95 °C, 20 s at 62 °C and 15 s at 72 °C).
Cell cycle analysis
When treated cells in six-well plates were 70–80% confluent, cells were collected by trypsinizing and centrifuging at 1000 r.p.m. for 3 min. Cells were resuspended in 1 ml PBS and collected by centrifugation at 1000 r.p.m. for 3 min, with this step being repeated one or two times. After two to three PBS washes, cells were fixed with 1 ml cold 75% ethanol (in PBS) and stored at −20 °C for at least 8–10 h. Fixed cells were centrifuged as described above, and the supernatant was discarded. Cells were washed with PBS one to times times as above. After removing the PBS, cells were resuspended in 500 μl staining buffer (50 μg/ml propidium iodide, 100 μg/ml RNase A and 0.2% Triton X-100 in PBS) and incubated at 4 °C for 30 min (with protection from light). The cell cycle status was analyzed by flow cytometry (BD, Franklin Lakes, NJ, USA).
Luciferase reporter assay
To construct luciferase reporters, fragments containing miRNA binding sites for the 3′-UTR of target genes were cloned into the 3′-UTR of Renilla luciferase in psiCHECK-2 vector (Promega, Madison, WI, USA) using the XhoI and NotI restriction sites. Corresponding recognition sequences for XhoI and NotI were added to the 5′ terminus of either the forward or reverse primer used for amplifying 3′-UTR fragments of target genes, respectively. To produce mutated plasmids, site-directed mutation was used to introduce base substitution to the miR-133 binding site of luciferase reporters using Site-Directed Gene Mutagenesis Kit (Beyotime). Supplementary Table S2 lists detailed information of these primers.
BHK-21 cells were aliquoted into 24-well plates 24 h before transfection. Constructs (0.2 μg) and miRNA mimics (0.4 μg) were co-transfected into cells with Lipofectamine 2000 (1.5 μl) (Invitrogen). Luciferase activity in treated cells was measured approximately 24–30 h after transfection with the Dual-Luciferase Reporter Assay System (Promega) using the manufacturer’s instructions, whereas luminescence was quantified by luminometer (VICTOR X2; Perkin-Elmer, Waltham, MA, USA). In each sample, the signal value of Renilla luciferase must be normalized to its signal value of firefly luciferase. The interaction of miRNAs and target sites was analyzed by comparing normalized luciferase activity between cells transfected with miRNA mimics and Duplex NC.
Western blotting
Antibodies used were FGFR1 (Flg (C-15): sc-121; Santa Cruz Biotechnology), PP2AC (2038; Cell Signaling), ERK1/2 (p44/p42 MAPK, 9102; Cell Signaling), p-ERK1/2 (phospho-p44/p42 MAPK, 9101; Cell Signaling), tubulin (AT819; Beyotime), HRP-linked anti-rabbit IgG (7074; Cell Signaling) and HRP-linked anti-mouse IgG (A0216; Beyotime).
Total protein from cells cultured in 6-cm-diameter dishes was extracted with 300 μl RIPA lysis buffer (P0013B; Beyotime) supplemented with 1% protease inhibitor and 1% phosphatase inhibitor. Protein concentration was measured with BCA Protein Assay Kit (P0012S; Beyotime). Also, 5% stocking gel and 10% running gel were used for SDS-PAGE. Expression analysis of tubulin, PP2AC and ERK1/2 was performed with 25 μg protein, whereas 50 μg of protein was used for the expression analysis of FGFR1 and p-ERK1/2. Proteins were transferred to PVDF membranes (Millipore, Billerica, MA, USA; 0.45 μm for FGFR1 and 0.22 μm for others). Membranes were blocked for 2 h at room temperature in blocking buffer (1 × TBS with 5% non-fat dry milk and 0.1% Tween-20). Next, the membrane was incubated at 4 °C overnight with primary antibody at a 1 : 1000 dilution (FGFR1 and p-ERK1/2) or 1 : 2000 dilution (tubulin, ERK1/2 and PP2AC). Membranes were washed three to four times (10 min for each time) with washing buffer (1 × TBS with 0.1% Tween-20) and were incubated at room temperature for 1–2 h with HRP-conjugated secondary antibody at 1 : 3000 dilution (HRP-linked anti-rabbit IgG) or at 1 : 2000 dilution (HRP-linked anti-mouse IgG). Membranes were washed three to four times again (10 min for each time) with washing buffer (1 × TBS with 0.1% Tween-20), incubated with ECL reagent (Thermo Scientific, Waltham, MA, USA) and exposed to X-films (Kodak, Rochester, NY, USA). Lastly, the results of immunoblotting were displayed on X-films by developing and fixing.
Immunostaining
Cells treated in 24-well plates were fixed with 4% formaldehyde for 10 min, washed three times in PBS (5 min for each time). Then, cells were incubated with 0.3% Triton X-100 in PBS for 20 min, washed two times in PBS (5 min for each time). After that, cells were incubated with 1% BSA in PBS for 1 h. Next, cells were incubated with primary antibodies (anti-myosin, M4276; Sigma, St. Louis, MO, USA) for 2 h and washed two times in PBS (5 min for each time), and then incubated with secondary antibodies (Alexa Fluor 488; a10680; Invitrogen) for 1 h and washed two times in PBS (5 min for each time). 4′, 6-Diamidino-2-phenylindole dihydrochloride (DAPI) was then added for 2 min. After several washes with PBS, 10 μl of the antifading solution was added to each well. Finally, images of samples were captured on a fluorescence microscope. At least three representative images per sample were scored for myotube area (area occupied by myotubes relative to the total area) by Image Pro Plus software (Media Cybernetics, Rockville, MD, USA).
Statistical analysis
The significance of the difference between case and control data was analyzed by Student’s t-test. P<0.05 was considered to be significant.
Abbreviations
- miRNA:
-
microRNA
- ERK1/2:
-
extracellular signal-regulated kinase 1/2
- FGFR1:
-
fibroblast growth factor receptor 1
- PP2AC:
-
protein phosphatase 2A catalytic subunit
References
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Sun W, Julie Li YS, Huang HD, Shyy JY, Chien S . MicroRNA: a master regulator of cellular processes for bioengineering systems. Annu Rev Biomed Eng 2010; 12: 1–27.
Wightman B, Ha I, Ruvkun G . Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993; 75: 855–862.
Lee RC, Feinbaum RL, Ambros V . The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993; 75: 843–854.
Stefani G, Slack FJ . Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol 2008; 9: 219–230.
Sokol NS, Ambros V . Mesodermally expressed Drosophila microRNA-1 is regulated by Twist and is required in muscles during larval growth. Genes Dev 2005; 19: 2343–2354.
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 2006; 38: 228–233.
Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A . Muscle-specific microRNA miR-206 promotes muscle differentiation. J Cell Biol 2006; 174: 677–687.
Anderson C, Catoe H, Werner R . MIR-206 regulates connexin43 expression during skeletal muscle development. Nucleic Acids Res 2006; 34: 5863–5871.
Naguibneva I, Ameyar-Zazoua M, Polesskaya A, Ait-Si-Ali S, Groisman R, Souidi M et al. The microRNA miR-181 targets the homeobox protein Hox-A11 during mammalian myoblast differentiation. Nat Cell Biol 2006; 8: 278–284.
Sun Q, Zhang Y, Yang G, Chen X, Cao G, Wang J et al. Transforming growth factor-beta-regulated miR-24 promotes skeletal muscle differentiation. Nucleic Acids Res 2008; 36: 2690–2699.
Wong CF, Tellam RL . MicroRNA-26a targets the histone methyltransferase Enhancer of Zeste homolog 2 during myogenesis. J Biol Chem 2008; 283: 9836–9843.
Wang H, Garzon R, Sun H, Ladner KJ, Singh R, Dahlman J et al. NF-kappaB-YY1-miR-29 regulatory circuitry in skeletal myogenesis and rhabdomyosarcoma. Cancer Cell 2008; 14: 369–381.
Mishima Y, Abreu-Goodger C, Staton AA, Stahlhut C, Shou C, Cheng C et al. Zebrafish miR-1 and miR-133 shape muscle gene expression and regulate sarcomeric actin organization. Genes Dev 2009; 23: 619–632.
Zhang D, Li XH, Chen CC, Li YY, Lei Z, Jing YY et al. Attenuation of p38-mediated miR-1/133 Expression Facilitates Myoblast Proliferation during the Early Stage of Muscle Regeneration. PLoS One 2012; 7: e41478.
Campbell JS, Wenderoth MP, Hauschka SD, Krebs EG . Differential activation of mitogen-activated protein kinase in response to basic fibroblast growth factor in skeletal muscle cells. Proc Natl Acad Sci USA 1995; 92: 870–874.
Janssens V, Goris J . Protein phosphatase 2A: a highly regulated family of serine/threonine phosphatases implicated in cell growth and signalling. Biochem J 2001; 353 (Part 3): 417–439.
Pearson G, Robinson F, Beers Gibson T, Xu BE, Karandikar M, Berman K et al. Mitogen-activated protein (MAP) kinase pathways: regulation and physiological functions. Endocr Rev 2001; 22: 153–183.
Bennett AM, Tonks NK . Regulation of distinct stages of skeletal muscle differentiation by mitogen-activated protein kinases. Science 1997; 278: 1288–1291.
Yang W, Chen Y, Zhang Y, Wang X, Yang N, Zhu D . Extracellular signal-regulated kinase 1/2 mitogen-activated protein kinase pathway is involved in myostatin-regulated differentiation repression. Cancer Res 2006; 66: 1320–1326.
Yaffe D, Saxel O . Serial passaging and differentiation of myogenic cells isolated from dystrophic mouse muscle. Nature 1977; 270: 725–727.
Huang TH, Zhu MJ, Li XY, Zhao SH . Discovery of porcine microRNAs and profiling from skeletal muscle tissues during development. PLoS One 2008; 3: e3225.
Mitchell P, Steenstrup T, Hannon K . Expression of fibroblast growth factor family during postnatal skeletal muscle hypertrophy. J Appl Physiol 1999; 86: 313–319.
Sheehan SM, Allen RE . Skeletal muscle satellite cell proliferation in response to members of the fibroblast growth factor family and hepatocyte growth factor. J Cell Physiol 1999; 181: 499–506.
Martelly I, Soulet L, Bonnavaud S, Cebrian J, Gautron J, Barritault D . Differential expression of FGF receptors and of myogenic regulatory factors in primary cultures of satellite cells originating from fast (EDL) and slow (Soleus) twitch rat muscles. Cell Mol Biol (Noisy-le-Grand) 2000; 46: 1239–1248.
Templeton TJ, Hauschka SD . FGF-mediated aspects of skeletal muscle growth and differentiation are controlled by a high affinity receptor, FGFR1. Dev Biol 1992; 154: 169–181.
Scata KA, Bernard DW, Fox J, Swain JL . FGF receptor availability regulates skeletal myogenesis. Exp Cell Res 1999; 250: 10–21.
Belevych AE, Sansom SE, Terentyeva R, Ho HT, Nishijima Y, Martin MM et al. MicroRNA-1 and -133 increase arrhythmogenesis in heart failure by dissociating phosphatase activity from RyR2 complex. PLoS One 6: e28324.
Weyman CM, Wolfman A . Mitogen-activated protein kinase kinase (MEK) activity is required for inhibition of skeletal muscle differentiation by insulin-like growth factor 1 or fibroblast growth factor 2. Endocrinology 1998; 139: 1794–1800.
Jones NC, Fedorov YV, Rosenthal RS, Olwin BB . ERK1/2 is required for myoblast proliferation but is dispensable for muscle gene expression and cell fusion. J Cell Physiol 2001; 186: 104–115.
Milasincic DJ, Calera MR, Farmer SR, Pilch PF . Stimulation of C2C12 myoblast growth by basic fibroblast growth factor and insulin-like growth factor 1 can occur via mitogen-activated protein kinase-dependent and -independent pathways. Mol Cell Biol 1996; 16: 5964–5973.
Miralles F, Ron D, Baiget M, Felez J, Munoz-Canoves P . Differential regulation of urokinase-type plasminogen activator expression by basic fibroblast growth factor and serum in myogenesis. Requirement of a common mitogen-activated protein kinase pathway. J Biol Chem 1998; 273: 2052–2058.
de Alvaro C, Martinez N, Rojas JM, Lorenzo M . Sprouty-2 overexpression in C2C12 cells confers myogenic differentiation properties in the presence of FGF2. Mol Biol Cell 2005; 16: 4454–4461.
Abraham D, Podar K, Pacher M, Kubicek M, Welzel N, Hemmings BA et al. Raf-1-associated protein phosphatase 2A as a positive regulator of kinase activation. J Biol Chem 2000; 275: 22300–22304.
Ory S, Zhou M, Conrads TP, Veenstra TD, Morrison DK . Protein phosphatase 2A positively regulates Ras signaling by dephosphorylating KSR1 and Raf-1 on critical 14-3-3 binding sites. Curr Biol 2003; 13: 1356–1364.
Jaumot M, Hancock JF . Protein phosphatases 1 and 2A promote Raf-1 activation by regulating 14-3-3 interactions. Oncogene 2001; 20: 3949–3958.
Kubicek M, Pacher M, Abraham D, Podar K, Eulitz M, Baccarini M . Dephosphorylation of Ser-259 regulates Raf-1 membrane association. J Biol Chem 2002; 277: 7913–7919.
Adams DG, Coffee RL Jr, Zhang H, Pelech S, Strack S, Wadzinski BE . Positive regulation of Raf1-MEK1/2-ERK1/2 signaling by protein serine/threonine phosphatase 2A holoenzymes. J Biol Chem 2005; 280: 42644–42654.
Flanagan-Steet H, Hannon K, McAvoy MJ, Hullinger R, Olwin BB . Loss of FGF receptor 1 signaling reduces skeletal muscle mass and disrupts myofiber organization in the developing limb. Dev Biol 2000; 218: 21–37.
Li J, Johnson SE . ERK2 is required for efficient terminal differentiation of skeletal myoblasts. Biochem Biophys Res Commun 2006; 345: 1425–1433.
Bischoff R, Holtzer H . Mitosis and the processes of differentiation of myogenic cells in vitro. J Cell Biol 1969; 41: 188–200.
Xie SS, Li XY, Liu T, Cao JH, Zhong Q, Zhao SH . Discovery of porcine microRNAs in multiple tissues by a Solexa deep sequencing approach. PLoS One 2011; 6: e16235.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33: e179.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 2001; 25: 402–408.
Acknowledgements
This work was supported by the National Outstanding Youth Foundation and Youth foundation of NSFC (31025026; 30901020); National Basic Research Program of China (973 Project, 2012CB124702); and the National High Technology Plan of China (2013AA102502).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by E Candi
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Feng, Y., Niu, LL., Wei, W. et al. A feedback circuit between miR-133 and the ERK1/2 pathway involving an exquisite mechanism for regulating myoblast proliferation and differentiation. Cell Death Dis 4, e934 (2013). https://doi.org/10.1038/cddis.2013.462
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.462
Keywords
This article is cited by
-
Genome-wide analysis of circular RNA-mediated ceRNA regulation in porcine skeletal muscle development
BMC Genomics (2023)
-
Regulation of muscle stem cell fate
Cell Regeneration (2022)
-
miR-133a silencing rescues glucocorticoid-induced bone loss by regulating the MAPK/ERK signaling pathway
Stem Cell Research & Therapy (2021)
-
LncRNA IMFlnc1 promotes porcine intramuscular adipocyte adipogenesis by sponging miR-199a-5p to up-regulate CAV-1
BMC Molecular and Cell Biology (2020)
-
Chromatin accessibility is associated with the changed expression of miRNAs that target members of the Hippo pathway during myoblast differentiation
Cell Death & Disease (2020)