Abstract
Spinal muscular atrophy (SMA) is a genetic disorder characterized by degeneration of spinal cord motoneurons (MNs), resulting in muscular atrophy and weakness. SMA is caused by mutations in the Survival Motor Neuron 1 (SMN1) gene and decreased SMN protein. SMN is ubiquitously expressed and has a general role in the assembly of small nuclear ribonucleoproteins and pre-mRNA splicing requirements. SMN reduction causes neurite degeneration and cell death without classical apoptotic features, but the direct events leading to SMN degeneration in SMA are still unknown. Autophagy is a conserved lysosomal protein degradation pathway whose precise roles in neurodegenerative diseases remain largely unknown. In particular, it is unclear whether autophagosome accumulation is protective or destructive, but the accumulation of autophagosomes in the neuritic beadings observed in several neurite degeneration models suggests a close relationship between the autophagic process and neurite collapse. In the present work, we describe an increase in the levels of the autophagy markers including autophagosomes, Beclin1 and light chain (LC)3-II proteins in cultured mouse spinal cord MNs from two SMA cellular models, suggesting an upregulation of the autophagy process in Smn (murine survival motor neuron protein)-reduced MNs. Overexpression of Bcl-xL counteracts LC3-II increase, contributing to the hypothesis that the protective role of Bcl-xL observed in some SMA models may be mediated by its role in autophagy inhibition. Our in vitro experimental data indicate an upregulation in the autophagy process and autophagosome accumulation in the pathogenesis of SMA, thus providing a valuable clue in understanding the mechanisms of axonal degeneration and a possible therapeutic target in the treatment of SMA.
Similar content being viewed by others
Main
Spinal muscular atrophy (SMA) is a genetic neuromuscular disorder characterized by the degeneration of motoneurons (MNs) in the anterior horn of the spinal cord, resulting in muscular atrophy and weakness. SMA is caused by homozygous disruption of the Survival Motor Neuron 1 (SMN1) gene by deletion, conversion or mutation. In humans, the SMN gene is present in multiple copies, one SMN1 (telomeric) and several SMN2 (centromeric) on chromosome 5q13. SMN1 expresses a full-length transcript and SMN2 expresses primarily a truncated isoform that is unable to compensate SMN1 deficiency in SMA. SMN2 gene encodes for 10% of full-length SMN, but results in inadequate levels of SMN. It is well known that SMN protein level is critical to disease onset and severity, and is determined in part by the SMN2 copy numbers.1, 2, 3
One of the best characterized SMN functions is the assembly of small nuclear ribonucleoprotein and pre-mRNA splicing requirements, but the protein probably also has a particular role in MN axons that results in appropriate levels of certain transcripts in the distal axon and may contribute to SMA pathogenesis.4, 5, 6, 7, 8 The direct events leading to MN degeneration in SMA are still unknown. Although apoptotic cell death has been proposed,9, 10 no direct evidences of MN apoptosis have been reported that demonstrate the involvement of this process in SMA etiology. In some neurodegenerative diseases, the cell death features inducing neuronal loss do not fulfill the criteria of apoptosis,11 contributing to the hypothesis that other cell death processes may be mediating neuronal degeneration in these disorders.12
Autophagy is a highly regulated, self-destructive process that is important for differentiation, homeostasis and survival under physiological and pathological conditions.13 Cytosolic proteins and organelles are sequestered by a double membrane and the resulting vacuoles (autophagosomes) are delivered to lysosomes for degradation. Autophagy can either be generally involved in the turnover of long-lived protein and organelles, or specifically target distinct organelles (for example, mitochondria). Excessive autophagic activity can lead to self-destruction and cell death, but insufficient activity (or imbalanced autophagic flux) may also contribute to cell death.
Alteration of the autophagy pathway is associated with many neurodegenerative diseases and spinal cord injury.14, 15, 16, 17, 18, 19 Although it is generally believed that autophagy is beneficial to neuronal survival, its specific role in neurodegeneration remains unclear. In particular, it is unclear whether autophagosome accumulation is protective or destructive. For instance, accumulation of autophagosomes in the neuritic beadings observed in several neurite degeneration models suggests a close relationship between the autophagic process and neurite collapse.20, 21
The present study explored autophagy in a cellular model of SMA. Our previous work demonstrated that murine survival motor neuron protein (Smn) knockdown causes neurite degeneration and late cell death in MNs. This phenotype was rescued by the overexpression of the anti-apoptotic protein Bcl-xL.22 Here, we demonstrate increased autophagosomes and the autophagy-related proteins Beclin1 and light chain (LC)3-II in Smn-reduced MNs. The autophagosomes are localized in MNs soma and neurites. Together, these results suggest a deregulation of the autophagy process in these cells. Moreover, the LC3-II protein increase is prevented by Bcl-xL overexpression, suggesting that the protective role of this protein observed in some SMA models can be mediated through its involvement in autophagy inhibition. Our present data suggest an increase of autophagosome accumulation in the pathogenesis of SMA, thus providing a valuable clue in understanding the mechanisms of axonal degeneration and a possible therapeutic target in the treatment of SMA.
Results
Smn knockdown causes LC3-II protein increase
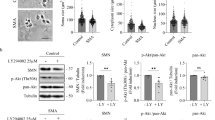
We developed an in vitro model of SMA using a lentiviral RNA interference method to downregulate the Smn protein level of isolated mouse spinal cord MNs,22 and have now used this model to analyze changes in the autophagy process in Smn-reduced MNs (Figures 1a and b). An overall indicator of autophagy impairment is the microtubule-associated proteinLC3-II. Conversion of LC3-I into LC3-II has been considered a general marker of autophagy initiation, and LC3-II level is correlated with autophagosome quantity.23 To analyze whether Smn reduction affects LC3-II protein levels, MNs were transduced with a lentivirus containing empty vector (EV) or short hairpin RNA sequences targeting specific sites of mouse Smn (shSmn),22 and maintained in the presence of the neurotrophic factors (NTFs) cocktail (1 ng/ml brain-derived neurotrophic factor; 10 ng/ml glial cell line-derived neurotrophic factor, 10 ng/ml ciliary neurotrophic factor, 10 ng/ml cardiotrophin-1 and 10 ng/ml hepatocyte growth factor). At 6, 8 and 12 days after transduction, cell lysates were collected and submitted to western blot using an anti-LC3 antibody. During SDS-polyacrylamide gel electrophoresis, LC3-II migrates faster than LC3-I, and the two forms can be distinguished even though both are recognized by anti-LC3 antibodies. Although LC3-II levels can be calculated as a function of LC3-I, it has been suggested that this relative analysis is not ideal for several reasons, including delipidation of LC3-II, and differences between tissues and cell lines in LC3-I levels.24 Thus, LC3-II levels as a function of actin or tubulin (loading control) are thought to be more reliable immunoblot measurements. Using this approach, we observed a significantly increased LC3-II level in shSmn cells after 6, 8 and 12 days of transduction, compared with the EV control (Figure 1c). We also analyzed LC3-II protein level in protein extracts at 4 days post-transduction, but observed no differences between controls and shSmn cells (data not shown). Protein extracts from non-transduced cultures were also submitted to western blot in order to discard changes in LC3-II level after lentiviral transduction; no differences were observed (Figure 1c). The increased LC3-II level in Smn-reduced MNs indicates increased autophagosome generation and/or accumulation. To localize the LC3-positive structures in cell somas and/or neurites, we analyzed the endogenous LC3 protein by immunofluorescence. MNs were plated on glass coverslips and transduced using the EV or shSmn constructs. Eight days after transduction, cells were fixed and LC3 immunostaining was performed. Figure 2 shows the distribution of endogenous LC3-positive spots in both MN soma (Figure 2a) and neurites (Figure 2b).
Effect of Smn reduction in LC3-II protein level. MNs were transduced with shSmn or EV lentiviral constructs or were left non-transduced (Φ) and maintained in the presence of the NTFs cocktail (10 ng/ml of ciliary neurotrophic factor, glial cell line-derived neurotrophic factor, cardiotrophin-1, hepatocyte growth factor and 1 ng/ml of brain-derived neurotrophic factor). (a) Representative microscopy images of 8 day-EV and shSmn-transduced cells maintained in the presence of NTFs: phase contrast (left) and GFP (right), indicates GFP-expressing cells in the same microscopic field. Scale bar, 30 μm. (b) Representative confocal images of 8-day EV or shSmn cultures. Cells were fixed and immunofluorescence with an anti-Smn antibody (blue) was performed. Scale bar, 15 μm. Protein extracts of 4-, 6-, 8- and 12-day-transduced cultures were probed with an anti-Smn antibody by western blot and reprobed with an anti-α-tubulin antibody, used as a loading control. Graph represents the expression of Smn and corresponds to the quantification of three independent experiments±S.E.M. (c and d) Protein extracts from 6-, 8- and 12-day EV or shSmn-transduced cultures (c); and 8 days EV or non-transduced (d) cultures were probed with an anti-LC3 antibody. Membranes were reprobed with an antibody against α-tubulin, used as a loading control, or an anti-Smn antibody as protein reduction control. Graphs in c and d represent the expression of LC3-II versus α-tubulin and correspond to the quantification of at least four independent experiments±S.E.M. Asterisks indicate significant differences using one-way ANOVA test and Bonferroni post-hoc multiple comparisons (*P<0.05; **P<0.005)
Cellular distribution of LC3-II-positive aggregates. Representative images of 8 days shSmn- or EV-transduced cultures. Cells were fixed and processed for immunofluorescence with an anti-LC3 antibody. (a) Images of cell soma and their emerging neurites. (b) Images of distant neurites. Arrows in a and b indicate the presence of LC3-positive aggregates, in a and b, Hoechst (blue), GFP (green). Scale bar: (a) 6 μm and (b) 12 μm. Images were acquired with an FV10I confocal microscope (Olympus), using the × 60 objective, and the same microscopy settings. Images were not submitted to any post-capture manipulation
An increase in autophagosomes can result from increased synthesis or from reduced degradation, which reflects an alteration of the autophagic flux. To determine whether increased autophagic synthesis or reduced autophagic flux caused the LC3-II increase in Smn-reduced MNs, cultures were treated with the lysosomal proteolysis inhibitor Baphilomycin A1 (BafA1).25 This treatment causes LC3-II increase when the autophagic flux is not altered.24 MNs were transduced and, 8 days after plating, were treated or not with BafA1 (50 nM during 4 h). Protein extracts were collected and submitted to western blot analysis using the anti-LC3 antibody. When the lysosomal proteolysis is inhibited there is an accumulation of autophagosomes in both conditions (i.e., LC3-II is increased in shSmn condition, compared with EV, but addition of BafA1 increases LC3-II in both conditions, as shown in Figure 3). Of the possible outcomes of autophagy modulation by a drug,24 this one indicates that Smn reduction increases autophagosome synthesis without affecting turnover.
Effect of BafA1 on LC3-II protein level. MNs were transduced with EV or shSmn, and 8 days later cells were treated with 50 nM BafA1 for 4 h. Cell extracts were collected and submitted to western blot using an antibody against LC3. The same membrane was reprobed with an anti-α-tubulin antibody as a loading control. Graph represents the expression of LC3-II versus α-tubulin and correspond to the quantification of three independent experiments±S.E.M. Asterisks indicate significant differences using one-way ANOVA test and Bonferroni post-hoc multiple comparisons (*P<0.05)
Smn reduction increases autophagic structures in MNs
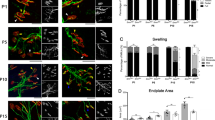
Autophagosomes are double-membraned vesicles that mediate the first step of autophagy by sequestering organelles, long-lived proteins, and/or portions of the cytoplasm. This material is finally degraded by fusion with the lysosomal compartment, forming the autolysosome.26, 27 Transmission electron microscopy is an important tool for detecting autophagosomes and can provide significant insight to the extent of on-going autophagy. We used this approach to analyze the presence of autophagosomes and autolysosomes in the cytoplasm and neurites of Smn-reduced MNs. The MNs were transduced with lentivirus containing EV or shSmn, and after 8 days, MNs were processed and analyzed. Non-transduced neurons were analyzed as a control of the lentiviral transduction process (data not shown). Smn reduction produced a marked increase in the number of autophagic profiles, including double-membraned autophagosomes and autolysosomes (Figure 4). These structures can be easily identified in MN soma and, on average from three independent experiments, 0.46±0.18 of autophagic compartments per MN soma were found in non-transduced controls (n=14) and EV (n=14), whereas 2.66±0.45 of these structures per MN soma were found in shSmn cultures (n=16; P<0.0005). These autophagic profiles were also found in neurites (Figures 4g and h). Normal endoplasmic reticulum, Golgi complex and mitochondria were found in EV- and shSmn-transduced neurons. These results suggest an increase of the number of autophagy profiles in Smn-reduced spinal cord MNs when compared with non-transduced and EV controls.
Ultrastructural changes in Smn-reduced spinal cord MNs. Mouse MNs were transduced with lentivirus containing the EV (a and b) or shSmn (c–h) construct. Cells were maintained in the presence of NTFs. Eight days after transduction, cultures were fixed and submitted to electron microscopy. Arrows indicate autophagic profiles present in cell somas (b–f) and neurites (g and h). Scale bar: (a) 5 μm and (b–h) 1 μm
Bcl-xL overexpression reduces LC3-II protein level in Smn-reduced MNs
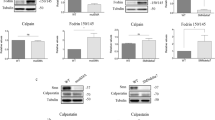
Smn protein knockdown causes neurite degeneration and non-apoptotic cell death of spinal cord MNs. This neurodegeneration can be prevented by Bcl-xL overexpression without affecting Smn protein level.22 It is known that Bcl-xL protein is an important factor in autophagy through its binding to Beclin1 and inhibiting Beclin1-mediated autophagy.28 To assess whether Bcl-xL overexpression reduces autophagy in our model, the level of LC3-II protein was analyzed in shSmn and EV cultures cotransduced with lentivirus carrying an expression construct containing the human Bcl-xL (hBcl-xL) gene. Eight days after cotransduction, western blot analysis of protein extracts showed that Bcl-xL was overexpressed. Overexpression did not change Smn protein level compared with controls (Figure 5a). Compared with the EV control, LC3-II level was 1.76±0.16 in shSmn, and 1.24±0.14 in hBcl-xL and shSmn cotransduced cultures, indicating that Bcl-xL significantly reduced the LC3-II protein levels (P<0.05). On the other hand, Bcl-xL overexpression did not reduce LC3-II protein in EV cells (1.05±0.11-fold increase compared with EV controls), indicating that Bcl-xL reduces the LC3-II increase caused by Smn knockdown.
Effect of Bcl-xL overexpression on LC3-II protein level. MNs were cotransduced with lentivirus containing the hBcl-xL plus shSmn or the EV; or transduced with EV or shSmn. (a) Protein extracts of 8 day-transduced cultures were probed with an anti-LC3 or an anti-Bcl-xL antibody. (b) Protein extracts of 8 days were probed with an anti-Beclin1 antibody or an anti-Smn antibody. Membranes were reprobed with an antibody against α-tubulin, used as a loading control. Graph values in a and b represent the expression of LC3-II or Beclin1, respectively, versus α-tubulin, and corresponds to the quantification of at least four independent experiments±S.E.M. Asterisks indicate significant differences using one-way ANOVA test and Bonferroni post-hoc multiple comparisons (*P<0.05)
Because Bcl-xL inhibits Beclin1-induced autophagy, we wanted to know whether Beclin1 protein levels were affected by Smn reduction. Total protein extracts of 8-day-transduced MNs were analyzed using an anti-Beclin1 antibody. Beclin1 protein was slightly but significantly increased in shSmn condition compared with the EV control (1.22±0.07-fold increase, P<0.05). To assess whether Bcl-xL overexpression prevented Beclin1 increase in Smn-reduced MNs, we cotransduced these cells with both lentiviral constructs. Western blot results indicated that Bcl-xL overexpression did not reduce Beclin1 protein expression in EV or shSmn conditions when compared with the non-Bcl-xL-transduced controls (1.14±0.02- and 1.18±0.2-fold increase, respectively) (data not shown). Taken together, these results show an increase of Beclin1 protein after Smn reduction, and that Bcl-xL overexpression prevents LC3-II, but not Beclin1, increase.
LC3-II increases in MNs from a mouse model of SMA
To determine whether changes in the level of LC3-II can also be detected in MNs from an in vivo model of SMA, we used the Smn(−/−); SMN2 (SMA type I mice mutant, mtSMA) mouse model for human type I SMA. Protein extracts were obtained from cultures of isolated spinal cord MNs of mutant (mtSMA) and wild-type (WT) animals. We dissected and genotyped E13 embryos by crossing two Smn(+/−);SMN2 mutants. After genotyping, MNs from WT and mtSMA embryos were purified and cultured (Figure 6). Eight days after plating, protein extracts were obtained and submitted to western blot using anti-LC3 and anti-Beclin1 antibodies. As shown in Figure 6, LC3-II protein was significantly increased in MNs from mtSMA (1.65±0.12-fold increase, P<0.05) cultures compared with the WT control. As we had observed in shSmn conditions, Beclin1 levels were slightly increased in mtSMA (1.23±0.1-fold increase; data not shown). Together, these results suggest that cultured MNs obtained from an in vivo model SMA show increased levels of autophagy markers LC3-II and Beclin1, compared with MNs cultured from their WT littermate embryos. This increase of LC3-II and Beclin1 is comparable to that observed in shSmn-transduced MNs, reinforcing the hypothesis that Smn-reduced MNs undergo changes related to autophagy.
LC3 protein levels in MNs from SMA mice model. (a) Representative images of 8 days cultures of isolated MNs from WT and Smn(−/−); SMN2 (mtSMA) embryos. Cells were cultured in the presence of the NTFs cocktail. Arrows indicate neurite degeneration. Scale bar, 30 μm. (b) Protein extracts of EV and mtSMA of 8 days cultures were probed with an anti-LC3 antibody by western blot analysis. Membranes were reprobed with an antibody against α-tubulin as a loading control. Graph values represent the expression of LC3-II versus α-tubulin and corresponds to the quantification of three independent experiments ±S.E.M. Asterisks indicate significant differences using one-way ANOVA test and Bonferroni post-hoc multiple comparisons (*P<0.05)
Discussion
In the present study, we described an increase in the levels of the autophagy markers in MNs obtained from SMA models. Smn protein reduction caused an increment of the autophagosome number as well as of the autophagy-related proteins Beclin1 and LC3-II. Our group’s main research agenda is to analyze the cellular and molecular events involved in MN degeneration caused by Smn protein reduction. We previously described that Smn knockdown causes neurite degeneration and late non-apoptotic cell death, both of which are prevented by Bcl-xL overexpression.22 It is known that Bcl-xL inhibits autophagy by binding to Beclin1, which is required for the initiation of autophagosome formation.28, 29
In order to study the molecular process that leads Bcl-xL protein to counteract the Smn knockdown degenerative process, we decided to analyze the autophagy in Smn-reduced MNs. We observed an increase of the autophagosome formation that can be prevented by Bcl-xL. Thus, we propose two promising avenues for further research: autophagy deregulation as a contributor to MN degeneration in SMA disease and the protective effect of the Bcl-xL pathway in Smn-reduced MNs.
Among the list of molecular and cellular approaches to assess autophagy deregulation, two are considered key methods: autophagosome detection by electron microscopy24, 27 and the presence of LC3-II.23, 30, 31 In our model, electron microscopy showed an increase of the autophagosome number and protein analysis demonstrated an increase of LC3-II levels. Both parameters can be associated with either enhanced autophagosome synthesis or reduced autophagosome turnover. To interpret the changes in levels of processed LC3-II, it is necessary to inhibit the degradation of autolysosome content. In the presence of inhibitors (such as BafA1), accumulation of LC3-II-positive autophagosomes would be evidence of efficient autophagic flux, whereas failure of LC3-II to increase would indicate a defect or delay earlier in the process, prior to degradation at the autolysosome.24, 27 From western blot results, we concluded that in our model, Smn reduction caused increased autophagosome production, but not alterations in the autophagic flux. LC3-II increase can be counteracted by Bcl-xL overexpression, which promotes morphological recovery of Smn-reduced MNs.22 These results together suggest that an increase of Beclin1-dependent autophagy could be one of the mechanisms responsible for MNs degeneration in the Smn knockdown model.
Beclin1 is a Bcl-2- and Bcl-xL-interacting protein essential for autophagosome formation and autophagy initiation.32, 33 Its BH3 domain binds to Bcl-2 and Bcl-xL,34 thus reducing Beclin1-dependent autophagy.28 The interaction between Beclin1 and its inhibitors is dynamic and subject to phosphorylation of either partner in the complex,35 and it has been proposed that Beclin1 phosphorylation and dissociation from Bcl-xL proteins initiates Beclin1-dependent autophagy.36 In our studies, Beclin1 protein level is slightly, but significantly, increased in Smn-reduced MNs, a change suggesting that Beclin1 and Beclin1-dependent autophagy may be mediating autophagosome formation and LC3-II increase in these cells. In some pathological models of neuronal injury, increased Beclin1 expression at the lesion site19, 37, 38 has been associated with increased autophagy and autophagic cell death in that region. Thus, Beclin1-dependent autophagy is involved in neuronal degeneration observed in several injury models, suggesting an important role of this cell death mechanism in damaged neurons.
Autophagy is essential for tissue homeostasis. When autophagosome formation is compromised or when autophagosomes accumulate, cell functions and survival may be reduced. In the nervous system, autophagy can be the main route for the degradation of toxic proteins. In some neurodegenerative disorders, especially those caused by aggregates of mutant forms of specific proteins, there is an accumulation of autophagosomes that may interfere with intracellular trafficking or may become a source of cytotoxic products.14, 39 On the other hand, mutations that perturb axonal transport can cause substantial autophagosome accumulation, which in turn can perturb axonal transport, leading to progressive neuronal degeneration.12, 40 Moreover, autophagosome accumulation can contribute to production of toxic aggregated proteins.41 In SMA pathogenesis, axonal deficiencies have been described in several in vitro and in vivo models.8, 22, 42, 43 However, there are no evidences of toxic aggregated protein accumulation in SMA neurons and non-neuronal cells. The origin of the axonal degeneration is not clear, although autophagosome accumulation or transport impairment may be contributing to this process. For example, autophagy inhibition by knocking down Beclin1 efficiently delays neurite degeneration after neurotrophic factor deprivation,20 which suggests an essential role of autophagy activation in inducing neurite degeneration after neuronal damage such as trophic support withdrawal. However, autophagy suppression induces degeneration of the axon terminals in non-damaged neurons, which is suggestive of autophagy involvement in axon homeostasis.44 Therefore, the existence of autophagy as a pro-death pathway is controversial, and the mechanism by which autophagy programs neurites to die is still unclear.
Our studies of Smn-reduced MNs reveal an increase of the autophagosome synthesis but not an alteration of autophagosome turnover. Therefore, stimulation of autophagosome production could be detrimental for the integrity of neurites and neurons. The implication is that drugs that induce autophagy should not be considered for use in the treatment of SMA.
In summary, our results indicate that Smn reduction causes changes in Beclin1-dependent autophagy and could be one of the contributors to neurite degeneration observed in MNs of SMA experimental models. These observations suggest a critical role of autophagy in neurite degeneration of damaged neurons and a potentially valuable clue in understanding the mechanism of axonal and dendritic degeneration, and a possible therapeutic target in the treatment of SMA.
Materials and Methods
Spinal cord MN isolation and culture
MN cultures were prepared from embryonic 12.5-day (E12.5) CD1 mouse spinal cord essentially as described.22, 45 Isolated cells were pooled in a tube containing culture medium and plated. All procedures were in accordance with the Spanish Council on Animal Care and approved by the University of Lleida Advisory Committee on Animal Services.
Isolated MNs were plated in four-well tissue culture dishes (Nunc, Thermo Fisher Scientific, Madrid, Spain) for electron microscopy (15 000 cells/well), immunofluorescence (10 000 cells/well) and western blot (50 000 cells/well) analysis. Culture medium was Neurobasal (Gibco, Invitrogen, Paisley, UK) supplemented with B27 (Gibco; Invitrogen), horse serum (2% v/v), L-glutamine (0.5 mM) and 2-mercaptoethanol (25 μM). Cells were plated with complete medium containing a cocktail of recombinant NTFs (1 ng/ml brain-derived neurotrophic factor, 10 ng/ml glial cell line-derived neurotrophic factor, 10 ng/ml ciliary neurotrophic factor, 10 ng/ml cardiotrophin-1 and 10 ng/ml hepatocyte growth factor; Peprotech, London, UK).
SMA animals
SMA type I mice FVB.Cg-Tg(SMN2)89AhmbSmn1tm1Msd/J were kindly provided by Dr. Josep E Esquerda (IRBLLEIDA-Universitat de Lleida). Heterozygous animals were crossed to obtain homozygous Smn−/−;SMN2+/+ (mtSMA). Littermates mtSMA and Smn+/+;SMN2+/+ (WT) were used for the experiments. For MNs purification E13 embryos were removed from the uterus and a piece was snipped from the head for genotyping. The REDExtract-N-Amp Tissue PCR Kit (Sigma, St Louis, MO, USA) was used for genomic DNA extraction and PCR setup, with the following primers: WT forward 5′-CTCCGGATATTGGGATTG-3′, SMA reverse 5′-GGTAACGCCAGGGTTTTCC-3′ and WT reverse 5′-TTTCTTCTGGCTGTGCCTTT-3′. After genotyping, WT and mtSMA animals were submitted to spinal cord dissection, MN isolation and culture as described before.
Western blot analysis
Western blots were performed as described.45, 46 Total cell lysates were resolved in SDS-polyacrylamide gels and transferred onto polyvinylidene difluoride Immobilon-P transfer membrane filters (Millipore, Billerica, MA, USA) using an Amersham Biosciences semidry Trans-Blot (Buckinghamshire, UK). The membranes were blotted with anti-SMN antibody (1 : 5000); anti-Bcl-xL antibody (1 : 2000; BD Transduction Laboratories, Franklin Lakes, NJ, USA); anti-LC3 antibody (1 : 1000); anti-Beclin1 antibody (1 : 1000; Cell Signaling Technology, Boston, MA, USA). To control the specific protein content per lane, membranes were reprobed with monoclonal anti-α-tubulin antibody (Sigma). Blots were developed using Super Signal chemiluminiscent substrate (Pierce, Rockford, IL, USA).
Plasmids and production of lentiviral particles
For RNA interference experiments, constructs were generated in pSUPER.retro.puro (OligoEngine, Seattle, WA, USA) using specific oligonucleotides (Invitrogen) targeting SMN sequence as described.22 pLVTHM, pSPAX2 and pMD2G were kindly provided by Dr. Trono (University of Geneva, Switzerland). Viruses at 4 × 105–1 × 106 TU/ml were used for the experiments. For lentiviral transduction, MNs were plated in four-well dishes. Medium containing lentivirus (2 TU/cell) was added 3 h later, and then changed after 20 h. In each experiment, green fluorescent protein (GFP)-positive cells were counted directly to monitor infection efficiency. Frequency of infection rose 99%.22, 47
For Bcl-xL overexpression, cDNA for human Bcl-xL was subcloned into pWPI as described previously.48
Electron microscopy
MNs were plated on round glass coverslips and transduced. Cultures were fixed for 1 h at 4°C in 1.6% glutaraldehyde in 0.5 M phosphate buffer (pH 7.3), washed, fixed again in aqueous 2% osmium tetroxide, stained in 2% uranyl acetate in 30% methanol and finally embedded in Eppon. Ultrathin sections were taken from selected areas containing MN cell bodies and neurites previously identified in toluidine blue-stained 1-mm-thick semithin sections. Ultrathin sections were stained with lead citrate and uranyl acetate and observed by electron microscopy (Zeiss EM910, Oberkochen, Germany): one researcher loaded the section, another examined it and a third counted the organelles, blinded to which condition was loaded.
Immunofluorescence
MNs were plated on round glass coverslips and transduced with the lentiviral constructs. After several days cells were fixed with ice-cold methanol, permeabilized with 0.1% Triton X-100 and incubated for 2 h in 1 ml 5% BSA in PBS. Primary antibody (LC3B antibody, Cell Signaling Technology) was added at 1 : 100 dilution to the 5% BSA/PBS and incubated overnight. Cells were washed and the secondary antibody ALEXA555 (Invitrogen) added at 1 : 400 dilution. Hoechst staining was performed to identify nuclear localization in MNs soma. Samples were mounted using Mowiol (Calbiochem (EMD Millipore), Darmstadt, Germany) plus 0.5% DABCO (Sigma) mounting medium. Microscopy observations were performed in FV10I Olympus confocal microscope (Tokyo, Japan).
Statistical analysis
All experiments were performed at least three times. Values were expressed as mean ±S.E.M. Differences between groups were assessed by one-way ANOVA of lentiviral-transduced cultures at each time point; if significant, post-hoc multiple comparisons were done using Bonferroni test; P-values <0.05 were considered significant.
Abbreviations
- SMA:
-
spinal muscular atrophy
- MN:
-
motoneurons
- SMN:
-
human survival motor neuron protein
- Smn:
-
murine survival motor neuron protein
- NTF:
-
neurotrophic factor
- GFP:
-
green fluorescent protein
- EV:
-
empty vector
- WT:
-
wild type
- mtSMA:
-
SMA type I mice mutant
References
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995; 80: 155–165.
Wirth B . An update of the mutation spectrum of the survival motor neuron gene (SMN1) in autosomal recessive spinal muscular atrophy (SMA). Hum Mutat 2000; 15: 228–237.
Sumner CJ . Therapeutics development for spinal muscular atrophy. NeuroRx 2006; 3: 235–245.
Zhang Z, Lotti F, Dittmar K, Younis I, Wan L, Kasim M et al. SMN deficiency causes tissue-specific perturbations in the repertoire of snRNAs and widespread defects in splicing. Cell 2008; 133: 585–600.
Cifuentes-Diaz C, Nicole S, Velasco ME, Borra-Cebrian C, Panozzo C, Frugier T et al. Neurofilament accumulation at the motor endplate and lack of axonal sprouting in a spinal muscular atrophy mouse model. Hum Mol Genet 2002; 11: 1439–1447.
Kong L, Wang X, Choe DW, Polley M, Burnett BG, Bosch-Marce M et al. Impaired synaptic vesicle release and immaturity of neuromuscular junctions in spinal muscular atrophy mice. J Neurosci 2009; 29: 842–851.
Murray LM, Comley LH, Thomson D, Parkinson N, Talbot K, Gillingwater TH . Selective vulnerability of motor neurons and dissociation of pre- and post-synaptic pathology at the neuromuscular junction in mouse models of spinal muscular atrophy. Hum Mol Genet 2008; 17: 949–962.
Rossoll W, Jablonka S, Andreassi C, Kroning AK, Karle K, Monani UR et al. Smn, the spinal muscular atrophy-determining gene product, modulates axon growth and localization of beta-actin mRNA in growth cones of motoneurons. J Cell Biol 2003; 163: 801–812.
Soler-Botija C, Ferrer I, Gich I, Baiget M, Tizzano EF . Neuronal death is enhanced and begins during foetal development in type I spinal muscular atrophy spinal cord 2002 Brain 125: 1624–1634.
Tsai MS, Chiu YT, Wang SH, Hsieh-Li HM, Lian WC, Li H . Abolishing Bax-dependent apoptosis shows beneficial effects on spinal muscular atrophy model mice. Mol Ther 2006; 13: 1149–1155.
Bredesen DE, Rao RV, Mehlen P . Cell death in the nervous system. Nature 2006; 443: 796–802.
Marino G, Madeo F, Kroemer G . Autophagy for tissue homeostasis and neuroprotection. Curr Opin Cell Biol 2011; 23: 198–206.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–1075.
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet 2004; 36: 585–595.
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D . Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 2004; 305: 1292–1295.
Boland B, Kumar A, Lee S, Platt FM, Wegiel J, Yu WH et al. Autophagy induction and autophagosome clearance in neurons: relationship to autophagic pathology in Alzheimer's disease. J Neurosci 2008; 28: 6926–6937.
Lee JA, Gao FB . Inhibition of autophagy induction delays neuronal cell loss caused by dysfunctional ESCRT-III in frontotemporal dementia. J Neurosci 2009; 29: 8506–8511.
Chen S, Zhang X, Song L, Le W . Autophagy dysregulation in amyotrophic lateral sclerosis. Brain Pathol 2012; 22: 110–116.
Kanno H, Ozawa H, Sekiguchi A, Itoi E . Spinal cord injury induces upregulation of Beclin 1 and promotes autophagic cell death. Neurobiol Dis 2009; 33: 143–148.
Yang Y, Fukui K, Koike T, Zheng X . Induction of autophagy in neurite degeneration of mouse superior cervical ganglion neurons. Eur J Neurosci 2007; 26: 2979–2988.
Yang Y, Xu K, Koike T, Zheng X . Transport of autophagosomes in neurites of PC12 cells during serum deprivation. Autophagy 2008; 4: 243–245.
Garcera A, Mincheva S, Gou-Fabregas M, Caraballo-Miralles V, Llado J, Comella JX et al. A new model to study spinal muscular atrophy: neurite degeneration and cell death is counteracted by BCL-X(L) overexpression in motoneurons. Neurobiol Dis 2011; 42: 415–426.
Mizushima N . Methods for monitoring autophagy. Int J Biochem Cell Biol 2004; 36: 2491–2502.
Menzies FM, Moreau K, Puri C, Renna M, Rubinsztein DC . Measurement of autophagic activity in mammalian cells. Curr Protoc Cell Biol 2012; Chapter 15 Unit15.16.
Yamamoto A, Tagawa Y, Yoshimori T, Moriyama Y, Masaki R, Tashiro Y . Bafilomycin A1 prevents maturation of autophagic vacuoles by inhibiting fusion between autophagosomes and lysosomes in rat hepatoma cell line, H-4-II-E cells. Cell Struct Funct 1998; 23: 33–42.
Fader CM, Colombo MI . Autophagy and multivesicular bodies: two closely related partners. Cell Death Differ 2009; 16: 70–78.
Barth S, Glick D, Macleod KF . Autophagy: assays and artifacts. J Pathol 2010; 221: 117–124.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Yip KW, Reed JC . Bcl-2 family proteins and cancer. Oncogene 2008; 27: 6398–6406.
Kimura S, Fujita N, Noda T, Yoshimori T . Monitoring autophagy in mammalian cultured cells through the dynamics of LC3. Methods Enzymol 2009; 452: 1–12.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. Embo J 2000; 19: 5720–5728.
Kihara A, Kabeya Y, Ohsumi Y, Yoshimori T . Beclin-phosphatidylinositol 3-kinase complex functions at the trans-golgi network. EMBO Rep 2001; 2: 330–335.
Erlich S, Mizrachy L, Segev O, Lindenboim L, Zmira O, Adi-Harel S et al. Differential interactions between Beclin 1 and Bcl-2 family members. Autophagy 2007; 3: 561–568.
Oberstein A, Jeffrey PD, Shi Y . Crystal structure of the Bcl-XL-Beclin 1 peptide complex: Beclin 1 is a novel BH3-only protein. J Biol Chem 2007; 282: 13123–13132.
Kang R, Zeh HJ, Lotze MT, Tang D . The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ 2011; 18: 571–580.
Zalckvar E, Berissi H, Mizrachy L, Idelchuk Y, Koren I, Eisenstein M et al. DAP-kinase-mediated phosphorylation on the BH3 domain of beclin 1 promotes dissociation of beclin 1 from Bcl-XL and induction of autophagy. EMBO Rep 2009; 10: 285–292.
Diskin T, Tal-Or P, Erlich S, Mizrachy L, Alexandrovich A, Shohami E et al. Closed head injury induces upregulation of Beclin 1 at the cortical site of injury. J Neurotrauma 2005; 22: 750–762.
Penas C, Font-Nieves M, Fores J, Petegnief V, Planas A, Navarro X et al. Autophagy, and BiP level decrease are early key events in retrograde degeneration of motoneurons. Cell Death Differ 2011; 18: 1617–1627.
Wong E, Cuervo AM . Autophagy gone awry in neurodegenerative diseases. Nat Neurosci 2010; 13: 805–811.
Nixon RA, Yang DS, Lee JH . Neurodegenerative lysosomal disorders: a continuum from development to late age. Autophagy 2008; 4: 590–599.
Yu WH, Cuervo AM, Kumar A, Peterhoff CM, Schmidt SD, Lee JH et al. Macroautophagy--a novel Beta-amyloid peptide-generating pathway activated in Alzheimer's disease. J Cell Biol 2005; 171: 87–98.
McGovern VL, Gavrilina TO, Beattie CE, Burghes AH . Embryonic motor axon development in the severe SMA mouse. Hum Mol Genet 2008; 17: 2900–2909.
Burghes AH, Beattie CE . Spinal muscular atrophy: why do low levels of survival motor neuron protein make motor neurons sick? Nat Rev Neurosci 2009; 10: 597–609.
Komatsu M, Wang QJ, Holstein GR, Friedrich VL Jr, Iwata J, Kominami E et al. Essential role for autophagy protein Atg7 in the maintenance of axonal homeostasis and the prevention of axonal degeneration. Proc Natl Acad Sci USA 2007; 104: 14489–14494.
Gou-Fabregas M, Garcera A, Mincheva S, Perez-Garcia MJ, Comella JX, Soler RM . Specific vulnerability of mouse spinal cord motoneurons to membrane depolarization. J Neurochem 2009; 110: 1842–1854.
Perez-Garcia MJ, Cena V, de Pablo Y, Llovera M, Comella JX, Soler RM . Glial cell line-derived neurotrophic factor increases intracellular calcium concentration. Role of calcium/calmodulin in the activation of the phosphatidylinositol 3-kinase pathway. J Biol Chem 2004; 279: 6132–6142.
Mincheva S, Garcera A, Gou-Fabregas M, Encinas M, Dolcet X, Soler RM . The canonical nuclear factor-{kappa}B pathway regulates cell survival in a developmental model of spinal cord motoneurons. J Neurosci 2011; 31: 6493–6503.
Gozzelino R, Sole C, Llecha N, Segura MF, Moubarak RS, Iglesias-Guimarais V et al. BCL-XL regulates TNF-alpha-mediated cell death independently of NF-kappaB, FLIP and IAPs. Cell Res 2008; 18: 1020–1036.
Acknowledgements
This work was supported by grants from GENAME (Defining targets for Therapeutics in SMA) to RMS; from Instituto de Salud Carlos III, Fondo de Investigaciones Sanitarias (PI11-01047), Generalitat de Catalunya (SGR740) and Consolider-Ingenio 2010 (CSD2007-00020) to RMS. AG holds a post-doctoral contract from Genoma España; SA holds a fellowship from Universitat de Lleida and AP holds a fellowship from Comissionat de Universitats i Recerca, Departament d’Innovació, Universitats i Empresa de la Generalitat de Catalunya and Fons Social Europeu. We thank Elaine Lilly, Ph.D. (Writer’s First Aid), for English language revision of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by A Verkratsky
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Garcera, A., Bahi, N., Periyakaruppiah, A. et al. Survival motor neuron protein reduction deregulates autophagy in spinal cord motoneurons in vitro. Cell Death Dis 4, e686 (2013). https://doi.org/10.1038/cddis.2013.209
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.209
Keywords
This article is cited by
-
Spinal Muscular Atrophy autophagy profile is tissue-dependent: differential regulation between muscle and motoneurons
Acta Neuropathologica Communications (2021)
-
Calpain system is altered in survival motor neuron-reduced cells from in vitro and in vivo spinal muscular atrophy models
Cell Death & Disease (2020)
-
AAV9 Vector: a Novel modality in gene therapy for spinal muscular atrophy
Gene Therapy (2019)
-
Calpain Inhibition Increases SMN Protein in Spinal Cord Motoneurons and Ameliorates the Spinal Muscular Atrophy Phenotype in Mice
Molecular Neurobiology (2019)
-
Neuronal activity regulates DROSHA via autophagy in spinal muscular atrophy
Scientific Reports (2018)