Abstract
Peroxisomes are a critical rheostat of reactive oxygen species (ROS), yet their role in drug sensitivity and resistance remains unexplored. Gene expression analysis of clinical lymphoma samples suggests that peroxisomes are involved in mediating drug resistance to the histone deacetylase inhibitor (HDACi) Vorinostat (Vor), which promotes ROS-mediated apoptosis. Vor augments peroxisome numbers in cultured lymphoma cells, concomitant with increased levels of peroxisomal proteins PEX3, PEX11B, and PMP70. Genetic inhibition of peroxisomes, using PEX3 knockdown, reveals that peroxisomes protect lymphoma cells against Vor-mediated cell death. Conversely, Vor-resistant cells were tolerant to elevated ROS levels and possess upregulated levels of (1) catalase, a peroxisomal antioxidant, and (2) plasmalogens, ether glycerophospholipids that represent peroxisome function and serve as antioxidants. Catalase knockdown induces apoptosis in Vor-resistant cells and potentiates ROS-mediated apoptosis in Vor-sensitive cells. These findings highlight the role of peroxisomes in resistance to therapeutic intervention in cancer, and provide a novel modality to circumvent drug resistance.
Similar content being viewed by others
Main
Peroxisomes are spherical organelles, which can form de novo from the endoplasmic reticulum (ER) via the concerted action of peroxins such as PEX3, 16, and 19, and divide via a fission-based mechanism, involving the PEX11 gene family.1, 2, 3 These organelles perform key roles in bile acid, ether phospholipid, and fatty acid metabolism.4, 5 For instance, peroxisomes perform β-oxidation of very-long-chain fatty acids into long-chain fatty acids,6, 7 which are oxidized to completion in neighboring mitochondria.8, 9, 10 Within the peroxisome, β-oxidation generates reactive oxygen species (ROS) such as hydrogen peroxide (H2O2), which is primarily quenched via peroxisomal catalase. Furthermore, peroxisomes are required for the initial steps of plasmalogen biosynthesis;4 plasmalogens are a class of vinyl-ether phospholipids, which can serve as sacrificial oxidants.11, 12 Multiple investigations highlight not only the ability of the peroxisome to quench self-generated ROS13, 14 but also ROS that originates from other organelles.15, 16 ROS-generating organelles include the mitochondria via Bax/Bak activation,17, 18 and ER via the unfolded protein response (UPR).19
Several chemotherapies trigger dysfunctions in the mitochondria and ER, causing apoptosis via overwhelming the cell with ROS.20, 21, 22 For instance, Vorinostat (Vor), a FDA-approved histone deacetylase inhibitor (HDACi) used in lymphoma treatment, has been well documented to trigger macromolecule degradation (autophagy),23 and mitochondrial-mediated apoptosis via ROS accumulation.24, 25 Furthermore, HDACi treatment of leukemic cells causes HSP90 hyperacetylation,26 leading to an enhanced UPR, and ROS-mediated apoptosis.27 Curiously, Vor can restore peroxisomal function in oligodendrocytes derived from patients with X-linked adrenoleukodystrophy (ABCD1/2 mutation), an inherited peroxisome disorder. Here Vor reverts oligodendrocytes toward baseline levels of peroxisomal β-oxidation.28 Despite the potential for Vor to correct defects associated with peroxisomal disorders,29 a link between Vor (or other therapies) and peroxisomes in cancer has not been established.
In this work, we provide evidence that acute Vor treatment induces the expression of peroxisomal proteins, thus increasing peroxisome proliferation in lymphoma model systems. Also, knockdown of peroxisomes via genetic silencing of PEX3 potentiates Vor-induced ROS-mediated apoptosis. In cells with acquired resistance to Vor, peroxisome levels are highly elevated, yet ROS levels, alongside ROS-mediated damage from other agents such as doxorubixin,23 and H2O2 are kept to a relative minimum. Importantly, knockdown of PEX3 or catalase overcomes Vor resistance. These data are further complemented by bioinformatics analyses of lymphoma cells from patients who were refractory to HDACi treatment, wherein a marked elevation of peroxisomal transcripts, including catalase, is demonstrated. Our findings demonstrate a role for peroxisomes in resistance to chemotherapy which could be further explored to develop adjunctive therapies that abrogate their tumor-promoting functions.
Results
HDAC inhibition drives peroxisome biogenesis in lymphoma cell models
Expression of peroxisomal ABC transporters PMP70 and ABCD2 in fibroblasts and oligodendritic cells can be induced by Vor.28, 29 We hypothesized that this regulation could also occur in Vor-treated lymphoma cells. qPCR analyses of Vor-treated U937, OCI-LY8, and SU-DHL4 lymphoma cell lines show transcriptional upregulation of peroxisome biogenesis factor PEX3, and of peroxisome fission and membrane transporter genes, PEX11B and PMP70, respectively (Figure 1a). Immunoblots of Vor-treated samples also show a time-dependent increase in expression of PEX3, PEX11B, and PMP70 (Figure 1b). The Vor-induced upregulation of peroxisomal transcripts is further supported by gene expression analyses of Vor-treated U937 cells and mined cDNA microarray data of non-lymphocytic cells, such as the leukemic lines HL-60 and NB4, as well as the breast cancer cell line MDA-MB-231. Here we observe a marked increase in the expression of transcripts corresponding to the gene ontology (GO) term ‘peroxisome organization’ (GO:0007031) (Supplementary Figure S1).
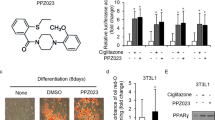
HDAC inhibition drives peroxisome biogenesis in lymphoma model systems. (a) PEX3,PEX11B, and PMP70 mRNA expression profiles relative to vehicle (DMSO) in Vor-treated U937 (2 μM, 12 h), OCI-LY8 (4 μM, 12 h), and SU-DHL4 (1 μM, 12 h) cells, versus 36B4 (housekeeping control). Graphs represent means±S.E.M. (unpaired t-test) of N=4 replicates (technical triplicate). (b) Immunoblots of PEX3, PEX11B, PMP70, and β-actin (loading control) for vehicle, and 6 and 12 h Vor-treated cells. (c) PMP70 immunofluorescence (TO-PRO-3 nuclear stain) in vehicle and Vor-treated (2 μM, 12 h) cells, puncta quantitation on right. Scale bar represents 10 μm. Graph represents means±S.E.M. (unpaired t-test) of N=2 with 5 images per condition and ~5 cells per image. (d) Left: schematic of plasmalogen biosynthesis. 1-O-alkyl Dihydroxyacetone phosphate (DHAP) is exported to the ER whereby additional enzymatic reactions occur. sn-1 and sn-2 denote carbon groups, whereas the head group is shown on bottom. Note: sn-1 position is larger/bold and colored blue to indicate the plasmalogen species being examined. Right: total PlsEtn levels (pmol) in vehicle and Vor-treated (2 μM, 12 h) U937 cells. Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=2 in technical triplicate. *P⩽0.05, **P⩽0.01, and ***P⩽0.001
To establish whether the Vor-induced expression of peroxisomal proteins leads to peroxisome assembly, we performed immunofluorescence for PMP70 puncta (a peroxisome marker) on lymphoma cells treated with and without Vor. After 12 h of Vor treatment, there is a significant increase in PMP70 puncta levels, indicating that Vor drives peroxisome proliferation (Figure 1c). A well-characterized function of peroxisomes is the synthesis of ether lipids, including plasmalogens.12 Plasmalogens contain a vinyl-ether bond that is preferentially oxidized compared to their diacyl phospholipid counterparts, enabling them to act as ‘sacrificial oxidants’ by potentially protecting other membrane lipids from oxidation.17 Thus, plasmalogens may be synthesized as an attempt to ‘quench’ the ROS-inducing effects of Vor. We next measured plasmalogen ethanolamine (PlsEtn) levels (pmol) via LC–MS/MS in lymphoma cells in the presence and absence of Vor.30 Our analyses show that PlsEtn levels are more abundant in U937 cells treated with Vor for 12 h, compared to vehicle-treated cells (Figure 1d; Supplementary Figure S1). Specifically, PlsEtn levels of 16:0 (16 carbons at the sn-1 position with no additional double bonds), 18:0, and 18:1 (18 carbons at the sn-1 position, one additional double bond) species are increased in Vor-treated versus vehicle-treated U937 cells (Figure 1d; Supplementary Figure S1).
PEX3 knockdown compromises peroxisome biogenesis and potentiates HDACi-induced cell death
We next investigated whether the ability of Vor to generate peroxisomes was linked to its ability to induce apoptosis. Cells with reduced PEX3 have compromised peroxisome biogenesis,31 thus we transfected U937 cells with scrambled siRNA (siSCR) and siRNAs directed against different regions of the PEX3 transcript (termed siPEX3-1 and siPEX3-2). Knockdown of PEX3 shows a marked reduction in Vor-induced peroxisome proliferation, as determined by immunofluorescence analyses of PMP70 puncta (Figure 2a; Supplementary Figure S2). Furthermore, immunoblot of the peroxins PEX16 and PEX19 demonstrates that PEX3 knockdown alone is sufficient to compromise peroxisome biogenesis (Figure 2b; Supplementary Figure S2).31 PEX3 knockdown in lymphoma cells also causes a significant reduction in cell proliferation compared to control cells (Figure 2c). When Vor is added to U937 cells upon PEX3 knockdown, we observe a time-dependent increase in HO-1 and CL-CASP3, versus Vor-treated siSCR cells (Figure 2d; Supplementary Figure S2), suggesting that peroxisomes protect lymphoma cells from accumulating ROS and Vor-mediated cell death. To further investigate this, we measured ROS levels using the redox-sensitive fluorescent dye, 2′,7′-dichlorofluorescin diacetate H2 (DCFDA).32 Here we observe a significant increase in DCFDA fluorescence when comparing siPEX3 to siSCR-transfected cells, and potentiated fluorescence emission upon Vor addition (Figure 2e), reminiscent of the trend observed for HO-1 in Figure 2d.
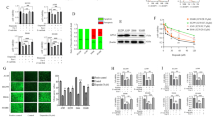
Knockdown of PEX3 compromises peroxisome biogenesis and potentiates Vor-induced cell death. (a) PMP70 immunofluorescence staining (DAPI nuclear stain) of siSCR and siPEX3-1 vehicle (DMSO) and Vor-treated (2 μM, 12 h) U937 cells. Vor was added 72 h post transfection. Graph represents means±S.E.M. (one-way analysis of variance (ANOVA), Tukey) of N=3 with 5 images taken per condition and ~4 cells per image. Scale bar represents 10 μm. *P⩽0.05 and ***P⩽0.001. (b) Immunoblot of PEX3, PEX16, PEX19, and β-actin (loading control) in vehicle, and 6 and 12 h Vor-treated (2 μM) U937 cells post siSCR and siPEX3-1 transfection. (c) Growth curves of siSCR and siPEX3-tranfected U937 cells up to 72 h post transfection. Graph represents means±S.E.M. (two-way ANOVA, Bonferroni) of N=3 (technical triplicate) ***P⩽0.001. (d) Immunoblot of HO-1, CL-CASP3, and β-actin (loading control) in vehicle, and 6 and 12 h Vor-treated (2 μM) U937 cells. (e) DCFDA relative mean fluorescence intensity (MFI) in vehicle and Vor-treated (2 μM, 12 h) U937 cells. Graph represents mean±S.E.M. (one-way ANOVA, Tukey) of N=2 (technical triplicate). **P⩽0.01 and ***P⩽0.001. (f) Percent apoptotic cells as measured by the sum of PI/Annexin V-FITC double-positive and Annexin V-FITC staining. Vor treatment was 2 μM for 18 h. Graph represents means±S.E.M. (one-way ANOVA, Tukey) of N=3 (technical triplicate). *P⩽0.05 and ***P⩽0.001
Control siRNA- and PEX3 siRNA-transfected cells were also subject to flow cytometric analyses after propidium iodide (PI) and Annexin V-FITC staining.33 We found an increase in basal apoptosis in cells transfected with siPEX3 alone (Figure 2f; Supplementary Figure S2). However, with the addition of Vor, we observe a further increase in apoptosis upon PEX3 knockdown, demonstrating an additive effect (Figures 2d and f; Supplementary Figure S2). Phase contrast images of PEX3 knockdown cells further support our FACS analysis (Supplementary Figure S2). Using a second lymphoma cell line, OCI-LY8, we also observe the greatest amount of apoptosis upon PEX3 knockdown and Vor treatment, compared to either treatment alone. (Supplementary Figure S2). These results were further supported by silencing another peroxisomal biogenesis factor PEX19 in U937 and OCI-LY8 cell lines (Supplementary Figure S3). Similar to the effects with PEX3 knockdown, we observe a potentiation in Vor-induced apoptosis when PEX19 is silenced in U937 and OCI-LY8 (Supplementary Figure S3). Together, these data indicate that peroxisomes are promoting the survival of lymphoma cells, and function to protect lymphoma cells from Vor-mediated cell death.
HDACi-resistant cells contain elevated levels of peroxisomes
We next investigated the role of the peroxisome in HDACi-resistant cells, using an isogenic model of Vor resistance we generated from U937 cells, termed B8.23 Peroxisomes are more abundant in B8 cells cultured in Vor, compared to parental U937 cells (or B8 vehicle), as determined by an increased number of PMP70 immunofluorescent puncta (Figure 3a). This finding was further supported by observing that B8 cells cultured in Vor express increased protein levels of PEX3, PEX11B, PMP70, and acyl-CoA synthetase long-chain family member 1 (ACSL1),34 compared to their parental counterparts and vehicle-cultured B8 (Figure 3b).
Elevated peroxisome levels promote HDACi resistance. (a) Left: PMP70 immunofluorescence staining (TO-PRO-3 nuclear stain) of vehicle (DMSO) and Vor-treated (2 μM, 12 h) U937 and B8 cells cultured in the presence (2 μM) and absence (DMSO) of Vor. Scale bar represents 10 μm. Right: puncta quantitation. Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=3 with 5 images taken per condition with ~4 cells per image. *P⩽0.05 and ***P⩽0.001. (b) Immunoblots of PEX3, PEX16, PEX19, PEX11B, PMP70, ACSL1, and β-actin (loading control) in vehicle, Vor-treated (2 μM) U937 cells at indicated times, and B8 cells cultured in the presence (2 μM, 1 week) and absence (DMSO) of Vor. (c) 16:0, 18:0, and 18:1 PlsEtn levels, relative to vehicle-treated U937 cells. Graphs represent means±S.E.M. (one-way analysis of variance, Tukey) of N=3 (technical triplicate). *P⩽0.05, **P⩽0.01, and ***P⩽0.001. (d) Immunoblots of PEX3, PEX16, PEX19, HO-1, CL-CASP3, and β-actin (loading control), 96 h post transfection with siSCR and siPEX3-1 in B8 cells (2 μM Vor). (e) PMP70 immunofluorescence staining of B8 siSCR versus siPEX3-1-transfected cells, 96 h post transfection with puncta quantitation on right. Graph represents means±S.E.M. (unpaired t-test) of N=3 with 5 images taken per condition and ~4 cells per image. **P⩽0.01. (f) % Apoptotic cells as measured by PI/Annexin V-FITC double-positive and Annexin V-FITC-positive cells corresponding to conditions in dand e. Graph represents means±S.E.M. (unpaired t-test) of N=3 (technical triplicate). ***P⩽0.001
Although the above data demonstrate an elevation in peroxisomes in B8, we next wanted to determine whether these peroxisomes were functional. Compared to vehicle and Vor-treated U937 cells, we observe the highest levels of total cellular PlsEtn in B8 cells (Figure 3c, left). Having profiled 16:0 18:0, and 18:1 PlsEtn levels (Supplementary Figure S4), we note that Vor-resistant cells contain significantly elevated levels of cellular PlsEtn 18:1 (Figure 3c, right). We suggest that the elevated levels of the 18:1 plasmalogen species in B8 serve to protect these cells against Vor-mediated oxidative damage. The additional double bond at the sn-1 position present in PlsEtn 18:1 could further enhance ROS scavenging, as polyunsaturated fats react with hydroxyl radicals and terminate as lipid peroxides.35, 36
Knockdown of peroxisomes, via PEX3 silencing, overcomes HDACi resistance
Compromising peroxisome function cooperates with Vor to induce cell death (Figure 2; Supplementary Figure S2), so we next investigated the contribution of increased peroxisomes to the resistance phenotype in B8 cells (Figures 3a–c). PEX3 knockdown in B8 cells, using either PEX3-1 and PEX3-2 siRNAs, results in reduced levels of PEX16 and 19, and an increase in HO-1 and CL-CASP3 (versus siSCR; Figure 3d). These data are further supported by reduced numbers of PMP70 immunofluorescent puncta upon PEX3 knockdown (Figure 3e; Supplementary Figure S4).
To investigate the role of peroxisome reduction on cell viability, flow cytometry of PI and Annexin V-FITC co-stained cells was performed (Figure 3f; Supplementary Figure S4). Upon PEX3 knockdown in Vor-resistant B8 cells, we observe an ~70% increase in cell death (18.92±1.022%), compared to siSCR (11.1±0.333%), indicating that Vor-resistant cells are partially dependent on peroxisomes for survival (Figure 3f; Supplementary Figure S4).
HDACi-resistant cells possess elevated catalase levels and effectively sequester H2O2
Considering that Vor induces apoptosis via ROS accumulation, and B8 cells survive amidst being cultured in Vor, we next investigated the effects of acute and chronic Vor treatment on ROS levels. The generation of ROS in B8 cells was determined via DCFDA flow cytometric analyses. We show that B8 cells maintain relatively low ROS levels compared to acute treatment of Vor-sensitive U937 cells (Figure 4a). This is complemented by data showing that Vor-treated U937 cells results in increased expression of HO-1 and γH2AX (ROS-mediated DNA damage marker), alongside an increase in apoptosis (CL-PARP), while B8 cells possess low levels of these cellular stress markers (Figure 4b).
Vor-resistant cells efficiently sequester ROS and possess elevated catalase levels. (a) Relative MFI (versus U937 veh.) of DCFDA fluorescence after vehicle (DMSO), and 6, 12, and 18 h Vor treatment in U937, and B8 chronically maintained in Vor (2 μM). Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=2 (technical triplicate). (b) Immunoblots of samples from a, probing CL-PARP, HO-1, γH2AX, and β-actin (loading control). (c) H2O2 decomposition assay. Relative (versus U937 veh.) changes in absorbance at 240 nm were monitored in vehicle, Vor-treated (2 μM) U937 cells for the indicated times, and B8 cells (2 μM Vor). Moles of H2O2 consumed was calculated via Beer’s law (see Materials and Methods). Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=3 (technical triplicate). (d) PI-stained (sub G1 population) U937 and B8 cells treated with vehicle, and 0.25 and 0.50 mM H2O2 for 24 h. Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=3 (technical triplicate). (e) RT-qPCR analyses of CAT versus 36B4 in vehicle, Vor-treated (2 μM) U937 for the indicated times, and B8 cells chronically in Vor. Graph represents means±S.E.M. (one-way analysis of variance, Tukey) of N=5 (technical triplicate). (f) Immunoblots of catalase and β-actin (loading control) corresponding to conditions in (e). (g) Immunofluorescence staining of catalase (TO-PRO-3 nuclear stain), comparing vehicle and Vor-treated U937 cells, and B8 cells. Scale bar represents 10 μm. For all statistical tests ***P<0.001
B8 cells maintained in Vor, an ROS-inducing agent, do not show signs of marked ROS-induced stress (Figure 4b). To determine how efficiently B8 cells sequester ROS, we monitored the breakdown of exogenous H2O2 with respect to time, compared to vehicle and Vor-treated U937 cells. Upon addition of H2O2 to U937 and B8 cells, we observe an increased rate of H2O2 breakdown in B8 cells, while no changes were found in Vor-treated U937 compared to vehicle (Figure 4c). We further support that Vor-resistant cells are better able to decompose ROS via monitoring the effect on cell viability from the addition of increasing doses of H2O2. This results in relatively minimal cell death in B8 cells, yet pronounced cell death in parental U937 cells (Figure 4d).
Peroxisomes are important in promoting the survival of Vor-resistant cells and their resistance to ROS-induced cell death. Thus, we hypothesized that cells with acquired resistance to Vor bypass ROS-induced cell death by expressing a peroxisomal enzyme that functions to counteract oxidative stress. Catalase is the most abundant peroxisomal antioxidant enzyme, thus we examined its expression in B8 cells compared to vehicle and Vor-treated U937. B8 cells possess a high level of catalase mRNA, while Vor treatment of U937 cells does not result in changes in catalase expression (Figure 4e). Immunoblots confirm that catalase levels are more abundant in B8 cells, compared to vehicle or Vor-treated U937 cells (Figure 4f). These findings are complemented by catalase immunofluorescence, which shows the highest level of puncta in B8 cells and no increase in puncta upon Vor treatment of U937 cells (Figure 4g; Supplementary Figure S5).
Catalase knockdown partially overcomes HDACi resistance
Next, we genetically knocked down catalase in U937 and B8 cells and profiled the rate of H2O2 decomposition in both systems. B8 cells transfected with siSCR display a rate of decomposition ~4.5-fold greater than B8 cells transfected with catalase siRNA (Figure 5a). Catalase is largely responsible for catalyzing the decomposition of H2O2 to water and oxygen,37 and B8 cells compared to their parental counterparts are relatively resistant to ROS-induced cell death in a catalase-dependent manner (Supplementary Figure S5). Therefore, we examined ROS-mediated damage and apoptosis markers upon catalase knockdown. Here we observe that catalase knockdown in B8 cells induces ROS-mediated damage (HO-1, γH2AX) and triggers apoptosis, as seen by increased CL-CASP3 levels (Figure 5b; Supplementary Figure S6). Upon catalase knockdown in Vor-resistant B8 cells, we observe an increase in apoptosis (16.97±0.36%), compared to siSCR (10.97±0.81%), indicating that Vor-resistant cells are in part reliant on catalase for survival (Figures 5c and d; Supplementary Figure S6). Furthermore, in Vor-treated U937 and OCI-LY8 cells we measured apoptosis via PI and Annexin V-FITC co-stain and found that catalase knockdown potentiates apoptosis (Supplementary Figure S7).
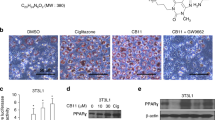
Catalase knockdown overcomes HDACi resistance. (a) H2O2 decomposition assay in siSCR- and catalase siRNA (siCAT)-1-transfected U937 and B8 cells. Absorbance at 240 nm is plotted as function of time (s). A plot of moles of H2O2 consumed/minute in siSCR- versus siCAT-1-transfected B8 cells is shown on right. Graphs represent means±S.E.M. (one-way analysis of variance, Tukey) of N=2 (technical triplicate). ***P⩽0.001. (b) Immunoblot of B8 cells (2 μM Vor) transfected with control (SCR) and catalase (CAT)-targeting siRNA collected 96 h post transfection. Shown are catalase, γH2AX, HO-1, CL-CASP3, and β-actin (loading control). (c) Flow cytometry scatter plots of B8 cells stained with Annexin V-FITC (horizontal axis) versus PI (vertical axis) post siSCR and siCAT transfection. (d) % Apoptotic cells as measured by PI/Annexin V-FITC double-positive and Annexin V-FITC-positive cells in B8 cells maintained in Vor post siSCR and siCAT-1 transfection. Graph represents means±S.E.M. (unpaired t-test) of N=3 (technical triplicate). ***P⩽0.001
HDACi-refractory patients possess elevated peroxisome levels
The above data suggest that we might exploit the dependence on peroxisomal redox proteins as a means of delaying or circumventing resistance to ROS-inducing chemotherapies.
To further address this, we examined peroxisomal transcript expression of purified lymphocytes isolated from biopsies of DLBCL patients who were treated with the HDACi panobinostat. We performed a gene set enrichment analysis (GSEA),38 which examines the expression of multiple genes corresponding to ontology pathways across two data sets; in this case HDACi responders versus non-responders. Using this approach, we analyzed the cDNA levels of transcripts corresponding to the KEGG term ‘peroxisome’ (map 04146) in patients who were refractory versus responsive to HDACi. Specifically, the map consists of 78 genes, of which 49 were detectable upon filtering a restricted set of ~10 000 expression probes. Expression profiles of HDACi-refractory to -responsive patients produced an enrichment score of 0.45 (positive correlation), and false discovery rate (FDR) q-value of 0.09, with 30 of the 49 genes possessing a core enrichment (Figure 6a). Peroxisomal genes associated with fatty acid metabolism such as HSD17B4(multifunctional enzyme type 2),PECR (peroxisomal trans-2-enoyl-CoA reductase),ECI2 (enoyl-CoA delta isomerase 2) and acetylation, and CRAT (carnitine O-acetyltransferase) are amongst the top 5% of enriched genes. Importantly, 3 of the 4 peroxisomal genes involved in plasmalogen synthesis, FAR1 and FAR2 (fatty acyl-CoA reductase 1 and 2) and GNPAT (glyceronephosphate O-acyltransferase) are amongst those with a high running enrichment score. CAT (catalase) is amongst the top 15% of all detectable genes amongst HDACi-refractory patients (Figure 6b). The HDACi panobinostat is subject to different kinetics and metabolism than vorinostat.39 However, we show that similar to Vor, panobinostat drives the production of peroxisomal components and it potentiates apoptosis upon PEX3 knockdown (Supplementary Figure S8). Furthermore, compared to U937 cells, B8 cells are not only cross-resistant to panobinostat23 but also express the highest amounts of peroxisomal proteins when cultured in panobinostat (Supplementary Figure S8).
HDACi-resistant patients possess elevated peroxisome transcripts. (a) GSEA plot of cDNAs from HDACi-refractory versus -responsive diffuse large B-cell lymphoma (DLBCL) patients. An enrichment score (ES) of 0.45 is shown, indicating a positive correlation between HDACi-refractory patients and the KEGG pathway ‘Peroxisome.’ (b) Chart of nominal P-value, FDR q-value, and familywise error rate (FWER) P-value. Gene list rank, metric score, and ES are tabulated for selected genes. (c) Strategies to overcome Vor resistance or to potentiate Vor-induced cell death. In terms of Vor resistance, knockdown of CAT or PEX3 results in increased cell death, while cell death is potentiated in the sensitive cells treated with Vor. Lightly colored peroxisomes indicate low catalase level. Round and shriveled cells portray cell survival and apoptosis, respectively
From our in vitro and patient-derived data, we propose a model whereby peroxisomes can be therapeutically downregulated to potentiate cell death in lymphoma, and forestall or overcome chemotherapy resistance (Figure 6c). When PEX3 is knocked down in HDACi-sensitive cells treated with Vor, less peroxisomes are present, ROS damage increases, and cell death is potentiated. When catalase is knocked down, this results in increased ROS and again a potentiation of Vor-induced cell death (Figure 6c). B8 cells, resistant to Vor, have the highest levels of functional peroxisomes, and maintain low levels of ROS. When PEX3 is knocked down, peroxisome levels decrease and cells undergo apoptosis. Upon catalase knockdown, B8 cells possess a reduced ability to quench Vor-induced (and exogenous) ROS, resulting in cell death (Figure 6c).
Discussion
We provide a comprehensive analysis of the cyto-protective function of peroxisomes in lymphoma by demonstrating that either reducing peroxisome number (PEX3 knockdown) or function (CAT knockdown) can potentiate Vor-mediated cell death and overcome Vor resistance. In the clinical setting, lymphomas that are refractory to HDACi show a gene signature that reflects the presence of increased peroxisome function. We hypothesize that this group of patients may show increased sensitivity to drugs that compromise peroxisome numbers or function.
Vor can induce the formation of peroxisomes, and we speculate that this may occur via a post-transcriptional mechanism. HDACi do not exclusively operate at the transcriptional level via targeting HDACs,40 but can also induce the acetylation of non-histone proteins.26 Specifically, lysine acetylation of proteins can enhance their stability, by opposing the effects of proteasome-targeted ubiquitination.41 Zhao et al.42 observed that cellular metabolism is largely regulated by protein acetylation, with virtually every enzyme associated with fatty acid metabolism shown to be acetylated in human liver tissue. Throughout fatty acid metabolism (oxidation), peroxisomes generate acetyl-CoA,43 which serves as a substrate for acetyltransferases, resulting in lysine acetylation.44, 45 Peroxisomes contain CRAT,46 which could acetylate peroxisomal matrix and membrane proteins in addition to carnitine. Thus, the mechanism by which HDACi regulate peroxisome abundance remains an area of current investigation in our lab.
HDACi treatment promotes peroxisome abundance. To demonstrate what role (increased) peroxisomes have in the cell we show that compromising peroxisome biogenesis and function potentiates Vor-induced apoptosis (Figures 2,3,5, and 6; Supplementary Figures S2–S8). Per the de novo biogenesis model, peroxisome formation begins with PEX3, or PEX16 budding from the ER and interacting with PEX19.47, 48 This is followed by expansion of the peroxisomal membrane and import of peroxisomal matrix proteins, resulting in a mature peroxisome.49, 50 Curiously, upon acute Vor treatment of U937 cells we do not observe increases in ACSL1 and catalase (Figures 3b and 4e–g). This leads us to hypothesize that peroxisome production in response to acute Vor treatment is an insufficient attempt to salvage the pro-apoptotic effect of the drug. However, with acquired resistance to Vor, we put forth that peroxisomes protect drug resistant cancer cells from ROS damage via engaging two antioxidant mechanisms: (1) upregulating catalase; and (2) increasing the levels of plasmalogens (PlsEtn). Interestingly, B8 cells possess levels of PlsEtn 16:0 and 18:0 that are not significantly different than those in Vor-treated U937 cells (Supplementary Figure S4). However, the overall significant increase in total cellular PlsEtn in B8 versus Vor-treated U937 is attributed to elevated levels of 18:1 PlsEtn in B8 (Figure 3c). The relative abundance of this species may cause the downregulation of other plasmalogens such as PlsEtn 16:0 and 18:0 that could be less-effective ROS sinks due to lack of an additional unsaturated bond.
The data we present here using cell models of lymphoma is supported in the clinical setting. We show that the GSEA of the peroxisome KEGG pathway ranks 13th of the 59 enriched (145 total) KEGG pathways in HDACi-refractory patients (Supplementary Table S1), and serves as a predictor of sensitivity versus de novo resistance to HDACi (Figures 6a and b). Specifically, in refractory versus responsive patients, three of four peroxisomal genes involved in plasmalogen synthesis, GNPAT,FAR1, and FAR2 are amongst those with a positive core enrichment (Figure 6b). Thus, we propose that blood biopsies could be used to assess circulating plasmalogens as a less-invasive diagnostic tool than tumor biopsies to monitor the acquisition of resistance or response to HDACi. This approach may not be far off the horizon, as plasmalogen levels in blood are already measured in certain diseases. Reduced plasmalogen levels in erythrocyte membranes are a diagnostic marker for the peroxisome biogenesis disorders, Zellweger spectrum and rhizomelic chondrodysplasia punctata. Investigations of Alzheimer’s patients have shown reductions in their blood plasmalogen levels that correlate with the progression of their neurological disease.51, 52 In the case of patients with lymphoma that are treated with HDACi, an increase from baseline in blood 18:1 PlsEtn levels, a surrogate marker for peroxisome proliferation, would be predicted to correlate with developing resistance to HDACi.
Finally, we have previously shown that mutations in myocyte enhancer factor 2B (MEF2B) were significantly associated with response to panobinostat.53 As MEF2B can bind HDACs,54 we were interested in testing whether any correlation exists between MEF2B mutational status and peroxisomal components. To this end, we interrogated our microarray data using GSEA and found that peroxisome signatures are enriched in the MEF2B wild-type patients (Supplementary Table S2). However, we only captured four MEF2B mutant tumors in our gene expression data set, of which only two responded to HDACi (Supplementary Table S2).
The role of peroxisomes in cancer and their contribution to therapy-resistant disease is waiting to be unveiled. Peroxisomes have a key role in cellular metabolism,55 and considering that a defining feature of cancer is metabolic dysregulation,56 elucidating the role of this critical metabolic organelle may help better understand the disease. Our data suggest that the design of compounds that target peroxisomes in cancer may be a worthy endeavor and should be tested in combination with existing chemotherapies and targeted therapies.
Materials and methods
Cell culture
U937, SU-DHL4, OCI-LY8, and B8 (Vor-resistant derived from U937) cells were maintained at cell densities ranging from 300 000 to 750 000 cells/ml in 75 cm2 vented flasks (Corning, Tewksbury, MA, USA) with 1640 RPMI (Wisent, Saint-Jean-Baptiste, QC, Canada), 10%(v/v) FBS (Wisent) and 0.5% (v/v) penicillin/streptomycin (Wisent). SU-DHL4, U937, and OCI-LY8 cells were treated with 1, 2, and 4μM Vor (Cayman chemicals, Ann Arbor, MI, USA), respectively, while B8 cells were constantly maintained in 2 μM Vor.
qPCR
Cultured cells were pelleted and RNA was isolated using the standard procedure of the TRIzol reagent (Thermo Fisher Scientific, Waltham, MA, USA). RNAs were then quantitated using a NanoDrop spectrophotometer (Thermo Fisher Scientific) and 1 μg of RNA was reverse transcribed using the iScript cDNA synthesis kit (Bio-Rad, Hercules, CA, USA) and PTC-100 thermocycler (Bio-Rad). RT-qPCR reactions were prepared using SYBR Green real-time PCR master mix (Applied Biosystems, Foster City, CA, USA), and samples (400 nM primer duplex) were run using a 7500 Fast Real-Time PCR system (Applied Biosystems). MIQE guidelines were followed for RT-qPCR with the primer sequences (IDT, Coralville, IA, USA),57 standard curve slopes, efficiencies (%), and R2-values as follows: CAT (forward 5′-GGTAACCCAGTAGGAGACAA-3′, reverse 5′-CGAGATCCCAGTTACCATCTT-3′, slope −3.247, 103.24%, R2=0.996); PEX3 (forward 5′-GCCTTAATGCAGCAACTGAA-3′, reverse 5′-AGCTTGTTTGAAGGCCTGTT-3′, slope −3.37, 98.052%, R2=0.994); PEX11B (forward 5′-GTAACTCAGCAGATGCCCTT-3′, reverse 5′-GCTCGATTGAGGTGACTAACA-3′, slope −3.442, 92.936%, R2=0.945); PMP70 (forward 5′-CGAGCATGATGGCCTACTT-3′, reverse 5′-TTTGCTCAGTTATTGTCATCTTACC-3′, slope −3.316, 100.04%, R2=0.997); 36B4 (forward 5′-TCCTCGTGGAAGTGACATCGT-3′, reverse 5′-CTGTCTTCCCTGGGCATCA-3′, slope −3.37, 98.052%, R2=0.994).
Immunoblots
Proteins from cell extracts were lysed in AMPK buffer (50 mM Tris-HCl, pH 7.4, 5 mM sodium fluoride, 5 mM sodium pyrophosphate, 1 mM EDTA, 1 mM EGTA, 250 mM mannitol, 1% v/v Tritox X-100, with protease and phosphatase inhibitors added fresh). Lysates were sonicated for 4 s at 50% power using a sonic dismembranator model 500 (Thermo Fisher Scientific). Acrylamide (19:1 acrylamide/bis) gels were run in SDS glycine running buffer (pH 8.3), with 5% stacking, and 8, 10, or 12% resolving gels, until adequate ladder (Kaleidoscope, Bio-Rad) separation was obtained. Proteins were then transferred onto polyvinylidene fluoride membranes (Roche, Laval, QC, Canada), blocked with 5% skim milk in 1 × TBST, and probed overnight at 4 °C with the following primary antibodies: Pex3 (Sigma, HPA042830); Pex11B (Abcam, Cambridge, UK, AB181066); PMP70 (Abcam, AB3421); Pex16 (Proteintech, Rosemont, IL, USA, 14816-1-AP); Pex19 (Abcam, 130372); HO-1 (Santa Cruz, Dallas, TX, USA, sc-10789); CL-CASP3 (Cell Signaling, Danvers, MA, USA, 9661); γH2AX (Cell Signaling, 2595); CL-PARP (Cell Signaling, 9541); ACSL1 (Cell Signaling, 9189); Catalase (Cell Signaling, 12980); and β-actin (Sigma, St. Louis, MO, USA, A5441). Membranes were briefly rinsed in 1 × TBST then washed twice for 15 min in 1 × TBST, followed by 1 h room temperature (RT) incubation with mouse IgG (GE Healthcare, Little Chalfont, UK) or rabbit IgG (GE Healthcare) secondary antibodies (1/3000 dilution) in 5% skim milk/1 × TBST. Membranes were then washed as mentioned for primary incubation. A chemiluminescence signal was produced via incubation with Amersham (GE Healthcare) or Immobilon (Millipore, Billerica, MA, USA) ECL western blot detection kit which was visualized using X-ray films (Thermo Fisher Scientific).
Immunofluorescence
A total of 750 000 cells were centrifuged at 100 × g for 5 min (media was aspirated and cells were washed twice in 1 ml of cold 1 × PBS. Cell pellets were then resuspended in 200 μl of cold 1 × PBS and 20 μl was dropped onto a cell suspension glass slide, then via a cytospin device (Hettich, Tuttlingen, Germany), centrifuged at 100 × g for 5 min. Samples were left to dry for ~20 min, and surrounding areas were circled with a wax pen. Cells were then fixed in 4% PFA for 20 min, then washed three times with 1 × PBS for 5 min at RT. Cell membranes were then permeabilized using 0.2% Triton X-100 for 10 min in 1 × PBS, then washed in 1 × PBS as mentioned above. Slides were then blocked with 10% BSA/1 × PBS for 1 h at RT and incubated with primary antibodies (1/800 catalase and 1/600 PMP70) in 2% BSA/1 × PBS overnight at 4 °C in a humid chamber. Following incubation, cells were washed three times with 1 × PBS for 5 min at RT, and incubated with 1:250 rabbit 594 (Invitrogen, Carlsbad, CA, USA) and 1:250 mouse 488 (Invitrogen) secondary antibodies for 1 h at RT in a humid, dark chamber. Slides were then washed three times with 1 × PBS at RT for 5 min, followed by incubation with 1:1000 TO-PRO or DAPI nuclear stain for 15 min in 1 × PBS. Cells were then washed 3 × with 1 × PBS, for 5 min each wash and mounted using ProLong gold mounting media (Life Technologies, Carlsbad, CA, USA). Glass coverslips were added on top and sealed with nitrocellulose-based lacquer. Slides were stored in the dark (4 °C) until viewing under a DM IL LED (Leica, Wetzlar, Germany) microscope and images were captured via an Infinity3 (Lumenera, Sarasota, FL, USA) camera. Coloring of images was performed using FIJI software (FIJI, Bethesda, MD, USA).
LC–MS/MS analysis of plasmalogen detection
Authentic standards of plasmalogens were purchased from Avanti Polar Lipids, Inc (Alabaster, Al, USA). The tetradeuterated internal standard lyso-PAF C-16-d4 was purchased from Cayman Chemical Co (Ann Arbor, MI, USA). HPLC-grade solvents (methanol, acetonitrile, chloroform, and water) and ammonium acetate were purchased from Thermo Fisher Scientific. Formic acid was purchased from Sigma-Aldrich (St. Louis, MO, USA). PBS was purchased from Thermo Fisher Scientific. Cell pellets were homogenized in PBS. An extraction solution of methanol containing 10 ng of the internal standards 16:0-D4 lyso-PAF (20.65 pmol) was added to 50 μg protein cell extract in a glass tube. The samples were incubated on a shaker at room temperature for 1 h. The samples were transferred to Corning Costar spin-X centrifuge tube filters (VWR, Radnor, PA, USA) and centrifuged for 5 min. The filtrates were then transferred to autosampler Verex vials (Phenomenex, Torrance, CA, USA) for analysis by LC–MS/MS.
A Waters (Brossard, QC, Canada) TQD interfaced with an Acquity UPLC system was used for positive ion electrospray (ESI)–MS/MS ionization. Plasmalogen species were detected by monitoring multiple reaction monitoring transitions representing fragmentation of [M+H]+ species to m/z 311, 339, 361, 385, 389, and 390 for compounds with 16:1, 18:1, 20:4. 22:6, 22:4, and 18:0, respectively, at the sn-2 position. Chromatographic resolution was achieved via the use of a 2.1 × 50 mm, 1.7 μm Waters Acquity UPLC BEH column. The solvent systems used were as follows: mobile phase A=54.5% water/45% acetonitrile/0.5% formic acid; and mobile phase B=99.5% acetonitrile/0.5% formic acid with both solutions containing 2 mM ammonium acetate. Injections of extracts were made with initial solvent conditions of 85% mobile phase A/15% mobile phase B. The gradient used was from 15 to 100% mobile phase B over a period of 2.5 min, held at 100% mobile phase B for 1.5 min before reconditioning the column back to 85% mobile phase A/15% mobile phase B for 1 min at a solvent rate of 0.7 ml/min. A column temperature of 35 °C and an injection volume of 5 μl were used.
Flow cytometry-based assays
All flow cytometry experiments were conducted on the LSRFortessa (BD Biosciences, San Jose, CA, USA). Cells were initially centrifuged at 100 × g for 5 min, washed twice in 1 × PBS, with 5 min spins at 100 × g, then stained with 5 μM H2-DCFDA (Sigma) for 30 min at 37 °C. Following staining, cells were washed twice in 1 × PBS and resuspended in 0.25 μg/ml PI (Sigma). A minimum of 10 000 gated events were collected per run and reads were performed within 45 min of collecting. For sub G1 population detection, see Dupéré-Richer et al.23 For apoptosis detection of non-fixed cells, PI/FITC-Annexin V kit (BD Pharmingen, San Jose, CA, USA) was used with 300 000 cells/tube, 1.5 μl of FITC-Annexin V, and 0.25 μg of PI, in 1 × binding buffer.
Transient siRNA knockdowns
A total of 2 × 106 cells for each condition were centrifuged at 100 × g, washed twice in PBS, and resuspended in 100 μl of solution containing 100 mM Na2HPO4, 9.375 mM KCl, and 13.75 mM MgCl2 (pH 7.75), placed in 2 mM gap electroporation cuvettes (Harvard Apparatus, Holliston, MA, USA) followed by addition of 10 μl siRNA (stocks at 100 μM). Cuvettes were then electroporated using program W-001 (U937 and B8) and X-001 (OCI-LY8) via the Amaxa Nucleofector device (Lonza, Basel, Switzerland), and contents were placed into six-well dishes with 5 ml of pre-warmed (37 °C) 1640 RPMI, 10% FBS, and 0.5% P/S. After overnight incubation, cells were spun down at 100 × g, media were aspirated, and resuspended in fresh RPMI as above. Equimolar amounts of scrambled negative control DsiRNA (IDT) was used for all transfections. The PEX3-1 siRNA duplex (IDT): 5′-rArCrUrGrCrArArArCrUrGrArArUrGrGrArUrCrUrGrUrCrCrGrUrUr-3′ and 5′-rCrGrGrArCrArGrArUrCrCrArUrUrCrArGrUrUrUrGrCrAGT-3′. The PEX3-2 siRNA duplex sequence (IDT): 5′-rGrArUrCrUrGrArArGrArUrAArUrArArGrUrUrUrCrArCAA-3′ and 5′-rUrUrGrUrGrArArArCrUrUrArUrUrArUrCrUrUrCrArGrArUrCrCrU-3′. PEX19-1 duplex sequence (IDT): 5′-rGrUrGrArArCrArGrUrGrUrCrUrGrArUrCrArUrGrUrGrAAA-3′ and 5′-rUrUrUrCrArCrArUrGrArUrCrArGrArCrArCrUrGrUrUrCrArCrCrA-3′. PEX19-2 duplex sequence (IDT): 5′- rGrArUrArUrGrArCrCrUrCrCrCrArArCrArArGrArArUrUCA-3′ and 5′-rUrGrArArUrUrCrUrUrGrUrUrGrGrGrArGrGrUrCrArUrArUrCrArC-3′. CAT-1 siRNA duplex sequence (IDT): 5′-rArCrCrArArCrUrGrGrGrArUrGrArGrArGrGrGrUrArGrUrCrCrUrU-3′ and 5′-rGrGrArCrUrArCrCrCrUrCrUrCrArUrCrCrCrArGrUrUrGGT-3′. CAT-2 siRNA duplex sequence (IDT): 5′-rArUrUrUrGrGrArGrCrArCrCrArCrCrCrUrGrArUrUrGrUrCrCUG-3′ and 5′- rGrGrArCrArArUrCrArGrGrGrUrGrGrUrGrCrUrCrCrArAAT-3′.
H2O2 decomposition assay
One million cells were centrifuged at 100 × g for 5 min, washed twice in 1 × PBS and pulsed for 4 s at 50 W in a sonic dismembranator model 500 (Thermo Fisher Scientific) in 100 μl 1 × PBS (without inhibitors). Samples were then quantitated via the Bradford method and 10 μg of protein extract was placed in 1 ml 1 × PBS, and used as a blank in 1 cm path length quartz cuvettes. For each condition, 5 μl of 3% (v/v) H2O2 (Sigma) was added to each cuvette and the absorbance at 240 nm (over the course of 60–120 s) was read on a DU 700 UV-VIS spectrophotometer (Beckman Coulter, Brea, CA, USA). Change in moles of H2O2 was calculated using Beer’s law, ΔA=ɛΔCl, with ΔA as absorbance, ɛ as extinction coefficient (43.6 M−1 cm−1 H2O2), C as concentration (M), and l as path length (cm). The change in concentration was determined and then multiplied by the volume in the cuvette to calculate the change in moles of H2O2.
Gene set enrichment analysis
GSEAs (GSEA v2.2.4, San Diego, CA, USA) were performed on the samples from the Q-CROC2 clinical trial in diffuse large B-cell lymphoma testing the efficacy of the deacetylase inhibitor panobinostat.53 We profiled gene expression data from responders and non-responders looking for enrichment in KEGG pathways (using c2.cp.kegg.v6.0.symbols curated data set). Using this unbiased approach, we found that of the 145 pathways, 59 were enriched in non-responders and 86 were enriched in responders. Supplementary Table S1 shows that in non-responders 2 pathways had an FDR <25% and a nominal P-value of <1%. Seven pathways had a nominal P-value of <5%. There were 16 pathways with an FDR <25% in responders, 7 with a nominal P-value <1% and 15 with a nominal P-value of <5%.
GSEA was also performed comparing MEF2B wild-type and mutant DLBCL samples for enrichment in peroxisome pathways (Supplementary Table S2). These pathways were enriched in wild-type MEF2B expressing DLBCL, although only expression data were available on only four MEF2B mutant samples.
Statistical analyses
For comparisons of two groups (e.g., U937 versus B8) and one variable (e.g., PMP70 puncta) the Student’s t-test was used (paired, two-tailed). In comparing more than two groups (e.g., U937+-Vor and B8) with one variable (e.g., catalase puncta) to each other, a one-way ANOVA was performed with Tukey’s comparison of all groups. For more than two groups and multiple variables (e.g., 16:0 PlsEtn and 18:1 PlsEtn) a two-way ANOVA with Bonferroni post-test was used. Prism software Graphpad 5.0 (Graphpad Software, La Jolla, CA, USA) was used to conduct all statistical tests.
Data availability
The data presented in this study are available from the authors upon request.
References
Kim PK, Mullen RT, Schumann U, Lippincott-Schwartz J . The origin and maintenance of mammalian peroxisomes involves a de novo PEX16-dependent pathway from the ER. J Cell Biol 2006; 173: 521–532.
Baudhuin P, Beaufay H, De Duve C . Combined biochemical and morphological study of particulate fractions from rat liver. Analysis of preparations enriched in lysosomes or in particles containing urate oxidase, D-amino acid oxidase, and catalase. J Cell Biol 1965; 26: 219–243.
Smith JJ, Aitchison JD . Peroxisomes take shape. Nat Rev Mol Cell Biol 2013; 14: 803–817.
Heymans HS, Schutgens RB, Tan R, van den Bosch H, Borst P . Severe plasmalogen deficiency in tissues of infants without peroxisomes (Zellweger syndrome). Nature 1983; 306: 69–70.
Kase BF, Bjorkhem I . Peroxisomal bile acid-CoA:amino-acid N-acyltransferase in rat liver. J Biol Chem 1989; 264: 9220–9223.
Wanders RJ, van Roermund CW, van Wijland MJ, Schutgens RB, Heikoop J, van den Bosch H et al. Peroxisomal fatty acid beta-oxidation in relation to the accumulation of very long chain fatty acids in cultured skin fibroblasts from patients with Zellweger syndrome and other peroxisomal disorders. J Clin Invest 1987; 80: 1778–1783.
Lazarow PB . Rat liver peroxisomes catalyze the beta oxidation of fatty acids. J Biol Chem 1978; 253: 1522–1528.
Jakobs BS, Wanders RJ . Fatty acid beta-oxidation in peroxisomes and mitochondria: the first, unequivocal evidence for the involvement of carnitine in shuttling propionyl-CoA from peroxisomes to mitochondria. Biochem Biophys Res Commun 1995; 213: 1035–1041.
Singh H, Derwas N, Poulos A . Very long chain fatty acid beta-oxidation by rat liver mitochondria and peroxisomes. Arch Biochem Biophys 1987; 259: 382–390.
Leighton F, Bergseth S, Rortveit T, Christiansen EN, Bremer J . Free acetate production by rat hepatocytes during peroxisomal fatty acid and dicarboxylic acid oxidation. J Biol Chem 1989; 264: 10347–10350.
Broniec A, Klosinski R, Pawlak A, Wrona-Krol M, Thompson D, Sarna T et al. Interactions of plasmalogens and their diacyl analogs with singlet oxygen in selected model systems. Free Radic Biol Med 2011; 50: 892–898.
Braverman NE, Moser AB . Functions of plasmalogen lipids in health and disease. Biochim Biophys Acta 2012; 1822: 1442–1452.
Fransen M, Nordgren M, Wang B, Apanasets O . Role of peroxisomes in ROS/RNS-metabolism: implications for human disease. Biochim Biophys Acta 2012; 1822: 1363–1373.
Schrader M, Fahimi HD . Peroxisomes and oxidative stress. Biochim Biophys Acta 2006; 1763: 1755–1766.
Willekens H, Chamnongpol S, Davey M, Schraudner M, Langebartels C, Van Montagu M et al. Catalase is a sink for H2O2 and is indispensable for stress defence in C3 plants. EMBO J 1997; 16: 4806–4816.
Sandalio LM, Romero-Puertas MC . Peroxisomes sense and respond to environmental cues by regulating ROS and RNS signalling networks. Ann Bot 2015; 116: 475–485.
Murphy MP . How mitochondria produce reactive oxygen species. Biochem J 2009; 417: 1–13.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 2001; 292: 727–730.
Tabas I, Ron D . Integrating the mechanisms of apoptosis induced by endoplasmic reticulum stress. Nat Cell Biol 2011; 13: 184–190.
Marchi S, Patergnani S, Pinton P . The endoplasmic reticulum-mitochondria connection: one touch, multiple functions. Biochim Biophys Acta 2014; 1837: 461–469.
Fulda S, Galluzzi L, Kroemer G . Targeting mitochondria for cancer therapy. Nat Rev Drug Discov 2010; 9: 447–464.
Banerjee S, Dixit A, Shridharan RN, Karande AA, Chakravarty AR . Endoplasmic reticulum targeted chemotherapeutics: the remarkable photo-cytotoxicity of an oxovanadium(IV) vitamin-B6 complex in visible light. Chem Commun (Camb) 2014; 50: 5590–5592.
Dupere-Richer D, Kinal M, Ménasché V, Nielsen TH, Del Rincon S, Pettersson F et al. Vorinostat-induced autophagy switches from a death-promoting to a cytoprotective signal to drive acquired resistance. Cell Death Dis 2013; 4: e486.
Hu Y, Lu W, Chen G, Zhang H, Jia Y, Wei Y et al. Overcoming resistance to histone deacetylase inhibitors in human leukemia with the redox modulating compound beta-phenylethyl isothiocyanate. Blood 2010; 116: 2732–2741.
Petruccelli LA, Dupéré-Richer D, Pettersson F, Retrouvey H, Skoulikas S, Miller WH et al. Vorinostat induces reactive oxygen species and DNA damage in acute myeloid leukemia cells. PLoS ONE 2011; 6: e20987.
Bali P, Pranpat M, Bradner J, Balasis M, Fiskus W, Guo F et al. Inhibition of histone deacetylase 6 acetylates and disrupts the chaperone function of heat shock protein 90: a novel basis for antileukemia activity of histone deacetylase inhibitors. J Biol Chem 2005; 280: 26729–26734.
Marcu MG, Doyle M, Bertolotti A, Ron D, Hendershot L, Neckers L . Heat shock protein 90 modulates the unfolded protein response by stabilizing IRE1alpha. Mol Cell Biol 2002; 22: 8506–8513.
Singh J, Khan M, Singh I . HDAC inhibitor SAHA normalizes the levels of VLCFAs in human skin fibroblasts from X-ALD patients and downregulates the expression of proinflammatory cytokines in Abcd1/2-silenced mouse astrocytes. J Lipid Res 2011; 52: 2056–2069.
Singh J, Khan M, Pujol A, Baarine M, Singh I . Histone deacetylase inhibitor upregulates peroxisomal fatty acid oxidation and inhibits apoptotic cell death in abcd1-deficient glial cells. PLoS ONE 2013; 8: e70712.
Bjorkhem I, Sisfontes L, Bostrom B, Kase BF, Blomstrand R . Simple diagnosis of the Zellweger syndrome by gas-liquid chromatography of dimethylacetals. J Lipid Res 1986; 27: 786–791.
Faust JE, Manisundaram A, Ivanova PT, Milne SB, Summerville JB, Brown HA et al. Peroxisomes are required for lipid metabolism and muscle function in Drosophila melanogaster. PLoS ONE 2014; 9: e100213.
Eruslanov E, Kusmartsev S . Identification of ROS using oxidized DCFDA and flow-cytometry. Methods Mol Biol 2010; 594: 57–72.
Vermes I, Haanen C, Steffens-Nakken H, Reutelingsperger C . A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labelled Annexin V. J Immunol Methods 1995; 184: 39–51.
Lageweg W, Wanders RJ, Tager JM . Long-chain-acyl-CoA synthetase and very-long-chain-acyl-CoA synthetase activities in peroxisomes and microsomes from rat liver. An enzymological study. Eur J Biochem 1991; 196: 519–523.
Merzliak MN, Iuferova SG, Sobolev AS . Role of superoxide anion-radicals and superoxide dismutase in the lipid photoperoxidation reactions of isolated chloroplasts. Biofizika 1977; 22: 846–849.
Fridovich I . The biology of oxygen radicals. Science 1978; 201: 875–880.
Jones P, Suggett A . The catalase-hydrogen peroxide system. A theoretical appraisal of the mechanism of catalase action. Biochem J 1968; 110: 621–629.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Afifi S, Michael A, Azimi M, Rodriguez M, Lendvai N, Landgren O . Role of histone deacetylase inhibitors in relapsed refractory multiple myeloma: a focus on vorinostat and panobinostat. Pharmacotherapy 2015; 35: 1173–1188.
Glozak MA, Sengupta N, Zhang X, Seto E . Acetylation and deacetylation of non-histone proteins. Gene 2005; 363: 15–23.
Caron C, Boyault C, Khochbin S . Regulatory cross-talk between lysine acetylation and ubiquitination: role in the control of protein stability. Bioessays 2005; 27: 408–415.
Zhao S, Xu W, Jiang W, Yu W, Lin Y, Zhang T et al. Regulation of cellular metabolism by protein lysine acetylation. Science 2010; 327: 1000–1004.
Poirier Y, Antonenkov VD, Glumoff T, Hiltunen JK . Peroxisomal beta-oxidation—a metabolic pathway with multiple functions. Biochim Biophys Acta 2006; 1763: 1413–1426.
Willadsen P, Eggerer H . Substrate stereochemistry of the acetyl-CoA acetyltransferase reaction. Eur J Biochem 1975; 54: 253–258.
Friedmann DR, Marmorstein R . Structure and mechanism of non-histone protein acetyltransferase enzymes. FEBS J 2013; 280: 5570–5581.
Kawamoto S, Ueda M, Nozaki C, Yamamura M, Tanaka A, Fukui S . Localization of carnitine acetyltransferase in peroxisomes and in mitochondria of n-alkane-grown Candida tropicalis. FEBS Lett 1978; 96: 37–40.
Agrawal G, Subramani S . Emerging role of the endoplasmic reticulum in peroxisome biogenesis. Front Physiol 2013; 4: 286.
Agrawal G, Fassas SN, Xia ZJ, Subramani S . Distinct requirements for intra-ER sorting and budding of peroxisomal membrane proteins from the ER. J Cell Biol 2016; 212: 335–348.
Guo T, Gregg C, Boukh-Viner T, Kyryakov P, Goldberg A, Bourque S et al. A signal from inside the peroxisome initiates its division by promoting the remodeling of the peroxisomal membrane. J Cell Biol 2007; 177: 289–303.
Kim PK, Mullen RT . PEX16: a multifaceted regulator of peroxisome biogenesis. Front Physiol 2013; 4: 241.
Wood PL, Mankidy R, Ritchie S, Heath D, Wood JA, Flax J et al. Circulating plasmalogen levels and Alzheimer Disease Assessment Scale-Cognitive scores in Alzheimer patients. J Psychiatry Neurosci 2010; 35: 59–62.
Schutgens RB, Heymans HS, Wanders RJ, van den Bosch H, Tager JM . Peroxisomal disorders: a newly recognised group of genetic diseases. Eur J Pediatr 1986; 144: 430–440.
Assouline SE, Nielsen TH, Yu S, Alcaide M, Chong L, MacDonald D et al. Phase 2 study of panobinostat with or without rituximab in relapsed diffuse large B-cell lymphoma. Blood 2016; 128: 185–194.
Lu J, McKinsey TA, Nicol RL, Olson EN . Signal-dependent activation of the MEF2 transcription factor by dissociation from histone deacetylases. Proc Natl Acad Sci USA 2000; 97: 4070–4075.
Lazarow PB . The role of peroxisomes in mammalian cellular metabolism. J Inherit Metab Dis 1987; 10 (Suppl 1): 11–22.
Ward PS, Thompson CB . Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 2012; 21: 297–308.
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009; 55: 611–622.
Acknowledgements
We are grateful to the patients that enrolled in QCROC-2 clinical trial NCT01238692, C Young and the LDI flow cytometry core facility, as well as M Witcher, N Johnson, C Autexier, LM Calderon, J Vagay, M Flores, and N Advincula for helpful discussions. We are thankful to members of the Lin lab for allowing use of their electroporation device, the Braverman, Miller and Mann labs for critical assessment of data, and CM Kraus for careful reading of this manuscript. This work was supported by the Canadian Institute for Health Research (CIHR; MOP#12683 and MOP#142281), The Samuel Waxman Cancer Research Foundation (WHM), Fonds de Recherche Santé Québec (FRSQ), and the McGill Integrated Cancer Research Training Program (MICRTP).
Author contributions
MSD designed and performed experiments, prepared figures, and wrote the manuscript. EDP performed all plasmalogen assays. ZH performed qPCR experiments. JNN, AMB, and FP aided in experimental design and editing of the manuscript. KKM oversaw the clinical trial and analyzed patient microarray data. DDR developed the Vor-resistant cell line. CG aided in fluorescence microscopy. NB provided tools for the plasmalogen assays and edited the manuscript. SVDR co-supervised the project, designed experiments, and edited the manuscript. WHM co-supervised the project, aided in experiment design, and edited the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by L Scorrano
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Dahabieh, M., Ha, Z., Di Pietro, E. et al. Peroxisomes protect lymphoma cells from HDAC inhibitor-mediated apoptosis. Cell Death Differ 24, 1912–1924 (2017). https://doi.org/10.1038/cdd.2017.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2017.115