Abstract
The transcriptional regulator p53 has an essential role in tumor suppression. Almost 50% of human cancers are associated with the loss of p53 functions, where p53 often accumulates in the nucleus as well as in cytoplasm. Although it has been previously suggested that amyloid formation could be a cause of p53 loss-of-function in subset of tumors, the characterization of these amyloids and its structure-function relationship is not yet established. In the current study, we provide several evidences for the presence of p53 amyloid formation (in human and animal cancer tissues); along with its isolation from human cancer tissues and the biophysical characterization of these tissue-derived fibrils. Using amyloid seed of p53 fragment (P8, p53(250-257)), we show that p53 amyloid formation in cells not only leads to its functional inactivation but also transforms it into an oncoprotein. The in vitro studies further show that cancer-associated mutation destabilizes the fold of p53 core domain and also accelerates the aggregation and amyloid formation by this protein. Furthermore, we also show evidence of prion-like cell-to-cell transmission of different p53 amyloid species including full-length p53, which is induced by internalized P8 fibrils. The present study suggests that p53 amyloid formation could be one of the possible cause of p53 loss of function and therefore, inhibiting p53 amyloidogenesis could restore p53 tumor suppressor functions.
Similar content being viewed by others
Main
p53 has been cast as a sentinel of the cell because it safeguards cells against stress and aberrancies, which threaten the cellular and genomic integrity.1, 2 Disruption in native p53 expression and activity, particularly due to mutation, has been linked to the incidence and progression of cancer.2, 3 Under cellular stress, p53 is primarily involved in transcriptional activity and hence found mostly in the nucleus.1, 4 However, cytoplasmic inclusions of wild-type (WT) and mutant p53 have been observed in several malignant cancers.5, 6 Sequestration of p53 in cytoplasm as large protein aggregates may lead to severe impairment of p53-mediated responses and might inevitably aggravate unregulated cell growth and subsequent tumorigenesis.5, 6 Several reports provide an account of abnormal p53 aggregation and amyloid formation in cancer cells/tissues.7, 8, 9, 10, 11 Amyloid formation is a result of anomalous protein folding, and their consequent aggregation,12, 13 which results in impairment of their regular functions and can have dire consequences for the cell. Amyloid forms of proteins have also shown the ability to ‘seed’ or initiate the aggregation of corresponding native protein molecules in the cellular milieu.14 More importantly, several amyloids possess prion-like ‘infectious’ properties15 wherein they can amplify themselves and transmit between cells, thus resulting in an extensive dissemination of the disease.16 In this context, it has been suggested that p53 aggregates possess prion-like properties in cancer.17, 18, 19
In this study, we present direct evidences of p53 amyloids in human and animal cancer tissues including its isolation and structural characterization. Using a cell model, we show functional inactivation as well as gain-of-tumorigenic functions upon p53 amyloid formation. Further, we observed prion-like properties of p53 amyloids in cells suggesting that this could be the probable mechanism of cancer propagation. Therefore, targeting p53 amyloid formation would be an important approach toward development of cancer therapeutics.
Results
Human and animal cancer tissues contain p53 amyloid
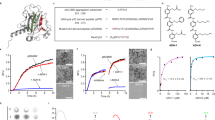
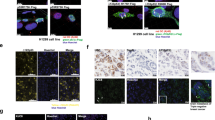
Previously, several reports have suggested the formation of p53 oligomers and amyloids in various tumor tissues7, 9, 10, 20 using amyloid oligomer-specific antibody A11.21 Amyloid-specific antibody OC22 and amyloid-specific dye Thio S, however, were used to detect p53 amyloids in basal cell carcinoma tissues sample.7 In this study, we used OC and Thio S dye to detect p53 amyloid in cancer tissues of human breast, human lung, human urothelial, mouse colon carcinoma and rat hepatocarcinoma. The H&E staining further confirmed the nature of cancer tissues (Supplementary Figure S1). Immunofluorescence co-localization experiments with anti-p53 DO-1 antibody and OC antibody or Thio S staining revealed co-localization of p53 with OC antibody (Figure 1a) as well as Thio S (Supplementary Figure S2) in all cancer tissues but not in the corresponding normal tissues (Supplementary Figure S3). Most of the human and animal cancer tissues also showed strong signals of amyloid oligomer-specific A11 binding (red), with a high degree of co-localization with p53 (green, Figure 1b), which was absent in corresponding normal tissues (Supplementary Figure S4). In contrast, mouse colorectal carcinoma tissues showed weak signals of A11 (red) and negligible co-localization with p53 staining. The data indicate that along with p53 amyloid fibrils, higher-order oligomers (which may not be cytotoxic) could be prevalent in the cancer tissues. Alternatively, p53 aggregates may bind to both OC as well as A11 antibody.
p53 amyloid formation in cancer tissues. (a) Immunofluorescence study showing co-localization of p53 antibody and OC antibody (specific to amyloids) in human and animal carcinomas, suggesting p53 are in amyloid state in these cancer tissues. Scale bars, 50 μm. (b) Immunofluorescence study showing co-localization of p53 antibody and A11 antibody (specific to oligomers) in human and animal carcinomas, suggesting p53 population in amyloid oligomer state in these cancer tissues. Scale bars, 50 μm
We also extracted the tissue amyloid fraction (TAF) from human breast cancer and lung cancer tissues, which showed fibrillar morphology under electron microscope (EM) (Figure 2a). Immunoelectron microscopy further ascertained the presence of p53 in TAF, when 10 nm colloidal gold-conjugated secondary antibody was used against p53 primary antibody. The data showed gold particles aligned along the length of fibrils in TAFs of breast and lung cancer (Figure 2b). The presence of p53 within the TAF fibrils was further confirmed by dot blot using p53-specific antibody (Figure 2c). The TAFs also bound to OC antibody (Figure 2c) and displayed a green-gold birefringence under cross-polarized light, when bound to Congo red (CR), supporting amyloid-like structure (Figure 2d).
Biophysical characterization of p53 amyloid isolated from cancer tissues. (a) Amyloid fibrils were isolated from human breast and lung cancer tissues and examined under TEM. (left) Morphology of amyloid fibrils isolated from breast cancer and (right) amyloid fibrils isolated from lung cancer tissues. Scale bars, 200 nm. (b) Immunoelectron microscopy showing 10 nm gold decorations due to p53 in fibrils isolated from human breast and lung cancer tissues. Scale bars, 200 nm. (c) Dot blot study of cancer TAF showing positive immunoreactivity of amyloid-specific antibody (OC) and p53-specific antibody (DO-1), confirming amyloid nature of fibrils and presence of p53. α-synuclein amyloid fibrils and cell lysate of GFP-p53 cells were used as positive controls for OC and p53 antibody, respectively. (d) Congo red binding study of TAFs from both breast cancer and lung cancer tissues showing green-gold birefringence under cross-polarized light (right). The bright-field images are also shown (left). (e) FTIR spectra of amyloid fibrils isolated from breast cancer and lung cancer tissues in the amide I region (1600–1700 cm−1). Amyloid fibrils isolated from breast cancer showing FTIR peaks at 1620 and 1638 cm−1 in the amide-I region, indicating presence of β-sheet secondary structure. Amyloid fibrils isolated from lung cancer tissues showing FTIR peaks at 1631 and 1618 cm−1, which indicates the presence of β-sheet structure. (f) X-ray diffraction pattern of fibrils isolated from human cancer tissues (both breast and lung carcinomas). The X-ray fiber diffraction showed the meridional reflection at ~4.7 Å and equatorial reflection at 9.3 Å for breast cancer TAF, characteristic of amyloid fibrils. The lung cancer TAF showed meridional reflections at 4.5 Å and equatorial reflection at 13.4 Å along with two additional reflections at 5.7 and 20 Å. The difference in diffraction patterns among the lung cancer and breast cancer TAFs could be due to the difference in the β-sheet spacing in the two TAFs as well as presence of protofibrils in lung cancer TAF fibrils
The FTIR study of TAFs revealed intense peaks at ~1620 and ~1638 cm−1 for lung as well as breast cancer TAF in the amide I region (Figure 2e), suggesting β-sheet-rich structure.23 The X-ray fiber diffraction study showed the meridional reflection at ~4.7 Å and equatorial reflection at 9.3 Å for breast cancer TAF, characteristic of amyloid fibrils24 (Figure 2f). The lung cancer TAF, however, showed meridional reflections at 4.5 Å and equatorial reflection at 13.4 Å along with two additional reflections (which could be due to the different β-sheet spacing24) at 5.7 and 20 Å. The study strongly suggests that though p53 forms amyloid fibrils in cancer tissues, the ultrastructure and the arrangement of these fibrils might vary depending on the in vivo conditions, under which they are formed.
Amyloid fibril formation by wild-type and mutant p53 core domain in vitro
Previous studies have showed p53 aggregation in vitro, where cancer-associated mutations enhanced the aggregation potential of the protein.9, 25, 26 Moreover, many previous studies also showed the aggregation and/or amyloid formation by p53 core domain in vitro (mostly in non-physiological conditions).9, 26, 27, 28, 29, 30, 31 To study the p53 aggregation and amyloid formation in physiologically relevant conditions, we incubated p53 core proteins (WT p53 and mutant (R175H)) for 6 days under various experimental conditions. The data showed amyloid fibril formation by both proteins in presence of TCEP (reduction of disulfide bond, which destabilizes the protein fold32) and glycosaminoglycans, chondroitin sulfate A (CSA is known to increase in cancer tissues33). This is evident by high ThT binding, β-sheet structure in CD, CR-binding (UV absorbance and fluorescence) and fibrillar morphology under EM (Supplementary Figures S5A–C and S6 A, B). The study also revealed the higher propensity of amyloid conversion of p53 core proteins in CSA.
The kinetics of aggregation using ThT binding and CD spectroscopy study showed faster aggregation and conformational transition by R175H mutant (Supplementary Figures S7A and B) compared to WT protein. After aggregation, amyloid-like fibrils were observed under EM (Figure 3a). This aggregation kinetics however is slow compared to the previous studies,9, 26, 30 which could be due to different experimental conditions used in the study. Moreover, the presence of salts can also delay the aggregation kinetics, as both proteins showed much faster aggregation and amyloid fibril formation (within 24 h) in absence of any salts (Supplementary Figure S8A–C).
Characterization of amyloid fibrils formed by p53 WT and mutant core protein in vitro. (a) Electron microscopy showing formation of amyloid-like fibrils by both proteins after incubation for 6 days in presence of CSA. Scale bars, 200 nm. (b) X-ray diffraction study of fibrils formed in presence of CSA showing meridional reflection at ~4.6 Å for both WT and mutant fibrils. The equatorial reflection for WT and mutant was 10.4 and 14.2 Å, respectively suggesting amyloid nature of these fibrils. (c) Proteinase K digestion of amyloid fibrils showing ~15 kDa protease resistant core for both the WT p53 core and mutant fibrils. (d) Equilibrium unfolding studies using guanidine hydrochloride (GdmCl) denaturation. The change in tryptophan fluorescence intensity was plotted versus concentrations of GdmCl showing mutant protein possess less stability than that of WT
The X-ray fibril diffraction of these in vitro fibrils (in presence of CSA) showed pattern corresponding to cross-β-sheet structure (Figure 3b) and displayed the characteristic green-gold birefringence upon binding to CR dye (Supplementary Figure S7C). Both p53 core aggregates also displayed β-sheet characteristics FTIR peaks23 at 1628 and 1634 cm−1 (Supplementary Figure S7D), showed protease resistant core of ~15 kDa (Figure 3c) and burial of tryptophan (Trp) (Supplementary Figure S7E). These studies suggest that in presence of CSA, p53 core domains formed well-defined amyloid fibrils. The dot blot analysis during aggregation using A11 antibody showed oligomeric species were formed at the early stage of aggregation process for both p53 core proteins (Supplementary Figure S7F). Similar results were also obtained with desalted proteins (Supplementary Figure S8D). The cytotoxicity study using MTT assay showed negligible toxicity of both fibrils, suggesting p53 fibrils might not be cytotoxic (Supplementary Figure S9A,B). The guanidine hydrochloride (GdmCl) denaturation using Trp fluorescence study showed mutant p53 core possesses lower stability (free energy of unfolding ~−1.8 kcal/mol) compared to WT core (~−2.6 kcal/mol; Figure 3d). Overall, the present data suggest that amyloid formation of p53 core is enhanced by R175H mutation, which destabilizes the protein fold and promotes amyloid formation.
Cellular model of p53 amyloid
To determine the p53 status in cells, we studied various cell lines SH-SY5Y, L-929, NIH-3T3, MCF-7 and MDA-MB-231. The immunofluorescence data using p53 antibody showed that except MCF-7 and MDA-MB-231, p53 staining was very faint, in other cell lines, indicating a very low basal level of p53 expression (Supplementary Figure S10). Since we observed detectable immunofluorescence of p53 in the nucleus of MCF-7 and MDA-MB-231 cells, we studied p53 aggregation status using OC, A11 antibodies and Thio S staining along with p53 staining. Negligible co-localization was observed for p53 staining with OC as well as with Thio S staining indicating absence of p53 amyloid fibrils in MCF-7 and MDA-MB-231 cells (Figure 4a). Interestingly, in case of MDA-MB-231, a few cells from a large population showed cytoplasmic p53 aggregates, which co-localized with OC and Thio S staining (Figure 4b). This indicates that a low population of cytoplasmic p53 in MDA-MB-231 cells might be in amyloid form. In contrast to OC staining, the A11 staining showed some co-localization of p53 in the nucleus of these cells suggesting amyloid-like oligomers in the nucleus, which is in accordance with previous studies.9, 10
p53 state in cancer cells. Two breast cancer cell lines MDA-MB 231 (p53 mutant) and MCF-7 (WT p53) were cultured and p53 status was checked by immunofluorescence study using p53 antibody, amyloid fibril specific antibody, OC, Thio S staining and amyloid oligomer-specific antibody, A11. (a) MDA-MB 231 and MCF-7 cells showing p53 localization within the nucleus. However, negligible co-localization was observed with OC, A11 and Thio S in the nucleus indicating presence of low amount of oligomeric and fibrillar species of p53. (b) Cytoplasmic aggregates of p53, which co-localized with OC and Thio S were seen in few cells from a large population of MDA-MB- 231. Scale bars are shown
Since most of the cells under study did not showed significant amount of p53 amyloids, we developed an 'in-cell' model of p53 amyloid formation in SH-SY5Y cells by exogenous addition of P8 (p53: 250-257) fibrils.34 These P8 fibril seeds the amyloid formation of WT and mutant p53 core domain in vitro (Supplementary Figure S11A-C). Upon treatment with P8 fibrils for 24 h, SH-SY5Y, L-929, NIH-3T3, MCF-7 and MDA-MB-231 cells showed stabilization of p53 in the cytoplasm as punctate structures (Supplementary Figure S12A). Similar observation was also seen using internalization of p53 core fibrils for SH-SY5Y, MCF7 and MD-MBA231 cells (Supplementary Figure S12B). For studying the functional consequences due to p53 amyloid formation, we chose SH-SY5Y cells, which do not show any significant signal of p53 unless treated with P8/core fibrils (Supplementary Figures S12 and S13A).
For detailed characterization of p53 aggregates due to P8 fibril internalization, the P8 fibril-treated cells were studied using OC antibody staining and Thio S staining. The p53 species were highly co-localized with both Thio S and OC (Figures 5a and b) suggesting p53 amyloid formation in cells. In contrast, much less co-localization with A11 antibody (Figure 5c) was observed indicating low levels of p53 oligomers in the cells. The immunoprecipitated p53 aggregates also bind to OC antibody (Figure 5d) further supporting their amyloid nature. Similar observation was also seen when p53 core fibrils were internalized in SH-SY5Y cells (Supplementary Figure S14). This templating effect is specific for internalized p53 fragment derived fibrils, as internalization of FITC labeled α-synuclein (α-Syn) fibrils seeds (green) did not induce the aggregation and cytoplasmic sequestration of p53 (red) in cells (Supplementary Figure S13B–D).
Cellular model of p53 amyloid using SH-SY5Y cells. (a–c) Aggregation and amyloid formation of p53 in SH-SY5Y cells. SH-SY5Y cells were treated with P8 fibrils and immunofluorescence study was done using p53 antibody (DO-1), amyloid-specific dye Thio S (a), amyloid-specific antibody, OC (b) and oligomer-specific antibody, A11 (c). The images show p53 aggregation and sequestration as punctate structure in the cytoplasm upon P8 fibril treatment. The fluorescence co-localization studies showing (a) Thio S binding and (b) OC antibody binding of p53 aggregates as represented in yellow color, suggesting amyloid nature of these p53 aggregates in cells. (c) The p53 aggregates however did not show significant immunoreactivity against amyloid oligomer-specific A11 antibody, suggesting absence of p53 oligomers in cell due to P8 fibril treatment. Scale bars, 50 μm. (d) The OC antibody-positive immunoreactivity by p53, which immunoprecipitated from cells (by DO-1 antibody) after treatment with P8 fibrils. The data confirming the amyloid nature of p53 aggregates in cells. Untreated cells (-P8) did not show any p53 immunoprecipitation. α-Syn fibrils were used as a positive control. The immunoprecipitaed p53 did not show any binding with A11 antibody. α-Syn oligomer was used as A11-positive control. (e) Immunofluorescence study of P8 fibril-treated cells using p53 antibody (green) and aggresome marker, Proteostat (red) showing co-localization of these two signals. (f) Quantification of aggresome formation by p53 amyloids using flow cytometric analysis. The data are showing negligible Proteostat signal in P8 untreated SH-SY5Y cell (Control), indicating that p53 was degraded in the proteasomes in those cells. Treatment with a proteasome inhibiting agent MG-132 showing cells with robust Proteostat signal confirming the presence of protein aggregates, which are not degraded due to the presence of non-functional proteasomes. Positive signal for Proteostat was also seen in P8 fibril-treated cells signifying the occurrence of p53 aggresomes. (g) Localization of nuclear (left) to cytoplasmic p53 aggregates (right) by GFP-p53 in SH-SY5Y cells upon P8 fibril treatment. These p53 aggregates appeared as cytoplasmic foci (green) displayed co-localization with amyloid antibody OC (red). Scale bars, 50 μm. (h) 'In-cell' fluorescence anisotropy decay kinetics showing faster GFP fluorescence anisotropy decay in P8 fibril-treated cells compared to untreated cells. This shows GFP are sufficiently close in GFP-p53 aggregates
We hypothesized that p53 aggregates are non-toxic to cells and that toxicity, if any, would be further suppressed by the formation of aggresomes,35 which are resistant to proteasomal degradation. Indeed, the p53 cytoplasmic aggregates induced by P8 fibrils formed aggresome as they bind aggresome marker, Proteostat, shown by immunofluorescence (Figure 5e) and by FACS analysis (Figure 5f). Cells treated with proteasome inhibitor MG132 were used as positive control.
To directly demonstrate the transition from nuclear localization to cytoplasmic sequestration of p53 aggregates, due to P8 fibril treatment, we used GFP-p53-transfected SH-SY5Y cells, which showed mostly nuclear localization of GFP-p53 (Figure 5g). Treatment with P8 fibrils resulted in the cytoplasmic sequestration of GFP-p53 aggregates, which showed co-localization with OC antibody (Figure 5g) as well as Proteostat dye (Supplementary Figure S15). The data indicate that the GFP-p53 aggregates in the cytoplasm are in amyloid form. The GFP-p53 aggregates in cells were further analyzed for homo - FRET of GFP fluorescence.36 The fluorescence anisotropy decay kinetics in P8 fibril-treated cells was significantly faster as compared to control cells (Figure 5h), suggesting homo-FRET (close intermolecular distance (<50 Å)) between GFP-p53 molecules in P8 fibril-treated cells. Overall these data suggest that P8 fibrils bind to native p53 protein in cells, causing their aggregation and amyloid formation in the cytoplasm and thereby preventing their degradation.
Loss-of-function of p53 due to amyloid formation
We hypothesized that amyloid formation by p53 in cells prevents its DNA-binding activity and in-turn its transcriptional activity in presence of various stresses. To study this, chromatin immuno-precipitation was carried out using anti-p53 antibody. We observed a marked reduction in the amount of p53 bound with DNA on P8 fibril treatment (Figure 6a), which is expected due to cytoplasmic sequestration of p53.
p53 loss of functions due to amyloid formation. (a) Chromatin immunoprecipitation (ChIP) assay showing negligible p53 pull down for P8 fibrils-treated cells compared to untreated cells, suggesting impairment of DNA binding due to p53 amyloid formation. WCE representing the whole-cell extract. (b) Effect of p53 amyloid formation on its transcriptional regulation. Quantitative real-time PCR depicting upregulation of pro-cancer genes MAPK1, CCND2 and downregulation of anti-cancer genes BAX and DDB2 due to amyloid formation of p53 in SH-SY5Y cell upon treatment with P8 fibrils. Error bars represent standard error. (c) Effect of p53 amyloid formation in drug-induced apoptosis. The cell death analysis of SH-SY5Y cells using Annexin V and PI showing ~40% apoptosis (early and late combined) in response to ActD. P8 fibrils treatment before ActD treatment (P8+ActD) show marked decrease in apoptotic populations (~20% of late and early apoptosis). (d) Effect of p53 amyloid formation on cell cycle arrest. Quantitative cell cycle analysis was done using SH-SY5Y cells showing ~48% arrested cells at G2/M phase of cell cycle when cells were treated with nocodazole. A substantial decrease in G2/M-arrested cells (reduced to 15% of total cells) was seen when cells were pre-treated with P8 fibrils before nocodazole treatment (P8+Noc). The untreated cells were used as control. (e) Cell motility using wound-healing assay showing more migration of SH-SY5Y cells that were treated with P8 fibrils compared to untreated cells. (f) Effect of p53 amyloid formation on adhesion of SH-SY5Y cells showing higher numbers of adhered cells for P8 fibrils-treated cells compared to untreated cells. This suggests more adhesivity of cells due to p53 amyloid formation. (g) Effect of p53 amyloid formation on transforming ability of SH-SY5Y cells. Soft agar colony formation assay of SH-SY5Y cells showing higher number and rapid colony formation by P8 fibrils treated cells compared to untreated cells. Scale bars, 500 μm
To examine the malfunction of transcriptional regulation due to p53 aggregation, we determined the expression of four transcription factors, which are either upregulated (BAX, DDB2) or downregulated (MAPK1 and CCND2) through p53 DNA-binding event.37 The qPCR analysis suggests that although BAX and DDB2 are upregulated in presence of stress (either actinomycin D (ActD) or nocodazole (Noc)) treatment), both are however, downregulated upon p53 amyloid formation (Figure 6b). In contrast, an increase in expression of the oncogenes MAPK1 and CCND2 occurs due to p53 amyloid formation both in presence and absence of stress (Figure 6b). Thus, once p53 forms amyloid, neither can it bind to p53 DNA-binding element nor is able to regulate transcriptional factors for tumor suppression. Moreover, amyloid formation may convert p53 from a tumor suppressor protein to an oncogenic protein as shown for mutant p53.38
To directly demonstrate the loss of p53 tumor suppressive functions due to amyloid formation, we treated SH-SY5Y cells with the DNA-damaging agent, ActD and the microtubule-destabilizing drug, nocodazole, under conditions of P8 fibril-induced amyloidogenesis of p53. FACS analysis revealed ~50% reduction of cell apoptosis (both early and late combined) due to p53 amyloid formation compared to control cells (Figure 6c). This is further supported by decrease in EtBr staining in the nucleus of cells bearing p53 amyloid with treatment of ActD (Supplementary Figure S16). We further studied the effect of p53 amyloid formation on cell cycle arrest in stressed cells. FACS analysis showed 48% population of nocodazole-arrested cells at the G2/M cell cycle stage in normal cells, which decreased to 15% population due to P8 fibril treatment (Figure 6d). Thus, the data confirms the loss of p53 normal function due to its amyloid formation in cells.
Gain-of-function of p53 on amyloid formation
We further hypothesize that p53 amyloid formation may introduce pro-metastatic properties in cells, such as enhanced motility, adhesion and higher transforming capability, reminiscent of cancer cells.38 Indeed, the wound-healing assay revealed an increase in motility of P8 fibril-treated cells as compared to control cells (Figure 6e and Supplementary Figure S17). P8 fibrils-treated cells also exhibited an increase in adhesion to collagen with higher number of focal adhesion complex formation (evident from FAK staining) as compared to control cells (Figure 6f and Supplementary Figure S18A–C). To test whether P8 fibril treatment can result in driving the cells towards a malignant phenotype, we carried out the soft agar assay. Our data confirm that P8-treated cells display enhanced ability to form colonies in soft agar as compared to control cells over the period of 9 days of incubation (Figure 6g and Supplementary Figure S18D). These gain-of-function phenotypes provide evidence that amyloid formation of p53 might be one of the primary causes of loss of cellular integrity and subsequent tumor formation in cancer.
Prion-like properties of p53 amyloid
It has been recently shown that many amyloid-associated neurodegenerative disorders are infectious,16, 39, 40 where amyloid fibrils behave similarly like prion. Since full-length p53 aggregates are able to internalize into cells,41 it was hypothesized that p53 aggregates might possess prion-like properties.17, 19, 41, 42 But thorough studies to establish the prion property of p53 amyloids are lacking. To test prion-like property of p53 amyloids, we demonstrate three distinct properties of p53 amyloid species; (i) internalization in cells, (ii) seeding capacity, where it templates the aggregation of endogenous p53 and (iii) cell-to-cell transmission (schematically represented in Figure 7a).
Internalization and seeding capacity of p53-derived amyloid species. (a) Schematic representation of possible prion-like propagation of p53 amyloids depicting internalization, templating (seeding) and cell-to-cell transmission. (b) Internalization and templating of p53 core fibrils. Rhodamine-labeled p53 core fibrils (Rh-p53C) were added to SH-SY5Y cells and incubated for 24 h. The p53 in the cells was analyzed using immunostaining with anti-p53 antibody. The punctate appearance of p53 signal (green) along with some co-localization (arrows) with Rh-p53C fibrils, indicating internalization and templating property of core fibrils. Scale bars are shown. (c) Internalization of GFP-p53 aggregates in to SH-SY5Y cells. This internalization was only seen when cells were treated with cell lysate of P8 fibrils treated cells (+P8), but not with lysate of untreated cells (−P8) (upper). Internalization of GFP-p53 amyloids quantified using FACS shows increase in the number of cells with GFP-p53 from 0 h to 48 h of culture (lower). (d) Seeding ability of internalized GFP-p53 amyloids on native cellular p53. Cells treated with P8 fibril-treated cell lysate (+P8) showing stabilization of native p53 signal in contrast to control cells (-P8). Scale bars are 50 μm
To demonstrate the internalization and templating capacity of p53 core fibrils, rhodamine-labeled p53 core fibrils were added to SH-SY5Y cells. The red signal inside the SH-SY5Y cells (Figure 7b), demonstrates the internalization of labeled core fibrils. Further after internalization, we observed stabilization of p53 as punctate appearance in cells suggesting p53 aggregation. The co-localization (yellow spot) data suggests that once internalized, core fibrils interact with native p53 expressed inside the cells and aggregate them, which are stable.
For demonstrating the internalization and templating capacity of full-length p53, we expressed GFP-p53 in SH-SY5Y cells and induced the GFP-p53 aggregation using P8 fibrils (Figure 5g). The experimental design is schematically demonstrated in Supplementary Figure S19A. Then, cell lysate with GFP-p53 aggregates was added to freshly cultured SH-SY5Y cells. Fluorescence microscopy data showed (Figure 7c and Supplementary Figure S19B) time-dependent internalization of GFP p53 aggregates into the cells, which was further confirmed by FACS analysis. The control experiment with cell lysate containing no p53 aggregates (without treatment of P8 fibrils) showed no detectable internalization of GFP p53 aggregates in the cells (Figure 7c and Supplementary Figure S19B). Similar observation was also obtained when GFP-p53 amyloids were induced by p53 core fibrils treatment (Supplementary Figure S19C).
The seeding ability of GFP-p53 amyloid after internalization in untransfected cells was tested by immunofluorescence study using p53 antibody (Figure 7d). The p53 staining showed aggregated punctate structure (red color) with a few yellow co-localization signals suggesting that once internalized, GFP-p53 amyloids caused the aggregation of native p53 in the target cells. In contrast, cellular p53 stabilization and aggregation was not observed when cells were treated with lysate containing non-aggregated GFP-p53.
To examine the cell-to-cell transmission, rhodamine-labeled p53 core fibrils were first internalized in SH-SY5Y cells for 24 h. These cells were then co-cultured with GFP expressing cells (Figure 8a, left). Fluorescence microscopy study revealed double-labeled cells (Figure 8a, right), suggesting transmission of p53 core fibrils from its host cells to GFP cells.
Cell-to-cell transmission of p53 amyloids. (a) Schematic representation of cell-to-cell transmission of p53 core fibrils (left). The rhodamine-labeled core fibrils were first internalized in SH-SY5Y cells. These cells were then co-cultured with another set of SH-SY5Y cells expressing GFP. The red-labeled core fibrils were visible in GFP expressing SH-SY5Y cells (right) suggesting inter-cellular transmission of core fibrils. Scale bars are shown. (b) Schematic representation of cell-to-cell transmission of full-length GFP-p53 amyloids (induced by P8 fibrils (left)). The amyloid formation of GFP-p53 in cells was first induced by the addition of P8 fibrils. Once P8 fibrils are internalized into GFP-p53 expressing cells, they template the amyloid formation of GFP-p53. These cells containing full-length GFP-p53 amyloids were then co-cultured with another set of SH-SY5Y cells expressing mCherry. The GFP-p53 signal was visible in mCherry containing cells (right), suggesting transmission of full-length GFP-p53 amyloids to neighboring mCherry-transfected cells. Scale bars are shown
To demonstrate cell-to-cell transmission of full-length p53 amyloids, cells containing GFP-p53 amyloid (due to P8 fibril treatment) were co-cultured with mCherry-expressed cells (for experimental schematic representation, see Figure 8b, left). Fluorescence microscopy study showed GFP-p53 signals were present in the population of mCherry cells after a period of 24 h of co-culture (Figure 8b, right). This suggests transmission of p53 amyloid from GFP cells to mCherry cells. The transmissibility of p53 amyloids suggest that they cross the cell barrier, propagate from one cell to another and induce the amyloid formation in neighboring cells.
Discussion
The presence of p53 amyloids in wide varieties of human and animal cancer tissues (Figure 1) suggest that p53 amyloid formation may be a potential cause of p53 loss of function in cancer7, 9, 10, 20 (Supplementary Figure S20). The functional loss of p53 mainly occurs through p53 mutations, which is augmented by the fact that 50% of human cancers are associated with p53 mutations.3, 43 Since p53 is an amyloidogenic protein,17, 34 it can aggregate to form amyloid in the cells, resulting in loss-of-functions, which may be further promoted by various factors including cancer-associated mutations,25 Zinc depletion44 and abnormal modification of the protein.11 The cytoplasmic p53 amyloid formation may hinder p53 entry into the nucleus as well as prevents its proteasome-mediated degradation,45 besides sequestering many other proteins including other tumor suppressors and p53 homologs and abolishes their native cellular functions.8 Moreover, if p53 amyloids are formed in the nucleus, it cannot bind to the cognate DNA sequence for transcriptional activities. Once p53 aggregates into amyloid, cells do not undergo apoptosis or cell cycle arrest, when exposed to DNA damaging or microtubule-destabilizing conditions (Figures 6c and d). This loss-of-function activity is further accompanied by downregulation of some anti-cancer genes and upregulation of some pro-cancer genes (Figure 6b). Interestingly, p53 amyloid formation in cells also introduced pro-metastatic properties in cells, such as enhanced motility, adhesion and a much higher transforming capability (Figures 6e and g), reminiscent of cancer cells.46 These gain-of-function phenotypes provide evidence that amyloid formation of p53 might be one of the primary causes of loss of cellular integrity and subsequent tumor formation in cancer.
Although amyloid fibrils are known to be cytotoxic and cause cell death in many amyloid diseases,12 p53 amyloids might not be cytotoxic (Supplementary Figure S9A and B). The toxicity of p53 amyloid could be further suppressed by formation of p53 amyloid-associated aggresomes (Figures 5e and f), which is known to be a cellular protective mechanism against the toxicity of protein aggregates.35
p53 amyloid possess ‘seeding’ capacity,41 indicating the possibility that once formed in minute amount inside cells, it can be amplified by transforming native p53 into amyloid state, similar to prions17, 18, 19 (Figure 7). This property can explain the ‘dominant-negative’ activity of p53 amyloids, which can sequester not only functional p53 but also other tumor suppressors and p53 homologs, p63 and p73,8 further weakening the cell’s defense against cancer (Supplementary Figure S20). Moreover, p53 amyloids can transmit from one cell to another in a prion-like manner (Figure 8), which can be severely detrimental as it could enable the widespread inactivation of p53 in cells thereby, transforming ‘the guardian of the genome’47 into a cancer-inducing prionogenic protein. Further, the prion-like ability of p53 amyloids might also enhance the metastatic potential of cancers.
Overall, our studies provide remarkable insight towards the mechanism of cancer progression by classifying cancer as an ‘amyloid disease’ and providing adequate ground to target anti-p53 aggregating agents48 for therapeutic intervention.
Materials and methods
Chemicals and reagents
All chemicals used for the experimental studies were purchased either from Sigma-Aldrich (St. Louis, MO, USA) or Merck (Darmstadt, Germany) and were of the highest purity. Double-distilled and de-ionized water was obtained using a Milli-Q system (Millipore Corp., Bedford, MA, USA). Peptides were custom synthesized using solid-state peptide synthesis method by USV Limited, Mumbai, India with >95% purity, which was checked by HPLC and MALDI-TOF mass spectrometry.
Cell lines and cancer tissues
All mammalian cell lines used in the study were obtained from the cell repository at the National Centre for Cell Science, Pune, India. The cell lines used were SH-SY5Y (human neuroblastoma), L-929 (mouse fibroblast), NIH-3T3 (mouse embryonic fibroblast), MCF-7 (human breast carcinoma cell line with WT p53) and MDA-MB-231 (human breast carcinoma cell line with mutant p53). These cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (Himedia, Mumbai, India). Antibiotic solution (1 ×, Himedia) was also added to the media and the cells were incubated at 37 °C in a humidified incubator at 5% CO2. Freshly frozen human breast and lung cancer and corresponding normal tissues were obtained from the Indian Council of Medical Research (ICMR) national tumor tissue repository at the Tata Memorial Hospital, Mumbai, India. Animal cancer tissues (formaldehyde fixed paraffin-embedded tissue blocks) were obtained from animal cancer biorepository at the Advanced Centre for Treatment, Research and Education in Cancer (ACTREC), India.
Hematoxylin and eosin (H&E) staining
Frozen tissue sections were deparaffinized using decreasing concentrations of the clearing agent xylene (100% xylene, followed by 50% xylene in 50% ethanol) and rehydrated in decreasing concentrations of graded ethanol (100%, 95%, 70% and 50%) and finally with distilled water. Sections were washed with distilled water for 10 min and stained with 0.5% hematoxylin prepared in distilled water for 2 min. The unbound stain was removed using 70 and 90% ethanol wash for 10 min each. Subsequently, 0.8% eosin in 95% ethanol was used to stain the sections for 1 min and kept in xylene for another 1 h. The sections were mounted using DPX mounting solution and observed under a DMi8 microscope (Leica, Wetzlar, Germany) with color HD camera.
Immunohistochemistry of tissues
Paraffin-embedded cancer tissue/biopsies were used for the immunohistochemistry study. All experimental procedures were approved by Institutional animal ethics committee, (Registration no. 65/PO/c/1999/CPCSEA), Indian Institute of Technology Bombay, Powai, Mumbai, India. Tissues were deparaffinized and rehydrated as mentioned above. Enzymatic antigen retrieval was performed on the sections with 0.05% trypsin treatment at 37 °C for 2 min. Sections were washed with TBST (Tris-buffered saline with 0.1% tween-20) at pH 7.4 and treated with 0.2% Triton X-100 in TBST for 10 min. Nonspecific antigenic sites were blocked using TBST containing 2% BSA. The sections were incubated with primary antibodies; mouse monoclonal anti-human p53 protein DO-1 (Santa Cruz Biotechnology, Dallas, TX, USA) antibody (1:200) and rabbit polyclonal oligomer-specific (A11)21 (1:500) or amyloid-specific (OC) antibody22 (1:500) overnight at 4 °C in a humidified chamber. Tissues were further incubated either with secondary antibody of goat anti-mouse FITC (1:500) or goat anti-rabbit Alexa Fluor-647 (Life Technologies, Thermo scientific, USA) for 2 h at room temperature in a humidified chamber.
For Thioflavin S (Thio S) staining, p53 staining was done using goat anti-mouse Alexa Fluor-555-conjugated secondary antibody (1:500 dilution). After antibody staining, the sections were stained with 0.6% Thio S (Sigma-Aldrich) for 2 min in dark. Sections were then washed two times with 50% ethanol followed by TBST and mounted with 1% DABCO (1,4-diazabicyclo-[2.2.2] octane, Sigma-Aldrich) in 90% glycerol and 10% phosphate-buffered saline (PBS). The sections were analyzed using Olympus FV-500 IX 81 confocal microscope (Shinjuku, Tokyo, Japan) and images were acquired using multi-channel image acquisition tool of Fluovision software (Zeiss, Oberkochen, Germany).
Extraction of amyloid fibrils from human cancer tissues
Amyloid fibril from human cancer tissues was isolated using previously published protocol49 with slight modification. Human cancer tissues (250–500 mg) were minced, homogenized for 15 min in 500 μl of 0.15 M NaCl (for 250 mg tissue) and centrifuged at 8600 × g for 1 h at 4 °C. The pellet obtained was re-homogenized in 500 μl of 0.15 M NaCl for 15 min followed by centrifugation at 8600 × g for 1 h at 4 °C. Further, the pellet obtained was homogenized in 500 μl of 0.05 M Tris-HCl, 3 mM NaN3, 0.01 mM CaCl2, pH 7.5. Collagenase type I powder (Himedia) was added at a ratio of 1:100 to the weight of the total tissue and incubated at 37 °C overnight. The mixture was centrifuged at 74 000 × g for 1 h at 4 °C. The pellet was re-homogenized in 300 μl of 0.15 M NaCl and re-centrifuged at 74 000 × g for 1 h at 4 °C. This procedure was repeated until the absorbance of the supernatant was below 0.3. The pellet was then homogenized in 200 μl distilled water and centrifuged at 38 000 × g for 1 h at 4 °C. The supernatant was collected and the homogenization of pellets was repeated until the supernatants turned clear. The cloudy supernatants containing the amyloid fibrils were pooled and NaCl was added for precipitation of fibrils to final concentration of 0.15 M NaCl. This mixture was centrifuged at 74 000 × g for 1 h at 4 °C. The final pellet containing amyloid fibrils was stored at 4 °C and suspended in PBS before use.
Transmission electron microscopy
An amount of 10 μl of TAF isolated from cancer tissues were directly spotted on copper coated formvar grids (Electron Microscopy Sciences, Hatfield, PA, USA), washed with Milli Q water and stained with 10 μl 0.1% uranyl formate (Electron Microscopy Sciences) solution for 20 min. Uranyl formate solution was freshly prepared and passed through 0.22 μm sterile syringe filters (Milipore, Billerica, MA, USA) before use.
For immunoelectron microscopy, the samples were spotted as mentioned above and then incubated with 10 μl anti-p53 DO-1 antibody (1:10) for 20 min. The grids were then washed with Milli-Q water and incubated with 10 μl anti-mouse 10-nm gold-labeled secondary antibody (1:10 dilution, Sigma-Aldrich) for 20 min. Further, the grids were again washed and stained with uranyl formate as mentioned above. The images were taken at different magnifications (× 26 000, × 43 000 and × 60 000) at 120 kV using a transmission EM (TEM) (FeiTecnai12 D312, Waltham, MA, USA).
For in vitro aggregated p53 samples (~70 μM), the incubated solutions (500 μl) were centrifuged at 40 000 r.p.m. for 45 min at 4 °C. The pellet obtained in each case was diluted with 50–70 μl of distilled water, which was spotted on the copper grids and stained with uranyl formate similarly to TAF samples. These grids were imaged using an EM (Philips CM-200, Amsterdam, Netherlands) at 200 kV with magnifications of × 6600. Images were recorded digitally by using the Keen View Soft imaging system (Olympus, Tokyo, Japan).
Dot blot
An amount of 4 μl samples were spotted on nitrocellulose membrane (Millipore) and allowed to air dry. The membranes were treated with blocking solution (5% non-fat skimmed milk powder in TBST) for 1 h at room temperature. These blots were washed with TBST and incubated with primary antibody (anti-p53, 1:200 dilution or OC, 1:500 dilution or A11 1:500 dilution) for 1 h at room temperature on a rocker. After incubation, the blots were washed three times with TBST for 10 min each, followed by 1 h incubation with horseradish peroxidase-conjugated secondary antibody (Calbiochem, San Diego, CA, USA, 1:200 dilution) at room temperature on a rocker. Nonspecific binding was removed by three washes of TBST, each for 10 min. The signals were detected with SuperSignal West Femto kit (Thermo Scientific).
Purification of wild-type and R175H mutant of p53 core domain (94-312 residues)
The WT p53 core protein was expressed and purified from E. coli BL21 (DE3) with N-terminal His6 tag attached to the protein. Briefly, p53 core protein expression was induced by 1 mM isopropyl β-D-1- thiogalactopyranoside (IPTG) at 25 °C for overnight. After induction, the cells were harvested and lysed in 50 mM sodium phosphate, pH 8.0 containing 0.3 M NaCl, 1 mg/ml lysozyme and 1 mM PMSF. The His-tagged p53 core protein was purified from the cell lysate using Ni2+ sepharose affinity chromatography (GE Healthcare, Arlington, TX, USA) and protein was eluted with 100 mM imidazole. Imidazole was then removed by using dialysis against 50 mM sodium phosphate buffer, 300 mM NaCl, pH 7.4. Thereafter protein fractions were checked with 12% SDS–polyacrylamide gel electrophoresis (SDS–PAGE). Proteins were lyophilized and stored at −80 °C for further use. For making desalted protein (used only for measuring amyloid formation kinetics) (Supplementary Figure S8), we used a 10 kDa membrane in a stirred ultrafiltration cell set-up (Amicon Bioseparations, Millipore) fitted to a nitrogen cylinder, to desalt the proteins according to manufacturer’s protocol (Millipore). Site directed mutagenesis was performed to obtain mutant p53 core (R175H). The WT p53 core plasmid (Addgene, Cambridge, MA, USA) was used as a template to clone the cancer-associated p53 mutation R175H. Mutant protein was then expressed in E. coli and purified similar to WT p53 core protein. Protein concentration was measured using UV absorption considering molar absorptivity of protein (ɛ) of 17420/M/cm.
Amyloid formation of p53 core and its R175H mutant in vitro
p53 aggregation was studied in presence and absence of different conditions such as different pH (pH 5.5 and 7.4), in presence of equimolar concentration of two glycosaminoglycans (heparin and chondroitin sulfate A), reducing agents DTT (dithiothreitol) and TCEP (tris(2-carboxyethyl)phosphine). The 70 μM protein in 50 mM sodium phosphate buffer, 0.3 M NaCl, pH 7.4 in presence and absence of equimolar concentration of various additives were placed into a rotating mixture and incubated at 37 °C with slight agitation (~50 r.p.m.). For aggregation study in presence of TCEP, 5% DMSO was present as used in the previous study.31 We also incubated p53 core proteins in presence of 5% DMSO as control. Aggregation of protein was monitored by CD spectroscopy and ThT fluorescence as according to previously established protocol.34 The morphology of the protein aggregates was imaged by TEM at the end of assembly reaction.
For kinetics of aggregation, the assembly reaction was initiated with p53 core and mutant at a concentration of ~70 μM (500 μl) dissolved in 50 mM sodium phosphate buffer, pH 7.4, containing 0.3 M NaCl, 0.01% sodium azide and equimolar concentration of CSA. The protein solutions were incubated at 37 °C with slight agitation and at regular intervals, ThT fluorescence and CD spectroscopy was performed. At the end of aggregation, the aggregates were characterized using various biophysical techniques. Morphology was analyzed using EM, amyloids characteristics was analyzed using CR binding by UV and birefringence method and structural characterization was done using FTIR and X-ray diffractions.
For measuring the kinetics of protein aggregation using desalted protein, 35 μM of proteins were incubated in 50 mM sodium phosphate buffer, pH 7.4, 0.01% sodium azide in presence of equimolar concentration of CSA. The kinetics of aggregation was measured as described above.
For seeding experiments with P8 fibrils, the P8 fibrils were prepared according to our published protocol34 and sonicated to make fibrils seed. P8 fibril seeds were added to 500 μl of WT p53 core (75 μM) and mutant protein (75 μM) in 50 mM sodium phosphate buffer, pH 7.4, 0.01% sodium azide such that final seed concentration was 0.5% (v/v) and 1% (v/v). The kinetics of amyloid formation by core proteins and in presence of P8 fibril seeds was studied by measuring CD and ThT fluorescence at regular intervals. For ThT binding, fluorescence at 480 nm was plotted against time of incubation.
Congo red binding study
Three different methods were used for studying CR binding to amyloid fibrils. For this, 0.2 mg/ml of CR (Sigma-Aldrich) solution was prepared as reported earlier.34 Briefly, 10 μl CR solution was added to 90 μl of incubated protein solution (~70 μM). CR absorbance spectra were recorded in the wavelength range of 300−700 nm using the V-650 spectrophotometer (Jasco, Easton, MD, USA). For CR fluorescence, 10 μl of CR solution was added to the 140 μl incubated proteins and the fluorescence was recorded immediately with excitation at 550 nm and emission in the range of 590−700 nm. The slit width used for both excitation and emission was of 5 nm. Fluoromax4 spectrophotometer (Horiba Scientific, Edison, NJ, USA) was used for fluorescence study.
To study CR birefringence, 10 μl of p53 aggregates (in vitro prepared or in vivo TAF) were mixed with an equal volume of CR solution (0.07% CR in 10% ethanol containing PBS, Sigma). This mixture was centrifuged at 12 000 r.p.m. for 1 min and the pellet was isolated. The pellet was re-suspended in 20 μl of milli Q water. The solution was spread on a glass slide and allowed to dry. The slides were analyzed using the DMi8 microscope (Leica) equipped with cross polarizer and a color HD camera.
Fourier transform infrared spectroscopy (FTIR) study
10 μl of in vitro p53 aggregates and in vivo TAF samples were spotted on a KBr pellet and analyzed by FTIR spectroscopy using a Vertex 80 FTIR system equipped with DTGS detector (Bruker, Ettlingen, Germany). Data corresponding region 1700−1600cm−1 were deconvoluted by Fourier self-deconvolution method. The deconvoluted spectra were then subjected to Lorentzian curve fitting procedure using Opus-65 software (Bruker, Leipzig, Germany) according to manufacturer’s instruction.
Proteinase K digestion
p53 fibril solution (100 μM) formed in the presence of CSA after 6 days incubation were subjected to proteinase K treatment. For this, proteinase K (from stock of 6.5 mg/ml) was added to 100 μM of p53 solution such that final proteinase K concentration became 6.5 μg/ml. After 30 min incubation at 37 °C, the reaction was stopped by adding SDS sample buffer and heating the protein solution at 100 °C for 10 min in a dry bath. Protein sample without proteinase K was considered as 0 min time point. The samples were then analyzed in 12% SDS–PAGE.
Acrylamide quenching of tryptophan (Trp) fluorescence using steady-state fluorescence spectroscopy
We used acrylamide quenching of Trp fluorescence for studying the changes in solvent accessibility of Trp in the WT p53 core and mutant proteins in their monomeric and fibrillar forms.50 For the fluorescence quenching experiment, 200 μl of 12 μ M protein samples (monomeric as well as fibrillar form) were taken in rectangular 10 mm quartz micro-cuvette (Hellma, Forest Hills, NY, USA). The steady-state Trp fluorescence spectrum was recorded using Jobin Yvon Fluoromax 4 fluorimeter by exciting at 280 nm. The excitation as well as emission slit width was set to 7 nm. The fluorescence emission spectrum was recorded in the range 300–500 nm. Thereafter, various concentration of acrylamide (from 5 M stock solution prepared in autoclaved Milli Q water) was added to the protein sample and the fluorescence emission spectrums were taken. All the fluorescence spectra were corrected against volume after addition of acrylamide solution. The Trp fluorescence at 350 nm was plotted against different concentration of acrylamide. F0/F was then plotted against each concentration of acrylamide, where F0 is the fluorescence intensity at emission maxima of Trp fluorescence spectra in the absence of acrylamide and F is the fluorescence intensity at emission maxima of Trp fluorescence spectra in the presence of each acrylamide concentration. The F0/F versus acrylamide concentration plots for each protein sample were compared to get information about the difference in solvent exposure of Trp in each protein sample.
MTT assay for p53 core and mutant fibrils
MTT assay is commonly used to test cellular toxicity of amyloid fibrils.51 To do so, SH-SY5Y cells were seeded in a 96-well plate (~1 × 104 cells per well) and incubated overnight. 100 μl of DMEM containing 10 μM p53 (core and mutant) fibrils were added to wells. Milli Q water, 10 μM Aβ(25–35) fibrils, 1% Triton X-100 and cell-free media were used as controls. The plate was then incubated for 24 h at 37 °C. After that, 10 μl of MTT dye (5 mg/ml, Sigma) was added to each well and incubated for 4 h. Subsequently, 100 μl of DMF-SDS mixture was added and incubated overnight. The absorbance at 560 nm and background scattering at 690 nm were measured in a SpectraMax M2 plate reader (Molecular Devices, LLC, Sunnyvale, CA, USA). The absorbance values at 560 nm were used to calculate the viability of cells in comparison to buffer control and plotted along with standard errors. Experiments were performed two times with three replicates for each sample.
Guanidine hydrochloride (GdmCl) denaturation study
We calculated the free energy of unfolding using Trp fluorescence. Chemical denaturation of proteins was performed by adding GdmCl solution (Pierce, Thermo Scientific) to each of WT p53 core and mutant protein. To do that, p53 proteins in MOPS buffer, pH 7.4 was mixed with increasing concentrations (0, 0.5, 1.0, 1.25, 1.5, 1.75, 2, 3, 4 and 6 M) of GdmCl such that final protein concentration was 10 μM. The protein mixtures were then incubated overnight for equilibrium unfolding at room temperature. After incubation, Trp fluorescence of these samples was measured with excitation at 290 nm and emission in the range of 300–500 nm, keeping the both excitation and emission slit width at 5 nm. The Trp fluorescence intensity at 350 nm was plotted against various concentration of GdmCl and used for free energy calculation (see Supplementary Text).
X-ray diffraction of p53 amyloid fibrils
For X-ray diffraction studies, p53 fibrils from tissue extracts and those formed in vitro were isolated as mentioned in the previous section and were spotted between two clean 0.7 mm capillary tubes. The dried fibril films were then used for X-ray diffraction studies using a Rigaku R Axis IV++ detector (Rigaku, Tokyo, Japan) as reported earlier.52 The data obtained were analyzed using Adxv software (Scripps Research Institute, CA, USA).
Preparation of α-Syn fibril seeds
Recombinant WT α-Syn was expressed and purified in Escherichia coli BL21 (DE3) as previously described.53 Soluble WT α-Syn was labeled with 3 molar excess of fluorescein-5-isothiocyanate (FITC 'Isomer I'), amine-reactive fluorescent dye as per manufacturer’s recommendation (Invitrogen, Carlsbad, CA, USA). The unreacted dye from the reaction was removed by extensive dialysis (~36 h) using mini dialysis unit (10 000 MWCO) (Millipore) in PBS (pH 7.4). The protein solution was then lyophilized and stored at −20 °C for further use. For labeled fibril preparation, soluble, low molecular weight of WT α-Syn and FITC labeled α-Syn in 20 mM glycine-NaOH buffer, pH 7.4 was mixed in 9:1 ratio and was incubated at 37 °C in rotating shaker (EchoTherm model RT11, Torrey Pine Scientific, Carlsbad, CA, USA) at ~50 r.p.m. for two weeks for the formation amyloid fibrils. The fibril was sonicated with 40% amplitude, 3 s on, 1 s off cycle for 3 min for making the fibril seeds. The morphology of the sonicated fibrils was examined using TEM and was used for cellular internalization.
Labeling of p53 core fibrils
For labeling of p53 core fibrils, freshly prepared core fibrils were labeled with 10 molar excess of Rhodamine dye as per manufacturer’s recommendation (Invitrogen). Labeled fibrils were centrifuged at 13 000 × g for 45 min followed by two times washing with Milli-Q. Fibrils were then sonicated with 40% amplitude, 3 s on, 1 s off cycle for 3 min for fibril making seeds.
In-cell seeding using amyloid fibrils
For in-cell seeding experiments, the SH-SY5Y cells were cultured with or without P8/core fibrils (10 μM final concentration, sonicated for 3 min, 20% amplitude, 3 s on/off pulse) for 24 h at 37 °C. The internalization of core fibril in the cells was confirmed by confocal microscopy. The aggregation of cellular p53 protein was analyzed by immunofluorescence using DO-1 anti-p53 antibody. For α-Syn fibril internalization, 1 μM α-Syn fibril seeds were added to the cells and incubated for 24 h. The coverslips containing cells were fixed and were mounted with 1% DABCO (Sigma-Aldrich). The slides were observed and imaged under Leica DMi8 microscope in DIC and in fluorescent mode.
Immunostaining of cells
Cells were washed with PBS (phosphate-buffered saline) at pH 7.4 and fixed using 4% paraformaldehyde for 20 min. The fixed coverslips were treated with 0.2% Triton X-100 in PBS for 10 min. Nonspecific antigenic sites were blocked using PBST containing 2% BSA. The coverslips were incubated with primary antibodies; mouse monoclonal anti-human p53 protein DO-1 (Santa Cruz Biotechnology) antibody (1:200) and rabbit polyclonal oligomer-specific (A11) (1:500) or amyloid fibrils specific (OC) antibody (1:500) overnight at 4 °C in a humidified chamber. The coverslips were washed three times with PBS and then further incubated either with secondary antibody of goat anti-mouse FITC (1:500) or goat anti-rabbit Alexa Fluor-647 (Life Technologies, Thermo scientific) for 2 h at room temperature in a humidified chamber.
For Thioflavin S (Thio S) staining, p53 staining was done using goat anti-mouse Alexa Fluor-555-conjugated secondary antibody (1:500 dilution). After antibody staining, the coverslips were stained with 0.6% Thio S (Sigma-Aldrich) for 2 min in dark. Coverslips were then washed two times with 50% ethanol followed by PBST and mounted with 1% DABCO (1,4-diazabicyclo-[2.2.2] octane, Sigma-Aldrich) in 90% glycerol and 10% PBS. The cells were analyzed using Olympus FV-500 IX 81 confocal microscope and images were acquired using multi-channel image acquisition tool of Fluovision software.
Immunoprecipitation of p53 from SH-SY5Y cells
SH-SY5Y cells were grown and treated with 10 μM P8 fibrils for 24 h. Immunoprecipitation was carried out as per manufacturer’s protocol in non-denaturing conditions (Abcam, Cambridge, MA, USA). Briefly, cells were lysed using lysis buffer (20 mM Tris-HCl, pH 8.0, 137 mM NaCl, 1% NP40, 2 mM EDTA and protease inhibitor cocktail) with constant agitation for 30 min at 4 °C and centrifuged at 12 000 r.p.m. for 20 min at 4 °C. Supernatant obtained was incubated with 1 μg of anti-p53 antibody (DO-1) for overnight at 4 °C under rotation. Further, 100 μl of sepharose G beads were added and protein mixtures were incubated for 4 h at 4 °C under rotary agitation and then centrifuged at 3000 r.p.m. for 20 s. Finally, proteins were eluted with 150 μl of 0.2 M glycine buffer, pH 2.6 by 10 min incubation with frequent agitation and centrifugation at 3000 r.p.m. for 2 min at 4 °C. The eluates were pooled and neutralized by adding equal volume of Tris HCl, pH 8.0.
Transfection of cells
SH-SY5Y cells were grown in a 24-well plate. Transfection was carried out with Lipofectamine plus reagent (Invitrogen) according to manufacturer’s instructions. To do that, cells were washed with PBS before transfection. For transfection, 1 μg of GFP-p53 (Addgene, plasmid#12091) or m-Cherry/CAG-GFP construct (kind gift from Prof. Jonaki Sen, IIT Kanpur, India) was added to mixture of 1 μl of Lipofectamine reagent and 25 μl of Opti-MEM medium and incubated for 10 min. This mixture was then added to the cells. After transfection, cells were incubated in complete media (DMEM with 10% FBS) for 48 h and observed for expression of GFP/mCherry. Only GFP-p53 transfected cells were then selected using 50 μg/ml G-418 in complete DMEM.
Proteostat binding assay and flow cytometry analysis
To study whether p53 aggregates/amyloids were sequestered into aggresomes, Proteostat (an aggresome binding dye) binding assay54 was performed using confocal microscopy and flow cytometry analysis. For immunofluorescence experiments, untransfected SH-SY5Y cells and GFP-p53 transfected SH-SY5Y cells were grown in a 24-well plate. Cells were treated with 10 μM P8 fibrils for 24 h. Cells treated with proteasome inhibitor (5 μM MG-132) for 24 h were used as positive control. Cells were fixed with 4% paraformaldehyde and permeabilized with 0.1% Triton X-100. 0.5 μl of Proteostat (5 μM, Enzo Life Sciences, Farmingdales, NY, USA) was added to 1 ml of assay buffer provided with the kit. The cells were stained and analyzed by confocal microscopy using Proteostat aggresome detection reagent (ENZ-51035-K100) as per manufacturer’s protocol.
For flow cytometric analysis, ~1 × 106 SH-SY5Y cells were stained using Proteostat aggresome detection reagent (ENZ-51035-K100, Enzo Life Sciences) as per manufacturer’s protocol. Cells were studied using a BD FACS Aria flow cytometer using Texas Red filter (BD Biosciences, San Jose, CA, USA). Fluorescence count from the cells (Y-axis) was plotted against cell number (X axis) to compare Proteostat binding in cells.
In cell fluorescence anisotropy decay
To study whether p53 are in close proximity to each other in p53 aggregates that observed in cells, fluorescence anisotropy decay studies were performed.36 To do that, GFP-p53 expressing SH-SY5Y cells were grown on glass-bottom 35 mm petri dishes. 10 μM P8 fibrils were added to induce aggregation of GFP-p53 expressed in cells. Untreated cells expressing GFP-p53 were used as control.
Homo-FRET between GFP-p53 in cells36 were measured using the time-resolved fluorescence microscope setup with a combination of a picosecond time-resolved fluorescence spectrometer and an inverted epifluorescence microscope. The instrumental setup is mentioned in the Supplementary Information.
Chromatin immunoprecipitation
Chromatin immunoprecipitation (ChIP) study is performed to determine p53 proteins’ ability to bind its cognate DNA sequence in cells.55 SH-SY5Y cells (2.5 × 105) incubated with or without P8 fibril for 24 h were used for ChIP study as described previously55 using 5 μg (25 μl) of p53 antibody (DO-1). DNA from all the samples (IPs and inputs) were purified using QIAquick PCR purification kit (QIAGEN, Valencia, CA, USA) according to the manufacturer's instructions and eluted with 50 μl of elution buffer. qPCR was performed using an Agilent qPCR machine in 20 μl SYBR Green reaction mixture. All the ChIP experiments were performed three times for which qPCR was performed in triplicates and the values were shown with error bars depicting standard deviation. Calculations for enrichment/input values were made using equation: ΔCT=CT(ChIP)−[CT(Input)−LogE (Input dilution factor)] where, E represents specific primer efficiency value; % Enrichment/Input=E−ΔCT.
Wound-healing assay
To determine the effect of p53 aggregation on cell motility (migration capability) wound-healing assay was performed.56 To do that, SH-SY5Y cells cultured on 12-well plate were treated with 10 μM P8 fibrils for 24 h. Untreated cells were used as controls. Cells were then washed with PBS, pH 7.4 and scratched with a sterile plastic tip. Wounds were made in duplicates. The peeled off cells were removed with two PBS washes and fresh media was added to the cells. Mitomycin C (0.5 μg/ml) was added as a cell proliferation inhibitor. Cells were further incubated for 48 h to allow wound coverage. Images at zero and final time-points were acquired under a bright-field microscope (Leica DMi1). Wound width was measured using the ImageJ software. Cell migration was calculated using the formula: % Cell Migration=100 × (0 time wound width- final wound width)/(0) time wound width).
Cell adhesion and focal adhesion complex formation studies
To determine the effect of p53 amyloid formation in cells for their adhesion capability,57 SH-SY5Y cells were treated with 10 μ M P8 fibrils for 24 h and plated on to glass coverslips in 24-well plates with a plating density of 4 × 104 per well. The cells were then allowed to adhere on to collagen (10 μg/cm2) -coated coverslips for 24 h. For evaluation of cell adhesion, the glass coverslips in 24-well plates were gently rinsed with PBS to remove unattached cells. Subsequently, the coverslips were imaged in phase contrast DMi1 microscope (Leica) with 10 × magnification. The number of adhered cells for each condition was quantified by manual counting of cells per frame for at least 10 frames per condition. Further, to analyze the distribution of focal adhesion complex in the adhered cells, immunostaining with anti FAK antibody was performed as previously described52 and imaged with fluorescence microscope (Zeiss Axiocam MRM, Oberkochen, Germany). The distribution of focal adhesions was calculated using image processing in Image J (NIH, NY, USA).
Soft agar assay
Soft agar assay was done to study the tumorigenic potential of transformed cells due to treatment of P8 fibrils.58 Untreated cells were used as control. To do that, 1.2% agarose was prepared in sterile water and maintained at 42 °C in a water bath. 2 × DMEM medium (GIBCO, Waltham, MA, USA) was prepared, filtered and also maintained at 42 °C. Both of these solutions were mixed in an equal ratio and 0.25 ml of this mixture was added to a 12-well cell culture plate and allowed to solidify at room temperature. SH-SY5Y cells (~2 × 106) were mixed in 1 ml of 0.3% agarose, prepared in DMEM medium and 0.25 ml was added on the solidified agar layer such that each well contained 7500 cells. The top layer was allowed to solidify at room temperature for 20 min. The plates were then incubated at 37 °C in a humidified incubator for 9 days and images were acquired using a bright-field microscope (Leica DMi1) periodically.
RNA isolation and qRT-PCR
SH-SH5Y cells were cultured and treated separately each with P8 fibrils (10 μ M), ActD (5 ng/ml), nocodazole (5 nM), P8 fibrils (10 μ M) followed by ActD (5 ng/ml) and P8 fibrils (10 μ M) followed by nocodazole (5 nM). Untreated cells (without any addition of compounds) were used as control. The treated cells were further used for RNA isolation using TriZol (Invitrogen) method accordingly to the manufacturer’s protocol. Concentration of the isolated RNA was measured in nanodrop spectrophotometer (Implen, West Village, CA, USA). cDNA synthesis was done with ProtoScript-II first strand cDNA synthesis kit (NEB) according to manufacturer’s protocol with a mixture of oligo-dT and random hexamer primers. Real-time PCR (qRT-PCR) was performed using SYBR Green method. Brilliant-III ultrafast SYBR Green qPCR master mix (Agilent Technologies, Santa Clara, CA, USA) was used according to manufacturer’s protocol on an Agilent MX3000P with ROX as passive dye.
Apoptosis and cell cycle arrest studies using flow cytometry
To study the effect of p53 amyloid formation in cells on its tumor suppressive functions (apoptosis and cell cycle arrest), studies were performed using flow cytometry and immunofluorescence. To do that, SH-SY5Y cells were cultured and treated with and without 10 μ M P8 fibrils for 24 h and subsequently with ActD (2 μg/ml) for another 24 h to analyze cell death by apoptosis. For studying apoptosis, treated/untreated cells were stained with Annexin V-FITC and PI (Annexin V-FITC Apoptosis Detection kit, Sigma Aldrich) according to manufacturer’s instructions and were quantified in a flow cytometer (FACS Aria, BD Biosciences) and analyzed using the BD FACS Diva software. For each sample, 20 000 cells were analyzed. For EtBr staining, similar treatments were done with SH-SY5Y cells, fixed and stained with anti-p53 antibody and 1 μg/ml EtBr solution for 5 min. The cells were mounted and imaged under an Olympus FV-500 IX 81 confocal microscope. Cells incubated with the fluorochrome conjugated secondary antibodies alone were used as negative controls.
For studying cell cycle arrest, cells were treated with 10 μ M P8 fibrils for 24 h and subsequently 5 nM nocodazole for another 24 h. Untreated cells and cells treated either with nocodazole or P8 fibrils were used as controls. Cells were trypsinized, washed with PBS and fixed by adding 70% cold ethanol. Cells were kept at 4 °C for 1 h and washed two times with PBS. 50 μl of RNase A solution (final concentration 0.5 μg/ml) was added to cells and incubated for 4 h at 4 °C to ensure that only DNA is stained. 200 μl propidium iodide in PBS (from 50 μg/ml stock solution) was added to cells to stain the DNA prior to analysis by flow cytometry. For each sample, 20 000 cells were analyzed.
Internalization and seeding behavior of cell-free GFP-p53 aggregates
To examine the seeding behavior of full-length p53 amyloids in cells, transfected SH-SY5Y cells (~1 × 106 cells) expressing GFP-p53 were first treated with 10 μ M P8/p53 core fibrils for 24 h. After 24 h, these cells were lysed using a sterile 21-gauge needle and the lysate was centrifuged at 1500 r.p.m. at 4 °C for 15 min. The supernatant containing aggregated GFP-p53 was then added to untransfected SH-SY5Y cells and were cultured for another 48 h. The internalization and seeding ability of p53 aggregates were studied using immunofluorescence and flow cytometry (only for P8 fibril-treated cells). As control, lysate was prepared from GFP-transfected SH-SY5Y cells without any treatment of P8/core fibrils and was added to untransfected cells. To determine the seeding ability of internalized GFP-p53 aggregates, native p53 in SH-SY5Y cells was immunostained, using anti-p53 DO-1 antibody (only for P8 fibril-treated cells).
Cell-to-cell transfer of p53 amyloids
To study the cell-to-cell propagation of p53 core fibrils, rhodamine-labeled core fibrils was first internalized in SH-SY5Y cells. For this, 1 × 104 SH-SY5Y cells were seeded onto 12 mm cover glass in a 24-well plate and cultured for 24 h. Subsequently, freshly prepared rhodamine-labeled core fibrils were added to cells at a concentration of 10 μM and incubated for 24 h. These treated cells were then thoroughly washed with DPBS (three times), trypsinized and mixed with trypsinized SH-SY5Y cells transfected with GFP plasmid. This mixture of cells were then co-cultured for 48 h. After 48 h, cells were washed once with DPBS and fixed with 4% paraformaldehyde at RT for 20 min. Fixed cells were then mounted on a glass slide and imaged under an Olympus FV-500 IX 81 confocal microscope for dual-labeled cells.
To study cell-to-cell propagation of full-length p53 amyloids, aggregation of GFP-p53 in transfected SH-SY5Y cells was induced by adding P8 fibrils. For this, GFP-p53 transfected SH-SY5Y cells (1 × 105) were seeded onto 12 mm cover glass in a 24-well cell culture plate and cultured for 24 h. Subsequently, these cells were treated with 10 μ M P8 fibrils for 24 h. These treated cells were trypsinized and were then co-cultured with another population of mCherry-transfected SH-SY5Y cells. After 24 h of co-culture experiment, cells were fixed with 4% paraformaldehyde and mounted on a glass slide, followed by imaging with confocal microscope Olympus FV-500 IX 81 for identifying dual-labeled cells.
Statistical analysis
The statistical significance was determined by one-way ANOVA followed by Newman–Keuls Multiple Comparison post hoc test; ∗P<0.01; ∗∗P<0.001.
References
Levine AJ . p53, the cellular gatekeeper for growth and division. Cell 1997; 88: 323–331.
Vousden KH, Lu X . Live or let die: the cell's response to p53. Nat Rev Cancer 2002; 2: 594–604.
Muller PAJ, Vousden KH . p53 mutations in cancer. Nat Cell Biol 2013; 15: 2–8.
Vogelstein B, Lane D, Levine AJ . Surfing the p53 network. Nature 2000; 408: 307–310.
Moll UM, LaQuaglia M, Benard J, Riou G . Wild-type p53 protein undergoes cytoplasmic sequestration in undifferentiated neuroblastomas but not in differentiated tumors. Proc Natl Acad Sci USA 1995; 92: 4407–4411.
Bosari S, Viale G, Bossi P, Maggioni M, Coggi G, Murray JJ et al. Cytoplasmic accumulation of p53 protein: an independent prognostic indicator in colorectal adenocarcinomas. J Natl Cancer Inst 1994; 86: 681–687.
Lasagna-Reeves CA, Clos AL, Castillo-Carranza D, Sengupta U, Guerrero-Muñoz M, Kelly B et al. Dual role of p53 amyloid formation in cancer; loss of function and gain of toxicity. Biochem Biophys Res Commun 2013; 430: 963–968.
Xu J, Reumers J, Couceiro JR, De Smet F, Gallardo R, Rudyak S et al. Gain of function of mutant p53 by coaggregation with multiple tumor suppressors. Nat Chem Biol 2011; 7: 285–295.
Ano Bom AP, Rangel LP, Costa DC, de Oliveira GA, Sanches D, Braga CA et al. Mutant p53 aggregates into prion-like amyloid oligomers and fibrils: implications for cancer. J Biol Chem 2012; 287: 28152–28162.
Levy CB, Stumbo AC, Ano Bom AP, Portari EA, Cordeiro Y, Silva JL et al. Co-localization of mutant p53 and amyloid-like protein aggregates in breast tumors. Int J Biochem Cell Biol 2011; 43: 60–64.
Nieva J, Song BD, Rogel JK, Kujawara D, Altobel L 3rd, Izharrudin A et al. Cholesterol secosterol aldehydes induce amyloidogenesis and dysfunction of wild-type tumor protein p53. Chem Biol 2011; 18: 920–927.
Chiti F, Dobson CM . Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 2006; 75: 333–366.
Maji SK, Wang L, Greenwald J, Riek R . Structure-activity relationship of amyloid fibrils. FEBS Lett 2009; 583: 2610–2617.
Harper JD, Lansbury PT Jr . Models of amyloid seeding in Alzheimer's disease and scrapie: mechanistic truths and physiological consequences of the time-dependent solubility of amyloid proteins. Annu Rev Biochem 1997; 66: 385–407.
Prusiner SB . Molecular biology and pathogenesis of prion diseases [review]. Trends Biochem Sci 1996; 21: 482–487.
Jucker M, Walker LC . Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 2013; 501: 45–51.
Silva JL, Gallo CVDM, Costa DCF, Rangel LP . Prion-like aggregation of mutant p53 in cancer. Trends Biochem Sci 2014; 39: 260–267.
Silva JL, Rangel LP, Costa DCF, Cordeiro Y, De Moura Gallo CV . Expanding the prion concept to cancer biology: dominant-negative effect of aggregates of mutant p53 tumour suppressor. Biosci Rep 2013; 33: e00054.
Costa DCF, de Oliveira GAP, Cino EA, Soares IN, Rangel LP, Silva JL . Aggregation and prion-like properties of misfolded tumor suppressors: is cancer a prion disease? Cold Spring Harb Perspect Biol 2016; 8: a023614.
De Smet F, Saiz Rubio M, Hompes D, Naus E, De Baets G, Langenberg T et al. Nuclear inclusion bodies of mutant and wild-type p53 in cancer: a hallmark of p53 inactivation and proteostasis remodelling by p53 aggregation. J Pathol 2017; 242: 24–38.
Kayed R, Head E, Thompson JL, McIntire TM, Milton SC, Cotman CW et al. Common structure of soluble amyloid oligomers implies common mechanism of pathogenesis. Science 2003; 300: 486–489.
Kayed R, Head E, Sarsoza F, Saing T, Cotman C, Necula M et al. Fibril specific, conformation dependent antibodies recognize a generic epitope common to amyloid fibrils and fibrillar oligomers that is absent in prefibrillar oligomers. Mol Neurodegener 2007; 2: 18.
Jackson M, Mantsch HH . The use and misuse of FTIR spectroscopy in the determination of protein structure. Crit Rev Biochem Mol Biol 1995; 30: 95–120.
Sunde M, Serpell LC, Bartlam M, Fraser PE, Pepys MB, Blake CC . Common core structure of amyloid fibrils by synchrotron X-ray diffraction. J Mol Biol 1997; 273: 729–739.
Bullock AN, Henckel J, DeDecker BS, Johnson CM, Nikolova PV, Proctor MR et al. Thermodynamic stability of wild-type and mutant p53 core domain. Proc Natl Acad Sci USA 1997; 94: 14338–14342.
Cino EA, Soares IN, Pedrote MM, de Oliveira GAP, Silva JL . Aggregation tendencies in the p53 family are modulated by backbone hydrogen bonds. Sci Rep 2016; 6: 32535.
Ishimaru D, Maia LF, Maiolino LM, Quesado PA, Lopez PCM, Almeida FCL et al. Conversion of wild-type p53 core domain into a conformation that mimics a hot-spot mutant. J Mol Biol 2003; 333: 443–451.
Ishimaru D, Andrade LR, Teixeira LS, Quesado PA, Maiolino LM, Lopez PM et al. Fibrillar aggregates of the tumor suppressor p53 core domain. Biochemistry 2003; 42: 9022–9027.
Butler JS, Loh SN . Folding and misfolding mechanisms of the p53 DNA binding domain at physiological temperature. Protein Sci 2006; 15: 2457–2465.
Wang G, Fersht AR . First-order rate-determining aggregation mechanism of p53 and its implications. Proc Natl Acad Sci USA 2012; 109: 13590–13595.
Wilcken R, Wang G, Boeckler FM, Fersht AR . Kinetic mechanism of p53 oncogenic mutant aggregation and its inhibition. Proc Natl Acad Sci USA 2012; 109: 13584–13589.
Burns JA, Butler JC, Moran J, Whitesides GM . Selective reduction of disulfides by tris(2-carboxyethyl)phosphine. J Org Chem 1991; 56: 2648–2650.
Suwiwat S, Ricciardelli C, Tammi R, Tammi M, Auvinen P, Kosma V-M et al. Expression of extracellular matrix components versican, chondroitin sulfate, tenascin, and hyaluronan, and their association with disease outcome in node-negative breast cancer. Clin Cancer Res 2004; 10: 2491–2498.
Ghosh S, Ghosh D, Ranganathan S, Anoop A, P SK, Jha NN et al. Investigating the Intrinsic aggregation potential of evolutionarily conserved segments in p53. Biochemistry 2014; 53: 5995–6010.
Kopito RR . Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol 2000; 10: 524–530.
Sharma P, Varma R, Sarasij RC, Ira, Gousset K, Krishnamoorthy G et al. Nanoscale organization of multiple GPI-anchored proteins in living cell membranes. Cell 2004; 116: 577–589.
Stracquadanio G, Wang X, Wallace MD, Grawenda AM, Zhang P, Hewitt J et al. The importance of p53 pathway genetics in inherited and somatic cancer genomes. Nat Rev Cancer 2016; 16: 251–265.
Muller PAJ, Vousden KH . Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell 2014; 25: 304–317.
Aguzzi A, Rajendran L . The transcellular spread of cytosolic amyloids, prions, and prionoids. Neuron 2009; 64: 783–790.
Eisenberg D, Jucker M . The amyloid state of proteins in human diseases. Cell 2012; 148: 1188–1203.
Forget KJ, Tremblay G, Roucou X . p53 aggregates penetrate cells and induce the co-aggregation of intracellular p53. PLoS One 2013; 8: e69242.
Rangel LP, Costa DCF, Vieira TCRG, Silva JL . The aggregation of mutant p53 produces prion-like properties in cancer. Prion 2014; 8: 75–84.
Joerger AC, Fersht AR . Structural Biology of the Tumor Suppressor p53. Annu Rev Biochem 2008; 77: 557–582.
Butler JS, Loh SN . Structure, function, and aggregation of the zinc-free form of the p53 DNA binding domain. Biochemistry 2003; 42: 2396–2403.
Zaika A, Marchenko N, Moll UM . Cytoplasmically "sequestered" wild type p53 protein is resistant to Mdm2-mediated degradation. J Biol Chem 1999; 274: 27474–27480.
Muller PA, Vousden Kh Fau - Norman JC, Norman JC . p53 and its mutants in tumor cell migration and invasion. J Cell Biol 2011; 192 (1540-8140 (Electronic)) 209–218.
Lane DP . p53, guardian of the genome. Nature 1992; 358: 15–16.
Soragni A, Janzen DM, Johnson LM, Lindgren AG, Thai-Quynh Nguyen A, Tiourin E et al. A designed inhibitor of p53 aggregation rescues p53 tumor suppression in ovarian carcinomas. Cancer Cell 2016; 29: 90–103.
Haltia M, Prelli F, Ghiso J, Kiuru S, Somer H, Palo J et al. Amyloid protein in familial amyloidosis (Finnish type) is homologous to gelsolin, an actin-binding protein. Biochem Biophys Res Commun 1990; 167: 927–932.
Lakowicz JR . Principles of Fluorescence Spectroscopy. Springer: New York, 2006.
Behl C, Davis JB, Lesley R, Schubert D . Hydrogen peroxide mediates amyloid β protein toxicity. Cell 1994; 77: 817–827.
Jacob RS, George E, Singh PK, Salot S, Anoop A, Jha NN et al. Cell adhesion on amyloid fibrils lacking integrin recognition motif. J Biol Chem 2016; 291: 5278–5298.
Singh PK, Kotia V, Ghosh D, Mohite GM, Kumar A, Maji SK . Curcumin modulates α-synuclein aggregation and toxicity. ACS Chem Neurosci 2013; 4: 393–407.
Shen D, Coleman J, Chan E, Nicholson TP, Dai L, Sheppard PW et al. Novel cell- and tissue-based assays for detecting misfolded and aggregated protein accumulation within aggresomes and inclusion bodies. Cell Biochem Biophys 2011; 60: 173–185.
Kaeser MD, Iggo RD . Chromatin immunoprecipitation analysis fails to support the latency model for regulation of p53 DNA binding activity in vivo. Proc Natl Acad Sci USA 2002; 99: 95–100.
Liang C-C, Park AY, Guan J-L . In-vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc 2007; 2: 329–333.
Friedl P, Wolf K . Tumour-cell invasion and migration: diversity and escape mechanisms. Nat Rev Cancer 2003; 3: 362–374.
Franken NAP, Rodermond HM, Stap J, Haveman J, van Bree C . Clonogenic assay of cells in vitro. Nat Protoc 2006; 1: 2315–2319.
Acknowledgements
We thank the ICMR (Indian Council of Medical Research) National Tumor Tissue Repository at the Tata Memorial Hospital, Mumbai, India for providing the human cancer tissue. Authors also acknowledge IIT Bombay central facilities for FACS, TEM, Confocal microcopy, FTIR and protein crystallography for performing experiments. SKM acknowledges IIT Bombay healthcare seed funding, Lady Tata memorial trust Young researcher fellowship and Department of Science and Technology (Grant number EMR/2014/001233) for funding. We acknowledge Prof. Charles Glabe, UC Urvine, USA for kind gift of OC, A11 antibody, Prof. Jonaki Sen, IIT Kanpur for GFP and mCherry plasmids and Addgene for p53 plasmids.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by JC Marine
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Ghosh, S., Salot, S., Sengupta, S. et al. p53 amyloid formation leading to its loss of function: implications in cancer pathogenesis. Cell Death Differ 24, 1784–1798 (2017). https://doi.org/10.1038/cdd.2017.105
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2017.105
This article is cited by
-
Physiology and pharmacological targeting of phase separation
Journal of Biomedical Science (2024)
-
Amyloid aggregates induced by the p53-R280T mutation lead to loss of p53 function in nasopharyngeal carcinoma
Cell Death & Disease (2024)
-
Amyloid-like p53 as prognostic biomarker in serous ovarian cancer—a study of the OVCAD consortium
Oncogene (2023)
-
p53 amyloid aggregation in cancer: function, mechanism, and therapy
Experimental Hematology & Oncology (2022)
-
Transition of amyloid/mutant p53 from tumor suppressor to an oncogene and therapeutic approaches to ameliorate metastasis and cancer stemness
Cancer Cell International (2022)