Abstract
Recombinant TRAIL and agonistic antibodies to death receptors (DRs) have been in clinical trial but displayed limited anti-cancer efficacy. Lack of functional DR expression in tumors is a major limiting factor. We report here that chromatin regulator KDM4A/JMJD2A, not KDM4B, has a pivotal role in silencing tumor cell expression of both TRAIL and its receptor DR5. In TRAIL-sensitive and -resistant cancer cells of lung, breast and prostate, KDM4A small-molecule inhibitor compound-4 (C-4) or gene silencing strongly induces TRAIL and DR5 expression, and causes TRAIL-dependent apoptotic cell death. KDM4A inhibition also strongly sensitizes cells to TRAIL. C-4 alone potently inhibits tumor growth with marked induction of TRAIL and DR5 expression in the treated tumors and effectively sensitizes them to the newly developed TRAIL-inducer ONC201. Mechanistically, C-4 does not appear to act through the Akt-ERK-FOXO3a pathway. Instead, it switches histone modifying enzyme complexes at promoters of TRAIL and DR5 transcriptional activator CHOP gene by dissociating KDM4A and nuclear receptor corepressor (NCoR)-HDAC complex and inducing the recruitment of histone acetylase CBP. Thus, our results reveal KDM4A as a key epigenetic silencer of TRAIL and DR5 in tumors and establish inhibitors of KDM4A as a novel strategy for effectively sensitizing tumors to TRAIL pathway-based therapeutics.
Similar content being viewed by others
Main
TNF-related apoptosis-inducing ligand (TRAIL) induces apoptosis selectively in many types of human tumor cells by binding to its cognate death receptors DR4/TRAIL-R1/TNFRSF10A and DR5/TRAIL-R2/TNFRSF10B located at the cell surface.1, 2, 3, 4, 5 The binding triggers configurational changes in the receptor trimer and leads to formation of the death-inducing signaling complex (DISC). Depending on cellular context, extrinsic and/or intrinsic pathway of apoptosis is induced with an initial activation of caspase-8 and -10 at DISC and a subsequent activation of the effector caspases such as caspase -3 or -7. The high-tumor cell selectivity and potency in induction of cell death by TRAIL pathway prompted development of therapeutics in the forms of recombinant soluble TRAIL and agonistic TRAIL-R antibodies.6, 7 However, recent clinical trials demonstrated very limited therapeutic benefits.8, 9 Major resistance mechanisms include low expression and/or function of the TRAIL receptors, overexpression of TRAIL decoy receptors (DcRs) and aberrant expression/activation of the anti-apoptotic proteins, such as c-Flip, Bcl-2 family members and IAPs.10, 11 Many synthetic or natural agents and cellular pathways have been identified in sensitizing cancer cells to TRAIL.12, 13 Given the limited stability and/or bio-distribution of the current TRAIL-based therapies, alternative strategies are being sought to induce tumor cell-endogenous TRAIL expression to block tumor growth. For example, a cell-based screening for TRAIL gene promoter activator identified small-molecule ONC210/TIC10. The subsequent encouraging results of ONC201 from xenograft tumor studies led to its recent entry into clinical efficacy evaluation.14, 15 ONC201 apparently induces TRAIL in cancer cells through inhibition of Akt and ERK kinase signaling and promoting nuclear translocation of Foxo3a. Interestingly, it can also upregulate DR5.14, 15
TRAIL and its receptor gene expression in tumor cells can be induced by activation of specific transcription factors and modulation of kinase signaling pathways.16, 17 In addition to Foxo3a, p53 was shown to bind to TRAIL promoter and mediate chemotherapeutics or radiation induction of TRAIL. Although NF-kappaB is well documented to mediate TRAIL resistance by up-regulation of a number of anti-apoptotic genes,18, 19 in certain context, NF-kappaB can also stimulate expression of TRAIL and other proapoptotic ligands.20, 21 Interferons can stimulate TRAIL expression in cancer cells through STAT1 or IRF-1. Interestingly, these factors were also shown to regulate the expression of DR4 and/or DR5.22 However, recent studies suggest that CHOP/DDIT3, a member of the C/EBP transcription factor family, is a major activator of the death receptor expression in cancer cell response to different forms of stress through activation of ERK1/2 and RSK2 signaling.22, 23 Moreover, TRAIL may be subject to epigenetic regulation. Treating AML cells with DNA methylation inhibitors or HDAC inhibitors strongly induces TRAIL and its dependent apoptosis.24, 25, 26 However, little is known about the major histone modifying or demodifying epigenetic regulators involved in regulation of TRAIL and its receptor gene expression.
Histone lysine demethylase 4A or KDM4A (a.k.a. JMJD2A) is a member of the Jumonji C (JmjC) domain-containing KDM4 subfamily of histone demethylases that consists of four paralogues, namely KDM4A, -4B, -4C and -4D. They are alpha-ketoglutarate and Fe(II)-dependent demethylases.27, 28 KDM4A-4D can demethylate the repressive histone marks H3K9me3 and-me2, whereas KDM4A-4C can demethylate H3K36me3 and -me2 as well. KDM4s are overexpressed in many types of human malignancies, including prostate, breast, lung, colorectal and lymphoma.28 KDM4s can stimulate or repress gene expression, control of chromatin accessibility during DNA replication and repair, and reprogram stem cell genome.29, 30, 31, 32 They can associate with members of the nuclear receptor superfamily such as androgen receptor (AR) and estrogen receptor (ER) to mediate their target gene activation in prostate and breast cancer cells.33, 34, 35, 36, 37 KDM4A can also repress expression of genes such as ASCL2 and p21 through association with co-repressor NCoR and HDACs.38, 39, 40 However, it is unclear whether its demethylase activity is involved in the repression.
KDMs are increasingly recognized as therapeutic targets for cancer and other diseases.41, 42, 43 However, few small molecules with high selectivity to specific KDMs and strong biological activities have been identified. We recently performed a structure-based, virtual screening for KDM inhibitors and identified compound-4 (NSC636819, abbreviated as C-4) that possesses a KDM4A/4B-selective inhibition activity and strongly inhibits prostate cancer cell proliferation.37 Here, we show that KDM4A is a key epigenetic silencer of TRAIL and DR5 expression and that its inhibitor C-4 potently induces both TRAIL and DR5 expression through switching a transcriptional repressor complex to an active one containing histone methylase MLL. Importantly, KDM4A inhibitor C-4 strongly inhibits tumor growth and effectively sensitizes tumors to TRAIL-inducing agent ONC201. Thus, our study reveals a novel function of KDM4A in cancer cell survival through suppressing TRAIL-DR5 pathway, and offers an epigenetic strategy in TRAIL-based cancer therapy.
Results
KDM4 inhibitor C-4 strongly induces the expression of both TRAIL and its receptor DR5 selectively in cancer cells
Our previous study revealed that inhibition of KDM4A/B with the small-molecule compound-4 (C-4) (Figure 1a) strongly up-regulated expression of a number of genes including DR5 in a prostate cancer cell line.37 This prompted us to examine whether the inhibitor induced cell death through activating TRAIL-initiated apoptosis pathway. First, we analyzed the effects of C-4 treatment on expression of TRAIL and its receptors. Interestingly, C-4 treatment of a castration-resistant prostate cancer cell C-4-2B resulted in dose-dependent increase of both TRAIL and its receptor DR5, whereas the treatment did not have any significant effect on the expression of DR4 or decoy receptors DcR1/TNFRSF10C and DcR2/TNFRSF10D (Figure 1b). C-4-2B cells are highly resistant to TRAIL induction of apoptosis (Supplementary Figure S1) and express functional AR and wild type p53. To determine whether C-4 induction of TRAIL and DR5 is dependent on AR and p53, we extended our analysis to other cancer cell lines, including breast cancer cell MDA-MB231 (AR−; p53 mutated) and lung cancer cells A549 (p53, wild type), H1299 (AR−; p53-null) and Calu-6 (p53, mutated). The analysis showed that KDM4A/B inhibitor C-4 induced TRAIL and DR5 expression (mRNA and protein) in both TRAIL-sensitive (Calu-6 and MDA-MB231) and TRAIL-resistant cancer cells (A549 and H1299) and that the induction is independent of p53 or AR status (Figures 1b and c; Supplementary Figure S2). Importantly, similar treatment of a number of normal or non-transformed human cells (HBE1, IMR90, PNT2, and MCF-10A) did not cause any significant change in expression of TRAIL and its receptors (Figures 1b and c; Supplementary Figure S3). The cell surface expression of DR4 or DR5 is a key determinant of tumor sensitivity to the TRAIL-DR targeted therapies.44 To examine whether KDM4A inhibitor C-4 induced DR5 expression at cell surface, we immunoprecipitated the receptors from the cell surface after cells were treated with C-4. We used an immunoprecipitation method45 instead of flow cytometry because of the auto-fluorescence activity of C-4. As shown in Figure 1d, C-4 treatment specifically increased the amount of DR5 protein on the surface of cells examined. We also detected significant increase of secreted form of TRAIL in the medium of cells treated by C-4 (Supplementary Figure S2). Taken together, these data indicated that pharmacological inhibition of KDM4A/KDM4B strongly induces the expression of TRAIL and DR5 in TRAIL-sensitive and -resistant cancer cells, but not in normal cells, and that the induction is independent of p53 status.
KDM4 inhibitor C-4 strongly induces the expression of both DR5 and TRAIL selectively in cancer cells. (a) Chemical structure of KDM4 inhibitor C-4. (b) qRT-PCR analysis of TRAIL and its receptor expression in C-4-2B, A549 and IMR90 cells treated with vehicle or indicated concentration of C-4 for 2 days. Significance was calculated using Student’s t-test. *P<0.05, **P<0.01. (c) Indicated cancer cells or non-malignant cells (PNT2) were treated with vehicle or C-4 for 3 days before collected for immunoblotting with indicated antibodies. (d) Cancer cells were treated with vehicle or C-4 (20 μM) for 3 days before collected for IP of isolated plasma membrane fractions. DR4 and DR5 were detected by immunoblotting analysis after IP
Silencing KDM4A, not KDM4B, de-represses TRAIL and DR5 expression
C-4 was identified as a selective inhibitor against the H3K9me3 demethylation activity of KDM4A and KDM4B.37 We thus examined whether KDM4A and/or KDM4B are involved in control of TRAIL and DR5 expression. Silencing KDM4A with specific siRNAs revealed that KDM4A knockdown strongly induced the mRNA and protein expression of TRAIL and DR5 in the cancer cells that displayed similar inductions by C-4 treatment (Figures 2a and c). In contrast, knockdown of KDM4B with specific siRNAs did not invoke any significant change in TRAIL or DR5 expression (Figures 2b and d). Like C-4 treatment, knockdown of KDM4A or KDM4B did not affect DR4 expression. To examine whether KDM4A gene silencing induced DR5 expression at cell surface, we performed flow cytometry analysis. As shown in Figure 2e, knockdown of KDM4A resulted in significantly enhanced cell surface expression of DR5 (Figure 2e). Together with data in Figure 1, the analyses revealed that KDM4A has a crucial role in repression of both TRAIL and its receptor DR5 expression, and that KDM4A inhibition via gene knockdown or pharmacological means can effectively upregulate their expression in cancer cells.
Silencing KDM4A, not KDM4B, de-represses TRAIL and DR5 expression cells were transfected with control or indicated KDM4A (a and c) or KDM4B (b and d) siRNA and cultured for 2 days. Total RNAs were prepared and subjected to qRT-PCR analysis for TRAIL and DR5 mRNA expression (a and b). For protein analysis, transfected cells were cultured for 3 days before collected for immunoblotting (c and d) or for flow cytometry analysis of cell surface expression of DR5. (e) DR5 cell surface expression quantification was carried out using geometric means of three independent experiments and plotted as fold change of surface expression. Significance was calculated using Student’s t-test. **P<0.01
KDM4A inhibition induces caspase-8 activation and apoptosis selectively in cancer cells
Given the induction of both TRAIL and DR5 by KDM4A inhibition, we thus examined whether inhibition of KDM4A can result in activation of the extrinsic or intrinsic TRAIL-DR apoptotic pathways, which involves cascades of events of caspase cleavage. Results in Figures 3a and b showed that both pharmacological and knockdown inhibition of KDM4A significantly activated a caspase cascade from the upstream initiator caspase-8 to the downstream effector caspase-7 and to PARP-1 cleavage. However, the inhibition did not affect the expression of the short or long form of c-FLIP, the expression of other anti-apoptotic proteins such as Mcl-1, CIAP1/2 and XIAP, the level of truncated Bid (tBid) (Figure 3b). Analysis of cell proliferation by cell counting and measurement of apoptosis by hoechst33342 staining or a cell death detection ELISA kit demonstrated that KDM4A inhibition strongly induced cell apoptosis and inhibited cell proliferation in various types of cancer cells (Figures 3c and d; Supplementary Figure S4) but had minimal effect on the survival of normal cells (Figure 3e). Collectively, these data suggest that inhibition of KDM4A efficiently induces cancer cell apoptosis primarily through activation of the extrinsic apoptosis pathway by TRAIL-DR5.
KDM4A inhibition induces caspase-8 activation and apoptosis specifically in cancer cells. (a) Cells were transfected with control or KDM4A siRNA and cultured for 3 days before collected for immunoblotting with specific antibodies against indicated proteins. (b) Cells were treated with vehicle or C-4 for 3 days before subjected to immunoblotting for indicated proteins analysis. (c–e) Cells were transfected with control or KDM4A siRNA (c) or treated with vehicle or C-4 (d,e) and cultured for 4 days before collected for apoptosis analysis by hoechst33342 staining. Representative images taken by fluorescent microscope are shown in c. Significance was calculated using Student’s t-test. *P<0.05, **P<0.01
Inhibition of KDM4A sensitizes TRAIL-resistant cells to TRAIL and TRAIL-inducer ONC201-induced apoptosis
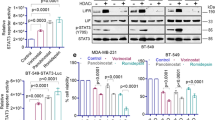
One major mechanism of resistance to TRAIL therapies appears to be the low cellular level and/or function of the TRAIL receptors.46 We then examined whether KDM4A inhibition-elevated DR5 and TRAIL could sensitize TRAIL-resistant cancer cells to exogenous TRAIL-induced apoptosis. Three TRAIL-resistant cancer cell lines (C-4-2B, A549 and H1299) were pretreated with a low concentration (5 μM) of KDM4A inhibitor C-4 for 48 h followed by treatment with recombinant TRAIL for another 24 h. Analyses of cell apoptosis demonstrated that although a low dose of either TRAIL or C-4 alone did not significantly affect the cell survival, their sequential, combinatorial treatment induced high levels of apoptosis in all three TRAIL-resistant cells (Figure 4a). Silencing of KDM4A also markedly increased apoptosis induced by the exogenous TRAIL (Figure 4b; Supplementary Figure S5), suggesting that C-4 sensitizes cells to TRAIL is through inhibition of KDM4A. In addition, we also found that knockdown of DR5 by siRNA effectively blocked cell death induced by KDM4A inhibitor C-4 in combination with TRAIL (Supplementary Figure S6). To further assess the role of TRAIL apoptosis pathway in KDM4A inhibition-induced apoptosis, we silenced TRAIL expression in cells before KDM4A inhibition via either siRNA knockdown or C-4. Results in Figures 4c and d indicated clearly that TRAIL silencing effectively blocked KDM4A inhibition-induced apoptosis. Therefore, the apoptosis induced by KDM4A inhibition was indeed initiated by TRAIL.
Inhibition of KDM4A sensitizes TRAIL-resistant cells to TRAIL and TRAIL-inducer compound ONC201-induced apoptosis. (a) TRAIL-resistant cancer cells (C-4-2B, H1299 and A549) were pretreated with 5 μM C-4 for 2 days and then treated with 10 ng/ml TRAIL for another 24 h, before collected for apoptosis analysis by hoechst33342 staining or for immunoblotting. (b) TRAIL-resistant cancer cells were transfected with control or KDM4A siRNA and cultured for 2 days and then treated with 10 ng/ml TRAIL for another 24 h. Apoptosis was analyzed by hoechst33342 staining. (c), TRAIL-sensitive cancer cells (MB231 and Calu-6) were transfected with KDM4A and/or TRAIL siRNA and cultured for 4 days before collected for apoptosis analysis by hoechst33342 staining or for immunoblotting. (d) Cells were transfected with control or TRAIL siRNA and cultured for 1 day and then treated vehicle or 5 μM C-4 for another 4 days. Apoptosis were detected by hoechst33342 staining. (e) A549 cells were treated with vehicle or ONC201 for 3 days before collected for immunoblotting. (f,g) A549 cells were treated with vehicle, C-4 (5 μM) and ONC201 (2.5 μM) for 4 days before collected for apoptosis analysis (f) and immunoblotting (g) as in a. Significance was calculated using Student’s t-test. **P<0.01
Recently, a small-molecule ONC201 was identified to be a potent inducer of TRAIL.14, 15 As shown recently in colon cancer cell HCT116,14 ONC201 also strongly induced TRAIL expression in a dose-dependent manner in lung cancer A549 cells (Figure 4e). Strikingly, treating cells with low doses of both ONC201 and C-4 resulted in pronounced cell death in the TRAIL-resistant cells whereas either alone elicited minimal effects on cell death (Figure 4f). As reported, ONC201 treatment strongly inhibited the phosphorylation of Akt and ERK as well as their target protein Foxo3a. Interestingly, C-4 did not appear to have a similar effect on Foxo3a phosphorylation. Furthermore, C-4 synergized with ONC201 to induced TRAIL expression (Figure 4g). Together, the data suggest that C-4 and ONC201 stimulate TRAIL expression through distinct mechanisms, which likely underlies the observed synergistic cell death effect.
KDM4A inhibition de-represses TRAIL and DR5 activator CHOP genes through switching histone modification complexes
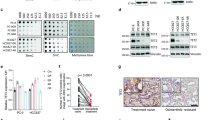
As KDM4A inhibition de-represses TRAIL and DR5 expression at mRNA level, we performed chromatin immunoprecipitation (ChIP) to examine whether KDM4A might bind to TRAIL and DR5 promoter to control their expression. Indeed, ChIP data indicated that KDM4A directly bind to TRAIL promoter. However, no significant KDM5 ChIP signal was detected at the DR5 promoter (Figure 5a; Supplementary Figure S7a). DR5 is transcriptionally regulated by factors including p53, NF-κB and the CEBP family member CHOP/DDIT3/GADD15321 (Supplementary Figure S7b). As shown in Figures 1 and 2, inhibition of KDM4A induced DR5 expression in the p53-null H1299 cell line, indicating a lack of p53 involvement in the DR5 induction. Likewise, silencing KDM4A did not alter the expression and activation of the major NF-κB components such as p105, p50, p52 and pS529-p65 (Supplementary Figure S8), thus leaving CHOP as the candidate of the downstream factor. Indeed, treatment of cells with KDM4A knockdown or its inhibitor C-4 markedly increased the mRNA and protein of CHOP (Figures 5b and c). Anti-KDM4A ChIP demonstrated that KDM4A binds to CHOP gene promoter (Figure 5d). Importantly, KDM4A inhibition also resulted in a significant increase of CHOP recruitment to the CHOP binding site at DR5 promoter (Supplementary Figure S9). Together, the data indicate that KDM4A binds to TRAIL and DR5 activator CHOP gene promoters to repress their expression and that KDM4A inhibition directly de-represses TRAIL and activates DR5 through de-repressing CHOP.
KDM4A inhibition de-represses expression of TRAIL and DR5 activator CHOP through switching histone modification complexes. (a) ChIP-qPCR analysis of relative KDM4A occupancy at the TRAIL promoter and distant region in A549 cells. (b,c) Cells were transfected with control or KDM4A siRN or treated with vehicle or C-4. After 2 or 3 days, cells were collected for qRT-PCR or immunoblotting, respectively. (d) ChIP-qPCR analysis of relative KDM4A occupancy at the CHOP promoter and distant region in A549 cells. (e,f) ChIP was performed with A549 cells treated with vehicle or 20 μM C-4. Indicated protein enrichments on promoter of TRAIL (e) or CHOP (f) were analyzed by qPCR. Significance was calculated using Student’s t-test. *P<0.05, **P<0.01
Although KDM4A catalyzes the demethylation of di- and trimethylated H3K9 and H3K36, the role of such activities in KDM4A-mediated transcriptional regulation is unclear.47, 48 Indeed, KDM4A knockdown has been shown to trigger an accumulation of H3K9me2/3 and H3K36me2/3 levels and a concomitant increase in its target ASCL2 and CHD5 gene expression.47, 48 Consistent with these studies, the inhibitor C-4 treatment increased the level of H3K9me2 and -me3 and H3K36me2 and -me3 at the KDM4A binding site of its target TRAIL and CHOP, indicating that the local KDM4A demethylase activity is indeed inhibited by C-4 (Figures 4e and f). To further understand the mechanism of TRAIL and CHOP de-repression due to KDM4A inhibition, we examined the potential effect of C-4 on the local function of KDM4A-associated repressor complex. As shown in Figures 5e and f, C-4 treatment for 24 h resulted in strong reduction of occupancy of co-repressor NCOR1/NCoR and its associated histone deacetylase HDAC1 at TRAIL and CHOP promoters, which was accompanied by a significantly increased level of H3K9ac and H3K27ac. Consistent with KDM4Abeing a H3K36 and H3K9 demethylase, the level of H3K36me2 and H3K9me3 was also increased as previously reported (Figures 5e and f). Further analysis showed that the increase of H3K9ac and H3K27ac is likely contributed by an increased recruitment of the histone acetylase CBP. Finally, KDM4A inhibition by C-4 led to an increased occupancy at the promoters of CTD phosphor-Ser5 form of RNA polymerase II, indicating an enhanced transcription initiation (Figures 5e and f). Together, the data suggest that the de-repression by C-4 inhibition of KDM4A involves dissociation of KDM4A from its target gene promoters, disruption of NCOR1-HDAC1 repressor complex and assembly of gene-activating histone acetylase complex.
Targeting KDM4A strongly suppresses tumor growth and sensitizes tumors to TRAIL-inducer ONC201
Given the remarkable induction of cell death by KDM4A inhibition, we next assessed the antitumor potential of targeting KDM4A. First, silencing KDM4A expression using two distinct shRNAs potently suppressed colony formation of both TRAIL-sensitive and resistant cancer cells (Supplementary Figure S10a). Consistent with the in vitro results, KDM4A knockdown effectively blocked tumor formation and growth in the xenograft models of MDA-MB231 breast cancer and A549 lung cancer (Figure 6a; Supplementary Figure S10b). Examination of the residual tumors revealed that tumors with KDM4A silenced displayed increased TRAIL and DR5 expression and caspase activation (Supplementary Figure S10c).
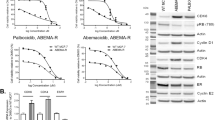
Targeting KDM4A strongly suppresses tumor growth and sensitizes tumors to TRAIL-inducer ONC201. (a) MDA-MB231 cells (2.5 × 106) infected with control or KDM4A shRNA lentivirus were injected into the mouse mammary fat pad. Tumor growth was monitored every week. Each data point is the mean value ± S.E.M. from 5 to 6 mice. *P<0.05, **P<0.01. (b,c) Nude mice bearing the A549 xenografts (n=6 mice per group) received vehicle, C-4 (intraperitoneally (i.p.); 20 or 40 mg/kg, 5 times a week), ONC201 (i.p.; 25 mg/kg, 1 time a week), or a combination of C-4 and ONC201, as indicated. Mean tumor volume ±S.E.M. and mean tumor weight ±S.E.M. were shown. Significance was calculated using Student’s t-test. *P<0.05, **P<0.01. (d), Immunoblotting of xenograft tumors after treatment with vehicle or C-4 for indicated protein expression. (e) Representative images of anti-TRAIL and anti-DR5 IHC analysis of xenograft tumor sections from mice treated with vehicle or SR2211
We then evaluated the antitumor activity of KDM4A inhibitor C-4 and TRAIL-inducer ONC201, either alone or in combination, in a lung cancer xenograft model. C-4, when administered five times a week at 20 and 40 mg/kg, dose-dependently inhibited the tumor growth. As reported previously, ONC201, at 25 mg/kg and once a week, also strongly inhibited the tumor growth (Figures 6b and c). Significantly, when a low dose of C-4 and a low dose of ONC201 were administered in combination to the tumor-bearing animals, a sustained blockade of tumor growth was observed, indicating that KDM4A inhibitor C-4 can sensitize tumors to TRAIL (Figures 6b and c). Monitoring of mice weight and general behaviors did not indicate any significant toxicity associated with the C-4 treatments (Supplementary Figure S11a). Importantly, C-4 did not induce the expression of TRAIL or DR5 in mouse tissues such as liver and lung (Supplementary Figure S11d). Notably, tumor expression of TRAIL, DR5 and the level of cleaved caspase-7 and H3K9me3 were strongly induced whereas the expression of KDM4A was markedly decreased in xenograft tumors treated with C-4 compared (Figures 6d and e). Taken together, these results indicate that KDM4-selective inhibitor C-4, either alone or in combination with TRAIL-inducer therapeutics ONC201, can be efficacious in induction of tumor TRAIL and DR5 expression and in stopping tumor growth.
Discussion
The expression of TRAIL and its receptors DR4 and DR5 in tumor cells is usually kept at low level, which constitutes one of the major TRAIL-resistance mechanisms. Here, we present several lines of evidence that histone demethylase KDM4A is a key epigenetic silencer of both TRAIL and DR5. C-4, a newly identified small-molecule inhibitor of KDM4A/B induces both TRAIL and DR5 in cancer cells and tumors in a dose-dependent manner. Intriguingly, the KDM4A inhibition-mediated induction of TRAIL and DR5 is not observed in the normal cells, suggesting a cancer cell-selective function of KDM4A in silencing TRAIL and DR5. One possible reason for the apparent cancer-specific function of KDM4A is its overexpression and hence a ‘gain-of-function’ in tumor cells. However, KDM4B, which is also overexpressed in tumors, does not seem to have similar silencing functions as indicated by our studies. Thus, it is likely that other factors such as transcription factors that bind to TRAIL and the DR5 activator CHOP gene promoters are also involved. Their cancer cell-specific interactions and functions with KDM4A (and presumably not with KDM4B) could mediate KDM4A recruitment to the promoters of CHOP and TRAIL. The recruited KDM4A can in turn recruit transcriptional repressor proteins such as the co-repressor NCOR1/NCoR and HDACs. Indeed, our data show that C-4 effectively disrupts the local chromatin association of NCoR-HDAC1 co-repressor complex, therefore strongly suggesting that KDM4A silences TRAIL and DR5 through the histone deacetylase co-repressor complex. Interestingly, however, inhibitor C-4 also strongly dissociates KDM4A protein from its target gene promoter, thus suggesting that its demethylase activity is required for its chromatin association and for its transcriptional silencing function. Further elucidation of the cancer cell-selective mechanism of KDM4A-mediated silencing of TRAIL and CHOP-DR5 can provide insights valuable for development of KDM4A inhibitor-assisted TRAIL therapy.
One significant observation of this study is that targeting KDM4A by the small-molecule inhibitor C-4 or shRNA sensitizes cells and tumors to TRAIL or TRAIL-inducer ONC201. In TRAIL-resistant cells, KDM4A inhibition can synergize with TRAIL or ONC201 in induction of apoptosis. Our further analyses suggest that the sensitization/synergy appears to be mediated by both the induced DR5 and the supply of exogenous TRAIL. First, in the TRAIL-resistant cells, DR5 knockdown significantly mitigates C-4 sensitization of the cells to the exogenous TRAIL. In agreement, TRAIL-sensitive cells express significant levels of endogenous DR5. Second, in the resistant cells, when the endogenously produced TRAIL and KDM4A are silenced, which results in just the up-regulated DR5, the cells remain very sensitive to the exogenous TRAIL. The fact that KDM4A inhibition alone, which induces the endogenous production of TRAIL and DR5, is insufficient to induce robust apoptosis in the TRAIL-resistant cells, also suggests that a high level of exogenous TRAIL is required. Nonetheless, we cannot rule out the possibility that, in the resistant cell, endogenously produced TRAIL can act differently from the exogenous TRAIL in induction of apoptosis.
Aside from the TRAIL pathway silenced by KDM4A but not KDM4B, KDM4A and KDM4B likely regulate both overlapping and distinct gene targets49 (e.g., cell cycle genes by both KDM4A and KDM4B;37, 50 c-Myc and ribosomal RNA genes by KDM4A and KDM4B;51, 52 B-Myb targets such as PLK1 by KDM4B43). It is well known that cancer cells are addicted to the high levels of those genes for their uncontrolled growth and survival. Therefore, inhibitors such as C-4, which can inhibit both KDM4A and KDM4B, could exert their cellular effects through above targets or other undefined ones. Therefore, further development of KDM4 inhibitors are warranted that can more selectively and potently induce TRAIL-DR pathway, in conjunction with suppressing cancer-addicted targets.
Materials and Methods
Cell culture and reagents
All cell lines come from ATCC. Human lung cancer cell line A549, H1299, Calu-6, HCC827 cells and prostate cancer cell line C4-2B cells were cultured in RPMI and breast cancer cell line MDA-MB231 cells in DMEM medium (CORNING-Cellgro, Manassas, VA, USA), supplemented with 1% penicillin/streptomycin and 5% (lung cancer cell lines) or 10% fetal bovine serum (Gemini Bio Products, West Sacramento, CA, USA) at 37 °C under 5% CO2 in a humidified incubator. Antibodies were used with the sources and dilution ratios indicated in parentheses as in Supplementary Table 2. NSC636819 (Compound-4 or C-4) used for cell culture experiments was obtained from the NCI/DTP Open Chemicals Repository. C-4 for animal experiments was purchased from ChemPartner. Other chemicals were purchased from Sigma-Aldrich (St. Louis, MO, USA) unless specified otherwise.
Apoptosis and cell growth assays
For apoptosis, cells were treated as indicated and processed for Hoechst33342 staining as previously described.53 Random fields (10 fields/condition) of typical apoptotic nuclear morphology (nuclear shrinkage, condensation, and fragmentation) with positive Hoechst staining were counted and averaged. For cell growth, cells were seeded in 6-well plates at 2 × 105 per well and treated as indicated. Total survival cell numbers were counted using a Coulter cell counter. The assays were performed in triplicates and the entire experiments were repeated more than three times.
ChIP assay
ChIP was performed essentially as described previously54 with the following modifications. The crude chromatin solutions were first cleared with protein A beads (Invitrogen, Carlsbad, CA, USA) that had been pre-coated with normal IgG for 2 h at 4 °C, and then incubated at 4 °C overnight with specific antibodies (Supplementary Table 3) before precipitation with protein A beads that had been pre-blocked with BSA and sonicated salmon sperm DNA. The precipitated DNA was analyzed by real-time PCR with SYBR green on an iCycler instrument. Enrichment of genomic DNA was presented as the percent recovery relative to the input. The primers are listed in the Supplementary Table 1.
qRT-PCR, subcellular fractionation, immunoprecipiation and immunoblot analysis
Total RNA was isolated and the cDNA was prepared, amplified and detected in the presence of SYBR as previously.54 The fluorescent values were collected and a melting curve analysis was performed. Fold difference was calculated as described previously.
The immunoprecipitation of plasma membrane for DR4 and DR5 was performed as previously described.45 Cells were treated with vehicle or C-4 for 3 days and collected for the antibody incubation, Cells were then lysed in hypotonic lysis buffer. The membrane fraction was isolated by centrifugation and suspended in 1 ml of hypotonic lysis buffer containing 1% Triton X-100. Cell membrane lysate was precipitated with protein G-Sepharose beads as previously described.45
Proteins in the lysates were analyzed by immunoblotting with specific antibodies (Supplementary Table 2). The PCR primers are listed in the Supplementary Table 1.
KDM4A shRNA lentivirus production and siRNA transfection
Lentiviral plasmids encoding shRNA targeting KDM4A (TRCN0000013493 and TRCN0000013497) were purchased from Sigma-Aldrich. Nontargeting control shRNA were used as described. Lentiviral particles were produced in 293T cells after co-transfection of the above lentivirus vectors, psPAX2 and pMD2.G in 10-cm dishes, as described.55 siRNAs for gene knockdown were purchased from Dharmacon. The siRNA target sequences for KDM4A are as follows: KDM4A#1,GCUGCAGUAUUGAGAUGCUAA; KDM4A#2, GCCUUGGAUCUUUCUGUGAA; Control, CAGUCGCGUUUGCGACUGG. Transfections were performed with OptiMEM (Invitrogen) and Dharmafectin 1 (Dharmacon, Lafayette, CO, USA), following the manufacturer’s instructions.
Flow cytometry
The expression of TRAIL receptors on cell surface membrane was assessed by flow cytometry using PE-conjugated antibodies (R&D Systems, Minneapolis, MN, USA) as previously described.56 Briefly, cells were collected with trypsin, washed and incubated with anti-DR5-PE (eBioscience, San Diego, CA, USA; flow cytometry, 10 μg/ml) or anti-DR4-PE (eBioscience; flow cytometry, 10 μg/ml) for 45 min at 4 °C in the dark. Cells were then washed three times with cold PBS and resuspended in 0.3 ml PBS for the analysis. Cells were analyzed using a FACS Canto (Becton Dickinson, Franklin Lakes, NJ, USA) and the resulting data were analyzed with FlowJo Version 8.8.7. (Ashland, OR, USA) Geometric means were used for the analyses.
ELISA
For soluble TRAIL in cell culture medium, cells were treated with vehicle or C-4 for 3 days. Medium were collected from cell culture plates. Soluble TRAIL was measured using Human TRAIL ELISA Kit (ab46074, Abcam, Cambridge, MA, USA) following the manufacturer’s instructions.
Colony formation
For colony formation assay, 800 cells per well were seeded in a 6-well plate and cultured for 10–14 days in a 37 °C incubator with medium changed every 3 days. The cells were fixed with 10% formalin and stained with 0.2% crystal violet (in 10% formalin) for 15 min. The numbers of cell colonies were counted after washes. The assays were performed in triplicates and the entire experiments were repeated more than three times.
Xenograft tumor models and treatments
For establishing the lung cancer xenograft tumors, 5–6-week-old, BALB/c nu/nu athymic nude mice (Harlan, Indianapolis, IN, USA) were injected with 2.5 × 106 A549 cells in 100 μl PBS/Matrigel (1:1) into the dorsal flank on both sides of the mice. When the tumors were ~50 mm3 in size, mice were randomized and treated i.p. with 100 μl of either vehicle (DMSO and sesame oil, 1:50, Sigma, St. Louis, MO, USA), 20 mg/kg compound-4 (C-4) (five times a week), 40 mg/kg C-4 (five times a week), 25 mg/kg ONC201 (one time a week) or a combination of 20 mg/kg C-4 (five times a week) and 25 mg/kg ONC201 (one time a week). To assess the effect of shRNA mediated silencing of KDM4A on A549 xenograft tumors, 2 × 106 cells (infected with lentivirus shControl or shKDM4A) were suspended in total of 100 μl PBS/Matrigel (1:1) and implanted subcutaneously into the dorsal flank on both sides of the mice. For breast cancer MDA-MB231 xenograft tumors, 5–6-weeks-old BALB/c nu/nu athymic mice were injected unilaterally with 2.5 × 106 cells (infected with lentivirus shControl or shKDM4A) in 100 μl PBS/Matrigel (1:1) into the abdominal fat pad by subcutaneous injection at the base of the nipple. Tumor growth was monitored by calipers with volume calculated using the equation π/6 (length × width2). Body weight during the course of the study was also monitored. At the end of the studies mice were killed and tumors were dissected and weighed. Tumors were collected and immediately stored in liquid nitrogen or fixed in formalin solution. The procedures were approved by the Institutional Animal Care and Use Committee of University of California Davis.
Statistical analysis
The data are presented as mean values±S.D. from at least three independent experiments. Data analysis was performed using two-tailed Student’s t-tests and P-values are shown. *P<0.05; **P<0.01; N.S., not significant.
Abbreviations
- TRAIL/TNFSF10TNF-related apoptosis-inducing ligand/tumor necrosis factor superfamily member 10; TNFRSF10A/DR4:
-
TNF receptor superfamily member 10a/death receptor 4
- TNFRSF10B/DR5:
-
TNF receptor superfamily member 10b/death receptor 5
- KDM4A/JMJD2A:
-
lysine demethylase 4 A
- KDM4B:
-
lysine demethylase 4B
- C-4:
-
compound-4/ NSC636819
- FOXO3a:
-
forkhead box O3A
- DDIT3/CHOP:
-
DNA damage inducible transcript 3
- HDAC:
-
histone deacetylase
- NCoR:
-
nuclear receptor co-repressor
- AR:
-
androgen receptor
- ER:
-
estrogen receptor
- IHC:
-
Immunohistochemistry
References
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Pan G, Ni J, Wei Y-F, Yu G-l, Gentz R, Dixit VM . An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 1997; 277: 815–818.
Wu GS, Burns TF, McDonald ER, Jiang W, Meng R, Krantz ID et al. KILLER/DR5 is a DNA damage-inducible p53-regulated death receptor gene. Nat Genet 1997; 17: 141–143.
de Miguel D, Lemke J, Anel A, Walczak H, Martinez-Lostao L . Onto better TRAILs for cancer treatment. Cell Death Differ 2016; 23: 733–747.
Graves Jonathan D, Kordich Jennifer J, Huang T-H, Piasecki J, Bush Tammy L, Sullivan T et al. Apo2L/TRAIL and the death receptor 5 agonist antibody AMG 655 cooperate to promote receptor clustering and antitumor activity. Cancer cell 2014; 26: 177–189.
Dida F, Li Y, Iwao A, Deguchi T, Azuma E, Komada Y . Resistance to TRAIL-induced apoptosis caused by constitutional phosphorylation of Akt and PTEN in acute lymphoblastic leukemia cells. Exp Hematol 2008; 36: 1343–1353.
Herbst RS, Eckhardt SG, Kurzrock R, Ebbinghaus S, O'Dwyer PJ, Gordon MS et al. Phase I dose-escalation study of recombinant human Apo2L/TRAIL, a dual proapoptotic receptor agonist, in patients with advanced cancer. J Clin Oncol 2010; 28: 2839–2846.
Kindler HL, Richards DA, Garbo LE, Garon EB, Stephenson JJ, Rocha-Lima CM et al. A randomized, placebo-controlled phase 2 study of ganitumab (AMG 479) or conatumumab (AMG 655) in combination with gemcitabine in patients with metastatic pancreatic cancer. Ann Oncol 2012; 23: 2834–2842.
Mizrahi K, Stein J, Pearl-Yafe M, Kaplan O, Yaniv I, Askenasy N . Regulatory functions of TRAIL in hematopoietic progenitors: human umbilical cord blood and murine bone marrow transplantation. Leukemia 2010; 24: 1325–1334.
Kang YC, Kim KM, Lee KS, Namkoong S, Lee SJ, Han JA et al. Serum bioactive lysophospholipids prevent TRAIL-induced apoptosis via PI3K//Akt-dependent cFLIP expression and Bad phosphorylation. Cell Death Differ 2004; 11: 1287–1298.
He K, Zheng X, Li M, Zhang L, Yu J . mTOR inhibitors induce apoptosis in colon cancer cells via CHOP-dependent DR5 induction on 4E-BP1 dephosphorylation. Oncogene 2015; 35: 148–157.
Henrich CJ, Brooks AD, Erickson KL, Thomas CL, Bokesch HR, Tewary P et al. Withanolide E sensitizes renal carcinoma cells to TRAIL-induced apoptosis by increasing cFLIP degradation. Cell Death Dis 2015; 6: e1666.
Allen JE, Krigsfeld G, Mayes PA, Patel L, Dicker DT, Patel AS et al. Dual inactivation of Akt and ERK by TIC10 signals Foxo3a nuclear translocation, TRAIL gene induction, and potent antitumor effects. Sci Transl Med 2013; 5: 171ra117.
Prabhu VV, Allen JE, Dicker DT, El-Deiry WS . Small-molecule ONC201/TIC10 targets chemotherapy-resistant colorectal cancer stem–like cells in an Akt/Foxo3a/TRAIL–dependent manner. Cancer Res 2015; 75: 1423–1432.
Allen JE, El-Deiry WS . Regulation of the human TRAIL gene. Cancer Biol Ther 2012; 13: 1143–1151.
van Roosmalen IAM, Quax WJ, Kruyt FAE . Two death-inducing human TRAIL receptors to target in cancer: Similar or distinct regulation and function? Biochem Pharmacol 2014; 91: 447–456.
Ricci MS, Kim S-H, Ogi K, Plastaras JP, Ling J, Wang W et al. Reduction of TRAIL-Induced Mcl-1 and cIAP2 by c-Myc or sorafenib sensitizes resistant human cancer cells to TRAIL-induced death. Cancer Cell 2007; 12: 66–80.
Anna Trauzold HW, Arlt Alexander, Schütze Stefan, Schäfer Heiner, Oestern Stefanie, Röder Christian et al. CD95 and TRAIL receptor-mediated activation of protein kinase C and NF-B contributes to apoptosis resistance in ductal pancreatic adenocarcinoma cells. Oncogene 2001; 20: 4270–4280.
Baetu TM, Kwon H, Sharma S, Grandvaux N, Hiscott J . Disruption of NF-κB signaling reveals a novel role for NF-κB in the regulation of TNF-related apoptosis-inducing ligand expression. J Immunol 2001; 167: 3164–3173.
Schlegel CR, Fonseca AV, Stocker S, Georgiou ML, Misterek MB, Munro CE et al. DAPK2 is a novel modulator of TRAIL-induced apoptosis. Cell Death Differ 2014; 21: 1780–1791.
Prasad S, Kim JH, Gupta SC, Aggarwal BB . Targeting death receptors for TRAIL by agents designed by mother nature. Trends Pharmacol Sci 2014; 35: 520–536.
Oh Y-T, Yue P, Zhou W, Balko JM, Black EP, Owonikoko TK et al. Oncogenic Ras and B-Raf proteins positively regulate death receptor 5 expression through co-activation of ERK and JNK signaling. J Biol Chem 2012; 287: 257–267.
Nebbioso A, Clarke N, Voltz E, Germain E, Ambrosino C, Bontempo P et al. Tumor-selective action of HDAC inhibitors involves TRAIL induction in acute myeloid leukemia cells. Nat Med 2005; 11: 77–84.
Lund P, Kotova I, Kedinger V, Khanwalkar H, Voltz E, Hahn WC et al. Transformation-dependent silencing of tumor-selective apoptosis-inducing TRAIL by DNA hypermethylation is antagonized by decitabine. Mol Cancer Ther 2011; 10: 1611–1623.
Soncini M, Santoro F, Gutierrez A, Frigè G, Romanenghi M, Botrugno OA et al. The DNA demethylating agent decitabine activates the TRAIL pathway and induces apoptosis in acute myeloid leukemia. Biochim Biophys Acta 2013; 1832: 114–120.
Pedersen MT, Helin K . Histone demethylases in development and disease. Trends Cell Biol 2010; 20: 662–671.
Berry WL, Janknecht R . KDM4/JMJD2 histone demethylases: epigenetic regulators in cancer cells. Cancer Res 2013; 73: 2936–2942.
Black JC, Allen A, Van Rechem C, Forbes E, Longworth M, Tschöp K et al. Conserved antagonism between JMJD2A/KDM4A and HP1γ during cell cycle progression. Mol Cell 2010; 40: 736–748.
Black Joshua C, Manning Amity L, Van Rechem C, Kim J, Ladd B, Cho J et al. KDM4A lysine demethylase induces site-specific copy gain and rereplication of regions amplified in tumors. Cell 2013; 154: 541–555.
Young LC, McDonald DW, Hendzel MJ . Kdm4b histone demethylase is a DNA damage response protein and confers a survival advantage following γ-irradiation. J Biol Chem 2013; 288: 21376–21388.
Chung YG, Matoba S, Liu Y, Eum JH, Lu F, Jiang W et al. Histone demethylase expression enhances human somatic cell nuclear transfer efficiency and promotes derivation of pluripotent stem cells. Cell Stem Cell 2015; 17: 758–766.
Shin S, Janknecht R . Activation of androgen receptor by histone demethylases JMJD2A and JMJD2D. Biochem Biophys Res Commun 2007; 359: 742–746.
Wissmann M, Yin N, Muller JM, Greschik H, Fodor BD, Jenuwein T et al. Cooperative demethylation by JMJD2C and LSD1 promotes androgen receptor-dependent gene expression. Nat Cell Biol 2007; 9: 347–353.
Coffey K, Rogerson L, Ryan-Munden C, Alkharaif D, Stockley J, Heer R et al. The lysine demethylase, KDM4B, is a key molecule in androgen receptor signalling and turnover. Nucleic Acids Res 2013; 41: 4433–4446.
Shi L, Sun L, Li Q, Liang J, Yu W, Yi X et al. Histone demethylase JMJD2B coordinates H3K4/H3K9 methylation and promotes hormonally responsive breast carcinogenesis. Proc Natl Acad Sci 2011; 108: 7541–7546.
Chu C-H, Wang L-Y, Hsu K-C, Chen C-C, Cheng H-H, Wang S-M et al. KDM4B as a target for prostate cancer: structural analysis and selective inhibition by a novel inhibitor. J Med Chem 2014; 57: 5975–5985.
Zhang D, Yoon H-G, Wong J . JMJD2A is a novel N-CoR-interacting protein and is involved in repression of the human transcription factor achaete scute-like homologue 2 (ASCL2/Hash2). Mol Cell Biol 2005; 25: 6404–6414.
Kim T-D, Shin S, Berry WL, Oh S, Janknecht R . The JMJD2A demethylase regulates apoptosis and proliferation in colon cancer cells. J Cell Biochem 2012; 113: 1368–1376.
Gray SG, Iglesias AH, Lizcano F, Villanueva R, Camelo S, Jingu H et al. Functional characterization of JMJD2A, a histone deacetylase- and retinoblastoma-binding protein. J Biol Chem 2005; 280: 28507–28518.
Wang L, Chang J, Varghese D, Dellinger M, Kumar S, Best AM et al. A small molecule modulates Jumonji histone demethylase activity and selectively inhibits cancer growth. Nat Commun 2013; 4: 2035.
Hamada S, Kim T-D, Suzuki T, Itoh Y, Tsumoto H, Nakagawa H et al. Synthesis and activity of N-oxalylglycine and its derivatives as Jumonji C-domain-containing histone lysine demethylase inhibitors. Bioorg Med Chem Lett 2009; 19: 2852–2855.
Duan L, Rai G, Roggero C, Zhang QJ, Wei Q, Ma SH et al. KDM4/JMJD2 histone demethylase inhibitors block prostate tumor growth by suppressing the expression of AR and BMYB-regulated genes. Chem Biol 2015; 22: 1185–1196.
Zhang B, Chen J-J, Shen JHC, Rivera Rosado L, Zhang Y, Di X . Mislocalization of death receptors correlates with cellular resistance to their cognate ligands in human breast cancer cells. Oncotarget 2012; 3: 833–842.
Ren Y-G, Wagner KW, Knee DA, Aza-Blanc P, Nasoff M, Deveraux QL . Differential regulation of the TRAIL death receptors DR4 and DR5 by the signal recognition particle. Mol Biol Cell 2004; 15: 5064–5074.
Dimberg LY, Anderson CK, Camidge R, Behbakht K, Thorburn A, Ford HL . On the TRAIL to successful cancer therapy[quest] predicting and counteracting resistance against TRAIL-based therapeutics. Oncogene 2013; 32: 1341–1350.
Mallette Frédérick A, Richard S . JMJD2A promotes cellular transformation by blocking cellular senescence through transcriptional repression of the tumor suppressor CHD5. Cell Rep 2: 1233–1243.
Klose RJ, Yamane K, Bae Y, Zhang D, Erdjument-Bromage H, Tempst P et al. The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine[thinsp]9 and lysine[thinsp]36. Nature 2006; 442: 312–316.
Das Partha P, Shao Z, Beyaz S, Apostolou E, Pinello L, Angeles Alejandro De L et al. Distinct and combinatorial functions of Jmjd2b/Kdm4b and Jmjd2c/Kdm4c in mouse embryonic stem cell identity. Mol Cell 53: 32–48.
Guo L, Li X, Huang JX, Huang HY, Zhang YY, Qian SW et al. Histone demethylase Kdm4b functions as a co-factor of C/EBP[beta] to promote mitotic clonal expansion during differentiation of 3T3-L1 preadipocytes. Cell Death Differ 2012; 19: 1917–1927.
Yang J, AlTahan AM, Hu D, Wang Y, Cheng P-H, Morton CL et al. The role of histone demethylase KDM4B in Myc signaling in neuroblastoma. J Natl Cancer Inst 2015; 107: djv080.
Salifou K, Ray S, Verrier L, Aguirrebengoa M, Trouche D, Panov KI et al. The histone demethylase JMJD2A/KDM4A links ribosomal RNA transcription to nutrients and growth factors availability. Nat Commun 2016; 7: 10174.
Kepp O, Rajalingam K, Kimmig S, Rudel T . Bak and Bax are non-redundant during infection- and DNA damage-induced apoptosis. EMBO J 2007; 26: 825–834.
Yang P, Guo L, Duan ZJ, Tepper CG, Xue L, Chen X et al. Histone methyltransferase NSD2/MMSET mediates constitutive NF-kappaB signaling for cancer cell proliferation, survival, and tumor growth via a feed-forward loop. Mol Cell Biol 2012; 32: 3121–3131.
Wang J, Zou JX, Xue X, Cai D, Zhang Y, Duan Z et al. ROR-[gamma] drives androgen receptor expression and represents a therapeutic target in castration-resistant prostate cancer. Nat Med 2016; 22: 488–496.
Chen JJ, Bozza WP, Di X, Zhang Y, Hallett W, Zhang B . H-Ras regulation of TRAIL death receptor mediated apoptosis. Oncotarget 2014; 5: 5125–5137.
Acknowledgements
This work was supported in part by grants from the NIH (R01CA206222) and US Department of Veterans Affairs, Office of R&D (I01BX002237) to H-WC, the National Natural Science Foundation of China (No. 81373655) and Natural Science Foundation of Guangdong Province, China (No. 2013B021800226) to HW, and the NIH (R01CA165263) to H-JK.
Author contributions
H-WC, HW and JW conceived the research. H-WC, JW, H-JK and HW designed the research. JW, L-YW, DC, ZD, YZ, PC and JXZ performed research. H-WC, JW, JXZ and HW analyzed data. H-WC, JW, HW, XC and H-JK wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by S Kaufmann
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, J., Wang, H., Wang, LY. et al. Silencing the epigenetic silencer KDM4A for TRAIL and DR5 simultaneous induction and antitumor therapy. Cell Death Differ 23, 1886–1896 (2016). https://doi.org/10.1038/cdd.2016.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2016.92
This article is cited by
-
Zinc finger myeloid Nervy DEAF-1 type (ZMYND) domain containing proteins exert molecular interactions to implicate in carcinogenesis
Discover Oncology (2022)
-
KDM4B is a coactivator of c-Jun and involved in gastric carcinogenesis
Cell Death & Disease (2019)
-
Elucidation for modulation of death receptor (DR) 5 to strengthen apoptotic signals in cancer cells
Archives of Pharmacal Research (2019)
-
Natural Agents-Mediated Targeting of Histone Deacetylases
Archivum Immunologiae et Therapiae Experimentalis (2018)
-
Consensus report of the 8 and 9th Weinman Symposia on Gene x Environment Interaction in carcinogenesis: novel opportunities for precision medicine
Cell Death & Differentiation (2018)